Concurrent EEG and Functional MRI Recording and Integration Analysis for Dynamic Cortical Activity Imaging
Summary
An EEG-fMRI multimodal imaging method, known as the spatiotemporal fMRI-constrained EEG source imaging method, is described here. The presented method employs conditionally-active fMRI sub-maps, or priors, to guide EEG source localization in a manner that improves spatial specificity and limits erroneous results.
Abstract
Electroencephalography (EEG) and functional magnetic resonance imaging (fMRI) are two of the fundamental noninvasive methods for identifying brain activity. Multimodal methods have sought to combine the high temporal resolution of EEG with the spatial precision of fMRI, but the complexity of this approach is currently in need of improvement. The protocol presented here describes the recently developed spatiotemporal fMRI-constrained EEG source imaging method, which seeks to rectify source biases and improve EEG-fMRI source localization through the dynamic recruitment of fMRI sub-regions. The process begins with the collection of multimodal data from concurrent EEG and fMRI scans, the generation of 3D cortical models, and independent EEG and fMRI processing. The processed fMRI activation maps are then split into multiple priors, according to their location and surrounding area. These are taken as priors in a two-level hierarchical Bayesian algorithm for EEG source localization. For each window of interest (defined by the operator), specific segments of the fMRI activation map will be identified as active to optimize a parameter known as model evidence. These will be used as soft constraints on the identified cortical activity, increasing the specificity of the multimodal imaging method by reducing cross-talk and avoiding erroneous activity in other conditionally active fMRI regions. The method generates cortical maps of activity and time-courses, which may be taken as final results, or used as a basis for further analyses (analyses of correlation, causation, etc.) While the method is somewhat limited by its modalities (it will not find EEG-invisible sources), it is broadly compatible with most major processing software, and is suitable for most neuroimaging studies.
Introduction
Electroencephalography (EEG) and functional magnetic resonance imaging (fMRI) can be viewed as neuroimaging modalities with complementary features. FMRI captures brain activity with large temporal scale, as hemodynamic signals indirectly measure the underlying neuronal activity with a poor temporal resolution (on the order of seconds)1,2. In contrast, EEG directly measures the dynamic electrophysiological activity of the brain with a very high temporal resolution (millisecond level), but poor spatial resolution3,4. These properties have led to multimodal approaches designed to optimize the favorable aspects of each individual method5. Simultaneous use of EEG and fMRI allows for the excellent temporal resolution of EEG to be combined with the high spatial accuracy of fMRI to overcome the limitations associated with unimodal fMRI or EEG.
Methods for EEG and fMRI integration begin with fMRI-informed EEG source localization6,7. This technique utilizes fMRI-derived spatial information to improve EEG source localization, however, one drawback is the potential spatial bias caused by the application of fMRI as a "hard-constraint" — fMRI-derived spatial information is considered an absolute truth. This poses two large issues that must be reconciled6–8. First, it must be considered that the use of a static map of Blood Oxygen Level Dependent (BOLD) contrasts may inadvertently strengthen any erroneous activity that falls within it, while damping true activity outside of it. Second, crosstalk from sources occurring outside of the BOLD activation map may influence the presentation of true activity within the results or cause erroneous activity. Despite this, the use of the high spatial resolution of fMRI to provide prior spatial knowledge remains a favorable solution5, as the modeling of the EEG inverse problem can be constrained both in the anatomical and functional senses.
In this paper, we demonstrate a spatiotemporal fMRI-constrained EEG source imaging approach that addresses the issue of temporal mismatch between EEG and fMRI by calculating the optimal subset of fMRI priors based on a hierarchical Bayesian model9. FMRI-priors are computed in a data-driven manner from particular windows of interest in the EEG data, leading to time-variant fMRI constraints. The proposed approach utilizes the high temporal resolution of EEG to compute a current density mapping of the cortical activity, informed by the high spatial resolution of fMRI in a time-variant, spatially selective manner that accurately images dynamic neural activity.
Protocol
The protocol presented here was designed and performed in accordance with all guidelines for ethical human research as set forth by the respective Institutional Review Boards of the University of Houston and the Houston Methodist Research Institute.
1. Simultaneous EEG/fMRI Recording
- Obtain informed consent from the participant. Explain to the participant the purpose and procedure of the study, as well as the important safety measures for the simultaneous EEG/fMRI data recording process.
- Prepare the EEG cap and check impedance outside the MRI-scanner room.
- Place an appropriately sized, passive, MRI-compatible EEG cap onto the subject's head. Position the electrodes as per the 10–20 international labeling system10.
- On the EEG recording software, check the impedance of the ground and reference electrodes. To do this, click on the 'impedance' tab and select the electrode type on the software user-interface (see Figure 1).
NOTE: Exact instructions here are specific to the software used herein (see Table of Materials), and may need to be adapted to other systems. - For each electrode, use a syringe to inject electrolyte gel into the electrode, then use a cotton swab to spread the gel to ensure skin-electrode contact.
- As the impedance decreases, continue to monitor the values using the appropriate software (adjust the impedance scale as necessary, depending on the setup) to monitor impedance level properly (see Figure 1). Continue until all electrodes reach impedance levels below 10 kΩ to ensure a high-quality signal.
NOTE: Per the materials listed and utilized here, it is considered unsafe to have any electrode with an impedance level above 50 kΩ in MR-environment11. This may change depending on the design of the chosen cap and MRI settings, so please consult with the equipment manufacturer and MRI technologists to ensure the safety of the experimental setup.
- As the impedance decreases, continue to monitor the values using the appropriate software (adjust the impedance scale as necessary, depending on the setup) to monitor impedance level properly (see Figure 1). Continue until all electrodes reach impedance levels below 10 kΩ to ensure a high-quality signal.
- Simultaneous EEG/fMRI hardware setup.
- Once the EEG cap preparation is done, have the subject moved to the MR-scanner with the hardware setup described in Figure 2.
NOTE: Some details of the figure may change, depending on the system in use. - Set up the experimental paradigm display. Use a monitor located in the observation room, behind the glass window facing the front of the MR-scanner (see Figure 2). Use a head coil viewing mirror to allow subjects to view the monitor screen without moving their head or eyes while lying down.
- Display a sample image on the computer screen to ensure that subjects can comfortably view the screen, and that the paradigm will display properly. Make any necessary hardware or software adjustments.
- Once the EEG cap preparation is done, have the subject moved to the MR-scanner with the hardware setup described in Figure 2.
- Experimental paradigm (see Figure 3).
- Instruct the subject to remain still, and perform an initial T1-weighted anatomical MRI scan. If possible, use a Field of View that reaches from the bottom of the cerebellum to the top of the head, including the skull and skin.
- Start recording the EEG data (see Figure 4).
- Simultaneously click the appropriate buttons to begin the MRI recording and initiate the paradigm of interest on the presentation software. Check the EEG data recording to ensure signal quality and, if desired, appropriate markers are being recorded.
- When using the set-up described here, first click "Run" in the presentation software and enter the subject number and trial number. The paradigm will initiate upon confirming these settings.
NOTE: The paradigm employed here consisted of 10 trials in which an emotionally motivated motor response was evoked by means of visual stimulus. For each trial, subjects were asked to first rest for 50 s watching a green screen, after which the image of an unpleasant (images corresponding to surprise, anger, or disgust) or not-unpleasant (images corresponding to happiness or neutrality) face12 was presented for 10 s. Five images from each category were presented in a randomized order, and subjects were asked to squeeze a ball upon identifying a face as unpleasant, and hold the squeeze until it disappeared. - Use a Gradient-recalled Echo Planar Imaging (GR-EPI) sequence for fMRI recording (recommended); customize to suit the equipment and paradigm.
NOTE: The sequence used herein included: Echo Time (TE) = 35 ms; Repetition Time (TR) = 1,500 ms; Slice Thickness = 5 mm; Flip Angle = 90 °; Pixel Spacing: 2.75 mm x 2.75 mm. It may be necessary to use an MRI sequence that lasts slightly longer than the display of the paradigm itself, to ensure that the full paradigm is recorded without clipping.
- When using the set-up described here, first click "Run" in the presentation software and enter the subject number and trial number. The paradigm will initiate upon confirming these settings.
2. Structural MRI Data Analysis and Forward Model Generation
- Apply full segmentation and reconstruction of various surfaces from the subject's T1-weighted anatomical MRI volume using the Freesurfer image analysis suite13,14.
NOTE: A folder containing all segmentation outputs will be generated by Freesurfer. - Generate a subject-specific 3-layer Boundary Element Method (BEM) geometrical model following the instructions provided at (https://martinos.org/mne/dev/manual/appendix/bem_model.html)15 Use the available Graphical User Interface (GUI) to ensure that there is no overlap in the layers.
- Open the Freeview Application. Click "File" >> "Load Surface". Navigate to the Subjects directory in the Freesurfer folder. Open the "BEM" folder. Open the "Watershed" folder. Load the four files found here ('outer_skin_surface', 'outer_skull_surface', 'brain_surface', and 'inner_skull_surface').
- Move the slice selection sliders and look for overlap in the yellow surface layers. If overlap does occur, double-check the MRI data for anatomical defects or errors, and use the GUI drawing tools to clarify the layers.
- Load the original MRI data in the Freeview application by clicking "File" >> "Load Volume". Navigate to the subject folder and open the "mri" folder. Then click on the "orig" directory and open the structural MRI data found there (should be in.mgz or.nii format). Click "OK".
- View the greyscale image of the head. Look at the different layers of grey and black around the brain. Ensure that these layers do not have any gaps or irregularities. Use the "Color Picker" tool and select a voxel from the layer to be corrected.
- Switch to the "Freehand Voxel Edit", and click to draw on the image. Use this to fill in any defects in the MRI image. Perform correction for all layers and MRI slices, where defects occur.
NOTE: the "Polywire" and "Livewire" voxel editing tools may also be used in place of the "Freehand".
- Generate the source space based on the geometry of the pial surface.
- Perform subject-specific EEG sensor alignment (e.g., translation and rotation) to the MRI space using the Freesurfer head model overlay (Figure 5). Save the transformation.
- Open the MNE_analyze application. Click on "File" >> "Load surface". Navigate to the folder containing the subject data and load the pial surface.
- Click "File" >> "Load Digitizer Data and select the EEG file of interest (should contain digitizer data). Click "View" >> "Show Viewer". Once the viewer GUI appears, click "Options" and make sure that the "Scalp" and "Digitizer data" options are chosen. Electrodes here are shown in red, with fiducial points in yellow.
- On the main window (not the viewer), select "Adjust" >> "Coordinate alignment". Using the 'Coordinate alignment GUI', use the arrow and L/R buttons to shift and rotate the EEG electrodes in the viewer. Adjust as much as necessary. Once the alignment is done, click "Save…" at the bottom of the 'Coordinate Alignment GUI' to save the alignment.
NOTE: Typically, an even distribution of electrodes throughout the scalp with good fiducial alignment is required.
- Generate the forward model by providing the subject-specific BEM model, the source space, and the EEG sensor transformation using MNE software15.
3. Functional MRI Data Analysis
- Perform first-level (individual subject) fMRI statistical analysis using the General Linear Model (GLM) method to acquire BOLD activation maps for the tasks of interest. Correct for multiple comparisons as necessary16, using the cluster-based approach that is built into the Freesurfer group-analysis pipeline.
- Perform group-level analysis on all subjects, if desired, to acquire the BOLD activation map for all subjects in standard space (MNI or Talairach).
NOTE: The University of Oxford Centre for Functional MRI of the Brain (FMRIB) Software Library (FSL)17 and Analysis of Functional Neuroimages (AFNI)18 packages both allow for the analysis of fMRI data on the same surfaces generated by Freesurfer, making them convenient for subsequent analysis. - Use the tksurfer visualization tool to perform region-of-interest (ROI) identification by loading the fMRI activation map (both individual-level and group-level), and setting the desired FDR-corrected threshold19 (p <0.05 is used here).
NOTE: The ROIs identified from individual-level activation maps will serve as the subject-specific fMRI-derived spatial priors for subsequent source localization.- Using the fMRI activation map on the gray matter layer, extract surface patches using a connected-labeling algorithm.
NOTE: Dulmage-Mendelsohn decomposition was used in this example. - Further sub-divide the patches based on the labeling of a predefined brain atlas, so that any patch of activity covering more than one region is split.
NOTE: The atlas used here was the DKT40 atlas20 (available from Freesurfer)21. Atlases can be specialized or chosen, based on experimental preferences.
- Using the fMRI activation map on the gray matter layer, extract surface patches using a connected-labeling algorithm.
- Project the acquired group-level ROIs (which are currently in standard space) back to the individual source spaces of each subject. After performing the individual subject's structural MRI segmentation (step 2.1), the coordinate transformations between the subject and standard space are provided in the lh.sphere.reg and rh.sphere.reg files, found in the "surf" folder of the subject's Freesurfer output folder.
NOTE: All subjects will thereby share the same set of ROIs, but in their own specific model. See Figure 6 for examples of the fMRI results and resultant ROIs.
4. EEG Data Analysis
NOTE: Details in this section may be specific to the software used (See Table of Materials for more details). Please refer to the appropriate documentation if using different software packages.
- Perform scanner gradient artifacts correction through template subtraction. For this, click on the "MR Correction" button in the "Special Signal Processing" menu, and select appropriate parameters in the EEG analysis software GUI (see Figure 7). Input the appropriate parameters to the chosen scanner sequence and experimental design.
NOTE: Primary parameters include: repetition time (TR) for the MRI scan, scan type (interleaved or continuous), MRI volume markers (or gradient detection method and gradient trigger), channels for correction, and artifact template. - Remove cardioballistic artifacts through template subtraction. For this, click on the "CB Correction" button in the "Special Signal Processing" menu, and select appropriate parameters in the analysis software GUI.
NOTE: Parameters necessary here include minimum and maximum heart rate, artifact template, ECG channel, template correlation, and channels for correction. - Apply filtration. Select the button for IIR filtration at the top of the analysis GUI, under "Data Filtration". For example, apply high-pass at 0.05 Hz, low-pass at 40 Hz, and a notch-filter at the power-line frequency (60 Hz), with a roll-off of 48 dB/Hz.
NOTE: After the application of a low-pass filter at a cutoff frequency of 40 Hz, the 60 Hz notch-filter is not strictly necessary, but is employed as a safeguard against any residual power-line frequencies that may have survived due to the roll-off at filter edges. - Perform ocular artifact correction, on top of the analysis GUI: select "Transformation" >> "Artifact Rejection/Reduction" >> "Ocular Correction ICA".
- Segment the EEG data into epochs based on the specified pre- and post-stimulus time, with respect to the event timing markers. To do this, select, "Transformation" >> "Segment Analysis Functions" >> "Segmentation", then choose the marker of interest and the time segment of interest.
NOTE: Segmentation lengths should be chosen to suit the paradigm and expected brain activity of interest. - Perform manual or semi-automatic artifact rejection: select "Transformation" >> "Artifact Rejection/Reduction" >> "Artifact Rejection". When prompted, define criteria for artifacts within the three tabs of the GUI and proceed as instructed on the GUI.
- In the 'Inspection Method' tab, select choose "automatically", "semi-automatically", or "manually select artifacts" (semi-automatic mode is recommended). Then select "mark" or "remove artifacts", and specify if the corrections are for a single channel.
- In the 'Channel Selection' tab, select the channels which will be corrected for artifacts.
- In the 'Criteria' tab, select the basis by which artifacts will be identified. Make selections here to fit experimental needs. Click "OK" after selecting criteria, and artifacts will be identified and/or rejected in accordance with the selections.
- Perform baseline correction and trial averaging (if applicable).
- To perform baseline correction: select "Transformation" >> "Segment Analysis Functions" >> "Baseline Correction". To average the segmented data: select "Transformation" >> "Segment Analysis Functions" >> "Average".
5. Spatiotemporal fMRI Constraints — EG Source Imaging
- Define window size and window overlapping size (the default setting calls for a 40 ms window size with 50% (20 ms) overlap).
- Select the subject-specific ROIs set (obtained in step 3) as the spatial prior set. For each EEG segment, the algorithm will then estimate a set of weights for the subset of spatial priors that maximizes the model evidence, and computes the source covariance matrix accordingly.
- Using the resulting source covariance matrix, perform source localization for the EEG segment being analyzed, yielding the source current density results.
- Perform steps 5.2 and 5.3 for all EEG segments and, if necessary, summarize the current density results for all time segments into one complete current density time-course by averaging the overlapping portion.
NOTE: This step will result in a current density time-course of cortical activity at every source point defined in step 2.3 (this number is typically on the order of several thousands) (Figure 8). - Extract the representative current density time-course at each of the ROIs.
- Select the preferred method for summarizing the time-courses from the multiple source points within an ROI into a single signal time-course: average, first eigenvariate, etc.
- Repeat steps 5.1 to 5.5 for all subjects.
Representative Results
EEG source localization at the basic level involves the solving of the forward and inverse problem. The components required to build and solve the forward problem are shown in Figure 5C. Using a subject-specific T1 image, three layers — brain, skull, and skin — were segmented and meshed. These layers served as the inputs to generate the BEM model. Similarly, the subject's grey-matter layer was segmented from the structural MRI and used to construct the source space. EEG sensor locations were co-registered onto the head model using a series of rigid geometrical transformations. When constructed, the forward model represents how electrical activity originating from any location on the source space would give rise to the potential measurements at each EEG sensor location on the scalp.
fMRI provides 3D images of brain functional activity with excellent spatial resolution and accuracy. Conventional fMRI analysis follows the GLM methods to identify the brain voxels significantly activated by a certain task. The typical result of this analysis is an fMRI activation map: a single brain map highlighting active voxels, which can be projected onto the gray matter surface, as shown in Figure 6A. We further divide the obtained activation maps into sub-maps, each acting as a potential spatial prior for localizing the scalp potentials measured by EEG in any particular time window (Figure 6B). Figure 8 represents the focused schematic of the spatiotemporal fMRI constrained source analysis described above. Only the appropriate partial set of the fMRI activation map is used to generate the EEG source reconstruction for the corresponding EEG data segment at the specified window size. As all EEG time-windows are analyzed, a complete reconstruction of the cortical activity is achieved in a spatiotemporally specific fashion that alleviates the spatial bias of applying the same fMRI priors at all EEG time points.
We further demonstrated a successful application of the spatiotemporal fMRI constrained source analysis method when applied to a visual/motor activation task study9, in which the sequence of brain activity from visual input to motor output was recovered with high spatiotemporal accuracy (Figure 9). While there is some dependence on the user's choice of window size, the reconstructed source imaging results were generally robust to moderate changes, as shown in Figure 10. To this end, the window size should be selected by the experimenter to best fit their particular study (i.e., a window size too large could prove to be erroneous for rapid activity or oscillations, while a window size too short may miss lower frequency signals) (Figure 10).
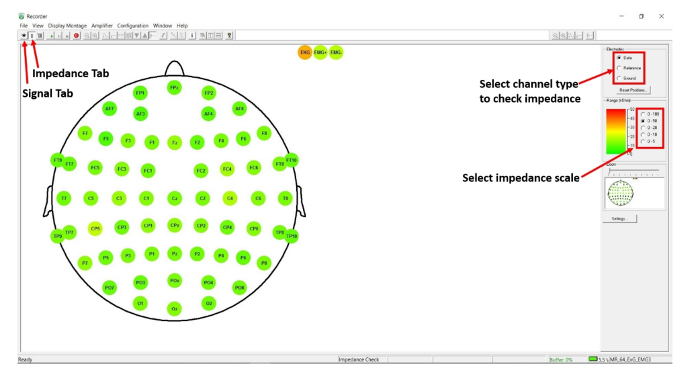
Figure 1: Scalp EEG impedance checking. Screenshot of the Recorder software user-interface, with arrows pointing to key icons in protocol step 1.2. Please click here to view a larger version of this figure.
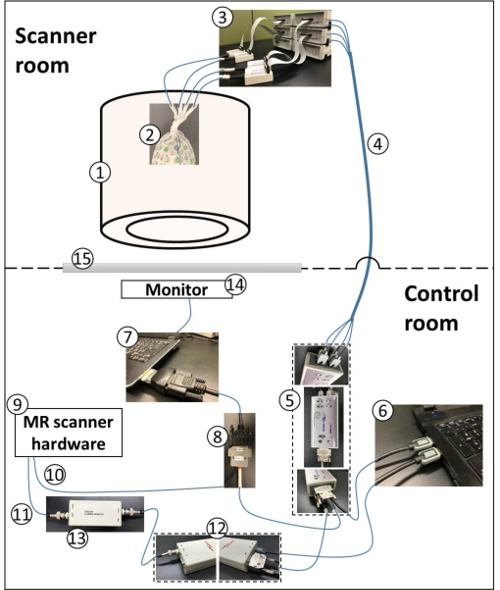
Figure 2: Schematic of simultaneous EEG/fMRI recording hardware setup — not drawn to scale. (1) Scanner; (2) participant wearing gelled EEG passive cap; (3) EEG amplifiers and Power Pack connected to the EEG cap; (4) optical fiber cables connecting the amplifiers to the USB 2 Adapter (also known as a BUA); (5) The BUA, an interface between amplifiers and the recording computer; (6) data acquisition computer; (7) Paradigm presentation computer, equipped with an express card to output event timing markers; (8) Transistor-transistor logic (TTL) trigger cable, delivering event timing markers from the presentation computer and the MR-scanner hardware to the BUA; (9) MR scanner hardware to provide timing markers at the start of (10) a new fMRI slice/volume acquisition and (11) clock synchronization signal; (12) Clock synchronization device, which provides synchronization between the clock of EEG amplifiers and the MR-scanner clock; (13) Interface module, interfacing between the MR-scanner and the clock synchronization device; (14) Monitor for the visual display of the experimental paradigm; (15) Glass window for viewing the scanner room from the control room. Please click here to view a larger version of this figure.
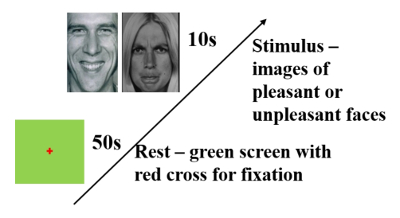

Figure 3: Experimental Paradigm. The subject was shown a series of visual stimuli, belonging to one of two categories: pleasant-face and unpleasant-face12. In each trial, a 50 s green screen baseline was first shown, followed by a randomly selected 10 s visual stimulus. The subject was to squeeze a rubber ball with his/her right hand for the entire duration of the stimulus shown, if the image was perceived as an unpleasant-face. Please click here to view a larger version of this figure.
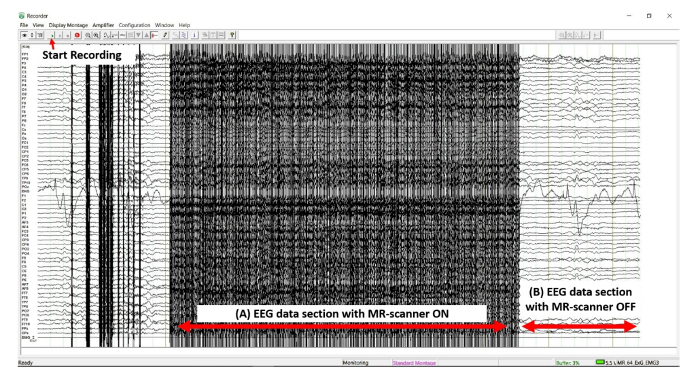

Figure 4: Screenshot of the EEG data recording. A representative section of the EEG data during the recording process. (A) Period of EEG data with fMRI pulse-sequence in effect, MR-scanner artifacts are pronounced. (B) Period of EEG data without fMRI pulse-sequence, no obvious MR-scanner artifacts are visible. Please click here to view a larger version of this figure.
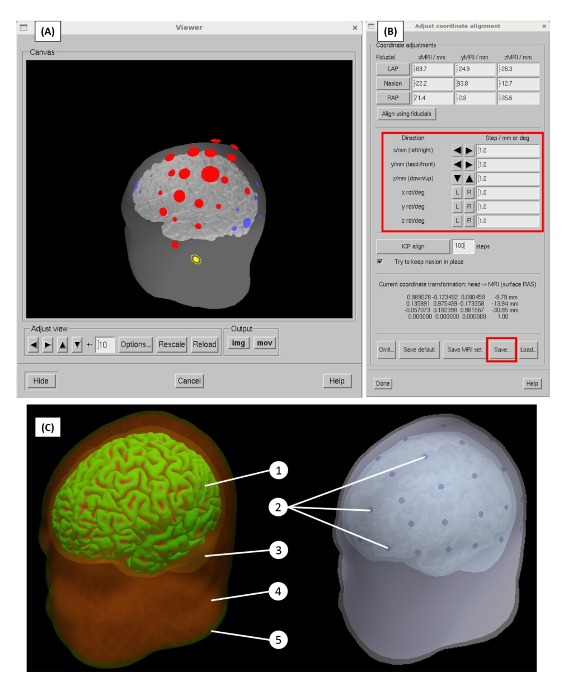
Figure 5: The forward model generation. (A) Alignment of EEG electrodes onto the head model space. Red and blue circles represent digitized EEG sensor locations, yellow circles represent the digitized EEG fiducial points: nasion, left preauricular, and right preauricular. (B) Options for the sensor alignment process, including manual transformation, such as the translation and rotation of the EEG sensor space (protocol step 2.4). (C) Subject specific BEM model generated, including 3 compartments: (3) brain, (4) skull, and (5) skin. The distributed source space on the surface of the (1) gray matter layer. (2) EEG sensor locations are aligned on the model. Please click here to view a larger version of this figure.
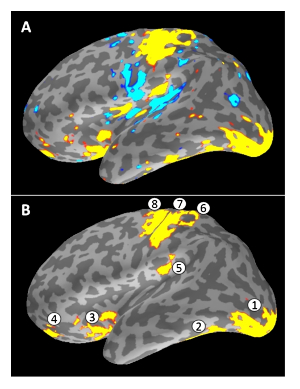
Figure 6: fMRI activation map and the extraction of regions of interest. (A) fMRI activation map shown on inflated surface for ease of inspection. Regions color coded in red and yellow are significantly activated (p-corrected<0.05). (B) 8 representative regions of interest extracted from the fMRI activation map. Note the atlas-based separation of motor activity into 3 priors. Please click here to view a larger version of this figure.
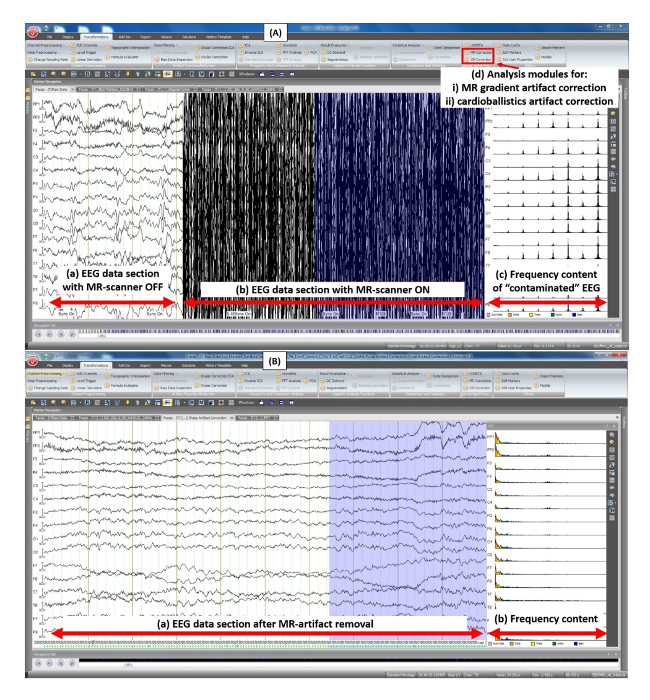
Figure 7: Screenshot of the Analyzer software user-interface — removal of MR-scanner artifacts.(A) Before scanner artifact correction: (a) EEG data section before the start of fMRI pulse-sequence; (b) EEG data section during fMRI pulse-sequence in effect, scanner artifacts are clearly visible; (c) the frequency content (FFT) of data section in (b); (d) Analyzer software's built-in analysis modules for scanner gradient-artifact correction and cardioballistic artifact correction. (B) After scanner artifact correction: (a) EEG data section after removal of MR-scanner artifacts; (b) the frequency content (FFT) of data section in (a). Please click here to view a larger version of this figure.
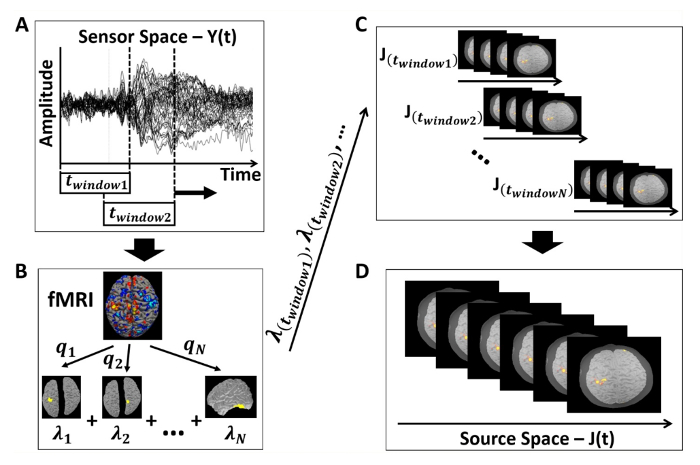
Figure 8: Overall schematic of the analysis process. (A) EEG data processing and window size selection. (B) fMRI data analysis, followed by the extraction of regions of interest to be used as spatial priors for the source analysis. (C) Source analysis performed at each EEG segment, specified by window sizes and percent overlap. (D) Complete reconstructed cortical activity over the time-course of interest. Please click here to view a larger version of this figure.
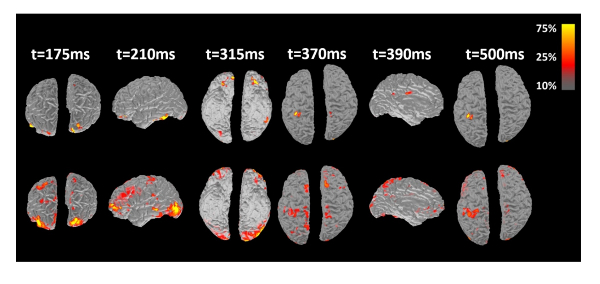
Figure 9: Reconstructed cortical activity of one representative subject underwent visual/motor activation paradigm. Source reconstruction results from contrasting two methods: spatiotemporal fMRI constrained (top) and time-invariant fMRI constrained source imaging (bottom). Figure reproduced with permission from reference9. Please click here to view a larger version of this figure.
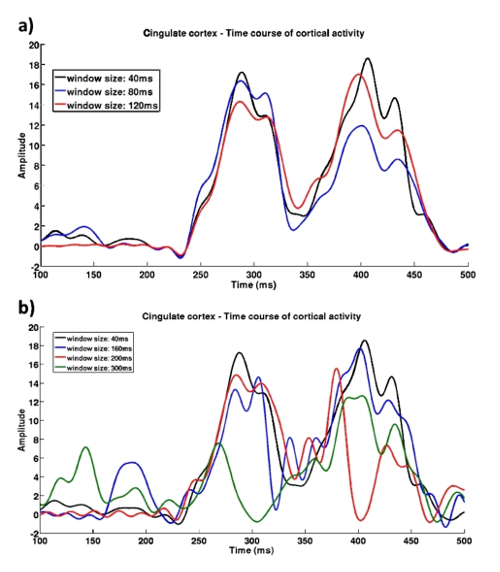
Figure 10: Reconstructed activity time-course at the cingulate cortex using different window sizes. (a) Activity time-courses reconstructed using smaller window sizes showed very similar results (correlation R >0.95). (b) Using larger window sizes resulted in high disparity (R <0.7). Figure reproduced with permission from reference9. Please click here to view a larger version of this figure.
Discussion
We have shown here the necessary steps to use the spatiotemporal fMRI constrained source analysis method for EEG/fMRI integration analysis. EEG and fMRI have become well established as the fundamental methods for non-invasively imaging brain activity, though they face difficulty in their respective spatial and temporal resolutions. While methods have been developed to capitalize on the favorable properties of each, current fMRI-constrained EEG source localization methods frequently rely upon simple fMRI constraints, which may be subject to biases and crosstalk that limit spatial accuracy (e.g., if true activity occurs outside of the MRI map, static constraints will result in the true source being diminished, while false peaks will be observed in nearby MRI-active regions. Similarly, erroneous or noise-based activity in an MRI-active region will be enhanced as if it were true). The spatiotemporal fMRI constrained approach has sought to improve on this using variable fMRI constraints in a two-layer hierarchical Bayesian model. The current source activity is estimated from EEG data in a sliding window manner. The fMRI activation map is first divided into multiple submaps, each acting as a possible spatial prior for the cortical sources. A subset of these spatial priors is selectively used as constraints to solve the EEG inverse problem. Thus, EEG and fMRI data are integrated in a spatially and temporally specific fashion. This effectively replaces the traditional fMRI activation map with a set of regions of interest that can be variably applied based on evidence from the EEG data, resulting in a data-driven approach that limits bias and error.
The methodology presented here is based on available methods (Freesurfer, FSL, etc.), and generates cortical models and processes EEG and fMRI data. While some of the procedures mentioned here do make use of specific software, most of these programs are freely available under GNU licensing. The exception to this would be BrainVision Analyzer, though separate methods may be used for this as well (specifically EEGLAB22 with the FMRIB plug-in for EEGLAB, provided by the FMRIB23,24). Similarly, the spatiotemporal fMRI-constrained EEG source imaging method makes use of a relatively simple data structure for its fMRI priors and atlases, allowing them to be imported from a number of sources, including other imaging suites, or user defined sources. The only limitation in this regard is fitting the desired layout to the subject model with appropriately assigned vertices.
The general processing parameters described above outline the methods typically employed in these experiments. Once more, it is notable that there are no serious technical limitations on the selection of these parameters — data filtration and adjustment methods can be added or removed from the pipeline to suit any experiment. More important is the selection of the window size, as this directly affects the calculation of the model evidence and consequent application of fMRI priors. While variations in window size from approximately 40–150 ms result in only minor shifts in the resultant waveforms, extension beyond this does pose a risk to stability and may cause certain regions to be co-active or masked inappropriately. More specifically, a larger window size may be more useful when low frequencies are of interest, while a smaller window size may be preferable when focusing on higher frequency oscillations. The overlap and shift of the sliding window should also be considered here, as it has an effect on the computational complexity of the process and may become prohibitive due to the resources needed for analysis. Regardless of the exact parameters selected, the following steps are considered critical in the process: 1) Obtaining anatomical MRI data and simultaneous EEG/fMRI data; 2) 3D model generation; 3) MRI data analysis; 4) removal of the MR-artifact from EEG data; 5) forward and inverse calculations; 6) ROI generation; 7) sliding-window selection of ROI priors and source localization. The procedure here presents the overall pipeline and method that we have developed and utilized to achieve favorable dynamic results. It should be noted that many of the details — exact localization methods, evidence calculation, statistical methods, EEG and fMRI parameters, etc. — can be modified to suit the user's preferences.
The spatiotemporal fMRI constrained source analysis method is considered a noteworthy step forward in the integration of EEG and fMRI, but it is subject to certain minor limitations. While we do see an increase in the quality of reconstructed deep-sources, this method is still subject to the overall limitations from its individual modalities; if a source is deep enough to be effectively invisible to EEG, it will not be captured by this method. Second, analysis focuses on 3D models of the pial surface, and will not reconstruct any interior regions, regardless of any fMRI-identified hemodynamic activity.
Using EEG in combination with divided and conditionally applied fMRI priors, we have generated an advanced, spatiotemporally specific imaging algorithm. Immediate results have shown that the algorithm has an increased capability for reconstructing deep sources, and is less susceptible to cross-talk than its counterpart, the traditional time-invariant fMRI constrained source imaging. Further, the method is largely customizable and can be suited for each application, or used as a basis for subsequent analyses. These properties give the spatiotemporal fMRI constrained source analysis method potential as both an independently capable analysis method, and a foundation for future research.
Disclosures
The authors have nothing to disclose.
Acknowledgements
This work was supported in part by NIH DK082644 and the University of Houston.
Materials
| BrainAmp MR Plus | Brain Products | Amplifiers for EEG recording, MR-compatible | |
| BrainAmp ExG MR | Brain Products | Amplifier for auxilary sensor (EMG), MR-compatible | |
| BrainAmp Power Pack | Brain Products | Provide power to amplifiers in the MR environment | |
| Ribbon Cables | Brain Products | Connects the Power Pack to Amplifiers | |
| SyncBox | Brain Products | Synchronize MR scanner clock with EEG amplifier clock | |
| BrainCap MR | Brain Products | Passive-electrode 64-channel EEG cap, MR-compatible | |
| BrainVision Recorder | Brain Products | EEG data recording software (steps 1.2-1.4.2) | |
| BrainVision Analyzer 2.0 | Brain Products | EEG analysis software (steps 4.1-4.6) | |
| USB 2 Adapter (also known as BUA) | Brain Products | Interface between the amplifiers and data acquisition computer | |
| Fiber Optic Cables | Brain Products | Connects the EEG cap in the MR scanner to the Recording Computer | |
| SyncBox Scanner Interface | Brain Products | Synchronize MR scanner clock with EEG amplifier clock | |
| Trigger Cable | Brain Products | Used to send scanner/paradigm triggers to the recording computer | |
| ABRALYT HiCl EEG Electrode Gel | EasyCap | Abrasive EEG gel for passive electrode in MR environment | |
| Ingenia 3.0T MR system | Philips | 3.0 T MRI system | |
| Patriot Digitizer | Polhemus | EEG channel location digitization | |
| MATLAB r2014a | MathWorks | Programming base for the DBTN algorithm (steps 3.3-3.4 and 5.1-5.7) | |
| Pictures of Facial Affect | Paul Eckman Group | A series of emotionally valent faces used as stimuli | |
| E-Prime 2.0 | Psychology Software Tools, Inc | Presentation Software (step 1.4.3) | |
| Bipolar skin EMG electrode | Brain Products | Used to detect muscle activity. | |
| POLGUI | MATLAB software for digitization | ||
| Freesurfer | Software used in steps 2.1-2.4, and steps 3.1-3.2 | ||
| MNE | Software used in step 2.5 |
References
- Belanger, M., Allaman, I., Magistretti, P. J. Brain energy metabolism: focus on astrocyte-neuron metabolic cooperation. Cell Metab. 14 (6), 724-738 (2011).
- Logothetis, N. K. The underpinnings of the BOLD functional magnetic resonance imaging signal. J Neurosci. 23 (10), 3963-3971 (2003).
- Michel, C. M., et al. EEG source imaging. Clin Neurophysiol. 115 (10), 2195-2222 (2004).
- Ramon, C., Schimpf, P. H., Haueisen, J. Influence of head models on EEG simulations and inverse source localizations. Biomed Eng Online. 5, 10 (2006).
- Mosher, J. C., Spencer, M. E., Leahy, R. M., Lewis, P. S. Error bounds for EEG and MEG dipole source localization. Electroencephalogr Clin Neurophysiol. 86 (5), 303-321 (1993).
- Hamalainen, M. S., Ilmoniemi, R. J. Interpreting magnetic fields of the brain: minimum norm estimates. Med Biol Eng Comput. 32 (1), 35-42 (1994).
- Pascual-Marqui, R. D. Review of methods for solving the EEG inverse problem. International journal of bioelectromagnetism. 1 (1), 75-86 (1999).
- Rosa, M. J., Daunizeau, J., Friston, K. J. EEG-fMRI integration: a critical review of biophysical modeling and data analysis approaches. J Integr Neurosci. 9 (4), 453-476 (2010).
- Nguyen, T., et al. EEG Source Imaging Guided by Spatiotemporal Specific fMRI: Toward an Understanding of Dynamic Cognitive Processes. Neural Plast. 2016, 10 (2016).
- Klem, G. H., Luders, H. O., Jasper, H. H., Elger, C. The ten-twenty electrode system of the International Federation. The International Federation of Clinical Neurophysiology. Electroencephalogr Clin Neurophysiol Suppl. 52, 3-6 (1999).
- Mullinger, K. J., Castellone, P., Bowtell, R. Best current practice for obtaining high quality EEG data during simultaneous FMRI. J Vis Exp. (76), (2013).
- Ekman, P., Friesen, W. V. . Pictures of Facial Affect. , (1976).
- Dale, A. M., Fischl, B., Sereno, M. I. Cortical surface-based analysis. I. Segmentation and surface reconstruction. Neuroimage. 9 (2), 179-194 (1999).
- Fischl, B., Sereno, M. I., Dale, A. M. Cortical surface-based analysis. II: Inflation, flattening, and a surface-based coordinate system. Neuroimage. 9 (2), 195-207 (1999).
- Gramfort, A., et al. MNE software for processing MEG and EEG data. Neuroimage. 86, 446-460 (2014).
- Hagler, D. J., Saygin, A. P., Sereno, M. I. Smoothing and cluster thresholding for cortical surface-based group analysis of fMRI data. Neuroimage. 33 (4), 1093-1103 (2006).
- Jenkinson, M., Beckmann, C. F., Behrens, T. E., Woolrich, M. W., Smith, S. M. Fsl. Neuroimage. 62 (2), 782-790 (2012).
- Cox, R. W. AFNI: software for analysis and visualization of functional magnetic resonance neuroimages. Comput Biomed Res. 29 (3), 162-173 (1996).
- Genovese, C. R., Lazar, N. A., Nichols, T. Thresholding of statistical maps in functional neuroimaging using the false discovery rate. Neuroimage. 15 (4), 870-878 (2002).
- Klein, A., Tourville, J. 101 labeled brain images and a consistent human cortical labeling protocol. Front Neurosci. 6, 171 (2012).
- Fischl, B., et al. Automatically parcellating the human cerebral cortex. Cereb Cortex. 14 (1), 11-22 (2004).
- Delorme, A., Makeig, S. EEGLAB: an open source toolbox for analysis of single-trial EEG dynamics including independent component analysis. J Neurosci Methods. 134 (1), 9-21 (2004).
- Iannetti, G. D., et al. Simultaneous recording of laser-evoked brain potentials and continuous, high-field functional magnetic resonance imaging in humans. Neuroimage. 28 (3), 708-719 (2005).
- Niazy, R. K., Beckmann, C. F., Iannetti, G. D., Brady, J. M., Smith, S. M. Removal of FMRI environment artifacts from EEG data using optimal basis sets. Neuroimage. 28 (3), 720-737 (2005).

