Growing a Cystic Fibrosis-Relevant Polymicrobial Biofilm to Probe Community Phenotypes
Summary
This protocol describes a cystic fibrosis (CF) lung-relevant four-species polymicrobial biofilm model that can be used to explore the impact of bacterial interspecies interactions.
Abstract
Most in vitro models lack the capacity to fully probe bacterial phenotypes emerging from the complex interactions observed in real-life environments. This is particularly true in the context of hard-to-treat, chronic, and polymicrobial biofilm-based infections detected in the airways of individuals living with cystic fibrosis (CF), a multiorgan genetic disease. While multiple microbiome studies have defined the microbial compositions detected in the airway of people with CF (pwCF), no in vitro models thus far have fully integrated critical CF-relevant lung features. Therefore, a significant knowledge gap exists in the capacity to investigate the mechanisms driving the pathogenesis of mixed species CF lung infections. Here, we describe a recently developed four-species microbial community model, including Pseudomonas aeruginosa, Staphylococcus aureus, Streptococcus sanguinis, and Prevotella melaninogenica grown in CF-like conditions. Through the utilization of this system, clinically relevant phenotypes such as antimicrobial recalcitrance of several pathogens were observed and explored at the molecular level. The usefulness of this in vitro model resides in its standardized workflow that can facilitate the study of interspecies interactions in the context of chronic CF lung infections.
Introduction
Strategies aimed at eradicating disease-causing microbes such as the ones detected in the cystic fibrosis (CF) airway, a multiorgan genetic disease, often fail1. That is, the presence of resilient biofilm-like microbial communities growing in the mucus-rich CF lung environment can cause chronic infections spanning multiple decades2. Furthermore, although the utilization of front-line antimicrobials (Abx) and modulators has resulted in improved outcomes in people with CF (pwCF), the "one bug, one infection" clinical approach typically employed against canonical CF pathogens such as Pseudomonas aeruginosa and Staphylococcus aureus has been ineffective in resolving hard-to-treat infections detected in the lungs of these individuals3,4,5,6,7.
Over the last two decades, multiple studies have changed our understanding of chronic CF lung disease8. That is, reports indicate that infections detected in the airways of pwCF are not solely caused by a single pathogen but are rather polymicrobial in nature8. Furthermore, although the pathogenesis of mixed species CF lung infections is still poorly understood, clinical evidence shows that pwCF that are co-infected with both S. aureus and P. aeruginosa in their airways have worsened lung function than individuals colonized by either of these two pathogens9.
Interspecies interactions among CF pathogens are hypothesized to be a reason why polymicrobial biofilm-based infections are not readily eradicated through the utilization of CF therapeutics, ultimately impacting patient outcomes3. Supporting this, in vitro studies have demonstrated that interactions between microbes such as P. aeruginosa, S. aureus, and others can impact clinically relevant phenotypes such as Abx responsiveness to front-line CF drugs, including vancomycin and tobramycin6,10,11. Therefore, revisiting the current treatment strategies aimed at eradicating CF pathogens in the context of mixed-species infections remains to be achieved.
The gold standard in the management of microbial-based CF lung infections heavily relies on the utilization of antimicrobial susceptibility testing (AST) to guide clinical interventions12. However, AST is typically done using bacterial monocultures grown in rich and well-mixed cultures, which is not reflective of the microbial growth conditions detected in the CF airway3,13. Given the significant disconnect between patient outcomes and treatment success, clinical reports now advocate for the development of novel approaches integrating critical features of the CF lung12,14.
Few studies have developed CF-relevant bacterial multispecies in vitro systems containing more than two pathogens15,16. However, these models (1) do not entirely reflect the polymicrobial and biofilm-like nature of the CF lung and (2) do not use nutritional and environmental conditions approximating the ones detected in the CF airway15,16. Through the mining of large CF-lung derived 16S rRNA gene data sets and computation, Jean-Pierre and colleagues recently developed an in vitro co-culture model integrating the above-mentioned features10. This system includes P. aeruginosa, S. aureus, Streptococcus spp. and Prevotella spp. grown in CF airway-like conditions and stable for up to 14 days. The authors also reported community-specific growth of Prevotella spp. and several changes in the Abx responsiveness of these CF pathogens through mechanisms that remain to be identified10. This tractable in vitro system also offered the possibility to delve into mechanistically focused questions related to Abx recalcitrance of a common variant of P. aeruginosa frequently detected in the airways of pwCF10. Therefore, the goal of this protocol is to provide the CF research community with a standardized experimental workflow to grow an in vitro biofilm that has the potential to be further expanded to tackle a number of CF-relevant questions.
Protocol
The details of all the reagents and equipment are listed in the Table of Materials.
1. Preparing artificial sputum medium
NOTE: Throughout this protocol, artificial sputum medium (ASM) is defined as the CF-relevant medium modified from SCFM2, previously published by Turner and colleagues17. Below is a description of the steps to prepare this medium. See Table of Materials to make stock solutions and storage conditions. This version of ASM differs from Turner et al.,17 as the buffering capacity in the original version does not allow for a stable pH over time. That is, fermentation activity carried out by Streptococcus spp. and anaerobes like Prevotella spp. results in the acidification of the growth medium (unpublished observations). Therefore, the 3-morpholinopropane-1-sulfonic acid (MOPS) concentration has been adjusted from 10 mM to 100 mM.
NOTE: Preparing a 2x ASM base (ASMb) stock solution facilitates the addition of components such as mucin, agar, water, etc., without impacting the final concentration of molecules present in ASM10,18. The 2x ASMb can be prepared in advance and stored at 4 °C for up to two weeks in a dark place. If the medium turns yellow, discard.
- To prepare 2x ASM base stock, in a clean beaker containing 400 mL of diH2O, add the following while stirring:
- Add 6.50 mL of a 0.2 M NaH2PO4.H2O stock; 6.25 mL of a 0.2 M Na2HPO4 stock; 348 µL of a 1 M KNO3 stock; 1.084 mL of a 0.25 M K2SO4 stock.
- Add 4.0 g of Yeast synthetic dropout without Tryptophan; 3.032 g of NaCl; 20.92 g of MOPS; 1.116 g of KCl and 0.124 g of NH4Cl.
- Add 9.30 mL of a 1 M L-Lactic acid stock; 2.70 mL of a 1.1 M glucose stock; 1.75 mL of a 1 M CaCl2.2H2O stock; 1.20 mL of a 0.25 M N-acetylglucosamine stock and 1.00 mL of a 3.6 mM FeSO4.7H2O stock.
- Add 660 µL of a 0.1 M Tryptophan stock; 606 µL of a 1 M MgCl2.6H2O stock; 0.60 g of deoxyribonucleic acid from herring sperm; 400 µL of a 250 mg/mL 1,2-dioleoyl-sn-glycero-3-phosphocholine (DOPC) stock.
- Once all the components have been added, adjust the pH to 6.80 with a 5 M NaOH stock. Complete to 500 mL with diH2O. Sterilize using a 0.22 µm filter, and store at 4 °C.
- To prepare a 10 mg/mL (2x) mucin stock, use a clean glass bottle containing 250 mL of diH2O and while stirring:
- Add 5.0 g of mucin from porcine stomach – Type II. Mix the suspension for 10 min. Complete to 500 mL with diH2O.
- Sterilize by autoclaving at 121 °C for 15 min. Keep at 4 °C until utilization.
- Reconstitute the ASM.
NOTE: This step needs to be performed on the day of the experiment.- Mix a 1:1 volume ratio of the 2x ASMb stock and the 10 mg/mL mucin stock.
- Vortex thoroughly before use.
2. Preparing polymicrobial community selective media
NOTE: Below are the quantities necessary to prepare a volume of 1 L of media.
- To prepare Mannitol Salt Agar (MSA) and Pseudomonas Isolation Agar (PIA), follow the manufacturer's instructions.
- To prepare Prevotella Selective Agar (PSA), in a clean glass bottle, add the following:
- 30.0 g of tryptic soy broth (TSB). 15.0 g of agar. 5.00 g of Yeast extract. 0.50 g of L-cysteine hydrochloride.
- 10.0 mL of a 0.5 mg/mL hemin stock. 100 µL of a 10 mg/mL menadione stock.
- Add 940 mL of diH2O and stir. Sterilize by autoclaving at 121 °C for 15 min.
- When cooled down (i.e., the bottle can be held barehanded), add while stirring: 50.0 mL of defribrinated sheep's blood. 2.00 mL of a 50 mg/mL kanamycin stock. 150 µL of a 50 mg/mL vancomycin stock. 500 µL of a 10 mg/mL polymixin B stock.
- To prepare Streptococcus Selective Agar (SSA), in a clean glass bottle, add the following:
- 30.0 g of TSB. 15.0 g of agar.
- Add 950 mL of diH2O and stir. Sterilize by autoclaving at 121 °C for 15 min.
- When cooled down (i.e., the bottle can be held barehanded), add while stirring: 50.0 mL of defribrinated sheep's blood. 1.00 mL of a 10 mg/mL polymixin B stock. 1.00 mL of a 10 mg/mL oxolinic acid stock.
NOTE: Pour in plates and keep at 4 °C for a maximum of 1 month. Before using, let the plates dry at room temperature up to 24 h in advance before they are inoculated. PSA has been validated with P. melaninogenica and Prevotella intermedia. SSA has been validated with S. sanguinis, and the Streptococcus milleri group (Streptococcus constellatus, Streptococcus intermedius, and Streptococcus anginosus, denoted as SMG). It is strongly recommended to de novo validate these selective media if new strains are to be tested.
3. Bacterial liquid media preparation and growth conditions
- To routinely grow P. aeruginosa and S. aureus, use a sterile rich medium (e.g., lysogeny broth (LB) or TSB).
- Start liquid overnights by picking a single colony previously grown on an LB/TSB agar plate for 20-24 h at 37 °C.
- Grow the broth cultures at 37 °C overnight with shaking at 250 rpm for 16-18 h.
- To grow Streptococcus spp., use sterile Todd Hewitt Broth supplemented with 0.5 % yeast extract (THB-YE).
- Start liquid culture overnight from a single colony previously grown on a TSB agar plate supplemented with 5% defibrinated sheep's blood (blood agar plate) for 24 h at 37 °C + 5% CO2.
- Grow the THB-YE cultures anaerobically or at 37 °C + 5% CO2 overnight without shaking for 18-22 h.
- To grow Prevotella spp., prepare 1 L of Prevotella growth medium (PGM) by adding the following to a clean bottle:
- 30.0 g of TSB. 5.00 g of yeast extract. 0.50 g of L-cysteine hydrochloride. 10.0 mL of a 0.5 mg/mL hemin stock and 100 µL of a 10 mg/mL menadione stock.
- Add 990 mL of diH2O. Sterilize by autoclaving at 121 °C for 15 min. Wrap the bottle in aluminum foil to protect it from light.
- Grow Prevotella spp. colonies on a blood agar plate incubated anaerobically at 37 °C for 48 h.
- Inoculate a single colony in PGM and grow overnight without shaking at 37 °C anaerobically for 24 h.
NOTE: PGM can be kept at room temperature for up to two weeks. Performing sterility tests on each growth medium is strongly recommended before use. If not using an anaerobic chamber, it might be necessary to start liquid overnights of Prevotella spp. from a patch of colonies.
4. Co-culture experiment preparation
NOTE: Figure 1 depicts a summarized experimental workflow. Setting up the experiment can be done in oxic conditions if the preparation time takes less than 1 h. If it is expected to take longer than that, it is strongly recommended to perform the experiment using an anaerobic chamber (with 10% CO2, 10% H2, and 80% N2 mixed gas). Anaerobic jars can also be used to incubate the experimental samples.
- Preparation of the monocultures and co-cultures
NOTE: Perform all the centrifugation steps at 10,000 x g for 2 min at room temperature. For the co-cultures, mix the bacterial samples just before inoculating the 96-well plates. Typically, a volume of 1 mL for P. aeruginosa and S. aureus strains are collected from the overnight cultures. For Streptococcus spp. and Prevotella spp., the overnight volumes may vary between 2-4 mL. These volumes can be adjusted depending on the number of conditions to be tested.- Using the bacterial liquid from the overnight cultures, perform the following steps:
- For P. aeruginosa and S. aureus strains, wash twice using sterile phosphate-buffered saline (PBS).
- For Streptococcus spp. and Prevotella spp., wash once with sterile PBS.
NOTE: Use a pipette to carefully discard supernatants, as the pellets can easily be lost. - After the washes, resuspend each pellet in 1 mL of 1x water-diluted ASMb.
- Make a 1:10 dilution of each sample in sterile PBS and measure the OD600.
- Adjust all the cultures to an OD600 of 0.2 in ASM.
- Using the OD600 = 0.2 suspensions, dilute in ASM the bacterial samples to a final OD600 of 0.01 (for each microbe) for the monocultures and co-cultures.
- Vortex thoroughly for 5 s.
NOTE: For example, to prepare a total of 1 mL of P. aeruginosa monoculture ASM suspension at OD600 = 0.01, add 50 µL of P. aeruginosa at OD600 = 0.2 to a volume of 950 µL of ASM. For the co-culture, take 50 µL of each P. aeruginosa, S. aureus, Streptococcus spp. and Prevotella spp. suspensions at an OD600 = 0.2 and add to 800 µL ASM. - Using a pipette, add 100 µl of the monoculture and co-culture suspensions in three separate wells of a sterile plastic flat bottom 96-well plate.
- Incubate the plate (without shaking) for 24 h at 37 °C in anoxic conditions.
NOTE: Other oxygen tensions (e.g., microxia, normoxia) can also be used here. However, the model has been developed and validated with anoxic conditions. Confirm the starting inocula by serially diluting the mono and co-culture bacterial suspensions, and plating on PIA, MSA, SSA, and PSA is strongly recommended. The targeted starting concentration for each bacterial species is 1 × 107 CFU/mL (P. aeruginosa), 3.5 × 106 CFU/mL (S. aureus), 1.2 × 106 CFU/mL (Streptococcus spp.), and 4.6 × 106 CFU/mL (Prevotella spp.). Bacterial inocula can also be modified, as described by Jean-Pierre and colleagues10.
- Using the bacterial liquid from the overnight cultures, perform the following steps:
- Replace the media after biofilm formation.
- After 24 h of incubation, remove the unattached (planktonic) cells by aspiration using a multichannel pipette.
- Replenish the pre-formed biofilms with 100 µL of fresh ASM or the desired treatment (e.g., antibiotics, metabolites, etc.).
- Incubate the plate for an additional 24 h at 37 °C in anoxic conditions.
NOTE: These steps can be done aerobically if performed quickly (i.e., less than 1 min). Otherwise, please use an anaerobic chamber.
5. Collecting and plating the samples
- After the additional 24 h of incubation, aspirate the non-attached (planktonic) cells with a multichannel pipette.
NOTE: The planktonic fraction can be kept, serially diluted, and plated on selective media or discarded. - Gently wash the biofilms twice using 125 µL of sterile PBS and discard the volumes.
- Add 50 µL of sterile PBS and detach the biofilms by gently scraping the cells from the plate using a 96-pin replicator.
- Transfer the resuspended biofilm cells to row A of a new sterile 96-well plate.
NOTE: The 96-well plate will contain sterile PBS in rows B to H. - Perform a 10x serial dilution of the planktonic (if needed) and biofilm fractions.
- Plate 3-5 µL of each dilution sample onto PIA, MSA, SSA, and PSA.
- Allow the inoculation spots to dry.
6. Incubating the selective plates and colony forming unit (CFU) counting
- For P. aeruginosa and S. aureus, incubate at 37 °C for 18-20 h.
- For Streptococcus spp, incubate at 37 °C in anoxic conditions for 24 h.
- For Prevotella spp, incubate at 37 °C in anoxic conditions for 48 h.
- After incubation, count the colonies (~10-35 per spot) and calculate the CFU per mL.
- Plot the data using a visualization tool.
Representative Results
As represented in Figure 2, several phenotypes were reported including (1) a reduction in the number of P. aeruginosa and S. aureus viable cell counts when grown a mixed planktonic communities compared to monoculture, (2) an increase in polymicrobial growth of S. sanguinis cells and, (3) mixed community growth of P. melaninogenica as previously reported by Jean-Pierre and colleagues10.
Furthermore, as depicted in Figure 1 the in vitro system can also be used to test molecules of interest (e.g., Abx, metabolites, etc.) once the biofilm communities have been established. That is, microbes grown in co-culture have either shown (1) an increase in susceptibility (P. aeruginosa–Figure 3A), (2) a decreased in sensitivity (S. aureus and S. sanguinis–Figure 3B,C) or (3) no changes (P. melaninogenica–Figure 3D) in the presence of the CF drug tobramycin versus monoculture.
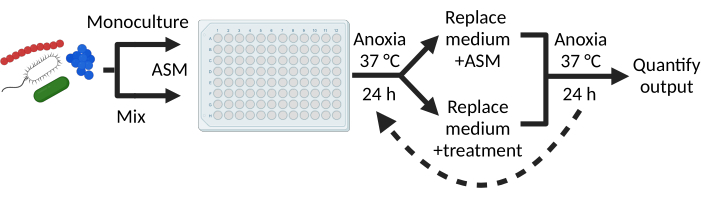
Figure 1: Experimental workflow. Growing monospecies and polymicrobial biofilms is done using a flat-bottom plastic 96-well plate into which microbes are cultivated in ASM for 24 h at 37 °C in anoxic conditions. After the initial incubation step, non-attached cells are aspirated, and pre-formed biofilms are either replenished with fresh ASM or treated with molecules of interest. This step can be repeated on many iterations depending on the hypothesis to be tested. At the end of the experiment, cells are collected, serially diluted, and plated on selective media for viable counts. Please click here to view a larger version of this figure.
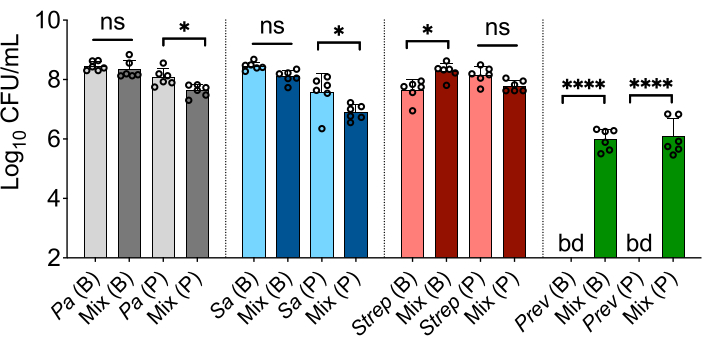
Figure 2: Detecting polymicrobial-specific phenotypes using in vitro modeling. Indicated are CFU/mL counts of P. aeruginosa (Pa), S. aureus (Sa), S. sanguinis (Strep) and, P. melaninogenica (Prev) bacterial species grown as a monoculture or in a mixed community (Mix) for biofilm (B) and planktonic (P) fractions. The presented data indicate the average from at least three technical replicates performed at least on six different days (n = 6) for each bar column. Statistical analysis was performed using ordinary one-way analysis of variance (ANOVA) and Tukey's multiple comparisons posttest with *P < 0.05; ****P < 0.0001, ns = non-significant. Error bars represent the standard deviation. bd = below detection. Limit of detection = 102 CFU/mL. The figure is modified from Jean-Pierre et al.10. Please click here to view a larger version of this figure.
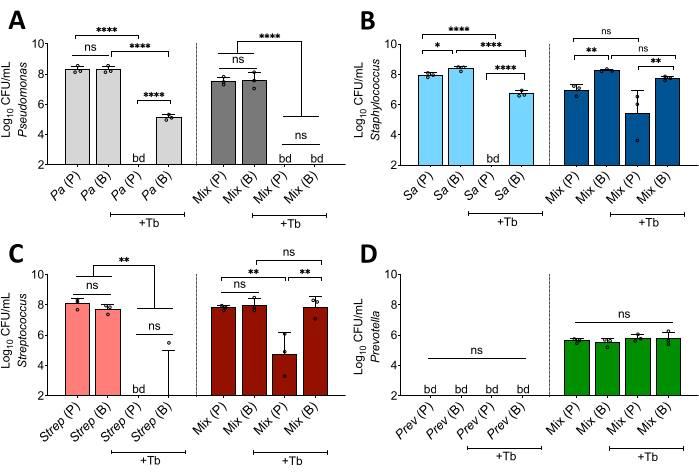
Figure 3: Interspecies interactions impact Abx susceptibility of CF pathogens. Indicated are CFU/mL counts of (A) P. aeruginosa (Pa), (B) S. aureus (Sa), (C) S. sanguinis (Strep),and (D) P. melaninogenica (Prev) bacterial populations grown as a monoculture or in a polymicrobial (Mix) in the absence or presence of tobramycin (+Tb) at a concentration of 100 µg/mL. The presented data indicate the average from at least three technical replicates performed at least on three different days (n = 3) for each bar column. Statistical analysis was performed using ordinary one-way analysis of variance (ANOVA) and Tukey's multiple comparisons posttest with *P < 0.05; **P < 0.01, and ****P < 0.0001. ns = non-significant. Error bars represent the standard deviation. bd = below detection. Limit of detection = 102 CFU/mL. The figure is modified from Jean-Pierre et al.10. Please click here to view a larger version of this figure.
Discussion
The usefulness of this clinically informed in vitro co-culture system resides in its capacity to detect polymicrobial-specific bacterial functions. That is, through the utilization of this model we have reported microbial phenotypes ranging from community-specific growth of Prevotella spp. (Figure 2) to changes in susceptibility to front-line CF Abx of P. aeruginosa, S. aureus and S. sanguinis (Figure 3). Moreover, even though data presented in Figure 2 are representative of laboratory strains, similar results have been obtained using various species/strains of each microbe including methicillin resistant S. aureus (MRSA) strains, the SMG, P. intermedia and clinical isolates of P. aeruginosa10.
Through the utilization of this co-culture system, we have previously reported that several LasR loss-of-function strains (both laboratory and clinical isolates) perform better in the presence of tobramycin when grown in a polymicrobial environment compared to monoculture10. Such observations are of particular relevance given that the presence of lasR variants in the CF lung correlates with poor outcomes in pwCF19. Furthermore, it was reported that MRSA and CF-relevant pathogenic streptococci (SMG) strains all show decreased susceptibility to tobramycin when grown in co-culture compared to their pure culture counterpart10. These in vitro results are particularly interesting given that studies looking at the post-treatment impact of tobramycin on the CF lung microbiome reveal little to no changes in the abundances of S. aureus and Streptococcus spp20,21. Taken together, this co-culture system might provide us with the opportunity to in vitro model clinical observations and to explore the mechanisms whereby CF pathogens are recalcitrant to drug therapy.
We argue that this co-culture system can be leveraged to test a large spectrum of CF-related questions ranging from virulence to microbe-microbe/host-microbe interactions. For example, Kesthely and colleagues have recently looked at changes in transcriptional responses of microbes grown in co-culture versus monocultures using the polymicrobial model described here22. Furthermore, although the development of novel modulators has resulted in improved outcomes in pwCF, chronic microbial-based infections in their airways persist4,5,7. Therefore, we propose that this in vitro community model represents an ideal platform to explore the mechanisms allowing these communities to thrive in the CF lung in the presence of these modulators.
The common limiting steps/technical hurdles encountered in using this model generally are (1) the rigourous and reproducible preparation of the monocultures and co-cultures mixes and, (2) well-to-well microbial contamination between adjacent samples containing different microbial species of the community. For the former point, restricting the number of experimental conditions (i.e., identifying the minimal number of conditions necessary to test a hypothesis while including all appropriate controls) is a strategy. Also, preparing the experiment using an anaerobic chamber can mitigate some of the issues (oxygen-mediated killing of Prevotella) that can arise while setting up the experiment in oxic conditions on the bench for too long. For the latter point, it is recommended to including several controls as checkpoints. Also, cross-contamination between well samples due to the presence volatile molecules/metabolites (either microbially produced or exogenously added) is also a potential risk to be considered. As such, shuffling the position of the samples between experimental biological replicates is a strategy.
A few limitation of this in vitro co-culture system include (1) the absence of host-derived factors (e.g., immune cells, host-derived metabolites, etc.) that are known to be important in the context of CF lung disease23, (2) the lack of spatial information on the polymicrobial biofilm architecture, (3) the lack of information on suspended biofilms and, (4) the absence of interkingdom interactions24,25,26. However, this polymicrobial model represents a good starting point to tackle these outstanding questions and to understand how such features might impact community-specific microbial phenotypes. Furthermore, the system described here has been designed to facilitate the creation of high throughput experiments (i.e., 96-well format). Finally, we acknowledge the work of previous groups that have built complex bacterial multispecies in vitro models which all contributed to the creation of new CF-relevant knowledge15,16.
Disclosures
The authors have nothing to disclose.
Acknowledgements
We would like to acknowledge Dr. George A. O'Toole and Dr. Thomas H. Hampton for their significant role in the design and development of the in vitro polymicrobial community model. We thank Dr. Sophie Robitaille for her helpful comments on the manuscript. This work was supported by a Cystic Fibrosis Foundation grant JEAN21F0 to F.J-P. Figure 1 of the manuscript was created using BioRender.
Materials
| 1,2-dioleoyl-sn-glycero-3-phosphocholine (DOPC) | Fisher Scientific | NC0387928 | Prepare a 250 mg/mL stock in chloroform directly in the bottle. Keep at -20°C. Final concentration 200 μg/mL in 2x ASM base. |
| 3-morpholinopropane-1-sulfonic acid (MOPS) | Sigma | M1254 | Final concentration 200 mM in 2X ASM base. |
| 96-pin replicator | Fisher Scientific | 05-450-9 | Disposable replicators can also be used. Cat #NC1567338. |
| 96-well sterile standard plate with lid | Fisher Scientific | 62406-081 | |
| Agar | Fisher Scientific | DF0145-17-0 | |
| AnaeroPack 2.5 L Rectangular Jar | Fisher Scientific | 23-246-385 | |
| CaCl2*2H2O | Fisher Scientific | C79-500 | Prepare a 1 M stock in water and filter sterilize with a 0.22 μm filter. Final concentration 3.5 mM in 2x ASM base. |
| Defribrinated sheep's blood | To prepare growth plates for Streptococcus and Prevotella. | ||
| Deoxyribonucleic acid from herring sperm | Sigma | D7290 | Final concentration 1.2 mg/mL in 2x ASM base. |
| FeSO4*7H2O | Fisher Scientific | AA1449830 | Prepare a 1 mg/mL stock (3.6 mM) in water and filter sterilize with a 0.22 μm filter. Keep wrapped in aluminum foil. Final concentration 0.0072 mM in 2x ASM base. |
| GasPaks | Fisher Scientific | B260678 | |
| Glucose | Fisher Scientific | D16-3 | Prepare as a 20% stock (1.1 M) in water and sterilize by autoclave. Final concentration 6 mM in 2x ASM base. |
| Hemin | Sigma | 51280 | Dissolve 100 mg of hemin in 2 mL of 1 M NaOH and then bring up volume to 200 mL with water. Store in an amber bottle or a regular bottle wrapped with aluminum foil at 4 °C. |
| K2SO4 | Fisher Scientific | P304-500 | Prepare a 0.25 M stock in water and filter sterilize with a 0.22 μm filter. Final concentration 0.54 mM in 2x ASM base. |
| Kanamycin | Prepare a 50 mg/mL stock solution in water. Filter sterilize with a 0.22 μm filter and store at 4 °C. | ||
| KCl | Fisher Scientific | P217-500 | Final concentration 30.0 mM in 2x ASM base. |
| KNO3 | Fisher Scientific | P263-500 | Prepare a 1 M stock in water and filter sterilize with a 0.22 μm filter. Final concentration 0.70 mM in 2x ASM base. |
| L-cysteine hydrochloride | Fisher Scientific | AAA1038922 | |
| L-Lactic acid | Sigma | L1750 | Prepare a 1 M stock in water and filter sterilize with a 0.22 μm filter. pH adjust to 7.0. Final concentration 18.6 mM in 2x ASM base. |
| L-Tryptophan | Sigma | T0254 | Prepare a 0.1 M stock in 0.2 M NaOH and filter sterilize with a 0.22 μm filter. Keep refrigerated at 4 °C wrapped in aluminum foil. Final concentration 0.132 mM in 2x ASM base. |
| Mannitol Salt Agar | Fisher Scientific | B11407 | Prepare using manufacturer's recommendations. |
| Menadione | Sigma | M5625 | Prepare a 10 mg/mL stock solution dissolved in 96-100% ethanol. Conserve at 4 °C wrapped in aluminium foil. Vortex before use. |
| MgCl2*6H2O | Fisher Scientific | AA12288A9 | Prepare a 1 M stock in water and filter sterilize with a 0.22 μm filter. Final concentration 1.21 mM in 2x ASM base. |
| Mucin from porcine stomach – Type II | Sigma | M2378 | Prepare a 10 mg/mL (2x) stock in water; mix thoroughly before autoclaving. Keep at 4 °C. |
| Na2HPO4 | Fisher Scientific | S375-500 | Prepare a 0.2 M stock in water and filter sterilize with a 0.22 μm filter. Final concentration 2.5 mM in 2x ASM base. |
| N-acetylglucosamine | Sigma | A8625 | Prepare a 0.25 M stock in water and filter sterilize with a 0.22 μm filter. Keep stock refrigerated at 4 °C. Final concentration 0.6 mM in 2x ASM base. |
| NaCl | Fisher Scientific | S271-500 | Final concentration 103.7 mM in 2x ASM base. |
| NaH2PO4*H2O | Fisher Scientific | S369-500 | Prepare a 0.2 M stock in water and filter sterilize with a 0.22 μm filter. Final concentration 2.6 mM in 2x ASM base. |
| NaOH | Fisher Scientific | S318-500 | Prepare a 5 M stock solution. Use to adjust pH of 2x ASM base stock. |
| NH4Cl | Fisher Scientific | A661-500 | Final concentration 4.6 mM in 2x ASM base. |
| Oxolinic acid | Prepare a 10 mg/mL stock solution 0.5 M NaOH. Filter sterilize with a 0.22 μm filter and store at 4 °C. | ||
| Polymixin B | Prepare a 10 mg/mL stock solution in water. Filter sterilize with a 0.22 μm filter and store at 4 °C. | ||
| Pseudomonas Isolation Agar | Fisher Scientific | DF0927-17-1 | Prepare using manufacturer's recommendations. |
| Todd Hewitt Broth | Fisher Scientific | DF0492-17-6 | Prepare using manufacturer's recommendations. |
| Tryptic Soy Broth | Fisher Scientific | DF0370-17-3 | Prepare using manufacturer's recommendations. |
| Vancomycin | Prepare a 50 mg/mL stock solution in water. Filter sterilize with a 0.22 μm filter and store at 4 °C. | ||
| Yeast Extract | Fisher Scientific | B11929 | |
| Yeast Synthetic Dropout without Tryptophan | Sigma | Y1876 | Final concentration 8.0 mg/mL in 2x ASM base. |
References
- Cogen, J. D., Nichols, D. P., Goss, C. H., Somayaji, R. Drugs, drugs, drugs: Current treatment paradigms in cystic fibrosis airway infections. J Pediatric Infect Dis Soc. 11 (2), S32-S39 (2022).
- O’Sullivan, B. P., Freedman, S. D. Cystic fibrosis. Lancet. 373 (9678), 1891-1904 (2009).
- Jean-Pierre, F., Vyas, A., Hampton, T. H., Henson, M. A., O’Toole, G. A. One versus many: Polymicrobial communities and the cystic fibrosis airway. mBio. 12 (2), e00006-e00021 (2021).
- Martin, C., et al. Longitudinal microbial and molecular dynamics in the cystic fibrosis lung after elexacaftor-tezacaftor-ivacaftor therapy. Respir Res. 24 (1), 317 (2023).
- Sosinski, L. M., et al. A restructuring of microbiome niche space is associated with elexacaftor-tezacaftor-ivacaftor therapy in the cystic fibrosis lung. J Cystic Fibros. 21 (6), 996 (2022).
- Orazi, G., O’Toole, G. A. 34;It takes a village": Mechanisms underlying antimicrobial recalcitrance of polymicrobial biofilms. J Bacteriol. 202 (1), e00530-e00619 (2019).
- Nichols, D. P., et al. Clinical effectiveness of elexacaftor/tezacaftor/ivacaftor in people with cystic fibrosis: A clinical trial. Am J Respir Crit Care Med. 205 (5), 529-539 (2022).
- Filkins, L. M., O’Toole, G. A. Cystic fibrosis lung infections: Polymicrobial, complex, and hard to treat. PLoS Pathog. 11 (12), e1005258 (2015).
- Limoli, D. H., et al. Staphylococcus aureus and Pseudomonas aeruginosa co-infection is associated with cystic fibrosis-related diabetes and poor clinical outcomes. Eur J Clin Microbiol Infect Dis. 35 (6), 947-953 (2016).
- Jean-Pierre, F., et al. Community composition shapes microbial-specific phenotypes in a cystic fibrosis polymicrobial model system. Elife. 12 (12), e81604 (2023).
- Toole Orazi, G., O’Toole, G. A. Pseudomonas aeruginosa alters Staphylococcus aureus sensitivity to vancomycin in a biofilm model of cystic fibrosis infection. mBio. 8 (4), e00873-e00917 (2017).
- Lipuma, J. J. The sense and nonsense of antimicrobial susceptibility testing in cystic fibrosis. J Pediatric Infect Dis Soc. 11 (2), S46-S52 (2022).
- Coenye, T. Biofilm antimicrobial susceptibility testing: Where are we and where could we be going. Clin Microbiol Rev. 36 (4), e0002423 (2023).
- Waters, V. J., et al. Reconciling antimicrobial susceptibility testing and clinical response in antimicrobial treatment of chronic cystic fibrosis lung infections. Clin Infect Dis. 69 (10), 1812-1816 (2019).
- Vandeplassche, E., et al. Antibiotic susceptibility of cystic fibrosis lung microbiome members in a multispecies biofilm. Biofilm. 2, 100031 (2020).
- Varga, J. J., et al. Antibiotics drive expansion of rare pathogens in a chronic infection microbiome model. mSphere. 7 (5), e0031822 (2022).
- Turner, K. H., Wessel, A. K., Palmer, G. C., Murray, J. L., Whiteley, M. Essential genome of Pseudomonas aeruginosa in cystic fibrosis sputum. Proc Natl Acad Sci U S A. 112 (13), 4110-4115 (2015).
- Clay, M. E., et al. Pseudomonas aeruginosa lasR mutant fitness in microoxia is supported by an anr-regulated oxygen-binding hemerythrin. Proc Natl Acad Sci U S A. 117 (6), 3167-3173 (2020).
- Hoffman, L. R., et al. Pseudomonas aeruginosa lasR mutants are associated with cystic fibrosis lung disease progression. J Cyst Fibros. 8 (1), 66-70 (2009).
- Heirali, A., et al. Sputum microbiota in adults with CF associates with response to inhaled tobramycin. Thorax. 75 (12), 1058-1064 (2020).
- Nelson, M. T., et al. Maintenance tobramycin primarily affects untargeted bacteria in the cf sputum microbiome. Thorax. 75 (9), 780-790 (2020).
- Kesthely, C. A., Rogers, R. R., El Hafi, B., Jean-Pierre, F., O’Toole, G. A. Transcriptional profiling and genetic analysis of a cystic fibrosis airway-relevant model shows asymmetric responses to growth in a polymicrobial community. Microbiol Spectr. 11 (5), e0220123 (2023).
- Jean-Pierre, F., Henson, M. A., O’Toole, G. A. Metabolic modeling to interrogate microbial disease: A tale for experimentalists. Front Mol Biosci. 8, 634479 (2021).
- Hendricks, M. R., et al. Respiratory syncytial virus infection enhances Pseudomonas aeruginosa biofilm growth through dysregulation of nutritional immunity. Proc Natl Acad Sci U S A. 113 (6), 1642-1647 (2016).
- Hogan, D. A., Vik, A., Kolter, R. A Pseudomonas aeruginosa quorum-sensing molecule influences Candida albicans morphology. Mol Microbiol. 54 (5), 1212-1223 (2004).
- Wu, Y., et al. Co-assembling living material as an in vitro lung epithelial infection model. Matter. 7 (1), 216-236 (2024).

.
