Monitoring Circadian Oscillations with a Bioluminescence Reporter in Organoids
Summary
Circadian rhythms, which exist in most organisms, regulate the temporal organization of biological processes. 3D organoids have recently emerged as a physiologically relevant in vitro model. This protocol describes the use of bioluminescent reporters to observe circadian rhythms in organoids, enabling in vitro investigations of circadian rhythms in multicellular systems.
Abstract
Most living organisms possess circadian rhythms, which are biological processes that occur within a period of approximately 24 h and regulate a diverse repertoire of cellular and physiological processes ranging from sleep-wake cycles to metabolism. This clock mechanism entrains the organism based on environmental changes and coordinates the temporal regulation of molecular and physiological events. Previously, it was demonstrated that autonomous circadian rhythms are maintained even at the single-cell level using cell lines such as NIH3T3 fibroblasts, which were instrumental in uncovering the mechanisms of circadian rhythms. However, these cell lines are homogeneous cultures lacking multicellularity and robust intercellular communications. In the past decade, extensive work has been performed on the development, characterization, and application of 3D organoids, which are in vitro multicellular systems that resemble in vivo morphological structures and functions. This paper describes a protocol for detecting circadian rhythms using a bioluminescent reporter in human intestinal enteroids, which enables the investigation of circadian rhythms in multicellular systems in vitro.
Introduction
Circadian clock
All organisms, from bacteria to mammals, have a complex and dynamic relationship with their environment. Within this relationship, adaptation to environmental changes is critical for the survival of organisms. Most organisms possess circadian rhythms that enable them to adapt and optimize their functions to diurnal cycles of approximately 24 h. The circadian clock is a hierarchical network of central and peripheral clocks that work in cooperation to maintain physiological homeostasis and keep organisms synchronized with daily changes1,2. In mammals, the central or master clock located in the suprachiasmatic nucleus (SCN) receives external cues, such as light, and transmits the information to the peripheral clocks via an advanced interplay of neural and humoral signaling pathways3. In addition to the central clock, peripheral tissues possess their own cell-autonomous circadian clock mechanism, maintained by a transcriptional-translational negative feedback loop (TTFL) regulating tissue-specific clock-controlled genes (CCGs)4,5. This molecular machinery produces approximately 24 h rhythmicity in cellular and physiological events, such as gene expressions, signaling pathways, immune responses, and digestion. The circadian clock is present in almost all mammalian cells, and it has been demonstrated that up to 50% of the genes' expression patterns exhibit circadian rhythmicity6. Considering the abundance of CCGs, disruption of this clock mechanism may result in critical physiological problems. Hence, investigations into circadian rhythms are necessary to elucidate essential biological mechanisms and develop novel therapeutic strategies.
Luciferase reporter system
In circadian studies, real-time monitoring is critical for a better understanding of cellular behaviors and responses because it allows tracking of temporal changes in gene expression and/or protein levels, providing insights into the molecular mechanisms regulated by the circadian clock. Furthermore, real-time monitoring enables researchers to study the effects of environmental changes on molecular mechanisms7,8. There are numerous techniques for real-time monitoring studies, including bioluminescence assay, which is widely used to track gene expression or protein levels over time. Bioluminescence assay is a method to detect biological processes using light production as a readout. In this assay, an oxidative enzyme that produces bioluminescence (e.g., luciferase) is either transiently or stably transfected into cells of interest, and the bioluminescence readout is measured in the presence of a substrate (e.g., luciferin) over time. For example, the luciferase enzyme produces bioluminescence by oxidizing the substrate luciferin in the presence of ATP9. Due to its short half-life, 3-4 h10, firefly luciferase is a powerful tool for circadian studies in terms of providing real-time dynamic monitoring with minimal background noise11,12,13. For the insertion of DNA with a luciferase-tagged promoter or open reading frame (ORF), the lentiviral gene delivery system is a reliable method that provides high transduction efficacy, stable integration, and low immunogenicity. Stable transduction of a bioluminescent reporter provides robust expression in dividing and non-dividing cells, generating consistent data for circadian studies14.
Organoid as a model
Traditional immortalized two-dimensional cell lines have been instrumental in biological studies ranging from uncovering fundamental molecular mechanisms of circadian rhythms to drug screening. Despite the convenience of utilizing homogenized cell lines, they lack multicellular structures and intercellular interactions. In contrast, organoids are in vitro 3D multicellular "organ-like" structures that mimic organ structure in a dish by displaying similarity to in vivo tissue architecture and multicellularity, including stem, progenitor, and differentiated cell types15,16. Possessing self-organization, multicellularity, and functionality features makes organoids a remarkable in vitro model representing the cellular and physiological processes occurring in real tissues17. Different types of organoids can be derived from pluripotent stem cells via directed differentiation or adult stem cells harvested from various organs, including the small intestine, brain, liver, lung, and kidney18,19. Since organoid structures possess a real tissue-like architecture and function with multicellularity and dynamic cell-to-cell interaction, they are superior to homogenized cell lines for understanding the cellular events occurring in in vivo tissues. Organoids are also easily manipulated and can be grown under controlled conditions, making them useful for circadian studies20.
The main purpose of this work is to introduce a real-time monitoring method utilizing a bioluminescence assay specifically tailored for studying circadian rhythms in multicellular 3D organoids. Real-time monitoring of cellular events using a bioluminescence assay technique has been widely performed for cell cultures lacking the multicellular complexity and intercellular communications that exist in real tissues. 3D organoids present unique opportunities to investigate the functions of circadian rhythms in multicellular systems in vitro. For example, one could investigate circadian rhythms in the organoids with altered cell compositions or organoids derived from patients' diseased tissues. This protocol enables the utilization of a bioluminescence assay to investigate different aspects of circadian rhythms in a more physiologically relevant in vitro model, organoids, which will help us better understand the roles of circadian rhythms in peripheral organs.
Protocol
All experiments using human tissues for the generation of HIEs were approved by an IRB at CCHMC (IRB#2014-0427). See the Table of Materials for details related to all materials used in this protocol.
NOTE: To illustrate the procedure outlined in this protocol, we utilized Bmal1-luc human intestinal enteroids (HIEs). These enteroids underwent stable lentiviral transduction22 with the pABpuro-BluF reporter plasmid, which contains the Bmal1 promoter fused to luciferase, showing the activity of the Bmal1 promoter21.
1. Lentiviral transduction
- Carry out the lentiviral transduction22 under stem cell-enriched conditions for HIEs. Specifically, ensure the formation of spheroids in HIEs within 3-4 days after passage in the intestinal growth media, which creates stem cell-enriched conditions.
- After the lentiviral transduction22, carry out antibiotic selection with 2 µg/mL puromycin, recover the puromycin-resistant HIEs, and passage them with relatively high density (i.e., 1 well into 2 wells) for 2-3x. If the growth rate of the HIEs is slow, add 10 µM ROCK inhibitors (Y-27632) and 10 µM GSK-3 inhibitor (CHIR-99021) to the intestinal organoid growth medium.
2. Preparation of organoids for bioluminescence assay
- Day 1: Passage HIEs to 35 mm round dishes.
- Briefly, for expansion of HIEs, mix the HIEs with growth factor-reduced (GFR) 3D culture matrix and seed them onto 24-well plates with three droplets of 10 µL of 3D culture matrix per well. Add 350 µL of the HIE growth medium and change it every other day.
NOTE: The density of HIE in the 3D culture matrix is critical for the growth of enteroids. Approximately, the number of spheroids should exceed more than 50 in each droplet. If the number is low, the growth rate of the HIE is negatively affected. - When HIEs grow sufficiently (approximately 7 days after the post-passage), collect HIEs from four wells into 1.5 mL microcentrifuge tubes.
- Wash the HIEs with the cold PBS using a benchtop mini centrifuge (2,000 × g) for 40 s quick spin to remove debris and excessive 3D culture matrix by pipet.
- Add 0.5 mL of the cold PBS into the pellet and apply physical turbulence by syringing up to 10x with a 1 mL insulin syringe to dissociate the HIEs.
- Spin down the dissociated HIEs with a benchtop mini centrifuge as in step 2.1.3 and gently remove the supernatant by pipet. Then, place the microcentrifuge tube with the HIE pellet on ice to cool down for approximately 3 min.
- Add GFR 3D culture matrix to the microcentrifuge tube and mix it with the HIE pellet.
NOTE: Calculate the amount of the 3D culture matrix based on the number of 35 mm round dishes that will be used for the experiment. We generally seed 30 µL of 3D culture matrix droplets per round dish. - Seed the mixture of HIEs and the 3D culture matrix on the center of the 35 mm round dish and incubate the dish at 37 °C for 20 min to solidify the 3D culture matrix.
NOTE: The density of seeding HIEs should be determined experimentally. Based on the intensity of bioluminescence, one can determine the number of HIEs during this process. - Add 2 mL of prewarmed intestinal organoid growth medium to each dish and incubate the dishes in a CO2 incubator until the next step.
- Briefly, for expansion of HIEs, mix the HIEs with growth factor-reduced (GFR) 3D culture matrix and seed them onto 24-well plates with three droplets of 10 µL of 3D culture matrix per well. Add 350 µL of the HIE growth medium and change it every other day.
- Day 3: Initiation of differentiation
- To initiate differentiation of the HIEs, remove the growth medium and add 2 mL of prewarmed differentiation medium.
NOTE: See Table 1 for the recipes of the HIE differentiation media23.
- To initiate differentiation of the HIEs, remove the growth medium and add 2 mL of prewarmed differentiation medium.
- Day 4: No additional process is required.
- Day 5: Replace the medium of the enteroids with 2 mL of prewarmed differentiation medium.
- Day 6: Clock synchronization and recording bioluminescence
- To synchronize the molecular clock of enteroids, add 2 µL of 100 µM dexamethasone (Dex) to make the final concentration 100 nM in each dish and then incubate the dishes in a CO2 incubator (37 °C, 5% CO2) for 1 h24.
- During the Dex resetting, prepare the incubating luminometer. Check the CO2 and temperature of the incubating luminometer and set to 5% and 37 °C, respectively.
NOTE: It is suggested to wipe the inside of the incubating luminometer with 70% alcohol before and after every use of the machine to keep it sterile. - After the clock synchronization, replenish the enteroids with 3 mL of prewarmed differentiation medium containing 200 µM D-luciferin.
NOTE: As luciferin is light-sensitive, the luciferin-containing medium should be wrapped with foil and prewarmed in a 37 °C heat bath before applying it to the dish. - Place the dishes in an 8-well dish table of the incubating luminometer and cover the sample dish table with its lid. Place two humidifier chambers with wet tissue or sponge on the top of the sample dish table. This is one-time process.
NOTE: After the experiment is started, the lid of the incubating luminometer should be kept closed until the experiment is terminated. Make sure to wet the tissues/sponges enough. - Close the lid of the machine and initiate the program to measure bioluminescence for the desired number of days and resolution.
- On the software, select the condition settings by clicking the Condition tab and adjust the settings on this panel: number of dishes, measurement time, background processing, and filter type.
NOTE: For this demonstration, we selected all dishes from A to H; default measurement time, which is determined by the software based on the number of dishes; waiting time for background processing, and filter type F0. Since there are three filter options, three luminescent colors can be measured simultaneously. - After entering all suitable settings based on the experiment, save the settings by clicking OK.
- To start running the experiment, click the Run tab | Start button (green triangle icon). Observe the data as raw, noise filtered, or detrended by making appropriate selections on the top of the screen of the software while running experiments. Click on Raw, Noise Filtered, or Detrended tabs to observe the selected data.
- Observe signals for up to 5 days.
NOTE: After 5 days of monitoring in an incubating luminometer, enteroids may begin to die due to media depletion. To prevent any possible circadian disruption during the experiment, we do not perform media change for the duration of the experiment. - At the end of the experiment, stop the run by clicking the Stop button (blue square icon), go to the File tab, and click Save as. Save the data in the Kronos format; export the data to a spreadsheet in the File section by clicking the File option and then selecting Export as excel format under the File for further analysis.
- On the software, select the condition settings by clicking the Condition tab and adjust the settings on this panel: number of dishes, measurement time, background processing, and filter type.
Representative Results
Bioluminescence recording was conducted to assess the circadian rhythmicity of human intestinal enteroids (HIEs) under two distinct conditions: stem cell-enriched conditions using intestinal organoid growth medium (Figure 3) versus differentiation-inducing conditions, which was achieved by replacing the intestinal organoid growth medium with a differentiation medium. On the day of the experiment, we synchronized the circadian clocks by performing a 1 h treatment with 100 nM Dexamethasone. Subsequently, we replaced the medium with either 3 mL of intestinal organoid growth medium or differentiation medium supplemented with 200 µM D-Luciferin and initiated the monitoring of bioluminescence activity. Following a recording period of 4-5 days, we conducted a Fast Fourier Transform (FFT) analysis to determine the period of the observed circadian rhythms. Figure 3 shows the temporal dynamics of Bmal1 promoter activity in the aforementioned conditions. Under differentiating conditions, Bmal1-luc bioluminescence exhibited robust circadian oscillations with a mean periodicity of 22.92 h. In contrast, HIEs demonstrated disrupted circadian rhythms in stem cell-enriched conditions (Figure 3). These data show that HIEs in stem cell-enriched conditions do not show robust circadian oscillations. In other words, one would need to assess the impact of growth conditions with respect to circadian rhythms before investigating circadian clock-dependent functions in organoids.
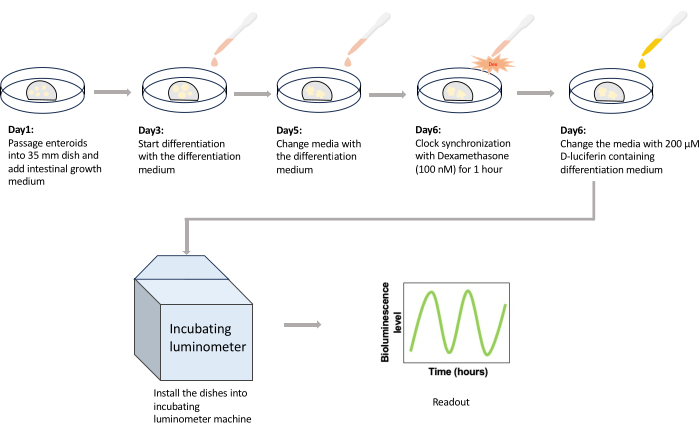
Figure 1: Schematic drawing of the protocol. HIEs are passaged into 35 mm dishes on day 1 and cultured with 2 mL of growth medium until day 3. On the third day after passage, the medium is replaced with 2 mL of prewarmed differentiation medium to initiate the differentiation of enteroids. Medium change is repeated with 2 mL of prewarmed medium on day 5 to prevent nutrition depletion. On day 6, enteroids are treated with 100 nM Dexamethasone for 1 h for circadian clock synchronization. After an hour, the media is replaced with 3 mL of 200 µM D-luciferin-containing medium, and the dishes are installed in the incubating luminometer. After recording the bioluminescence expression for 4-5 days, the experiment is terminated, and generated data are exported as spreadsheets for further analysis. Abbreviation: HIE = human intestinal enteroid. Please click here to view a larger version of this figure.
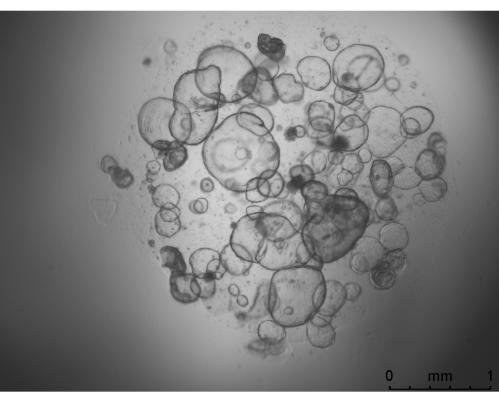
Figure 2: A representative image for HIEs under growth medium. The image was taken by a widefield microscope with a 25x objective. Scale bar = 1 mm. Abbreviation: HIE = human intestinal enteroid. Please click here to view a larger version of this figure.
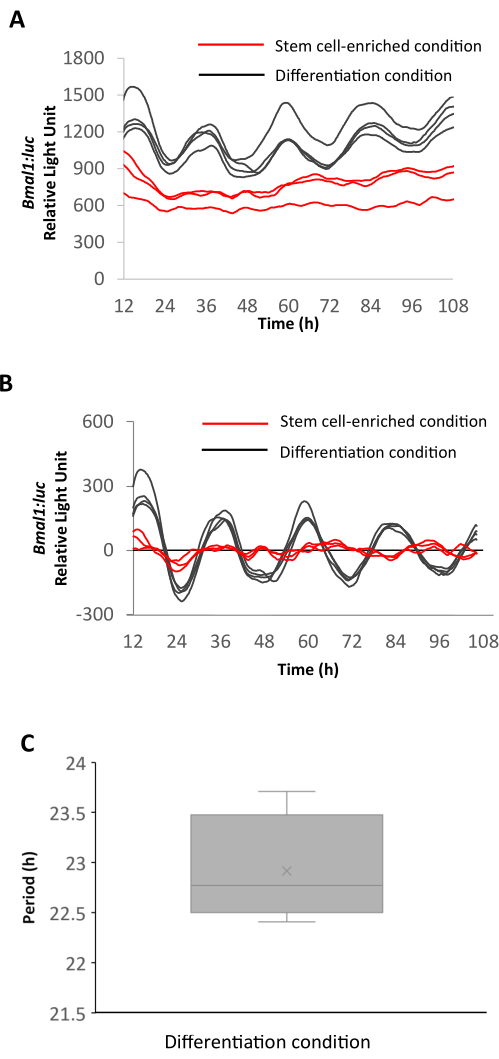
Figure 3: Bioluminescence recordings from HIEs from stem cell-enriched and differentiation conditions. (A) Raw and (B) detrended bioluminescence recording data of Bmal1-luc HIEs under stem cell-enriched (red) and differentiation (black) conditions. (C) Period of Bmal1-luc oscillations in HIEs under differentiation conditions. The period is calculated with FFT analysis; n = 3 for Bmal1-luc HIEs under stem cell-enriched condition and n = 4 for Bmal1-luc HIEs under differentiation condition. Error bars indicate SD. Abbreviations: HIE = human intestinal enteroid; FFT = Fast Fourier Transform. Please click here to view a larger version of this figure.
| Differentiation Media Recipes | ||||
| Name of Reagent | Company | Catalog Number | Final Concentration | Comments |
| Advanced DMEM/F12 | Life Technologies | 12634-028 | Add as the last component. If you prepare 50 mL media, after adding all other components, fill the rest up to 50 mL with Advanced DMEM/F12 at the end. | |
| A 83-01 | Sigma Aldrich | SML0788 | 500 nM | |
| B-27 Supplement (50x) | Gibco | 17504-044 | 1x | |
| Gastrin I Human | Sigma Aldrich | G9020 | 10 nM | |
| GlutaMAX | Gibco | 35050061 | 1x | |
| HEPES | Gibco | 15630080 | 10 mM | |
| N-2 Supplement (100x) | Gibco | 17502-048 | 1x | |
| N-Acetyl-L-cysteine | Sigma Aldrich | A9165 | 1 mM | |
| NOGGIN Conditioned Medium | 5% | Made in the lab | ||
| Penicillin-Streptomycin | Gibco | 15140122 | 100 U/mL | |
| Recombinant murine EGF | PeproTech | 315-09 | 50 ng/mL | |
| R-SPONDIN Conditioned Medium | 10% | Made in the lab |
Table 1: Composition of HIE differentiation media23.
Discussion
Bioluminescence assay offers several advantages for the investigation of circadian rhythms, which requires data collection from long-term time course experiments. First, it enables researchers to monitor the gene expression or protein of interest as cells move and proliferate. Without making unnecessary adjustments or disrupting the cells' functions, interested cellular events or gene expression can be recorded using bioluminescence readout, which gives reliable real-time data. Importantly, the bioluminescence assay generates minimal background noise compared to fluorescence assays because it is performed in constant darkness25. Furthermore, bioluminescence assay is preferable for long-term monitoring because the continuous excitation for fluorescence assay over several days may cause high phototoxicity and photobleaching. However, due to no light excitement, the bioluminescence assay provides a dimmer signal than fluorescence proteins26. The short half-life of luciferase makes it preferable for circadian studies, and it can further be shortened by adding additional destabilizing sequences12.
Despite offering several advantages for live-cell monitoring, bioluminescence assays have some limitations. Although the bioluminescence assay provides a robust readout for temporal gene expression or protein monitoring from a population of cells, its spatial resolution is limited, making it less feasible for experiments collecting protein localization data from single cells. Considering its advantages and disadvantages, the bioluminescence assay is superior to other methods, especially for circadian studies; it offers less toxicity and high sensitivity, making it ideal for long-term monitoring.
Several critical factors must be considered during the experimental preparation phase. Primarily, the implementation of a suitable selection strategy for lentiviral transduction is critical; in our case, we performed 2 µg/mL puromycin selection for 2 weeks. Inadequate execution of selection may result in insufficient numbers of enteroids harboring the reporter plasmid, yielding inconsistent luciferase expression levels. Optimal seeding density is equally important; excessive density may induce premature nutrient depletion during the experiment since there is no medium replacement after the initiation of bioluminescence recording. Conversely, low density of HIEs may result in reduced bioluminescence output, thereby influencing the robustness of readout. Ensuring a suitable environment for enteroids during monitoring is fundamental for the reliability of experimental outcomes. Maintaining optimal conditions including CO2, temperature, and humidity levels, is critical for the maintenance of HIEs, promoting the robustness of cellular processes within the enteroids. Deviations from optimal temperature and CO2 levels can affect cellular metabolism and signaling pathways, potentially influencing circadian rhythms. Adequate humidity levels are equally important, preventing desiccation and maintaining the structural integrity of enteroids. Thus, meticulous attention to environmental conditions is critical for the accuracy of the experimental model and the circadian studies involving enteroids.
Clock synchronization is essential for circadian studies as it aligns the internal circadian clock of individual cells, ensuring they are in a unified and coordinated state. This synchronization is necessary for precise observation and analysis of circadian rhythmicity at the level of an entire culture. In this protocol, we performed clock synchronization with 100 nM Dexamethasone (Dex). There might be some concerns about Dex's effect on the cell structure or function. Earlier research suggests that corticosteroids enhance epithelial barrier function in enteroids27. However, it is important to highlight that this phenotype is observed after treating enteroids with 10 µM prednisolone for 12 h. We expect that a short 1 h treatment of dexamethasone will not induce significant changes with respect to the structure and function of cells within enteroids.
Monitoring the bioluminescence activity of the organoid culture instead of the homologous single-cell culture possesses unique strengths. First, multiple cell types interact with each other in an organoid culture as in in vivo tissue, whereas a typical 2D cell culture monitors interested gene expression in a kind of isolated environment consisting of a single cell type. Organoid culture provides more realistic results with multicellular interactions and signaling pathways. Cell-to-cell interaction is critical, especially for the differentiation and intercellular pathway studies, which require heterogeneous cell culturing28. Our results indicate that HIEs in the differentiated condition show robust circadian oscillations but disrupted circadian rhythms in the stem cell-enriched condition. These data suggest potential remodeling of circadian clock machinery from adult stem cells to differentiated cells. In summary, organoids with a circadian bioluminescent reporter enable investigation of circadian rhythms in a multicellular in vitro system uncovering functions of circadian rhythms in a more physiologically relevant manner.
Disclosures
The authors have nothing to disclose.
Acknowledgements
The human intestinal enteroids were obtained from the lab of Dr. Michael Helmrath at Cincinnati Children's Hospital Medical Center (CCHMC). This work was supported by R01 DK11005 (CIH) and the University of Cincinnati Cancer Center Pilot Funding. We are grateful for imaging support from the University of Cincinnati Live Microscopy Core (NIH S10OD030402).
Materials
| 35 x 10 Falcon tissue culture dishes | Fisher Scientific | 08-772A | |
| A 83-01 | Sigma Aldrich | SML0788 | |
| Advanced DMEM/F12 | Life Technologies | 12634-028 | |
| B-27 Supplement (50x) | Gibco | 17504-044 | |
| BD Micro-Fine IV Insulin Syringes | Fisher Scientific | 14-829-1Bb | Mfrn: BD 329424 |
| CHIR99021 | Cayman Chemical | 13122 | GSK-3 inhibitor |
| Dexamethasone | Sigma Aldrich | D4902-500MG | |
| D-Luciferin (potassium salt) | Cayman Chemical | 14681 | |
| Gastrin I Human | Sigma Aldrich | G9020 | |
| GlutaMAX | Gibco | 35050061 | |
| Growth Factor reduced (GFR) Matrigel | Corning | CB-40230C | |
| HEPES | Gibco | 15630080 | |
| IntestiCult Organoid Growth Medium (Human) | Stemcell Technologies | 06010 | Consist of IntestiCult OGM Human Basal Medium, 50 mL and Organoid Supplement, 50 mL. Mix both as 1:1 ratio to use as intestinal organoid growth medium |
| Kronos Dio Luminometer Machine | ATTO Corporation | AB-2550 | |
| N-2 Supplement (100x) | Gibco | 17502-048 | |
| N-Acetyl-L-cysteine | Sigma Aldrich | A9165 | |
| pABpuro-BluF reporter plasmid | Addgene | 46824 | |
| PBS without Calcium and Magnesium | Corning | 21-040-CV | |
| Penicillin-Streptomycin | Gibco | 15140122 | |
| Recombinant murine EGF | PeproTech | 315-09 | |
| Y-27632 | R&D Systems | 1254/10 | ROCK inhibitor |
References
- Cox, K. H., Takahashi, J. S. Circadian clock genes and the transcriptional architecture of the clock mechanism. J Mol Endocrinol. 63 (4), R93-R102 (2019).
- Ayyar, V. S., Sukumaran, S. Circadian rhythms: Influence on physiology, pharmacology, and therapeutic interventions. J Pharmacokinet Pharmacodyn. 48 (3), 321-338 (2021).
- Reppert, S., Weaver, D. Coordination of circadian timing in mammals. Nature. 418 (6901), 935-941 (2002).
- Takahashi, J. Transcriptional architecture of the mammalian circadian clock. Nat Rev Genet. 18 (3), 164-179 (2017).
- Wolff, C. A., et al. Defining the age-dependent and tissue-specific circadian transcriptome in male mice. Cell Rep. 42 (1), 111982 (2023).
- Ruben, M. D., et al. A database of tissue-specific rhythmically expressed human genes has potential applications in circadian medicine. Science Trans Med. 10 (458), 8806 (2018).
- Huang, S., Lu, Q., Choi, M. H., Zhang, X., Kim, J. Y. Applying real-time monitoring of circadian oscillations in adult mouse brain slices to study communications between brain regions. STAR Protoc. 2 (2), 100416 (2021).
- Yagita, K., Yamanaka, I., Emoto, N., Kawakami, K., Shimada, S. Real-time monitoring of circadian clock oscillations in primary cultures of mammalian cells using Tol2 transposon-mediated gene transfer strategy. BMC Biotechnol. 10 (3), (2010).
- Choy, G., et al. Comparison of noninvasive fluorescent and bioluminescent small animal optical imaging. Biotechniques. 35 (5), 1022-1030 (2003).
- Leclerc, G. M., Boockfor, F. R., Faught, W. J., Frawley, L. S. Development of a destabilized firefly luciferase enzyme for measurement of gene expression. Biotechniques. 29 (3), 590-598 (2000).
- Ramanathan, C., Khan, S. K., Kathale, N. D., Xu, H., Liu, A. C. Monitoring cell-autonomous circadian clock rhythms of gene expression using luciferase bioluminescence reporters. J Vis Exp. (67), e4234 (2012).
- Yoo, S. H., et al. PERIOD2::LUCIFERASE real-time reporting of circadian dynamics reveals persistent circadian oscillations in mouse peripheral tissues. Proc Natl Acad Sci U S A. 101 (15), 5339-5346 (2004).
- Welsh, D. K., Yoo, S. H., Liu, A. C., Takahashi, J. S., Kay, S. A. Bioluminescence imaging of individual fibroblasts reveals persistent, independently phased circadian rhythms of clock gene expression. Curr Biol. 14 (24), 2289-2295 (2004).
- Tiscornia, G., Singer, O., Verma, I. Production and purification of lentiviral vectors. Nat Protoc. 1 (1), 241-245 (2006).
- Lee, S., Hong, C. I. Organoids as model systems to investigate circadian clock-related diseases and treatments. Front Genet. 13, 874288 (2022).
- Zachos, N. C., et al. Human enteroids/colonoids and intestinal organoids functionally recapitulate normal intestinal physiology and pathophysiology. J Biol Chem. 291 (8), 3759-3766 (2016).
- Bartfeld, S., Clevers, H. Stem cell-derived organoids and their application for medical research and patient treatment. J Mol Med. 95 (7), 729-738 (2017).
- Clevers, H. Modeling development and disease with organoids. Cell. 165 (7), 1586-1597 (2016).
- Rosselot, A. E., et al. Ontogeny and function of the circadian clock in intestinal organoids. EMBO J. 41 (2), 106973 (2021).
- Corrò, C., Novellasdemut, L., Li, V. S. W. A brief history of organoids. Am J Physiol Cell Physiol. 319 (1), C151-C165 (2020).
- Brown, S. A., et al. The period length of fibroblast circadian gene expression varies widely among human individuals. PLos Biol. 3 (10), e338 (2005).
- Van Lidth de Jeude, J. F., Vermeulen, J. L. M., Montenegro-Miranda, P. S., Vanden Brink, G. R., Heijmans, J. A protocol for lentiviral transduction and downstream analysis of intestinal organoids. J Vis Exp. (98), e52531 (2015).
- Criss, Z. K., et al. Drivers of transcriptional variance in human intestinal epithelial organoids. Physiol Genomics. 53 (11), 486-508 (2021).
- Izumo, M., Sato, T. R., Straume, M., Johnson, C. H. Quantitative analyses of circadian gene expression in mammalian cell cultures. PLoS Comput Biol. 2 (10), e136 (2006).
- Hara-Miyauchi, C., et al. Bioluminescent system for dynamic imaging of cell and animal behavior. Biochem Biophys Res Commun. 419 (2), 188-193 (2012).
- Tung, J. K., Berglund, K., Gutekunst, C. A., Hochgeschwender, U., Gross, R. E. Bioluminescence imaging in live cells and animals. Neurophotonics. 3 (2), 025001 (2016).
- Xu, P., Elizalde, M., Masclee, A., Pierik, M., Jonkers, D. Corticosteroid enhances epithelial barrier function in intestinal organoids derived from patients with Chron’s disease. J Mol Med (Berl). 99 (6), 805-815 (2021).
- AlMusawi, S., Ahmed, M., Nateri, A. S. Understanding cell-cell communication and signaling in the colorectal cancer microenvironment. Clin Transll Med. 11 (2), 308 (2021).

