Preparing and Injecting Embryos of Culex Mosquitoes to Generate Null Mutations using CRISPR/Cas9
Summary
CRISPR/Cas9 is increasingly used to characterize gene function in non-model organisms. This protocol describes how to generate knock-out lines of Culex pipiens, from preparing injection mixes, to obtaining and injecting mosquito embryos, as well as how to rear, cross, and screen injected mosquitoes and their progeny for desired mutations.
Abstract
Culex mosquitoes are the major vectors of several diseases that negatively impact human and animal health including West Nile virus and diseases caused by filarial nematodes such as canine heartworm and elephantasis. Recently, CRISPR/Cas9 genome editing has been used to induce site-directed mutations by injecting a Cas9 protein that has been complexed with a guide RNA (gRNA) into freshly laid embryos of several insect species, including mosquitoes that belong to the genera Anopheles and Aedes. Manipulating and injecting Culex mosquitoes is slightly more difficult as these mosquitoes lay their eggs upright in rafts rather than individually like other species of mosquitoes. Here we describe how to design gRNAs, complex them with Cas9 protein, induce female mosquitoes of Culex pipiens to lay eggs, and how to prepare and inject newly laid embryos for microinjection with Cas9/gRNA. We also describe how to rear and screen injected mosquitoes for the desired mutation. The representative results demonstrate that this technique can be used to induce site-directed mutations in the genome of Culex mosquitoes and, with slight modifications, can be used to generate null-mutants in other mosquito species as well.
Introduction
Culex mosquitoes are distributed throughout the temperate and tropical regions of the world and transmit several deadly viruses including West Nile virus1, St. Louis encephalitis2 as well as filarial nematodes that cause canine heartworm3 and elephantiasis4. Members of the Culex pipiens complex, which includes Cx. quinquefasciatus, Cx. pipiens pipiens and Cx. pipiens molestus, show striking variations in many aspects of their biology. For example, while Cx. quinquefasciatus and Cx. pipiens molestus are incapable of entering an overwintering dormancy5,6, Cx. pipiens pipiens display robust seasonal responses and enter diapause in response to short days7,8. Additionally, Cx. pipiens molestus tend to be more anthropophilic while Cx. pipiens and Cx. quinquefasciatus are more zoophilic6. However, in the United States and throughout many other places in the world, these species interbreed, which has strong implications for disease transmission as hybrids of the Cx. pipiens pipiens and Cx. pipiens molestus are opportunistic feeders and will bite both birds and humans9, thereby serving as bridge vectors for West Nile virus. Studying these and other fascinating aspects of the biology of Culex mosquitoes has been hampered, in part, because Culex mosquitoes are slightly more difficult to rear in the lab than Aedes mosquitoes, which produce quiescent and desiccation-resistant eggs10 and because functional molecular tools are not as well developed for Culex species.
CRISPR/Cas9 genome editing is a powerful technology that has been used to evaluate the biology of several important mosquito species11,12,13, including the Southern house mosquito, Culex quinquefasciatus14,15,16. This technology, developed by Jennifer Doudna and Emmanuelle Charpentier, exploits a natural bacterial defense against viruses by bacterially-derived, CRISPR-associated endonucleases (Cas proteins; see review by Van der Oost et al.17). When injected into animal embryos, the Cas9 proteins in combination with an appropriate guide RNA can produce double-stranded breaks within the genome. This is most frequently done by using the Cas9 protein that is complexed with guide RNAs, which directs endonuclease activity to a specific region of the genome. After the Cas9 protein has created a site-specific double-stranded break, the cellular machinery attempts to repair the break using one of two mechanisms. The first entails ligating the two ends together through non-homologous end joining (NHEJ), which is error-prone and often produces out-frame insertions and deletions in the genome that can result in non-functional proteins, thereby generating a knock-out mutation. Alternatively, the cellular machinery might use homology-directed repair (HDR) by finding similar sequences to correctly repair the break. The similar sequence may be provided by the second chromosome within the organism (see review18). However, if the repaired sequence exactly matches the original sequence, the Cas9 protein will be able to again cut the DNA. Alternatively, researchers can also include a donor plasmid that contains homologous sequences on either side of the cut site of the target sequence with an alternative repair sequence—often a fluorescent marker protein, modified version of the original gene, or other modification—that can be copied and inserted into the genome, or “knocked-in.”
Timing is critical when injecting embryos, and this is especially the case when using CRISPR/Cas9 genome editing to create mutations in insects. This is because the Cas9 protein and gRNAs have the greatest capacity to generate mutations only when the embryo is in its syncytial state, before cellular membranes have formed and when multiple nuclei are accessible within the embryo. For mosquitoes, nuclei reach the periphery ~2-4 hours after oviposition, depending on temperature19, and therefore successful microinjection must occur before this time. Additionally, the Cas9 protein will cut any nuclear DNA that it can access, such that the individual resulting from the injection will contain a mosaic of cells, some having the desired mutation, and others not. In order for these mutations to be successfully inherited, the Cas9 protein must cut DNA that resides in the germline that will give rise to the future eggs and sperm. To ensure that mutations are generated in the germline it is best to inject all materials close to the location of the pole cells within the embryo, which are the progenitors of the insect germline. The pole cells are located near the posterior end of Culex embryos20. In addition to injecting embryos, it is imperative to develop a careful plan for crossing and screening offspring in order to detect the desired mutation.
This protocol describes how to generate gRNAs and complex them with Cas9 protein to prepare injection mixes, as well as how to induce female mosquitoes of Culex pipiens to lay eggs and how to prepare and inject those eggs for CRISPR/Cas9-mediated genome editing. Additionally, we describe how to rear, cross and screen injected embryos and their progeny to confirm that the desired mutation has been obtained. Using this protocol, we generated null mutations for a gene of interest, cycle, in the Buckeye strain of Culex pipiens. This strain was originally established in 2013 from field-collected mosquitoes in Columbus, Ohio and is maintained by the Meuti lab. This protocol can be used for additional studies that require CRISPR/Cas9 genome editing in Culex mosquitoes, as well as other mosquito species, and, more generally, is relevant to employing CRISPR/Cas9 genome editing to any insect species.
Protocol
In most research institutions, an approved Biosafety Protocol must be in place before transgenic insects are generated or maintained to ensure that genetically modified organisms will not escape or be removed from the laboratory facility. Additional government regulations might also apply. Before beginning a project of this nature, check all institutional policies and procedures to determine what documents and approvals are required.
1. Designing gRNAs and preparing injection mixes
- Design 2-5 gRNAs for each target gene as some gRNAs work better than others. gRNAs that target a gene of interest can be either manually designed or free programs, such as Chop-Chop, can be used.
- Design and order primers that will amplify 100-200 bp fragments around each gRNA. Ideally, the cut site should be located approximately in the middle third to half of the PCR product. Note how many bp on each site of the cut will be produced.
- Obtain gRNAs.
- Purchase gRNAs from commercial companies such as Integrated DNA Technologies, Fisher Scientific, Genescript and others. This is the easiest option.
- Alternatively, create gRNAs in the lab using PCR amplification to generate the templates for subsequent isothermal synthesis. See the protocol described by Port et al.21 and Kistler et al.22.
- Provide gRNAs via plasmid that is injected into embryos. In this case, the gRNA should be driven by the U6 promotor (see 23,24 for details).
- Obtain the Cas9 protein.
- Order Cas9 commercially from several companies including Fisher Scientific. This is the easiest option.
- Make Cas9 protein and purify in the lab according to Basu et al.25 and Kistler et al.22.
- Inject Cas9 mRNA into embryos, where it will later be translated into active Cas9 protein. Cas9 mRNA can either be synthesized in the lab using in vitro transcription22 or purchased from vendors such as Sigma.
NOTE: The translated Cas9 protein must contain a nuclear localization signal (NLS) on the C-terminus, and therefore the NLS must also be present in the injected Cas9 mRNA. - Express Cas9 protein in vivo through plasmid-based expression. In this case, it is best to have the expression of Cas9 on the plasmid driven by an embryonic promoter that has been well-characterized in the target species.
NOTE: To date, embryonic promoters have not been well-characterized in Culex mosquitoes, and therefore injecting purified protein or Cas9 mRNA is recommended.
- Complex the gRNAs with the Cas9 protein, adapted from Kistler et al.22.
- If injecting Cas9 protein with the gRNA together (recommended), ensure that the final concentrations of Cas9 protein is 300 ng/µL and the final concentration of a single gRNA is 80 µg/µL. Mix the protein and an individual gRNA in a 0.2 mL PCR tube.
- Incubate for 20 min on ice.
- Testing gRNAs with an in vitro assay (e.g., Guide-it sgRNA Screening Kit).
- Amplify the fragments around the cut site for each gRNA using PCR.
- Run the PCR products on a gel to ensure that they are the correct size. Then, purify PCR products and sequence them to ensure that they target the correct gene region.
- Mix 200 ng of the leftover purified PCR product with 1 µL of Cas9 reaction buffer, 1 µL of BSA, 1.5 µL of complexed Cas9:gRNA and bring the total reaction volume to 15 µL. Incubate for 5 min at 37°C.
- Run the product on a gel again. If the Cas9 protein successfully cut the PCR product, the primary band should appear to be more faint and additional smaller bands should be present. Note the reduction in size of the original PCR product, and, if desired, calculate the reduction in band intensity to approximate the cutting efficiency.
- Preparing the final injection mix
- For each gRNA that successfully cut its targeted PCR product, mix volumes of each Cas9 and gRNA into a 0.5 mL tube such that the final volume remains 300 ng/µL of Cas9 protein and 80 ng/µL of each gRNA.
- Store the complexed Cas9 and gRNAs at -80 °C until they are needed for injection.
2. Pulling and beveling needles
NOTE: Successful injections and survival of embryos requires sharp needles (Figure 1).
- Use either 1 mm outer diameter borosilicate or quartz glass microcapillaries. Siliconize the microcapillaries in a chemical fume hood before being pulled.
- In brief, place 3 mL of Sigmacote into a glass beaker and vacuum draw into a second beaker containing the glass microcapillaries for at least 4 h, allowing the siliconizing solution to vaporize and settle on all surfaces of the microcapillaries.
- Afterwards, slowly allow the vacuum chamber to equalize to ambient pressure by gradually opening the stopcock valve and allowing air to return into the chamber. The needles are then ready to be pulled.
- Always read the instruction manual of needle pullers before use.
NOTE: The instructions below detail how to use a P-2000 Laser Needle Puller to pull glass needles. It is important to note that all needle pullers even of the same brand are not calibrated to pull exactly the same needle across different units. The key to getting good needles is to start at a given set of parameters then pull needles using parameters that are above and below these values. Test the needles at each of these parameters by evaluating how well the needle pierces embryos and how well the injection mix flows from the needle. Refine the parameters until a needle is obtained that is easy to open or bevel, that pierces embryos smoothly and that delivers injection mix easily.
- Pulling needles using the P-2000 laser needle puller
- Turn on the laser needle puller and allow the laser to warm up for 15 min before the first pull.
- Program the machine for the type of glass microcapillary tube (Remember any needle puller parameters are a starting point since needle pullers are not calibrated to produce the same needle across all pullers):
Quartz: Heat = 730; Fil = 4; Vel = 40; Del = 122; Pul = 156 - After programming the needle puller, load a microcapillary and clamp it in place, taking care to only touch the end of the microcapillary tube and not the area that will be heated by the laser.
- After clamping the microcapillary, place thumb and forefinger on the finger bars and then release the pull bars by pressing the spring releases one at a time.
- Squeeze thumb and forefinger to pull bars fully together.
- Position the other side of the microcapillary tube in its clamp such that a small portion of the tube extends from both sides. Tighten the clamps making sure that both clamps are finger-tight and that they hold the “pull bars” firmly in place.
- Close the protective shroud on the puller.
- Press the Pull button and the laser will start to heat the glass, indicated by the red “laser on” light located under the shroud. The pull is completed when the pull bars are completely separated, accompanied by a metallic thud.
- Ensure that the “laser on” light is not illuminated before lifting the shroud.
- Hold the small end of the microcapillary tube that extends beyond the clamp with thumb and forefinger and loosen the clamp. Then lift the microcapillary tube out of the clamp.
- Use forceps to carefully place the microcapillary tube into a square Petri dish lined with double-sided tape. Ensure when placing the needle into the holding box that the back end of the needle touches first and the needle is gently lowered into place so that the sharp tip is not damaged.
- Pulling needles using a PC-10/PC-100 needle-puller
- Set up PC-10 puller using 4 weights and set the temp for 50.4 °C for a single-step pull.
- Place a borosilicate glass microcapillary into the top clamp. Then raise the weighted bottom clamp into position and clamp the bottom portion of the microcapillary, making sure that the capillary is in position to pull two good needles. This is a rather long pull cycle, but it produces good needles for beveling.
- Beveling needles.
NOTE: This method of wet beveling gives real time feedback on relative opening size of the needle.- Assemble the abrasive plate and retaining ring of the needle beveler. Ensure that the gray side of the abrasive plate is oriented upward between the upper and lower retaining ring. Tighten the screws on the retaining ring, alternating from screw to screw to ensure that each screw is tightened evenly. The abrasive plate and retaining rings will hereafter be referred to as the grinding assembly.
- Place 13 drops (~ 520 µL) of pedestal oil onto the optical flat surface, and then place the grinding assembly on top of the plate. Place the assembly beneath a dissecting microscope, and then turn on the beveler. A little extra oil compared to Sutter’s recommendation is used, as this keeps the abrasive plate spinning for a longer period when used in conjunction with this wet beveling method.
- Add 1% Photo-Flo solution to the grinding surface and move the wick into position so that it is immersed in the Photo-Flo solution.
- Turn on the air at the main source. Then place a pulled borosilicate needle into the holder and tighten the retaining ring. Turn on the air at the regulator and increase until the pressure has reached 26 PSI.
- Watching from the side, lower the needle using the coarse adjustment knob until it almost touches the Photo-Flo liquid surface.
- Adjust the microscope to locate the needle in the field of view. Start at the lowest magnification and increase the magnification. Carefully adjust the position of the needle using the coarse adjustment knob until it just touches the surface of the Photo-Flo liquid.
- Using the coarse adjustment knob and continuing to look under the microscope, lower the needle until it is close to the abrasive surface but not touching it.
- Using the fine adjustment knob, carefully and slowly lower the needle, paying close attention to the needle and the shadow it leaves on the abrasive surface of the beveler. When the needle and its shadow look like they are about to touch the abrasive surface, continue to lower the needle even more slowly and carefully.
- Alternate between letting the needle “bevel” by lowering it on to the abrasive surface, and then raising it. Before raising the needle, quickly note the setting on the micrometer on the fine adjustment knob so that the needle can be lowered to the same position or, if necessary, to a slightly lower position. Remember to keep the needle below the Photo-Flo liquid layer. Once the needle has been raised above the abrasive surface, quickly stop and restart the rotation of the beveler to check for bubbles. If the needle has been adequately beveled, air bubbles should flow out from the needle and into the Photo-Flo. If the plate is not stopped, bubbles will likely not become visible until the needle opening is too large.
- If no bubbles appear when the needle is raised above the abrasive surface and the beveler has stopped, repeat the beveling procedure, lowering the needle to a slightly lower position on the micrometer located on the fine adjustment knob, raising the needle, stopping the beveler plate and checking for air bubbles. Repeat until a small but steady stream of bubbles appear.
- Once air bubbles are present, check the relative opening size by stopping the abrasive plate and lowering the air pressure, noting the PSI at which bubble formation stops, as this is a relative indication of the size of the opening of the beveled needles. The lower the pressure at which air stops escaping from the needle tip, the larger the needle opening. Needles beveled to the point at which bubbles stop escaping from the needle at 18-20 PSI indicates that the needle has a relatively small opening and should cause minimal damage to an embryo when used for microinjections.
- After checking the needle opening, increase the air pressure to 26 PSI to prevent liquid from entering the needle. Then raise the needle up and out of the Photo-Flo layer using the coarse adjustment knob. It is important to note that as long as air is flowing out of the needle the Photo-Flo does not enter the needle, and therefore the inside of the needle is protected from contamination.
- Once the needle is out of the Photo-Flo, turn off the air and remove the needle from the needle holder.
- Using forceps, place the newly beveled needle into a Petri dish that has either double sided tape or modeling clay to keep it in place. Repeat the process until 10-20 beveled needles have been produced.
3. Blood-feeding the parental generation of mosquitoes
- Rear larvae of Cx. pipiens mosquitoes under unambiguous long day conditions (>13 h of light/day) at 25-27 °C to ensure that mosquitoes do not enter their overwintering dormancy or diapause.
- Seven to ten days after peak adult emergence, prepare the Hemotek Membrane feeding system by plugging the heating units into the power supply. Adjust the temperature of each unit to 37 °C (human internal body temperature) or 41 °C (chicken internal body temperature) using the adjustment screw on the top of the unit. Ensure that the correct temperature has been reached using an electric thermometer.
- Prepare the blood-meal reservoir for feeding.
- Cut one square of parafilm for each blood-feeding cylinder, and rub parafilm against bare feet. This provides scents that are attractive to the mosquitoes.
- Stretch the parafilm as thinly as possible, and wrap the edges tightly around the meal reservoir. If desired, secure in place with a rubber O-ring.
- Add approximately 200 µL of 0.1 M ATP solution to 15 mL of fresh or thawed whole, sodium citrate-treated chicken blood. The ATP helps stimulate blood feeding, and sodium citrate prevents the blood from coagulating.
- Hold the reservoir so that the parafilm side is facing down, and carefully use a disposable pipette to fill the reservoir. Each reservoir holds ~3 mL of blood. Seal the filling ports with plastic plugs.
- Screw the meal reservoirs into the heating units and place on top of the mosquito cage such that the parafilm side is facing down and the mosquitoes can feed through the mesh.
- Cover the cages with black trash bags to simulate dusk and blow vigorously into each cage as the increased CO2 concentration stimulates blood-feeding.
- Keep the feeder on the cage for 2-8 h to maximize blood feeding. Check periodically and note the proportion of females that have taken a blood meal (evident by their red and distended abdomens).
4. Inducing egg-laying in adult mosquitoes
- Four to five days after blood feeding, prepare a 50 mL conical tube for mosquito oviposition.
- Create a ~20 mm hole near the bottom of the conical tube.
- Cover the hole with two squares (35 mm2) of dental dam rubber, one with a horizontal slit and another with a vertical slit. Use tape to attach the squares of dental dam to the conical tube.
- Place a 4.25 cm piece of filter paper onto a 35 mm x 10 mm Petri dish and wet the filter paper with 750 µL of distilled water.
- Invert the conical tube onto the filter paper and twist back and forth a few times to ensure that it is secure.
- Use a mouth aspirator to carefully remove ~10 females of Cx. pipiens from their cage and transfer them to the prepared conical tube.
- Place an opaque cylinder over the conical tube to provide darkness to the mosquitoes as this stimulates egg-laying, and wait 20-25 minutes.
5. Micro-manipulating freshly-laid Culex eggs
- Prepare the slide for attaching the mosquito embryos for microinjection.
- Attach a piece of double-sided tape to the width of a cover glass.
- Attach a thin strip of transparent medical dressing (e.g., Tegaderm, 2-3 mm wide) to the double-sided tape such that it runs parallel to the short end of the cover glass and is positioned approximately 5 mm from the end of the cover glass.
- Remove excess double-sided tape with a sharp razor blade, and remove 2-3 mm of medical dressing and double-sided tape from each end of the cover glass. This allows a layer of oil to remain on the slide after the eggs have been mounted to the cover glass.
- Using a wax pencil, draw a “D” shape around the double-sided tape and strip of medical dressing. This will act as a barrier to hold the water around the eggs after injection.
- After the slide has been prepared and 20-25 minutes have elapsed, remove the opaque tube and determine whether or not the mosquitoes have laid any eggs.
- If not, cover them for another 5-10 minutes.
- If the mosquitoes have laid eggs, carefully but quickly lift the conical tube and cover with a cap. Place the mosquitoes in a separate cage, and record the time that eggs were collected on a spreadsheet. Then add 50-100 µL of DI water to the filter paper containing the egg rafts.
- Under a dissecting microscope line up the eggs.
- Position a piece of filter paper (10 mm x 30 mm rectangles cut from coarse filter paper with fast flow rate) under a 24 mm x 40 mm cover glass such that the cover glass covers 50-60% of the left hand side of the filter paper.
- Wet the filter paper with 50 µL of DI water. The water will spread under the cover glass slowly, creating a reservoir to keep the filter paper and the eggs wet during micromanipulation. A meniscus will form between the cover glass and the filter paper when the correct amount of water is applied.
- Separate the eggs from their egg rafts using a fine paint brush, and position so that the narrow, posterior end of the embryo is touching the cover glass (left) and the wider, anterior end is to the right.
- Align eggs for 20 min, carefully observing the change of their color from a milky white to a very light gray. Routinely inspect the water film under the cover glass to ensure that there is an adequate amount of water. If the eggs appear to be drying out, add a small amount of DI water (20-30 µL) to the filter paper.
- After 20 min, discard all remaining eggs.
- Transfer the eggs to the microinjection slide.
- Use a small (10 mm x 30 mm) piece of coarse filter paper to wick water from the side of the saturated filter paper, and remove the wicking filter paper with forceps. Repeat 2-3 times to ensure that the filter paper with the eggs is as dry as possible.
- Lay a fresh, dry rectangle of filter paper and hold it in place with a pair of forceps. Carefully pull away the cover glass to the left, away from the aligned eggs.
- Dry the excess water on the stage without disturbing the small filter paper that contains the aligned eggs. Continue to dry the wet filter paper with the eggs several more times, pressing on the small rectangles of filter paper with thumbs and/or forceps, to ensure that the eggs are as dry as possible before transferring them to the injection slide.
- Using forceps, remove the paper backing off of the strip of medical dressing from the mounting slide.
- Hold the mounting slide with a thumb and middle finger and the exposed medical dressing facing down, and lower at a 45° angle onto the line of aligned eggs. Position the slide such that the wider, anterior end of the eggs (left) is hanging slightly off the end of the medical dressing, as this enhances the ability of the Culex larvae to successfully hatch from the mounted eggs.
- Press index finger to the underside of the cover glass, while continuing to hold the cover glass between the thumb and middle finger, to ensure that all of the aligned eggs are firmly attached to the medical dressing.
- Immediately invert the cover glass so that the eggs are facing up, and apply a thin layer of oil Halocarbon oil (2-3 drops; ~20 µL) to the eggs. This prevents the eggs from desiccating during microinjection. Remove any eggs that are not attached to the medical dressing or are not covered by the oil.
- Place the glass cover with the mounted eggs facing up inside of a square Petri dish with a large square of wet filter paper for transport for microinjection.
6. Injecting Culex embryos
- Back-filling the injection needles with the injection mix
- Choose a needle filler or gel loading tip that will fit easily into the back end of the selected injection needle, and will have a bit of space between the end of the needle filler and the injection needle.
- Carefully place a single and small drop of injection mix (~0.5-1 µL) into the injection needle, using the needle filler to back-fill the needle. Place the drop of injection mix near where the injection needle begins to taper.
- Attach the injection needle to the micro-injector.
- Place the slide containing the embryos onto the stage of the microscope.
- Opening the needle
NOTE: If using beveled needles, this is not necessary.- Use a starting injection pressure of ~30 PSI and adjust as necessary to deliver an appropriate volume of injection mix.
- Under a dissection microscope, position the needle above the embryo, and carefully use the micromanipulator to lower the needle onto the embryo. Once the needle is barely touching the embryo, quickly move the embryo perpendicular to the long axis of the needle.
- To determine if the needle was successfully opened, press the injection trigger and see whether any bubbles/injection mix escape from the needle. If not, repeat moving the embryo against the needle until the needle is open and injection mix escapes when the trigger is pressed.
- Injecting the embryos
- Position the first embryo in line in the center of view in the microscope and ensure that it is in focus.
- Place the needle over the first egg, also within the center of the field of view within the microscope.
- Progressively increase the magnification on the microscope, carefully lowering the needle to keep it just above the first embryo, centered and in focus. Proceed until the microscope has reached its highest magnification and both the embryo and the needle are in focus.
- Carefully move the embryo on the stage slightly to the left and lower the needle slightly so that the needle and the embryo are now in the same plane of view.
- Using the microscope stage to move the embryo, carefully and gently touch the embryo to the tip of the needle. The narrow, posterior end of the embryo should deflect slightly. Adjust the height of the needle if it is too high or too low.
- Using the microscope stage, move the posterior end of the embryo onto the needle and observe the needle penetrating the chorion of the mosquito egg. The needle should just penetrate the membrane in the approximate location of the pole cells within Culex embryos. Once this occurs, pull the injection trigger 1-3 times to shoot injection mix into the embryo. Deliver a small amount of injection mix, just enough that a small clearing in the embryoplasm can be detected.
- Using the stage, move the embryo to the right, away and off the tip of the needle. If the injection went well, little to no fluid should leak from the embryo.
- Move the stage down so that the next embryo in the line is positioned in front of the needle. Then again, move the stage to the left, positioning the embryo onto the injection needle and pull the injection trigger.
- If injection mix does not appear to be coming out and/or if a large amount of fluid escapes from the embryo after the needle is removed, replace the injection needle and start again.
- Destroy any un-injected or damaged eggs.
- Tending to the embryos post-injection
- After all embryos on the slide have been injected, wash off the Halocarbon oil by applying reverse osmosis water with a squirt bottle, such that the stream of water is directed onto the top of the glass coverslip above the injected embryos and allowing the water to flow down the coverslip and over the embryos to rinse the oil away. Make sure to not direct the water stream directly onto the embryos as this may damage or detach them from the medical dressing. This process removes most, but not all, of the Halocarbon oil.
- Cover the slide with 150 µL of DI water.
- Place the slide with the injected embryos on wet filter paper inside of a square Petri dish.
- Place the Petri dish containing the injected embryos in an environmental chamber set to 25-27 °C, 70-80% RH with a long day photoperiod (>13 h light/day).
7. Rearing injected embryos (F0) to adulthood and setting up crosses
- Examine the slides of injected embryos daily, expecting that 1st instar larvae should emerge 1.5 days after injection.
- Count the number of 1st instar larvae, and place 100-200 larvae into a shoebox-sized Tupperware container with 450 mL of DI water, and 50 mg of ground fish food. At this time, create 1-5 times more containers containing wild-type or un-injected larvae that will be used to cross with the injected mosquitoes.
- Feed the larvae daily, slightly increasing the amount of food they receive each day until reaching 300-500 mg on the 6th day of larval life.
- Once pupae appear, use a transfer pipette to place one pupae into a 2 mL microcentrifuge tubes filled 50-75% full of DI water. Repeat for all pupae, both the pupae that were injected as embryos as well as uninjected, wild-type pupae. Carefully press the lids so that they are slightly ajar, preventing the mosquitoes from escaping but still allowing for a small amount of gas exchange.
- Once the adult mosquitoes emerge inside their microcentrifuge tubes, release injected female mosquitoes into a cage containing virgin wild-type male mosquitoes, achieving a ratio of at least 1 wild-type male for every injected female. Similarly, release injected male mosquitoes into a cage containing virgin wild-type females, achieving a ratio of approximately 5-10 virgin wild-type females for every injected male. A higher ratio of wild-type females to injected males is recommended as each male mosquito can mate multiple times. Therefore, having higher numbers of wild-type females that mate with injected males increases the number of F1 progeny that can be produced.
8. Obtaining and rearing F1 mosquitoes and screening them for mutations
- Seven to ten days after adult emergence, offer a blood meal to the females in each cage as described above (step 3).
- Four days after blood-feeding, place oviposition water into each cage. Each egg raft represents the progeny of a single female mosquito, and therefore should be placed into its own plastic, shoebox-sized rearing container shortly after oviposition. Label each container with a unique identifier, and also indicate that the larvae belong to the F1 generation for the gene of interest, whether the male or female parent was injected, the date the pan was established, rearing conditions, and the experimenter’s initials.
- Monitor the pans of larvae daily. As soon as larvae hatch in the pans, provide 50 mg of ground fish food. Increase the food daily for 6 days as described above (step 7.3).
- Transfer individual pupae to 2 mL microcentrifuge tubes containing between 1-1.5 mL of DI water, label each tube, and crack the lids.
- Once the adult mosquitoes emerge, carefully open the 2 mL microcentrifuge tube, covering with an index finger, and using the other hand, place a 3 dram glass vial over the top of the microcentrifuge tube. The mosquito’s natural instinct should be to fly up and into the glass vial. Once this happens, slide a finger over the opening of the glass vial, and quickly invert it, placing a small ball of cotton that has been slightly wetted with 10% sucrose solution into the top of the vial. This prevents the mosquito from escaping and provides a necessary food and water source.
- Label both the tube containing the pupal exuvia and the glass vial containing the adult mosquito to indicate the larval pan from which it originated, the sex of the mosquito, and the unique identifier for that individual. Ex: II.C.M32, where “II” represents that the male parent was injected, “C” represents the pan/egg raft from which the individual originated, and “M32” designates that was the 32nd male to emerge from that larval rearing container.
- Place adult mosquitoes within their individual glass vials back into the environmental chamber, and add a small amount (~20 µL) of 10% sucrose solution to their cotton daily. Be sure not to overfeed or saturate the cotton as the mosquitoes will get stuck in the sucrose solution and die.
- Screening mosquitoes for the desired mutation
- Extract genomic DNA from 5-10 pupal exuvia from each egg raft/pan of larvae using the Phire Direct PCR Kit according to the manufacturer’s instructions, as well as 1-2 wild-type individuals which will serve as a control.
NOTE: As the offspring from each raft are likely full siblings, screening 5-10 larvae from each pan will determine whether the mutation has been passed on to the offspring. If none of the screened larvae from one pan have the desired mutation, do not screen additional adults. If some of the larvae have the mutation, then every individual from that pan should be screened. - Using the previously designed PCR primers (step 1.2), amplify the fragment around the predicted location of your mutation using the Phire PCR kit in both the wild-type and F1 progeny and run PCR fragments on a 3-4% agarose gel to determine if there are any visible insertions or deletions (indels).
NOTE: Any individuals that have the desired mutation will be heterozygous and should display 2 bands, one wild type and one either slightly shorter or longer at the desired position. To distinguish these, compare the position and thickness of the bands of the wild type individuals with those in the bands in the F1 progeny. Ideally 2 discrete bands will be visible, but if the indel is very small, the band might appear to be wider and/or blurrier. - Purify the PCR product either by excising the band containing the suspected indels (recommended) or by purifying the remaining PCR product that was not loaded into the gel. Submit the purified PCR for sequencing to determine whether the desired mutation is present.
- Extract genomic DNA from 5-10 pupal exuvia from each egg raft/pan of larvae using the Phire Direct PCR Kit according to the manufacturer’s instructions, as well as 1-2 wild-type individuals which will serve as a control.
9. Obtaining and rearing F2 and F3 mosquitoes and screening them for mutations
- Release all F1 adult mosquitoes for which the desired mutation was found by screening its pupal exuvia from their individual glass vials and into a cage containing wild-type, un-injected mosquitoes of the opposite sex. Backcrossing for this second generation should remove all deleterious effects of injection and inbreeding.
- Blood feed the cages of the mutant F1 mosquitoes and their wild-type counterparts as previously described. Four to five days later, collect the resulting F2 egg rafts.
- Label and rear the F2 larvae as described above, placing each egg raft into a separate, labeled container. Upon pupation, again separate individual pupae into individual 2 mL microcentrifuge tubes, and transfer each adult into separate glass vials capped with sucrose-soaked cotton upon emergence. Then screen the pupal exuvia of each F2 individual the desired mutation as previously described (steps 8.5-8.8).
NOTE: Here all individuals that have the desired mutation will be heterozygous, and therefore the PCR product should ideally produce 2 distinct bands when run on a 3-4% agarose gel. - Once mutant F2 larvae have been identified, place all males and females that have the mutation into the same cage to mate. Blood feed 7-10 days after adult emergence, and 4-5 days later, collect the F3 egg rafts.
- Place each F3 egg raft into a separate larval rearing container, and label and feed as described above.
- Separate the F3 pupae into individual 2 mL microcentrifuge tubes. Once adults emerge, transfer them to individual glass vials capped with 10% sucrose-soaked cotton. Screen the pupal exuvia as previously described. This time, approximately 25% of the offspring in each pan should be homozygous for the null mutation. Place these individuals into a single cage, and use them to create and maintain the newly-generated mutant line of mosquitoes.
NOTE: If a low number of homozygous mosquitoes are present, heterozygous F3 males and females can be crossed with each other to generate additional homozygous-null mosquitoes in the F4 generation.
Representative Results
Using the described protocol, we were able to successfully inject embryos of Cx. pipiens, and observed a high rate of survival among the injected embryos (~55%, Figure 1). Earlier trials had a lower percentage of survival, likely because the anterior of the egg follicle was attached to the medical dressing strip, preventing mosquito larvae from escaping from the chorion and successfully swimming into the water. Ensuring that the anterior end extends beyond the strip of medical dressing greatly increased larval survival, and results in high quality offspring that are capable of developing to adulthood and reproducing (Figure 2B).
Subsequent experiments and screening indicate that the null mutation for the circadian clock gene cycle (cyc; KM355981) made it into the germline of the injected embryos (Figure 3). Given that our primers amplified a large region of the cyc gene near the cut-sites, it was difficult to observe two separate bands in heterozygous, F1 mosquitoes. This demonstrates the utility of designing primers that will amplify fragments that are ~100-200 bp around the cut site, and also the importance of sequencing all suspected mutant mosquitoes. Sequencing revealed that approximately 10% of the screened mosquitoes showed a small insertion or deletion near the Cas9 cut site of the first guide RNA that we had designed (Figure 3), suggesting that this was a highly effective gRNA and that we successfully targeted the pole cells during microinjections. The F1 individuals successfully mated and produced viable offspring (F2 generation), some of which also contained the mutation.

Figure 1: Example of a beveled borosilicate needle. This needle was fabricated from a siliconized borosilicate microcapillary tube, pulled using a PC-10 vertical needle puller, and then beveled on a BV-10 Beveller. The needle is viewed under ~63x magnification. Please click here to view a larger version of this figure.
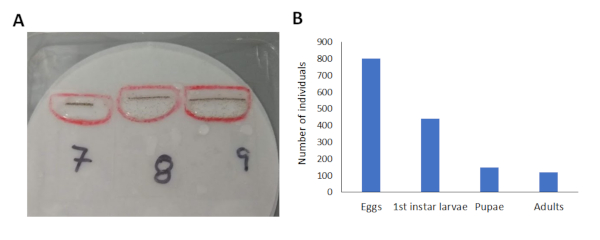
Figure 2: Survival of the F0 generation across their development. (A) Image of 1st instar larvae that hatched from 3 slides containing injected embryos. (B) Graphical representation of the number of individuals at each life stage. Of the 801 embryos that were injected, 441 1st instar larvae hatched, and of these 149 individuals reached pupation, resulting in a total of 121 adults (73 males and 43 females). Please click here to view a larger version of this figure.
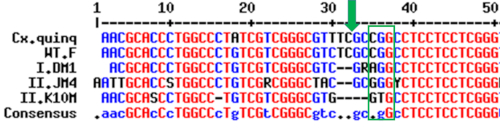

Figure 3: Representative results showing the transmission of the desired mutation. The top two sequences represent the sequence of the cycle gene in Culex quinquefascaitus (Cx.quinq; XP_001865023.1) and in wild type females of Cx. pipiens (WT.F; KM355981). The sequences below represent sequences in three male F1 offspring (I.DM1; II.JM4; II.K10M). The green box represents the PAM sequence recognized by the Cas9 protein (CGG) while the green arrow represents where the Cas9 protein cleaved genomic DNA. These results show that short deletions of 2 or 4 nucleotides were inherited in the F1 males. Please click here to view a larger version of this figure.
Discussion
This protocol presents methods to introduce specific mutations into the genome of Culex mosquitoes and can be used to edit the genome of other mosquitoes as well. The protocol is significant in that it provides specific details of not only how to prepare the injection materials, but also a detailed video overview of how to induce mosquitoes to lay eggs, as well as how to prepare and inject those eggs. We also summarize of how to take advantage of the biology of female Cx. pipiens to lay eggs in individual rafts and thereby screen a smaller proportion of offspring from each female for desired mutations. The methods presented here have been optimized for Cx. pipiens but can be adapted, with small adjustments, to edit the genomes of other mosquitoes or insects. Additionally, the micromanipulation and microinjection protocols described here are amenable to injecting several different materials into insect embryos, including transposons, dsRNA or other endonucleases such as TALENs or Zinc-Finger nucleases. Moreover, the beveling protocol generates microinjection needles with variable opening sizes therefore creating needles that can be used for injecting a wide variety of injection materials. The open size of the needle can be inferred by slowly decreasing the pressure of air after the needle has been beveled, and noting the pressure at which bubbles of air stop flowing from the tip of the newly opened needle. The less air pressure that is needed to produce bubbles indicates that the needle has a larger the opening size, and conversely, higher air pressures indicate that the needle has a smaller opening size. Needles with a larger opening sizes are better for injecting larger particles and more viscous materials, but will likely cause greater damage to the embryo and therefore reduce survival.
Microinjection experiments are in many ways a race against the clock. First, one must inject individual embryos within the first 2 hours of oviposition, before the chorion hardens and blastoderm formation occurs. Therefore, it is imperative to note when the mosquitoes were first placed in the oviposition chamber, how long it takes to manipulate the eggs for injection (we recommend no more than 20-30 minutes), and how long it takes to inject all embryos. Time is also limited when screening adult mosquitoes for mutations as Cx. pipiens are quite sensitive to their surroundings and mosquitoes generally only survive for 3-5 days in individual glass vials. Therefore, it is imperative to screen mosquito pupal exuvia as quickly as possible. Alternatively, one could also extract genomic DNA from a single mosquito leg14. In order to do so, however, the mosquitoes should first be anesthetized on ice. As we were weary of how the cold treatment and loss of limbs might impact mosquito survival and their ability to mate and lay viable eggs, we decided to instead screen the discarded pupal exuvia for mutations. The level of gDNA within the exuvia is likely lower than inside a mosquito leg, and this does add extra effort and time to separate mosquitoes at the pupal stage, but also ensures that all of the adults are unmated when they are released into their appropriate cages, which is absolutely vital. We feel that the results are worth it, but encourage others to consider if removing a leg might be a better course of action for their experimental needs.
The best way to expedite screening for mutations is to inject a plasmid containing a genetic marker, such as a fluorescent protein, flanked by homologous sequences on either side of the cut site. Rates of knock-in mutations using homology directed-repair (HDR) are generally lower than non-homologous end-joining (NHEJ), knock-out mutations (HDR=0.71%; NHEJ=24.87%;22). Therefore, the insertion of the marker will likely occur with a much lower frequency, but the time-savings offered by being able to quickly screen mosquito larvae under a fluorescent microscope may be worth the risk, depending on the project. Additionally, rates of HDR and knock-in mutations are enhanced when one also injects a plasmid containing the genetic sequence of the Cas9 protein, driven by the a well-characterized embryonic promotor, and the gRNA driven by the U6 promotor24,26. This is likely because early in embryonic development, when nuclei are rapidly dividing (S phase of the cell cycle), the error-prone but expedient NHEJ mechanism seems to be the preferred method of repairing double-stranded breaks (reviewed18). However, as embryogenesis progresses the cellular machinery is primed to use the more accurate HDR mechanism to repair double-stranded breaks (late S phase and G2 phase). Injecting a plasmid that contains the Cas9 sequence early in embryonic development allows the Cas9 protein to reach most target cells while the embryo is still in its syncytial state and before cellular membranes have formed, but the slight delay required to transcribe and translate the Cas9 protein seems to increase the likelihood that the endonucleasewill be active at a time when the developing embryo is more likely to use HDR to accurately repair double-stranded breaks in genomic DNA. For example, Lin et al.27 discovered that rates of HDR were enhanced to ~33% when the Cas9 protein complexed with gRNA were injected in human cell lines arrested in the M phase of the cell cycle. An additional benefit of delivering Cas9 and gRNAs on plasmids is that they are less viscous and therefore less likely to clog the needle during injections (R. Harrell, personal observation). However, until embyronic promoters are characterized and validated in Culex mosquitoes, we recommend injecting Cas9 protein or mRNA as described here.
Additionally, given the amount of time that one must dedicate to rearing and screening mosquitoes, we recommend having at least one full-time technician, student or researcher devoted to this task. We also recommend strategically considering how many null mutants one needs for a given experiment and the importance of genetic diversity within the null line. While we were able to identify mosquitoes in the F1 and F2 generations that had the mutation, only a handful of the F2 mosquitoes survived to adulthood the heterozygous mutant mosquitoes failed to produce viable offspring. We think that this is much less to do with the mutation, and more likely is a result of the lengthy period of time that both male and female mosquitoes were confined to glass tubes while we were screening them. This prolonged period of isolation most likely interfered with their ability to successfully mate and, in the case of females, obtain a blood meal and lay viable eggs. Outcrossing for a single generation and/or having optimized screening techniques in place should prevent this issue from occurring in the future. Therefore, by following this protocol we are confident that researchers will be able to successfully generate null lines of Culex mosquitoes and, with minor adjustments, additional insects.
Disclosures
The authors have nothing to disclose.
Acknowledgements
We thank Dr. David O’Brochta and all members of the Insect Genetic Technologies Coordination Research Network for the help and training that they provide to us and others on the implementation of genetic technologies. We especially thank Channa Aluvihare for optimizing the micromanipulation protocol to allow Culex embryos to be injected and hatch. We also thank Devante Simmons and Joseph Urso, undergraduate students working in the Meuti lab, for their assistance caring for and screening transgenic mosquitoes, and Zora Elmkami from the ITF for assistance rearing and prepping mosquitoes for injection. This work was supported by an Interdisciplinary Seeds grant from the Infectious Diseases Institute at OSU provided to MEM.
Materials
| Artificial Membrane Feeder | Hemotek | SP5W1-3 | Company location: Blackburn, UK |
| ATP | Invitrogen | 18330019 | Company location: Carlsbad, CA, USA |
| Borosilicate glass mirocapillary tubes, 1 mm outer diameter | World Precision Instruments | 1B100-6 | Company Location: Sarasota, FL, USA |
| BV10 Needle Beveler | Sutter Instruments | BV-10-B | Company Location: Nobato, CA, USA |
| Whatman Circular filter paper (12.5 cm) | Sigma Aldrich | WHA1001125 | Company Location: St. Louis, MO, USA |
| Conical tube (50 mL) | Thermo Fisher Scientific | 339652 | Company Location: Waltham, MA, USA |
| Fisherbrand course filter paper with fast flow rate | Thermo Fisher Scientific | 09-800 | Company Location: Waltham, MA, USA |
| Cover glass (24 x 40 mm) | Thermo Fisher Scientific | 50-311-20 | Company Location: Waltham, MA, USA |
| Dental dam | Henry Schein Inc | 1010171 | Company Location: Melville, NY USA |
| Scotch double-sided tape | Thermo Fisher Scientific | NC0879005 | Company Location: Waltham, MA, USA |
| FemtoJet 4i microinjector | Eppendorf | 5252000021 | Company Location: Hamburg, Germany |
| Glass vial (2 dram) | Thermo Fisher Scientific | 033401C | Company Location: Waltham, MA, USA |
| Halocarbon oil | Sigma Aldrich | H8898-50ML | Company Location: St. Louis, MO, USA |
| P-2000 Laser Needle Puller | Sutter Instruments | P-2000/G | Company Location: Nobato, CA, USA |
| Parafilm | Thermo Fisher Scientific | 50-998-944 | Company Location: Waltham, MA, USA |
| PC-100 Weighted Needle Puller | Narishige | PC-100 | This is compatabile with the earlier PC-10 model, which has been discontinued. Company Location: Amityville, NY, USA |
| Phire Direct PCR Kit | Thermo Fisher Scientific | F140WH | Company Location: Waltham, MA, USA |
| Kodak Photo-Flo (1%) | Thermo Fisher Scientific | 50-268-05 | Company Location: Waltham, MA, USA |
| Quartz glass mirocapillary tubes, 1 mm outer diameter | Capillary Tube Supplies Limited | QGCT 1.0 | Company Location: Cornwall, UK |
| Guide-it™ sgRNA Screening Kit | Takara, Bio USA | 632639 | This kit allows you to determine if gRNAs cut DNA sequences in vitro. Company Location: Mountain View, CA, USA |
| Sigmacote | Sigma Aldrich | SL2-100ML | Company Location: St. Louis, MO, USA |
| Small petri dishes (35X10 mm) | Thermo Fisher Scientific | 50-190-0273 | Company Location: Waltham, MA, USA |
| Sodium citrate chicken blood | Lampire biologicals | 7201406 | Company Location: Everett, PA, USA |
| Fisherbrand Square petri dish (10 cm x 10 cm) | Thermo Fisher Scientific | FB0875711A | Company Location: Waltham, MA, USA |
| Tegaderm | Henry Schein Inc. | 7771180 | Company Location: Melville, NY USA |
| Tropical fish food | Tetramin | N/A | |
| Whatman filter paper | Thermo Fisher Scientific | 09-927-826 | Company Location: Waltham, MA, USA |
| Whatman filter paper, 4.25 cm | Sigma Aldrich | 1001-042 | Company Location: St. Louis, MO, USA |
References
- Hamer, G. L., et al. Culex pipiens (Diptera: Culicidae): a bridge vector of West Nile virus to humans. Journal of Medical Entomology. 45 (1), 125-128 (2008).
- Bailey, C. L., et al. Isolation of St. Louis encephalitis virus from overwintering Culex pipiens mosquitoes. Science. 199 (4335), 1346-1349 (1978).
- Sabry, M. A new realistic index of experimental transmission efficiency for Bancroftian filariasis. The Journal of Tropical Medicine and Hygiene. 94 (4), 283-290 (1991).
- Cancrini, G., et al. Aedes albopictus and Culex pipiens implicated as natural vectors of Dirofilaria repens in central Italy. Journal of Medical Entomology. 44 (6), 1064-1066 (2007).
- Wilton, D. P., Smith, G. C. Ovarian diapause in three geographic strains of Culex pipiens (Diptera: Culicidae). Journal of Medical Entomology. 22 (5), 524-528 (1985).
- Mattingly, P. F. The Culex pipiens complex. Transactions of the Royal Entomological Society of London. 102 (7), 331-342 (1951).
- Eldridge, B. F. The effect of temperature and photoperiod on blood-feeding and ovarian development in mosquitoes of the Culex pipiens complex. The American Journal of Tropical Medicine and Hygiene. 17 (1), 133-140 (1968).
- Spielman, A., Wong, J. Environmental control of ovarian diapause in Culex pipiens. Annals of the Entomological Society of America. 66 (4), 905-907 (1973).
- Fritz, M. L., Walker, E. D., Miller, J. R., Severson, D. W., Dworkin, I. Divergent host preferences of above-and below-ground Culex pipiens mosquitoes and their hybrid offspring. Medical and Veterinary Entomology. 29 (2), 115-123 (2015).
- Meola, R. The influence of temperature and humidity on embryonic longevity in Aedes aegypti. Annals of the Entomological Society of America. 57 (4), 468-472 (1964).
- Dong, S., Lin, J., Held, N. L., Clem, R. J., Passarelli, A. L., Franz, A. W. Heritable CRISPR/Cas9-mediated genome editing in the yellow fever mosquito, Aedes aegypti. PloS one. 10 (3), (2015).
- Kyrou, K., et al. A CRISPR-Cas9 gene drive targeting doublesex causes complete population suppression in caged Anopheles gambiae mosquitoes. Nature Biotechnology. 36 (11), 1062-1066 (2018).
- Liu, T., et al. Construction of an efficient genomic editing system with CRISPR/Cas9 in the vector mosquito Aedes albopictus. Insect Science. 26 (6), 1045-1054 (2019).
- Itokawa, K., Komagata, O., Kasai, S., Ogawa, K., Tomita, T. Testing the causality between CYP9M10 and pyrethroid resistance using the TALEN and CRISPR/Cas9 technologies. Scientific Reports. 6, 24652 (2016).
- Li, M., Li, T., Liu, N., Raban, R. R., Wang, X., Akbari, O. S. Methods for the generation of heritable germline mutations in the disease vector Culex quinquefasciatus using clustered regularly interspaced short palindrome repeats-associated protein 9. Insect Molecular Biology. 29, 214-220 (2019).
- Anderson, M. E., et al. CRISPR/Cas9 gene editing in the West Nile Virus vector, Culex quinquefasciatus Say. PloS one. 14 (11), (2019).
- Van der Oost, J., Jore, M. M., Westra, E. R., Lundgren, M., Brouns, S. J. CRISPR-based adaptive and heritable immunity in prokaryotes. Trends in Biochemical Sciences. 34 (8), 401-407 (2009).
- Heyer, W. D., Ehmsen, K. T., Liu, J. Regulation of homologous recombination in eukaryotes. Annual Review of Genetics. 44, 113-139 (2010).
- Monnerat, A. T., et al. Anopheles albitarsis embryogenesis: morphological identification of major events. Memorias do Instituto Oswaldo Cruz. 97 (4), 589-596 (2002).
- Davis, C. W. C. A comparative study of larval embryogenesis in the mosquito Culex fatigans Wiedemann (Diptera: Culicidae) and the sheep-fly Lucilia sericata Meigen (Diptera: Calliphoridae). Australian Journal of Zoology. 15 (3), 547-579 (1967).
- Port, F., Chen, H. M., Lee, T., Bullock, S. L. Optimized CRISPR/Cas tools for efficient germline and somatic genome engineering in Drosophila. Proceedings of the National Academy of Sciences. 111 (29), 2967-2976 (2014).
- Kistler, K. E., Vosshall, L. B., Matthews, B. J. Genome engineering with CRISPR-Cas9 in the mosquito Aedes aegypti. Cell Reports. 11 (1), 51-60 (2015).
- Gokcezade, J., Sienski, G., Duchek, P. Efficient CRISPR/Cas9 plasmids for rapid and versatile genome editing in Drosophila. G3: Genes, Genomes, Genetics. 4 (11), 2279-2282 (2014).
- Hwang, W. Y., et al. Efficient in vivo genome editing using RNA-guided nucleases. Nature Biotechnology. 31 (3), 227-229 (2013).
- Basu, S., et al. Silencing of end-joining repair for efficient site-specific gene insertion after TALEN/CRISPR mutagenesis in Aedes aegypti. Proceedings of the National Academy of Sciences. 112 (13), 4038-4043 (2015).
- Hammond, A., et al. A CRISPR-Cas9 gene drive system targeting female reproduction in the malaria mosquito vector Anopheles gambiae. Nature Biotechnology. 34 (1), 78-83 (2016).
- Lin, S., Staahl, B. T., Alla, R. K., Doudna, J. A. Enhanced homology-directed human genome engineering by controlled timing of CRISPR/Cas9 delivery. eLife. 3, 04766 (2014).

