Evaluation of Polymeric Gene Delivery Nanoparticles by Nanoparticle Tracking Analysis and High-throughput Flow Cytometry
Summary
A protocol for nanoparticle tracking analysis (NTA) and high-throughput flow cytometry to evaluate polymeric gene delivery nanoparticles is described. NTA is utilized to characterize the nanoparticle particle size distribution and the plasmid per particle distribution. High-throughput flow cytometry enables quantitative transfection efficacy evaluation for a library of gene delivery biomaterials.
Abstract
Non-viral gene delivery using polymeric nanoparticles has emerged as an attractive approach for gene therapy to treat genetic diseases1 and as a technology for regenerative medicine2. Unlike viruses, which have significant safety issues, polymeric nanoparticles can be designed to be non-toxic, non-immunogenic, non-mutagenic, easier to synthesize, chemically versatile, capable of carrying larger nucleic acid cargo and biodegradable and/or environmentally responsive. Cationic polymers self-assemble with negatively charged DNA via electrostatic interaction to form complexes on the order of 100 nm that are commonly termed polymeric nanoparticles. Examples of biomaterials used to form nanoscale polycationic gene delivery nanoparticles include polylysine, polyphosphoesters, poly(amidoamines)s and polyethylenimine (PEI), which is a non-degradable off-the-shelf cationic polymer commonly used for nucleic acid delivery1,3 . Poly(beta-amino ester)s (PBAEs) are a newer class of cationic polymers4 that are hydrolytically degradable5,6 and have been shown to be effective at gene delivery to hard-to-transfect cell types such as human retinal endothelial cells (HRECs)7, mouse mammary epithelial cells8, human brain cancer cells9 and macrovascular (human umbilical vein, HUVECs) endothelial cells10.
A new protocol to characterize polymeric nanoparticles utilizing nanoparticle tracking analysis (NTA) is described. In this approach, both the particle size distribution and the distribution of the number of plasmids per particle are obtained11. In addition, a high-throughput 96-well plate transfection assay for rapid screening of the transfection efficacy of polymeric nanoparticles is presented. In this protocol, poly(beta-amino ester)s (PBAEs) are used as model polymers and human retinal endothelial cells (HRECs) are used as model human cells. This protocol can be easily adapted to evaluate any polymeric nanoparticle and any cell type of interest in a multi-well plate format.
Introduction
The determination of the number of plasmids complexed per nanoparticle is important to design effective nanoparticle-based gene delivery strategies, particularly for co-delivery of multiple plasmids to the same cell target, as often is required in stem cell reprogramming studies12. Few approaches to calculate the number of plasmids associated with a single nanoparticle have been described, and each approach has drawbacks in the techniques used for estimation13-16. Quantum dot (QD) labeling combined with TEM has been used to estimate plasmids per particle in chitosan-based nanoparticles. Estimation with this QD technique is complicated due to the need to label the DNA, which may alter its self-assembly properties; the possibility that encapsulated unlabeled DNA is not directly detected; potentially overlapping plasmids and QDs in the 2D TEM images of particles; and other simplifying assumptions13. An alternative approach that is applicable when ordered microdomains exist in the particles has been used to study Lipopolyamine-DNA complexes via cryo-transmission electron microscopy (cryo-TEM), X-ray scattering, and dynamic light scattering (DLS)14,15 . Unfortunately, materials such as the polymeric nanoparticles investigated here are not applicable with this method. In another study, Collins et al. used a flow particle image analysis technique to study (Lys)16-containing peptide/DNA complexes; however, their method can only evaluate larger, micron-sized particles16. Thus, we recently developed a novel and flexible assay to quantify the number of plasmids per nanoparticle11.
Protocol
1. Cell Seeding
- Do not allow cells to grow to overconfluency. Use early passage cells when transfecting primary cells.
- Twenty-four hours prior to transfection, trypsinize the cells, count the cells using a hemocytometer, and dilute the cell suspension with media to achieve the desired cell density (cells/volume). Seed cells into clear tissue culture-treated flat-bottom 96-well plates using a reservoir and multichannel pipettes. The chosen density should give 70-80% confluency on the day of transfection. For example, as displayed in Table 1, cells were diluted to 25 to 50 cells/μl for the transfection data shown here.
2. Cell Transfection
- Dilution of polymer and DNA stocks. Thaw polymer and DNA stock solutions at room temperature (RT). Dilute polymer stock solution and DNA stock solution, separately in clear 96-well plates using a twelve-channel pipette, with the appropriate solvent to concentrations required to obtain the desired polymer weight to DNA weight ratios (wt/wt). In this case, the chosen solvent is 25 mM sodium acetate buffer (pH=5.2).
- DNA dilution. Typically, DNA stored at 1 mg/ml is diluted in sodium acetate buffer to a concentration of 0.03 to 0.06 mg/ml in a clear non-tissue culture-treated 96-well plate (one well for a single formulation). Table 2 shows the typical DNA dilution protocol for a single formulation used to transfect four replicate wells in a 96-well plate seeded with cells from Step 1.
- Polymer dilution. The 100 mg/ml polymer/DMSO solution is diluted in sodium acetate buffer according to the concentration required to obtain the desired polymer to DNA wt/wt ratio. The range of wt/wt ratios typically used for gene delivery with poly(beta-amino ester)s (PBAEs) is 20 to 100. PBAE polymers are first diluted to 10 mg/ml, followed by the dilution protocol as shown in Table 3. The polymer dilutions can be performed in a clear non-tissue culture-treated 96-well plate that matches the sample orientation of the DNA dilution plate.
- Nanoparticle formation. Add the PBAE solution to an equal volume of the plasmid DNA solution using a twelve-channel pipette and mix vigorously. Let the mixture incubate at RT for 10 min to allow self-assembly.
- Nanoparticle transfection. Following self-assembly, 20 μl nanoparticles are added per well to the culture medium dropwise using the twelve-channel pipette. Replicate wells are left untreated or are transfected with commercially available reagents as controls. The transfected cells are incubated at 37 °C for two to four hours and then the wells are replaced with fresh media (100 μl/well). The choice of incubation time will depend on the cell line, culture conditions, and transfection system. The difference in transfection efficiency and cell toxicity as a result of varying incubation period will be quantified by the analysis protocol described in Step 3.
3. Analysis of Transfection Efficiency and Cell Toxicity
Transfection efficiency is analyzed visually with a fluorescence microscope and quantified using a flow cytometer forty-eight hours post-transfection. Cell toxicity is analyzed visually with a fluorescence microscope and quantified using the CellTiter 96 AQueous One assay twenty-four hours post-transfection.
- Fluorescence microscopy. Visually analyze the wells for the expression of the transfected reporter gene (for example, EGFP or DsRed) using the appropriate fluorescence channel. Acquire images for each well, choosing fields of view that appropriately represent the transfection efficiency for the particular formulations (Figure 1). Make a note of any cell toxicity observed visually.
- Flow Cytometry.
- Use multichannel pipettes to prepare the 96-well plate for flow cytometry. Wash with PBS, trypsinize with 30 μl/well of trypsin/EDTA, neutralize with 170 μl FACS buffer (PBS + 2% FBS), pipette up and down in each well, including around edges, and transfer the entire 200-μl volume of each well into corresponding wells in a round-bottom or V-bottom 96-well plate.
- Centrifuge the plate at 130 x g for 5 min at 4 °C.
- Remove 170 μl media from each well and pipette to resuspend cells while avoiding bubbles. To use a viability stain like propidium iodide (PI), add 10 μl of FACS buffer+PI (50:1 FACS buffer:PI dilution; PI stock concentration 1 mg/ml) to each well. Immediately place on ice and cover from light.
- Start the C6 Accuri flow cytometer, the HyperCyt 96-well attachment, and the HyperView software. Use HyperView Design and Protocol tabs to choose the appropriate plate type, plate layout, and other settings such as shake/sip/rinse/shake time. For the above protocol, the following parameters were used, a pre-plate prime for one minute, and a pre-plate shake for 30 sec at 2,400 rpm. Sip time was set for 15 sec at 15 rpm, a probe rinse every well for 2 sec, and an inter-well shake every 12 wells for 12 sec at 2,400 rpm. The inter-well shake is important to keep the cells dispersed in the media, as well as for improved ability to process the data by including timed breaks between groups of wells (in this case 12 sec).
- Use HyperView Well Identification tab to process the data and separate into the appropriate well (Figure 2). In order to identify the individual wells, it can help to use the software’s noise filter, as well as including a base even filter under the advanced settings.
- Quantify percentage of positively transfected live cells by appropriate gating to separate different populations (Figure 3). First view the flow data on a FSC vs. SSC scatter plot in order to separate cells from debris (Figure 3A). If propidium iodide (PI) was used, gate the individual cell population and view that data on a FSC vs. FL3 plot in order to separate live cells from dead cells and gate the live cells. In order to quantify the number of transfected cells, view the live cell population on the appropriate channels, for example, GFP can be seen in FL1. Using the untreated population, gate the untreated cells in order to identify the background signal (Figure 3B). The positively transfected cells can then be isolated using this gate and by viewing the transfected populations using both scatter and contour plots (Figure 3C and 3D). There may be a continuum of positively transfected cells, as different cells will be transfected to different degrees of fluorescence.
- CellTiter 96 AQueous One assay
- Transfect another plate in exactly the same manner for cell toxicity measurements.
- Premix 1 ml of the CellTiter 96 AQueous One assay solution with 10 ml fresh media. Remove media from cells and use a multichannel pipette to add 110 μl of the premixed solution+media per well.
- Measure the absorbance at 490 nm every one hour interval from 1 to 4 hr until the absorbance from the well with untreated condition is in the linear range of the assay. Include four control wells with only solution+media for a background absorbance reading.
- Average the absorbance values from the four replicate wells per condition and subtract the average background absorbance from each average. Normalize the corrected average for each treated condition to the corrected average of the untreated condition to determine the percent cell viability.
4. Nanoparticle Sizing with NTA and Plasmid Per Particle Calculations
- Prime the fluidics system of the Nanosight NS500 by running the diluent pump forward at approximately 1/5th the max speed and the sample pump backward at approximately 1/10th the max speed. Continue until the fluidics system is flushed with diluent.
- Prepare the nanoparticles to be analyzed. In the case of PBAEs, separately dilute the stock plasmid DNA (1 mg/ml) and stock PBAE (100 mg/ml) in sodium acetate buffer (25 mM, pH 5) in Eppendorf tubes.
- Dilute the plasmid DNA concentration to 0.06 mg/ml. Polymer concentration can vary depending on the required weight-weight ratio of polymer to DNA (for example, for 60 wt/wt particles, polymer is diluted to 3.6 mg/ml).
- Add the polymer solution to an equal volume of the plasmid DNA solution and mix vigorously. Incubate the mixture at room temperature for 10 min to allow self-assembly.
- Dilute the nanoparticle solution 100-fold into PBS in order for the nanoparticle concentration to be in the appropriate range for NTA and to obtain a final volume of at least 500 μl.
- Load the sample into the Nanosight by placing the loading tube into the sample Eppendorf tube (Figure 4A). Make sure not to introduce air bubbles.
- In capture mode, increase the camera level until particles can be seen. Adjust the focus so that the particles look smooth (Figure 4A and 4B).
- While in standard mode, adjust the camera level past the point that all particles can be seen on screen. Then decrease the camera level to the lowest level such that the particles can all still be observed. If an intermediate camera level is needed, go into advanced mode and modify the camera shutter and gain to the appropriate levels.
- Visually check to make sure there are between 20 – 100 particles on the screen. An ideal number for nanoparticle tracking is approximately 50 particles. If there are too many or too few, flush the NS500, adjust the dilution into PBS, and re-load the sample.
- Adjust the capture duration according to the standard mode table. Typically 30 – 60 sec is an appropriate capture length (Figure 4B).
- To load the next sample, flush the system, then re-load the new sample.
- Once videos are captured, proceed to the processing stage by opening a video file.
- There are a number of parameters that can be tuned in order to best process a video (Figure 4C). The goal is to select parameters that best capture each particle on the screen, as indicated by a red cross mark on the screen over each particle (Figure 5).
- Increase the screen gain to better see the particles. Under the standard or advanced mode, select the auto adjust for the parameters by clicking the appropriate boxes. In the case that the auto settings do not adequately select particles, unclick the box and manually set the parameters (Figure 4C).
- Once all the particles are picked on screen, click the process button of the software to process the video file (Figure 4C). This provides the particle size distribution, size averages, as well as particle concentration.
- In order to make certain that the particle concentration is accurate, change the PBS dilution, such as by increasing dilution by 2x, and repeat the above procedure. Measuring multiple dilutions of the same sample is recommended to ensure that the concentration measurement of the sample is accurate. A good range for measurement is 107-109 particles/ml.
- Verify that all plasmids are incorporated into nanoparticles by running gel electrophoresis of the nanoparticles and evaluating whether any free DNA is present. If not all plasmids are encapsulated, use standard techniques to measure DNA absorbance in order to quantify total encapsulated plasmids.
- To calculate the average number of plasmids per particle, divide the total encapsulated plasmid concentration by the NTA measured particle concentration. In order to estimate the plasmid per particle sample distribution, first use the NTA particle size histogram to calculate the volume fraction of each 1-nm bin of particle. Multiply this volume fraction distribution by the total plasmid amount to obtain the number of plasmids in each bin. Divide these numbers by the number of particles in each bin to obtain the number of plasmids-per-particle for each particle size (Figure 6).
Representative Results
Figure 1 shows a fluorescence microscopy image of an example of a successful transfection of HRECs with the EGFP plasmid. The brightfield image is helpful to ensure that cells maintain their usual morphology. Additionally, cell viability assays, such as MTS or similar assays, can be used to assess the nanoparticle toxicity7. Flow cytometry, as described, can be used to quantify the transfection efficiency. When using the HyperCyt multi-well plate attachment, the data will need to be processed appropriately in order to correctly identify the wells. As can be seen in the right panel of Figure 2, when the cell counts are good (thousands of cells per well), and the fluidics are operating correctly, the individual wells are easier to pick out both manually and by the software. However, if the cell counts are too low or there is a problem with the fluidics, it becomes much more difficult to identify the individual wells (left panel of Figure 2), and the experiment likely needs to be repeated. Replacing the tubing of the Hypercyt can often fix problems with the sample flow. Once the individual well data are obtained, most common flow cytometry software can be used to analyze the exported .FCS files. In Figure 3, FlowJo is used to gate the positively transfected cells by comparing to the untreated wells.
The PBAE nanoparticles are usually between 100 – 200 nm in size as measured by the Nanosight NTA. When performing NTA, it is important that the number of nanoparticles on the screen be between 20 – 100 so that the software will be able to accurately track the particles. Figure 5A is an example of too many particles, while Figure 5B shows an example of an appropriate number. Processing the captured video should be done such that the observed particles onscreen are picked up by the software, represented with the red cross hairs. An example of when the threshold for picking up particles is too low can be seen in Figure 5C, while an example of a better threshold level is seen in Figure 5D. A different dilution of the sample can be performed to make sure that the sample is in the correct concentration range. The new particle concentration given by the Nanosight should match the new dilution. Once the size and particle concentration are obtained, the plasmid per particle average and distribution can be calculated. Example results can be seen in Figure 6. The experimental timeline is shown in Figure 7.
| Wells/Plate | Volume/Well (μl) | Cells/Well |
| 96 | 100 | 2,500 to 5,000 |
Table 1. Typical cell plating protocol for a 96-well plate format.
| Wells/Plate | Volume/Well (μl) | Particle Volume/Well (μl) | DNA/Well (μg) | DNA (μg/ μl) | DNA 1 μg/μl stock (μl) | NaAc (μl) |
| 96 | 100 | 20 | 0.6 | 0.06 | 3 | 47 |
Table 2. Typical DNA dilution protocol for a 96-well plate format.
| Polymer:DNA (wt/wt) | Particle Volume/Well (μl) | # Replicate Wells | DNA/ Well (μg) | Polymer/Well (μg) | Polymer/ 10 μg/μl stock (μl) | NaAc (μl) |
| 20 | 20 | 4 | 0.6 | 12 | 6 | 44 |
| 40 | 20 | 4 | 0.6 | 24 | 12 | 38 |
| 60 | 20 | 4 | 0.6 | 36 | 18 | 32 |
| 100 | 20 | 4 | 0.6 | 60 | 30 | 20 |
Table 3. Typical polymer dilution protocol for a 96-well plate format.
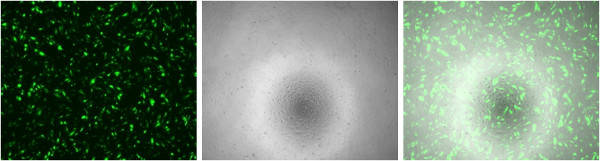
Figure 1. Single color channel fluorescence imaging of HRECs transfected with PBAE. (Left) GFP fluorescence, colored green; (Middle) Brightfield image; (Right) Composite image.
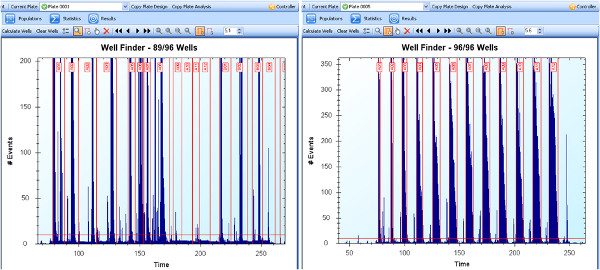
Figure 2. Hypercyt software well identification step after data collected. (Left) Example of problematic data due to low counts or issue with the fluidics; (Right) Example of clean data, with easily identified wells. Click here to view larger figure.
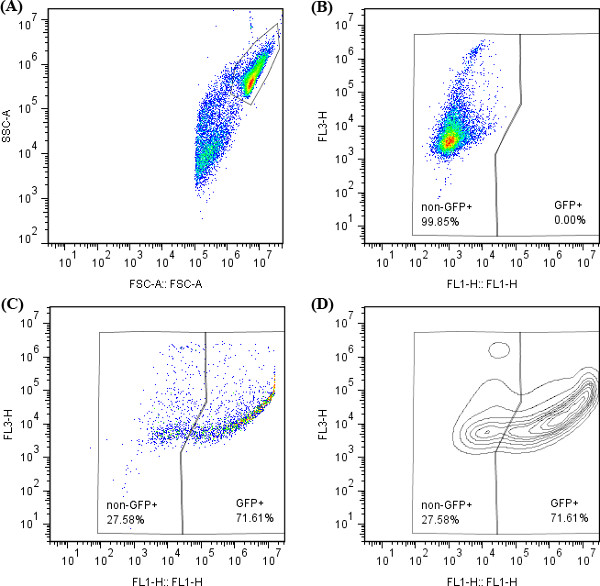
Figure 3. FlowJo gating for cells transfected with EGFP plasmid. (A) FSC vs. SSC for untreated cells; (B) FL1 vs. FL3 for untreated cells; (C, D) FL1 vs. FL3 for cells transfected with PBAE. Both pseudo-color density (C) and (D) contour plots are useful to determine the location to draw gates.
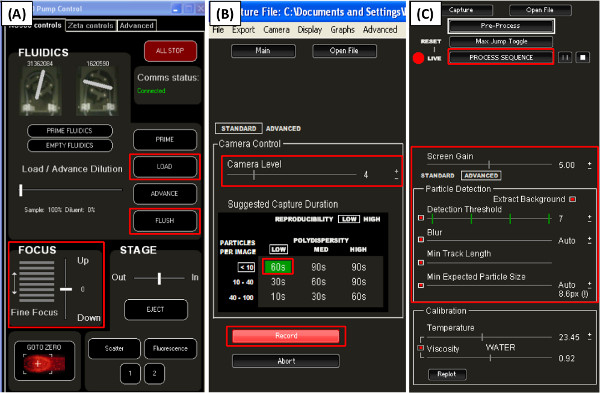
Figure 4. Screenshot of parts of the Nanosight nanoparticle tracking analysis software, version 2.2. (A) The fluidics control; (B) Capture mode, used to capture video of the nanoparticles; (C) Processing mode available after opening a previously captured video. Red boxes highlight functions discussed in the protocol. Click here to view larger figure.
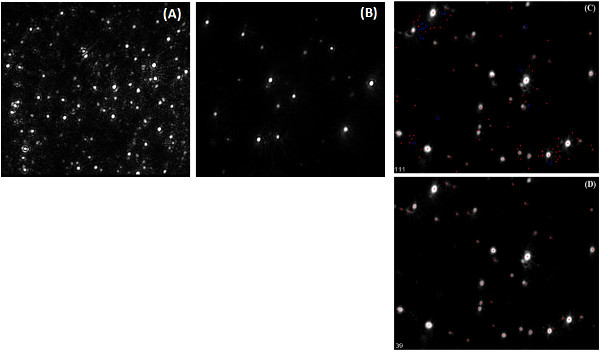

Figure 5. Example of Nanosight video capture and analysis. Screenshots of sample before video capture for sample that is not diluted enough (A), and with appropriate dilution (B); Screenshots of analysis mode with particle detection threshold set too low (C) and set appropriately (D), with red cross hairs identifying all particles appropriately. Click here to view larger figure.
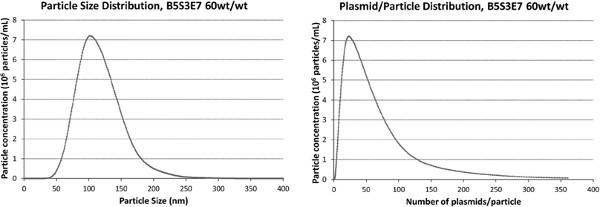
Figure 6. Size distribution and plasmid per particle distribution data of PBAE (B5S3E7, 60:1 polymer to DNA wt/wt) based nanoparticles analyzed using nanosight tracking analysis technique. Reprinted from [14] Small, 8, Bhise, N.S., Shmueli, R.B., Gonzalez, J., and Green, J.J. A novel assay for quantifying the number of plasmids encapsulated by polymer nanoparticles, 367-373, Copyright 2012, with permission from Wiley-VCH. Click here to view larger figure.
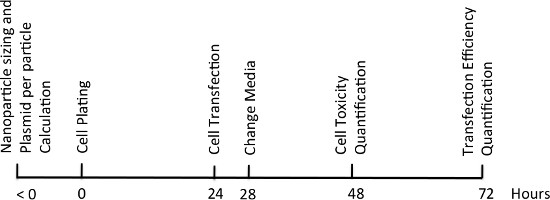
Figure 7. Experimental timeline.
Discussion
The protocols above describe methods of evaluating the transfection efficacy of nanoparticle formulations, as well as a way to characterize the particle size and DNA loading of the nanoparticles. The number of plasmids per particle is an important parameter that can help predict the effectiveness of the particle and can also be used for dose determination. Nanoparticle tracking analysis can be performed in a range of different aqueous solutions, such as those differing in salt concentration. Often this characterization is performed in PBS, to mimic physiological saline. While sizing in PBS can give a good estimate of the size of the nanoparticles in media or physiological saline, the amount of serum present can affect the size and stability of nanoparticles17. Therefore, particle characterization in various concentrations of serum can be important for certain applications as well. In order to characterize the particles in serum, an additional step is recommended due to the high background scattering of serum proteins. In this case, the particles should be fluorescently tagged, so that through the use of the fluorescence filter, the particles can be specifically tracked distinctly from serum proteins.
The plasmid per particle quantification is general enough to be used with different nanoparticle formulations, including other polymeric or inorganic particulate systems. Due to the different light scattering behavior of different materials, the video capture and processing parameters may need to be modified. Additionally, the sensitivity of the NTA may change depending on the material. The plasmid per particle protocol described above is a fast and useful method for characterizing nanoparticles.
Disclosures
The authors have nothing to disclose.
Acknowledgements
The authors thank the TEDCO MSCRF (2009-MSCRFE-0098-00) and NIH R21CA152473 for support.
Materials
| Reagent | |||
| Phosphate Buffered Saline, 1x (PBS) | Invitrogen | 10010 | |
| EGM-2MV BulletKit | Lonza | CC-3202 | |
| Trypsin | Invitrogen | 25300 | |
| Sodium acetate buffer | Sigma-Aldrich | S7899 | Dilute to 25mM in deionized water |
| Dimethyl sulfoxide | Sigma-Aldrich | 276855 | |
| pEGFP DNA | Elim Biopharmaceuticals | NA | |
| DsRed DNA | Addgene | 21718 | |
| PEI, branched | Sigma-Aldrich | 408727 | |
| CellTiter 96 AQueous One | Promega | G3580 | |
| Materials | |||
| Clear flat bottom 96-well plate, sterile | Sarstedt | 82.1581.001 | |
| Clear round bottom 96-well plate, sterile | Sarstedt | 82.1582.001 | |
| 12-channel Finnpipette | Thermo Scientific | NA | 5-50 and 50-300 μl |
| Fluorescence Microscope | Zeiss | NA | Model number: AX10 |
| C6 Accuri flow cytometer | BD Biosciences | NA | |
| HyperCyt attachment | Intellicyt | NA | |
| NS500 | Nanosight | NA |
References
- Putnam, D. Polymers for gene delivery across length scales. Nature Materials. 5, 439-451 (2006).
- Sheyn, D., et al. Genetically modified cells in regenerative medicine and tissue engineering. Advanced Drug Delivery Reviews. 62, 683-698 (2010).
- Boussif, O., et al. A versatile vector for gene and oligonucleotide transfer into cells in culture and in vivo: polyethylenimine. Proc. Natl. Acad. Sci. U.S.A. 92, 7297-7301 (1995).
- Green, J. J. Rita Schaffer Lecture: Nanoparticles for Intracellular Nucleic Acid Delivery. Ann. Biomed. Eng. 40, 1408-1418 (2011).
- Lynn, D. M., Langer, R. Degradable poly(beta-amino esters): Synthesis, characterization, and self-assembly with plasmid DNA. J. Am. Chem. Soc. 122, 10761-10768 (2000).
- Sunshine, J. C., Peng, D. Y., Green, J. J. Uptake and transfection with polymeric nanoparticles are dependent on polymer end-group structure, but largely independent of nanoparticle physical and chemical properties. Mol. Pharm. , (2012).
- Shmueli, R. B., Sunshine, J. C., Xu, Z., Duh, E. J., Green, J. J. Gene delivery nanoparticles specific for human microvasculature and macrovasculature. Nanomedicine: Nanotechnology, Biology, and Medicine. , (2012).
- Bhise, N. S., et al. The relationship between terminal functionalization and molecular weight of a gene delivery polymer and transfection efficacy in mammary epithelial 2-D cultures and 3-D organotypic cultures. Biomaterials. 31, 8088-8096 (2010).
- Tzeng, S. Y., et al. Non-viral gene delivery nanoparticles based on poly(beta-amino esters) for treatment of glioblastoma. Biomaterials. 32, 5402-5410 (2011).
- Sunshine, J., et al. Small-Molecule End-Groups of Linear Polymer Determine Cell-Type Gene-Delivery Efficacy. Adv. Mater. 21, 4947 (2009).
- Bhise, N. S., Shmueli, R. B., Gonzalez, J., Green, J. J. A novel assay for quantifying the number of plasmids encapsulated by polymer nanoparticles. Small. 8, 367-373 (2012).
- Yu, J., et al. Human induced pluripotent stem cells free of vector and transgene sequences. Science. 324, 797-801 (2009).
- Ho, Y. P., Chen, H. H., Leong, K. W., Wang, T. H. Evaluating the intracellular stability and unpacking of DNA nanocomplexes by quantum dots-FRET. Journal of Controlled Release: Official journal of the Controlled Release Society. 116, 83-89 (2006).
- Kreiss, P., et al. Plasmid DNA size does not affect the physicochemical properties of lipoplexes but modulates gene transfer efficiency. Nucleic Acids Research. 27, 3792-3798 (1999).
- Pitard, B., et al. Virus-sized self-assembling lamellar complexes between plasmid DNA and cationic micelles promote gene transfer. Proc. Natl. Acad. Sci. U.S.A. 94, 14412-14417 (1997).
- Collins, L., Kaszuba, M., Fabre, J. W. Imaging in solution of (Lys)(16)-containing bifunctional synthetic peptide/DNA nanoparticles for gene delivery. Biochimica et Biophysica Acta. 1672, 12-20 (2004).
- Green, J. J., et al. Biodegradable polymeric vectors for gene delivery to human endothelial cells. Bioconjug. Chem. 17, 1162-1169 (2006).

