Outer-Boundary Assisted Segmentation and Quantification of Trabecular Bones by an Imagej Plugin
Summary
We present a workflow for segmenting and quantifying trabecular bones for 2D and 3D images based on the bone's outer boundary using an ImageJ plugin. This approach is more efficient and accurate than the current manual hand-contouring approach, and provides layer-by-layer quantifications, which are not available in current commercial software.
Abstract
Micro-computed tomography (micro-CT) is routinely used to assess bone quantity and trabecular microstructural properties in small animals under different bone loss conditions. However, the standard approach for trabecular analysis of micro-CT images is slice-by-slice semi-automatic hand-contouring, which is labor intensive and error prone. Described here is an efficient method for automatic segmentation of trabecular bones according to the bone’s outer boundaries, where trabecular bones can be identified and segmented automatically with accuracy with less operator bias when appropriate segmentation parameters are set. To profile satisfactory segmentation parameters, an image stack of segmentation results is displayed, where all possible combinations of the segmentation parameters are changed one by one in sequence, and segmentation results with associated parameters can easily be visually checked. As a quality-control feature of the plugin, simulated standard objects are quantified where the measured quantities can be compared with theoretical values. Layer-by-layer quantification of trabecular properties and trabecular thicknesses are reported by such a plugin, and the distributions of such properties within the selected regions can be profiled easily. Although layer-by-layer quantification retains more information about trabecular bones and facilitates further statistical analysis of structural changes, such measures are unavailable from the output of current commercial software, where only a single quantified value for each parameter is reported for each sample. Therefore, the described workflows are better approaches for analyzing trabecular bones with accuracy and efficiency.
Introduction
Micro-CT analysis of trabecular bones is the standard approach for tracking morphological changes of the bones in small animals under different bone loss conditions1,2,3, where several variables related to the structures of the bones are reported4. However, such parameters are not evenly distributed in the metaphysis of long bones5, and only a summarized or averaged value is reported for each structural variable of each sample by current commercial micro-CT machines6,7, though a single value cannot fully represent the characteristics of the measured parameter in the analyzing region. Layer-by-layer quantification of trabecular bones not only retains more information for each variable, but also enables the profiling of the distributions of such variables in the analyzing region, facilitating subsequent statistical analysis of structural changes under different conditions5. Therefore, the goal of this method is quantifying trabecular bones of micro-CT scans at each slice level, which is not available in any commercially available micro-CT analysis package currently.
To efficiently segment trabecular bones slice-by-slice, automatic segmentation methods are desirable. However, the current standard technique for micro-CT analysis is based on manual interactive contouring followed by semiautomatic interpolation to separate trabecular bones from the cortical compartments, which is labor intensive, error-prone, and associated with substantial operator bias8,9,10. Automatic segmentation methods11,12 were reported, but such methods are only optimal in regions with good separation between trabecular bones and cortical bones, but not in regions without clear separations. Moreover, different segmentation parameters are required for different samples12, and it is tedious to manually select satisfactory segmentation parameters applicable to groups of bone samples by trying various parameter combinations12, even though the segmentation process is automatic when all related parameters are set. As the bone outer boundary has the greatest contrast with the scanning background and the metaphyseal cortical shells of long bones show few changes in the chosen analyzing region, segmentation methods according to the outer-boundary contour of long bones can reliably and accurately separate trabecular bones from cortical shells. The advantage of such a segmentation method is that the segmentation is based on the difference between the background and the bone's outer boundary, but not on the differences between trabecular and cortical bones6,12,13, therefore it is generally easy to find a combination of segmentation parameters that is satisfactory for a group of bone samples, facilitating more reliable analysis of trabecular changes between different groups.
At each slice level, area, perimeter, and two-dimensional (2D) thickness are reported for 2D analysis, while volume, surface, and three-dimensional (3D) thickness are reported in 3D quantifications. Such information is generally not reported by current image analysis tools, indicating that the reported procedures can be applied to general images where such information is desired.
Protocol
Procedures involving animal subjects were conducted in accordance with the Guide for the Care and Use of Laboratory Animals (NIH publication, 8th edition, 2011), and have been reviewed and approved by the Institutional Animal Care and Use Committee of Wuhan University.
1. Software Installation
- Install ImageJ software. Download the Windows version of the ImageJ (version 1.51p) software bundled with 64-bit Java from https://imagej.nih.gov/ij/. Extract the downloaded software into a folder, which will be subsequently referred to as the "ImageJ Directory".
NOTE: Trabecular analysis plugins require 64-bit java run time (version 1.8) and a 64-bit windows operating system, preferably a 64-bit Windows 7 operating system. - Install trabecular analysis plugins. Request trabecular analysis plugins from http://www.bomomics.com, and unzip the contents into ImageJ's plugins directory, which is "ImageJ Directory/plugins".
NOTE: The plugin can be obtained either as a free version, where the measurements for 5 specified adjacent slices are reported, or as a commercial version, where a range of slices can be specified and measured.
2. Prepare 3D Dataset for Trabecular Analysis
- Scan rat femurs with a micro-CT machine following the standard scanning protocol5 and then save the data in a format that can be imported into ImageJ, e.g., tiff format. If multiple bone samples were scanned simultaneously in a single sample tube, first import the data into ImageJ, and then separate each individual bone by cropping off other samples using ImageJ's image processing tools. Subsequently, save the resulting image in a format that can be imported into ImageJ later.
NOTE: A representative sample image file used in the analysis is included (Supplementary File 1). - Simulate standard 2D and 3D objects.
- Open ImageJ software. Under the Plugins | BoMomics | Simulate Objects menu, click the Circle button, and enter 200 as the diameter in the popup window, then click OK to generate the simulated circle with a diameter of 200 pixels (Figure 3B). Save the generated circle in tiff format.
- Open ImageJ software. Under the Plugins | BoMomics | Simulate Objects menu, click the Square button, and enter 200 as the side length in the popup window, then click OK to generate the simulated square with a side length of 200 pixels (Figure 3B). Save the generated square in tiff format.
- Open ImageJ software. Under the Plugins | BoMomics | Simulate Objects menu, click the Rectangle button, and enter 200 as the width and 100 as the height in the popup window, then click OK to generate the simulated rectangle with a width of 200 pixels and a height of 100 pixels (Figure 3B). Save the generated rectangle in tiff format.
NOTE: A circle (diameter: 200 pixels), a square (side length: 200 pixels), and a rectangle (width: 200 pixels; height: 100 pixels) are saved for the subsequent analyses. - Open ImageJ software. Under the Plugins | BoMomics | Simulate Objects menu, click the Cube button, and enter 30 as the side length in the popup window, then click OK to generate the simulated cube. Finally, click the Plugins | 3D | Volume Viewer to view the generated cube and save it in tiff format (Figure 3C).
- Open ImageJ software. Under the Plugins | BoMomics | Simulate Objects menu, click the Cuboid button, and enter 80 as the length, 40 as the width, and 30 as the height in the popup window, then click OK to generate the simulated cuboid. Finally, click the Plugins | 3D | Volume Viewer to view the generated cuboid and save it in tiff format (Figure 3C).
- Open ImageJ software. Under the Plugins | BoMomics | Simulate Objects menu, click the Sphere button, and enter 30 as the diameter in the popup window, then click OK to generate the simulated sphere. Finally, click the Plugins | 3D | Volume Viewer to view the generated sphere and save it in tiff format (Figure 3C).
- Open ImageJ software. Under the Plugins | BoMomics | Simulate Objects menu, click the Cylinder button, and enter 30 as the diameter and 100 as the height in the popup window, then click OK to generate the simulated cylinder. Finally, click the Plugins | 3D | Volume Viewer to view the generated cylinder and save it in tiff format (Figure 3C).
NOTE: A sphere (diameter: 30 pixels), a cube (side length: 30 pixels), a cuboid (length: 80 pixels; width: 40 pixels; height: 30 pixels), and a cylinder (diameter: 30 pixels; height: 100 pixels) are saved for the subsequent analyses.
3. Profiling Analysis Parameters
- Open ImageJ software and open or import a scanned image.
- Slide the bottom scrollbar to choose one slice, then click the Image | Adjust | Threshold button. In the popup Threshold window, adjust the minimum and maximum threshold values by manual inspection to ensure that bones are well separated from the background, and record the minimal threshold value as the cortical bone threshold value.
NOTE: As different micro-CT manufacturers use different scaling factors for storing X-ray attenuation coefficients in the images, the actual threshold value should be determined empirically either by manual inspection or following the specific manufacturer's recommendations. In our practice, setting the minimum threshold for ISQ files produced by micro-CT machines to 6,000-7,000 can reliably separate bones from the scanning background. - Click the Plugins | BoMomics | Trab Param Profiling button. In the popup window, set Slice Index to the position of the representative slice, and set the cortical bone ("Cort Bone"), range ("Range"), and step ("Step") values for calculating a set of cortical thresholds for profiling segmentation parameters, where the cortical bone threshold value is acquired by step 3.2 (Figure 1). The default range and step values of 2,000 and 400 work for most micro-CT images. Set the Slice Index to 5 for the example dataset, and keep the default values for other settings.
Note: "Cort Bone" is the threshold for cortical bones, and the threshold value used for parameter profiling changes from the lowest threshold value to the highest threshold value with the increment of Step value, where the lowest threshold value is the value of Cort Bone – Range, and the highest threshold value is Cort Bone + Range. If the set values for Cort Bone, Range, and Step are 6,000, 1,000, and 500, respectively, then the lowest threshold value for cortical bones is 6,000-1,000=5,000, the highest threshold value is 6,000+1,000=7,000, and the thresholds used in the analyses are 5,000, 5,500, 6,000, 6,500, and 7,000. - Set noise diameter ("Noise Dia."), step ("Step"), and range ("Range") values for specifying a set of noise values in the analyses, and the hole diameter ("Hole Dia."), step ("Step"), and range ("Range") values for calculating a set of hole values. In general, the default setting works for most micro-CT bone samples, where noise, step, and range are 5, 5, 2, respectively, and hole diameter, step, and range are 15, 5, 2, respectively. Keep the default settings for the example dataset.
Note: "Noise Dia." is the diameter for the noise suppression filter, and the "Hole Dia." is the diameter for holes within the cortical bones. The noise and hole values used for parameter profiling can be calculated similarly as cortical threshold values described above, using the specified noise/hole, range, and step values. Due to the variations in scanning parameters when acquiring the images, noise and hole parameters should be determined empirically according to the image quality, as no general ranges of noise and hole parameters are good for all scans. The lowest value for threshold, noise, or holes must be greater than or equal to 0, and if the calculated lowest value is negative using the supplied parameters, the specific lowest value is set to 0. - Click OK to perform parameter profiling. Visually check the segmentation results in the Parameter Profiling Results window, and select a slice layer so that the bone outer boundary is outlined quite accurately (Figure 1B). Subsequently, retrieve the profiling parameters from the entry in the Parameter Profiling Results table corresponding to the chosen slice layer (Table 1).
4. Trabecular Analysis
- Segmentation of trabecular bones
- Open ImageJ software, then open or import a scanned image.
- Click the Plugins | BoMomics | Trab Segmentation button, and fill in appropriate analysis parameters. Set "Start", "Outline Boundary", "Trab. Bones", "Noise Reduction Dia.", "Hole Filling Dia.", and "Cortical Thickness Dia." to 5, 7,200, 7,000, 6, 12 and 25, respectively.
NOTE: "Start" and "End" specify the selected slice range for segmentation of trabecular bones, "Outline Boundary" is corresponding to the profiled "Cort Bone" parameter, "Noise Reduction Dia." to "Noise Dia." Parameter, and "Hole Filling Dia." to "Hole Dia." parameter. "Cortical Thickness Dia." is the specified thickness for excluding the outer cortical bones. "Trab. Bones" is the threshold for extracting trabecular bones (Figure 2), where the settings are determined using profiling analysis parameters command as described in step 3.5. - Click OK to perform trabecular segmentation. Visually check the segmentation results in the Trab Segmentation Results window (Figure 2B). Save the extracted trabecular bones shown in the Segmented Trabecular Bones window in a tiff format (Figure 2B), which can be further analyzed by other software.
- Analysis of trabecular bones.
- Open ImageJ software, then open or import a scanned image.
- Click the Plugins | BoMomics | Trab Analysis button, and fill in appropriate analysis parameters, such as "Start", "Outline Boundary", "Trab. Bones", "Noise Reduction Dia.", "Hole Filling Dia.", and "Cortical Thickness Dia." as described above (Figure 3A), where the settings are determined using the profiling analysis parameters command as described in step 3.5. Set "Start", "Outline Boundary", "Trab. Bones", "Noise Reduction Dia.", "Hole Filling Dia.", and "Cortical Thickness Dia." to 5, 7,200, 7,000, 6, 12 and 25, respectively.
NOTE: In the free version of the plugin, five adjacent slices starting from the specified "Start" slice index are selected for measurement, while in the commercial version, an arbitrary number of slices can be specified by the user. - Select one or more choices in Results Reporting section for the parameters to be measured, where the trabecular bone volume (BV), total volume (TV) of the selected region, and thicknesses measured either two-dimensionally (2D) or three-dimensionally (3D) are available for selection through three checkboxes, namely "BV TV Only", "2D", and "3D". Select the checkboxes "2D" and "3D", then click "OK" to perform trabecular analysis (Figure 3A, Table 2).
NOTE: When "BV TV Only" is checked, regardless of the selection statuses of "2D" and "3D", raw measures of BV, TV, and Intensity are reported, and the segmented trabecular bones are extracted and displayed in a new window, which can be saved and further analyzed by other software. When "2D" is checked, raw measures of BV, TV, Intensity, and thicknesses measured two-dimensionally at each slice level using the plate model are reported. If "3D" is checked, three-dimensional thickness for each voxel is calculated directly without any model assumption, then raw measures of BV, TV, Intensity, and three-dimensional thicknesses sampled at each slice level are reported. However, if no checkbox is selected, trabecular bones are segmented using the above set of parameters, but no measurement is reported.
5. Quantifying Simulated Objects
- Open ImageJ software, then open a simulated image. Here, open the simulated sphere with a diameter of 30 pixels as an example.
- Select the Plugins | BoMomics | Trab Analysis button and fill in appropriate analysis parameters as described previously. Keep the default values for "Start", "End", "Outline Boundary", "Trab. Bones", and set "Noise Reduction Dia.", "Noise Reduction Dia.", "Hole Filling Dia.", and "Cortical Thickness Dia." to 0 (Figure 3).
Note: For simulated objects, there is no unintended signal noise, and there may be no corresponding cortical shells. Therefore, parameters for such values should be set appropriately (default settings are to zero). For 3D quantifications, 30 slices before and after the specified "Start" slice are processed using the free version of the plugin, while only slices in the range specified by "Start" and "End" slices are analyzed by the commercial version of the plugin. - In the Results Reporting section, select "2D" and "3D" for the parameters to be measured, and click OK to perform trabecular analysis for the simulated object (Table 3).
6. Calibration of Trabecular Measures and Data Presentation: Profile the Distributions of Trabecular Measures in the Selected Analyzing Region
- Get the calibration information from the scanned dataset, according to the micro-CT vendor's instructions.
NOTE: Only raw measures of BV, TV, Intensity, and thicknesses are reported by the plugin. To get measures comparable to reports generated by other software, result calibration is desired. - Open Microsoft Excel, and open the reported results table. Calculate calibrated bone volume (BV), total volume (TV), bone mineral content (BMC), bone volume fraction (BV/TV), and bone mineral density (BMD) in new excel columns using the reported raw BV, TV, and Intensity values according to the following expressions.
- NOTE: Scanning resolution (resolution, µm), gray scaling (scaling), density units (mg HA/cm3), density slope (slope), and density intercept (intercept) can be extracted from the scanned micro-CT image or meta files, such as the Scanco ISQ file. Therefore, the calibrated measures are calculated as follows:
Suppose BV, TV and Intensity are raw measures, BVc and TVc are calibrated values, resolution, scaling, slope, intercept are from calibration files of micro – CT images.
BVc = BV × resolution3 [μm3]
TVc = TV × resolution3 [μm3]
BV / TV = BV ÷ TV
BMC = (Intensity ÷ scaling × slope – BV × intercept) × resolution3 × 10-12 [mg HA]
BMD - BMC ÷ TVc × 1012 [mg HA / cm3] - Create an XY (Scatter) plot for calibrated measures (Y) against slice layers (X) using Microsoft Excel software (Figure 4).
Representative Results
The trabecular analysis plugin is designed to automatically segment and quantify trabecular bones with accuracy. Initially, bone outer boundary is detected and delineated followed by a hole-filling operation where any holes within bone outer cortical shells are filled. Then an erosion operation is performed to exclude the outer cortical bones and get the segmented trabecular bones. Finally, measures of trabecular bones in the segmented region are quantified.
As micro-CT images are inherently noisy, segmentations using predefined arbitrary parameters often fail to accurately identify the bone's outer boundaries. It is tedious and labor intensive to try a lot of parameter combinations for selecting satisfactory segmentation parameters. Therefore, a parameter profiling plugin is provided for changing the parameters one by one in the set range automatically to assist in selecting satisfactory parameter combinations, which also facilitates selecting a common set of parameters for a group of bone samples. Figure 1A shows the settings used for profiling good segmentation parameters. When the parameters for cortical bone threshold (Cort Bone), range of threshold to be profiled (Range), and increment amount for threshold (Step) at each step are specified, a series of thresholds to be profiled are generated. Subsequently, a series of noise and hole values are generated similarly by setting the corresponding parameters. Finally, trabecular bones are segmented by changing the parameters one at a time for all the possible parameter combinations. Figure 1B shows the representative segmentation results for different parameter combinations. Obviously, some parameter combinations are better than others at delineating a bone's outer boundaries, and more than one parameter combination shows satisfactory segmentation results. After visually checking the segmentation results, parameter values for satisfactory segmentations can be retrieved from the profiling results table (Table 1).
To quantify measures of trabecular bones, trabecular segmentation and analysis are performed. Figure 2A shows the setting dialog for trabecular segmentation, where trabecular bones in the selected region are segmented and extracted (Figure 2B), and segmentation results using the supplied parameters can be checked slice-by-slice visually. Subsequently, trabecular bones are analyzed with satisfactory parameters (Figure 3A). Depending on the selection statuses of reporting options, raw quantifications of bone volume (BV), total volume (TV), sum of gray values (Intensity), and thicknesses measured either two dimensionally or three dimensionally (Table 2) are reported. Finally, calibration information is extracted from the scanned micro-CT dataset and calibrated measures of BV, TV, BMC, BV/TV, and BMD are calculated followed by profiling their distributions in the selected analyzing region layer-by-layer against the layer positions (Figure 4).
As a quality-control feature of the plugin, quantification of simulated objects is supported. Simulated standard objects with known dimensions are quantified by the plugin for comparison with the theoretical values or measures from other software, such as commercial software supplied with micro-CT machines or BoneJ14, a free open source plugin for bone image analysis. For analyzing simulated objects, the noise settings are set to zero, as simulated images are considered high quality images without any noise. Objects with various thicknesses were simulated and results are shown (Figure 3, Table 3). For simulated standard 2D objects, such as circles, squares and rectangles, exact values for area (TV or BV) and thickness are reported (Table 3). For 3D objects, exact thickness measures for cubes, spheres, and cuboids are reported, however, the thicknesses for cylinders are not accurate for voxels near cylinder's both ends, while the voxel thicknesses in the middle slices of the cylinders are exactly as predicted. This is a feature of the underlying thickness measurement algorithm, wherein the voxel's thickness for each object is determined by the diameter of the largest sphere, or the side length of the largest cube, which contains this voxel and is completely inside the object. Therefore, thicknesses of objects made of different spheres and cubes can be measured accurately, while cylinders can only be measured accurately in the middle slice's radius-distance away from both ends.
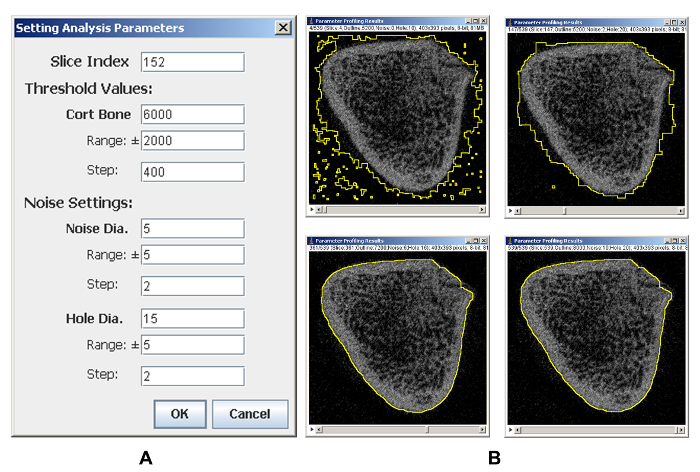
Figure 1: Representative results of parameter profiling analysis. (A) The parameters setting page. (B) Representative results of parameter profiling analysis. Some parameter combinations are better than others for detecting the outer boundaries of bones. Please click here to view a larger version of this figure.
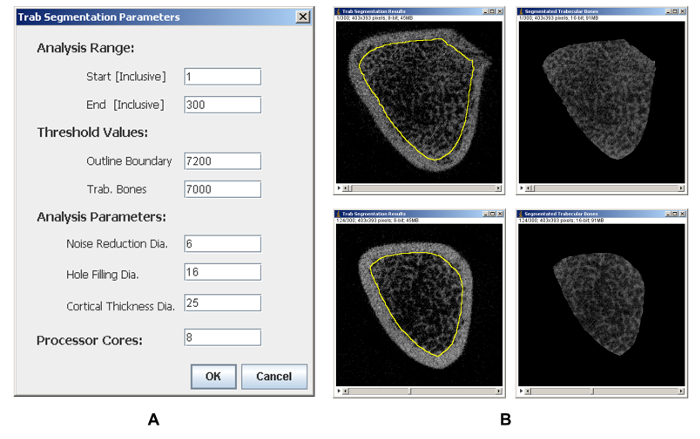
Figure 2: Representative results of trabecular segmentation analysis. (A) The parameters setting page. (B) Representative segmentation results of trabecular bones at different layers. Please click here to view a larger version of this figure.
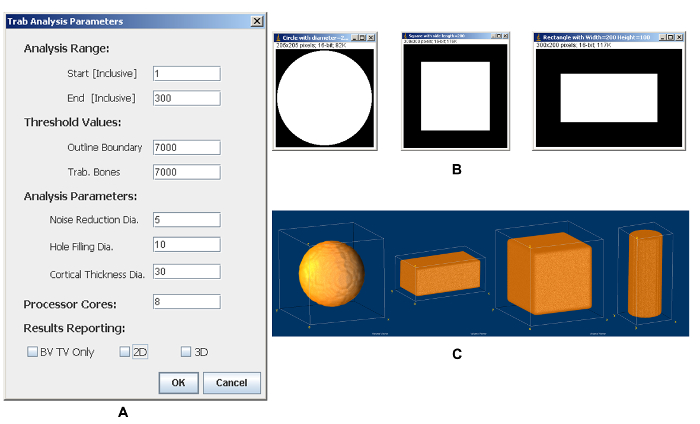
Figure 3: Trabecular analysis. (A) The parameters setting page. (B) Representative results of simulated 2D objects. (C) Representative results of simulated 3D objects visualized by 3D volume viewer. Please click here to view a larger version of this figure.
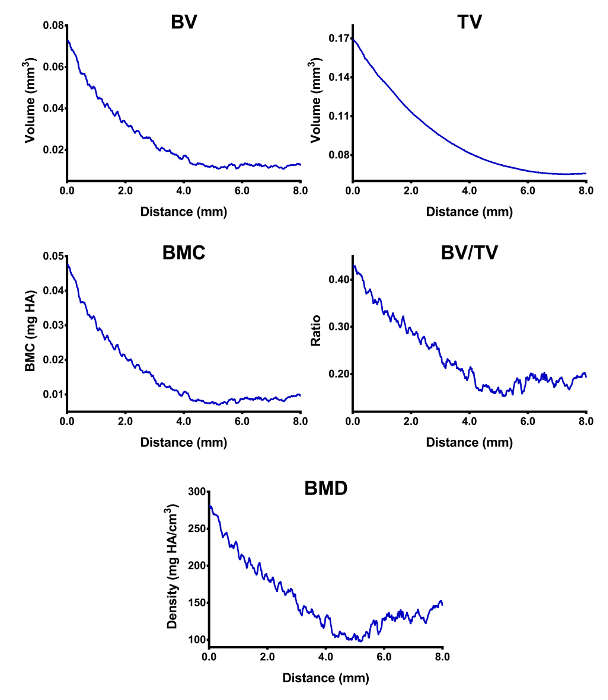
Figure 4: Distributions of trabecular bone measures in the selected analyzing region. The horizontal axis represents the relative distance to the starting slice layer in the analyzing region. Values in the Y-axis are calibrated trabecular measures in the analyzing region. Please click here to view a larger version of this figure.
| Slice | Outline Boundary | Noise Reduction Dia. | Hole Filling Dia. |
| 4 | 5200 | 0 | 10 |
| 147 | 5200 | 2 | 20 |
| 361 | 7200 | 6 | 16 |
| 539 | 8000 | 10 | 20 |
Table 1: Representative results of parameter profiling analysis.
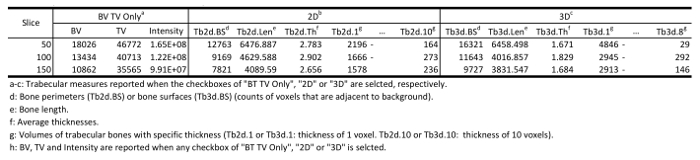
Table 2: Representative results of trabecular analysis.
| Object | Dimension | Volumeb | Surfacec | Thickness |
| Square | 200 X 200 | 40000 | 796 | 200 |
| Rectangle | 200 X 100 | 20000 | 596 | 100 |
| Circle | Dia: 200 | 31428 | 796 | 200 |
| Cube | 30 X 30 X 30 | 27000 | 5048 | 30 |
| Cuboid | 80 X 40 X 30 | 96000 | 13008 | 30 |
| Sphere | Dia: 30 | 14328 | 3944 | 30 |
| Cylinderd | Dia:30; H: 100 | 71600 | 12800 | 27.84 |
| Cylindere | Dia:30; H: 100 | 51552 | 9552 | 30 |
| a: Results are in raw voxels. Dia: diameter; H: height. | ||||
| b: Volume (3D) or area (2D). | ||||
| c: Surface (3D) or perimeter (2D). | ||||
| d: Measurs from slice 1 to slice 100 are used for analysis. | ||||
| e: Measurs from slice 15 to slice 85 are used for analysis. | ||||
Table 3: Quantification results for simulated objects.
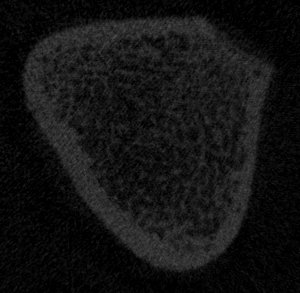
Supplementary File 1. Sample bone. Please click here to download this file.
Discussion
This study describes an ImageJ plugin for analyzing trabecular bones, which is automatic, efficient, and user friendly. The plugin can also be used to quantify any 2D or 3D object for layer-by-layer measures of areas, volumes, and thicknesses. Currently, only a single measured value for each trabecular parameter is reported for each sample by standard micro-CT analysis, which cannot fully represent the characteristics of the measured entity in the selected analyzing region. The described plugin reports layer-by-layer quantities of each parameter for each sample, which fully retains the distribution information of the measured parameter in the selected analyzing region, and therefore more advanced and sensitive statistical approaches are applicable for analyzing such data.
With the standard, semi-automatic hand-contouring segmentation method, substantial variation between different hand-contouring slices were observed with operator bias8,9,10, warranting a more uniform and automatic method for trabecular segmentation. As the contrast is maximal between the scanning background and the bone outer boundary, outer-boundary assisted trabecular segmentation is performed by the plugin, where the bone's outer boundary can be automatically detected with accuracy when appropriate threshold, noise, and hole settings are provided. To facilitate the determination of segmentation parameters, a range of parameter combinations are profiled one by one using a representative slice from the analyzing region, and satisfactory segmentation results are checked visually. Subsequently, trabecular bones are analyzed with the profiled parameters. When appropriate segmentation parameters are set, and the plugin has been successfully applied to analyze trabecular bones from rat distal femurs treated with different bone anabolic agents5, reproducible results are reported for the same image without operator bias.
The plugin currently can only process one sample at a time. If multiple samples are present in the image, all samples are quantified as a single object without discrimination. Therefore, preprocessing is required if multiple samples are scanned simultaneously in a single sample tube. The quantification outputs are raw measures of pixel counts or gray values, which should be calibrated manually using appropriate calibration information.
The most critical step in this protocol is selecting appropriate parameters for trabecular analysis. Generally, groups of bone samples are analyzed using identical segmentation parameters to make the results comparable. In such a case, care should be taken to ensure that segmentation results for all samples are examined visually for potential errors before further analyses are conducted.
A major limitation of this technique is that the noise reduction settings are applied to cortical bones only for contouring a bone's outer boundary, but not on extracted trabecular bones. As various filtering strategies have been reported15 as optimal for trabecular analysis under different conditions, no single filter is good for all image samples. Therefore, there is no built-in filtration step before quantification of trabecular bones. To be compatible with the output by software with built-in image filtrations, extracted trabecular bones can be processed similarly first using various imaging tools, and then imported into ImageJ and quantified by the plugin subsequently. Moreover, different image formats are generated by different micro-CT vendors, and most of them cannot be imported or opened by ImageJ directly, thus modules for importing various image formats into ImageJ for further analysis are desired. Another limitation is that the 3D thicknesses in slices near the ends of the selection range are not accurate if part of the objects are outside of the selection range, which is also true for any available 3D quantification packages, therefore, a larger range of slices should be quantified, and only the middle portion of selected slices should be used for further analysis. Such an approach can only be available when measured quantities are reported layer-by-layer as shown in this plugin, but not for measurements that only a single value is reported for each quantified parameter in the selection range.
Disclosures
The authors have nothing to disclose.
Acknowledgements
This work was partially supported by grant NFSC 81170806. The authors would like to thank the micro-CT core facility of School of Stomatology, Wuhan University for helping scan and analyze the rat femurs.
Materials
| ImageJ | NIH | imagej | Any version with a java 1.8 run time |
| trabecular analysis plugin | Bomomics | bomomics | free or commercial version |
| Micro CT scanner | Scanco | μ-50 | micro CT from any vendor |
| Computer System | Lenovo | any brand | |
| Windows Operating System | Microsoft | Windows 7 x64 | any 64-bit Windows operating system |
| Office Software | Microsoft | Office 2010 | any speadsheet software that has xy chart function |
References
- Ruegsegger, P., Koller, B., Muller, R. A microtomographic system for the nondestructive evaluation of bone architecture. Calcif Tissue Int. 58 (1), 24-29 (1996).
- Muller, R., Ruegsegger, P. Micro-tomographic imaging for the nondestructive evaluation of trabecular bone architecture. Stud Health Technol Inform. 40, 61-79 (1997).
- Clark, D. P., Badea, C. T. Micro-CT of rodents: state-of-the-art and future perspectives. Phys Med. 30 (6), 619-634 (2014).
- Bart, Z., Wallace, J. Microcomputed Tomography Applications in Bone and Mineral Research. Advances in Computed Tomography. 2, 121-127 (2013).
- Ji, Y., Ke, Y., Gao, S. Intermittent activation of notch signaling promotes bone formation. Am J Transl Res. 9 (6), 2933-2944 (2017).
- Jiang, Y., Zhao, J., White, D. L., Genant, H. K. Micro CT and Micro MR imaging of 3D architecture of animal skeleton. J Musculoskelet Neuronal Interact. 1 (1), 45-51 (2000).
- Laib, A., et al. 3D micro-computed tomography of trabecular and cortical bone architecture with application to a rat model of immobilisation osteoporosis. Med Biol Eng Comput. 38 (3), 326-332 (2000).
- Cole, H. A., Ichikawa, J., Colvin, D. C., O’Rear, L., Schoenecker, J. G. Quantifying intra-osseous growth of osteosarcoma in a murine model with radiographic analysis. J Orthop Res. 29 (12), 1957-1962 (2011).
- Jensen, M. M., Jorgensen, J. T., Binderup, T., Kjaer, A. Tumor volume in subcutaneous mouse xenografts measured by microCT is more accurate and reproducible than determined by 18F-FDG-microPET or external caliper. BMC Med Imaging. 8, 16 (2008).
- Soviero, V. M., Leal, S. C., Silva, R. C., Azevedo, R. B. Validity of MicroCT for in vitro detection of proximal carious lesions in primary molars. J Dent. 40 (1), 35-40 (2012).
- Kohler, T., Stauber, M., Donahue, L. R., Muller, R. Automated compartmental analysis for high-throughput skeletal phenotyping in femora of genetic mouse models. Bone. 41 (4), 659-667 (2007).
- Buie, H. R., Campbell, G. M., Klinck, R. J., MacNeil, J. A., Boyd, S. K. Automatic segmentation of cortical and trabecular compartments based on a dual threshold technique for in vivo micro-CT bone analysis. Bone. 41 (4), 505-515 (2007).
- Dougherty, G. Quantitative CT in the measurement of bone quantity and bone quality for assessing osteoporosis. Med Eng Phys. 18 (7), 557-568 (1996).
- Doube, M., et al. BoneJ: Free and extensible bone image analysis in ImageJ. Bone. 47 (6), 1076-1079 (2010).
- Bouxsein, M. L., et al. Guidelines for assessment of bone microstructure in rodents using micro-computed tomography. J Bone Miner Res. 25 (7), 1468-1486 (2010).

