A Peptide Microarray Technology for Specificity Profiling of Antibodies
Abstract
Source: Cornett, E. M., et al. Analysis of Histone Antibody Specificity with Peptide Microarrays. J. Vis. Exp. (2017).
This video demonstrates a technique that applies peptide microarray technology to profile the specificity of antibodies recognizing post-translational modifications (PTMs) on histone peptides. A microarray slide containing immobilized fluorophore-conjugated histone peptides with known PTM combinations is treated with target PTM-specific antibodies. This process helps identify the antibody's specificity for the target and its potential cross-reactivity with non-target PTMs.
Protocol
1. Designing the Array Slide and Source Plate Layout
- Under the 'Plan a slide layout' heading on the ArrayNinja interface, click on the link that corresponds to the microarray printer being used.
NOTE: ArrayNinja has been programmed to mimic the robotic movement of two commonly used microarray printers (see Table 1). Compatibility with the other arrayers can be configured upon request. - Click inside the 'Load plate' dialog box, type "empty", and click 'enter.' See Figure 1 for a screenshot of the ArrayNinja design module.
- Adjust the spot diameter to 275 µm and spot spacing to 375 µm. Adjust the remaining settings (Plate Blocks/plate row, Total plate Rows, replicates, features in y, Super Arrays, SuperA fudge; see Figure 1) to customize how the features will appear on the microarray slide.
NOTE: The spot diameter is determined by the size of the microarray pin. As these settings are adjusted, the cartoon slide will update in real-time. Use this cartoon to preview how each setting is modifying the final slide layout. - After the layout of features on the slide is finalized, mouse over each unique feature and enter the feature identifier in the pop-up dialog box.
NOTE: Feature identifiers can consist of numbers, letters, or combinations. This is only required for unique features, and a dialog box will not appear when replicates are selected. - After all unique features have been assigned an identifier, click 'populate.' Enter a name for the slide layout and click 'print your plate' to save. A new page will open displaying the number of 384-well source plates needed to fabricate the chosen slide design and a table mapping the physical location of each feature to be loaded in the source plate(s).
NOTE: Remember this name, as it will be used to recall this design when analyzing microarray data. Clicking 'print your plate' saves the layout within ArrayNinja.
2. Fabricating Microarrays
- Preparing the Source Plate
- Use the map generated with ArrayNinja to create the 384-well source plate(s).
NOTE: Detailed descriptions of peptides queried on this platform can be found elsewhere. - Deposit 1 – 2 µL of each feature (e.g., biotinylated histone peptide) into the correct well of the 384-well source plate(s).
NOTE: Biotinylated histone peptides are typically deposited from 200 – 400 µM stock solutions, which equates to 10 to 25-fold molar excess of the peptide to streptavidin binding sites in a single array spot. This is calculated using the equation:
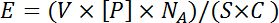
where V is the volume delivered by a pin, [P] is the concentration of the feature being printed, NA is Avogadro's number, and S is the surface area of a spot. C is the coverage of the slide, expressed as the number of streptavidin molecules per unit area multiplied by three (the average number of available streptavidin binding sites). V and C are obtained by the respective manufacturer's. Other features may require different concentrations depending on the size of the biomolecule where crowding may be a concern. A range of printing concentrations for each new type of feature should be tested empirically to determine the optimum printing concentration. - Dilute each feature 10-fold with 1x printing buffer supplemented with 1% bovine serum albumin (BSA). Spin the source plate(s) at 500 x g for 2 min at room temperature.
NOTE: The inclusion of fluorescein-labeled biotin (5 µg/µL) in the printing buffer is recommended as a spotting control and as a visual aid to facilitate proper array alignment during analysis (see Figure 3).
- Use the map generated with ArrayNinja to create the 384-well source plate(s).
- Printing Protocol (Figure 2A – B).
- Prepare the arrayer by emptying the waste collection container and filling the wash solution container and humidifier container with sterile distilled water.
- Enter the parameters used to design the slide in ArrayNinja (Figure 1) into the microarray printer control program.
- Using the microarray printer control program, set the washing procedure for a 1 s wash with one immersion. Set the post-wash settings to re-dip the pins 5 times following each wash. Set the humidity to 60%.
NOTE: The optimal wash configuration may vary depending on the microarray printer being used. The optimal post-wash pin dip configuration may vary depending on the microarray printer being used. - Insert functionalized slides (e.g., streptavidin-coated glass) into the substrate platens and place all platens into the platen elevator. Insert the source plate(s) into the plate holder(s) and place into the source plate elevator.
- Click 'print'. Monitor the printing process for a few rounds of feature deposition to ensure all wash and dip settings are correct. When the print run is complete, remove the substrate platens from the arrayer.
NOTE: When large print runs prohibit completion of blocking steps within one day, printed slides can be incubated in a humidified chamber at 4 °C overnight. Incubate the slides next to a small beaker of water inside a cardboard box sealed with plastic wrap. - Block the slides with blocking buffer for 30 min at room temperature with mixing.
- Wash the slides 2 x 10 min at room temperature in phosphate buffered saline (PBS), pH 7.6 with mixing. Dry the slides by spinning in a microarray slide centrifuge for 30 s at room temperature.
NOTE: For processing a large number of slides at once, a high-throughput microscope slide washing chamber can be used, allowing 50 slides to be washed in parallel. - For slides designed to be partitioned with wax, proceed to section 2.1. For all other designs, store slides at 4 °C protected from light and moisture.
NOTE: Printed biotinylated histone peptides are stable for at least 6 months when stored this way.
2. Partitioning Microarray Slides
- Hydrophobic Wax Pen (Figure 2C)
- Apply wax around the areas that contain features using a wax pen. Allow wax to air dry for 5 min before proceeding to section 3.
NOTE: After blocking, the array spots can be very difficult to visualize by eye. The slide design from ArrayNinja can be printed to scale and used as a guide for applying wax.
- Apply wax around the areas that contain features using a wax pen. Allow wax to air dry for 5 min before proceeding to section 3.
- Silicon gasket (Figure 2D)
- Peel the clear film off the back of the array gasket and place adhesive side down onto the microarray slide.
- Hold the gasket in place for 5 s prior to proceeding to section 3.
- Wax Imprint (Figure 2E)
- For slides designed to be partitioned by wax, print a test slide on plain glass using 10% BSA. Use this test slide to optimize the wax imprinter guides or array settings to ensure all features will be within the wax-mold chambers.
- Heat the microarray wax imprinter to 85 °C until all the wax is completely melted, approximately 30 min.
- Insert the slide with the printed side facing down and push the slide all the way to the right guide on the microarray imprinter. Pull up the lever to bring the mold into contact with the surface of the slide. Hold for 2 s.
NOTE: The hold time can be altered to achieve optimal wax border thickness. - Quickly remove the slide and visually inspect the wax borders to ensure all wells are enclosed and that the borders are not so thick that they encroach on the spots. Store slides at 4 °C protected from moisture and light.
NOTE: The hold time can be increased or decreased to obtain thicker or thinner borders.
3. Hybridizing a Histone PTM Antibody with a Peptide Microarray
- Prepare hybridization buffer (PBS, pH 7.6, 5% BSA, 0.1% Tween-20).
- Equilibrate the slide in hybridization buffer using a hybridization vessel. Completely cover the entire slide in hybridization buffer and incubate for 30 min at 4 °C on an orbital shaker at low speed.
- Prepare a solution containing diluted histone PTM antibody in hybridization buffer.
NOTE: In the example data, both histone Antibody #1 and Antibody #2 were diluted 1:1,000 in hybridization buffer. A dilution range similar to that used for immunoblotting is recommended as a starting point. - Incubate the array with the antibody solution for 1 h at 4 °C. Remove the antibody solution and wash the array 3 times for 5 min at 4 °C with cold PBS, pH 7.6.
- Prepare a 1:5,000 – 1:10,000 dilution of fluorescent dye-conjugated secondary antibody in hybridization buffer.
- Incubate the array with secondary antibody solution for 30 min at 4 °C protected from light. Remove the secondary antibody solution and wash the microarray slide 3 times for 5 min at 4 °C with PBS, pH 7.6. Dip the microarray in a 50-mL conical tube containing 0.1x PBS, pH 7.6 to remove excess salt at room temperature. Dry the slide in a microarray slide centrifuge at room temperature.
- Scan the slide with a microarray scanner at 25 µm resolution or higher, following the microarray scanner manufacturer's recommended scanning protocol.
NOTE: If fluorescein-labeled biotin tracer is present, scan both the green channel (ex: 488 nm, em: 509 nm) and the channel that corresponds to the fluorescent dye-conjugated secondary antibody, typically red (ex: 635 nm, em: 677 nm). The goal of scanning is to obtain single channel .tif files that can be merged into a single .png file.
Representative Results

Figure 1: ArrayNinja Design Module. A screen shot of the ArrayNinja design module is shown in the dotted line. The control panel (top) shows all of the parameters that can be altered on the microarray printer. As these parameters are adjusted, the cartoon image of the slide layout (bottom left) updates in real time. After the layout is set, the user can mouse over individual spots to enter unique feature identifiers. ArrayNinja constructs from this user input a map of the position of each feature in the source plate(s) (bottom right) needed to fabricate a specified microarray slide layout.
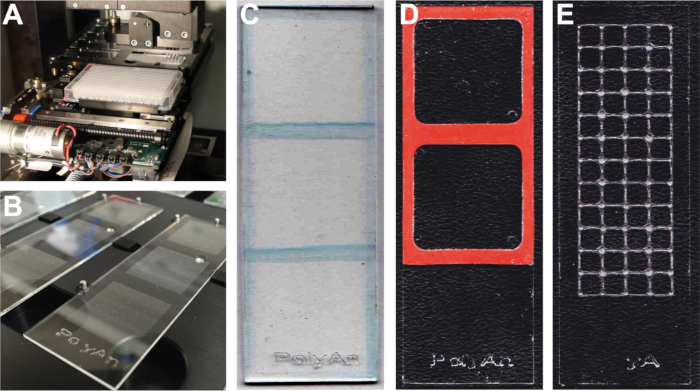

Figure 2: Microarray Fabrication. (A) Histone peptide microarray fabrication on streptavidin-coated microscope slides using a contact microarray printer. (B) Microarrays fabricated with 3 subarrays of a 48 x 48 grid of peptide features. Separation of (C) 3 subarrays with a hydrophobic wax pen, (D) 2 subarrays with a silicon adhesive, and (E) 48 subarrays with a wax imprint. All microarrays shown are fabricated using 25 x 75 mm microscope slides.
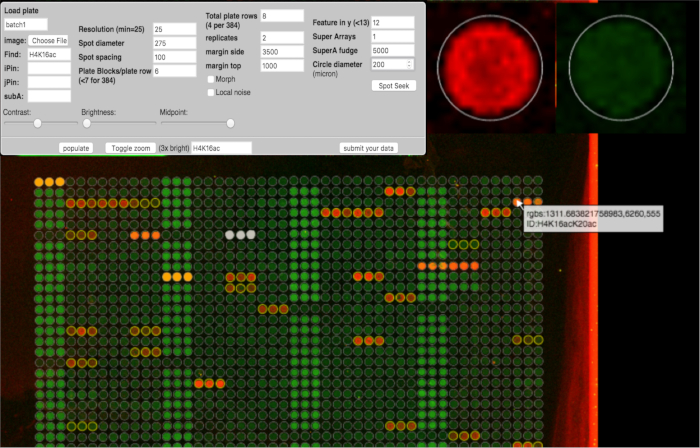
Figure 3: ArrayNinja Analysis Module. A screen shot of the ArrayNinja analysis module is shown. The control panel (top left) shows all of the parameters that can be adjusted to visualize the array, find spots, and align a grid over the array image. Hovering the mouse over a feature shows a zoomed-in view (top right) and displays a popup that contains the identification information associated with that feature (bottom). Reference spots selected for background correction are orange. Features to be excluded from downstream analysis are white. ArrayNinja contains a text-based search feature that highlights matching features in yellow, as shown in the example for H4K16.
Disclosures
The authors have nothing to disclose.
Materials
| Printing Buffer | ArrayIt | PPB | |
| BSA | Omnipure | 2390 | |
| Streptavidin-coated glass microscope slides | Greiner Bio-one | 439003-25 | |
| polypropylene 384 well plate | Greiner Bio-one | 784201 | |
| Biotin-fluorescein | Sigma | 53608 | |
| contact microarray printer | Aushon | 2470 | Aushon 2470 Microarray Printer |
| contact microarray printer | Gene Machines | OmniGrid 100 | OmniGrid Microarray Printer |
| PBS | Invitrogen | 14190 | |
| Blocking Buffer | ArrayIt | SBB | |
| Hydrophobic wax pen | Vector Labs | H-4000 | ImmEdge Hydrophobic Barrier PAP Pen |
| Silicon Gasket | Grace Bio-labs | 622511 | |
| Hybridization Vessel | Thermo Scientific | 267061 | or similar vessel |
| Fluorescent-dye conjugated secondary antibody | Life Technologies | A-21244 | Alexa Fluor 647 (anti-rabbit) |
| Fluorescent-dye conjugated secondary antibody | Life Technologies | A-21235 | Alexa Fluor 647 (anti-mouse) |
| Wax Imprinter | ArrayIt | MSI48 | |
| Tween-20 | Omnipure | 9490 | |
| Microarray Scanner | Innopsys | InnoScan 1100AL | or equivalent microarray scanner |
| EipTitan Histone Peptide Microarray | Epicypher | 112001 | |
| AbSurance Pro Histone Peptide Microarray | Millipore | 16668 | |
| MODified Histone Peptide Array | Active Motif | 13001 | |
| Histone Code Peptide Microarrays | JPT | His_MA_01 | |
| Wax | Royal Oak | GulfWax | for wax imprinter |
| Humidified Microarray Slide Hybridization Chamber | VWR | 97000-284 | |
| High throughput microscope slide washing chamber | ArrayIt | HTW | |
| Microscope slide centrifuge | VWR | 93000-204 | |
| Antibody 1 | Abcam | 8898 | |
| Antibody 2 | Millipore | 07-473 | |
| Biotinylated histone peptide | EpiCypher | 12-2001 | Example peptide. Similar peptides with various modifications are available from several commercial sources. |
| ImageMagick | https://www.imagemagick.org/script/index.php | ||
| ArrayNinja | https://rothbartlab.vai.org/tools/ |

