An Automated Culture System for Use in Preclinical Testing of Host-Directed Therapies for Tuberculosis
Summary
Rapid and efficient quantification of intracellular M. tuberculosis growth is crucial for pursuing improved therapies against tuberculosis (TB). This protocol describes a broth-based colorimetric detection assay using an automated liquid culture system to quantify Mtb growth in macrophages treated with candidate host-directed therapies.
Abstract
Mycobacterium tuberculosis (Mtb), the causative agent of tuberculosis (TB), was the most significant infectious disease killer globally until the advent of COVID-19. Mtb has evolved to persist in its intracellular environment, evade host defenses, and has developed resistance to many anti-tubercular drugs. One approach to solving resistance is identifying existing approved drugs that will boost the host immune response to Mtb. These drugs could then be repurposed as adjunctive host-directed therapies (HDT) to shorten treatment time and help overcome antibiotic resistance.
Quantification of intracellular Mtb growth in macrophages is a crucial aspect of assessing potential HDT. The gold standard for measuring Mtb growth is counting colony-forming units (CFU) on agar plates. This is a slow, labor-intensive assay that does not lend itself to rapid screening of drugs. In this protocol, an automated, broth-based culture system, which is more commonly used to detect Mtb in clinical specimens, has been adapted for preclinical screening of host-directed therapies. The capacity of the liquid culture assay system to investigate intracellular Mtb growth in macrophages treated with HDT was evaluated. The HDTs tested for their ability to inhibit Mtb growth were all-trans Retinoic acid (AtRA), both in solution and encapsulated in poly(lactic-co-glycolic acid) (PLGA) microparticles and the combination of interferon-gamma and linezolid. The advantages of this automated liquid culture-based technique over the CFU method include simplicity of setup, less labor-intensive preparation, and faster time to results (5-12 days compared to 21 days or more for agar plates).
Introduction
Mycobacterium tuberculosis (Mtb), the causative agent of TB, was the most significant infectious disease killer globally in 20191. To evade host defenses, Mtb subverts the mycobactericidal activity of innate immune cells such as macrophages and dendritic cells (DCs), allowing it to persist intracellularly and replicate2. The lack of an effective vaccine to prevent adult pulmonary TB and the increasing emergence of drug-resistant strains highlight the urgent need for new therapies.
Adjunctive host-directed therapies (HDT) could shorten treatment time and help overcome resistance3. Preclinical assessment of HDT candidates in vitro to determine mycobactericidal activity within macrophages often relies on the quantification of Mtb growth by colony-forming units (CFU) on solid agar plates. This is a slow, labor-intensive assay that does not lend itself to rapid screening of drugs. Commercially available automated, broth-based microbial detection systems are more commonly used in clinical microbiology laboratories for detection and drug susceptibility testing of Mtb and other mycobacterial species in clinical specimens4. These instruments measure growth indirectly based on the bacterial metabolic activity leading to physical changes in the culture media (change in CO2 or O2 levels or pressure) monitored over time5. The readout is time to positivity (TPP), which has previously been shown to correlate with Mtb CFU in sputum specimens of TB patients in response to treatment6,7 and in lysates of infected murine lung and spleen8. In addition, liquid culture detection systems have been used to measure the effect of conventional pathogen-directed therapies on the growth of mycobacteria in axenic culture and cultured macrophages9,10. The instrument has also been used to investigate the innate ability of dendritic cells and of alveolar macrophages to control intracellular growth of Mtb11,12. This experimental protocol demonstrates that a liquid culture diagnostic system can be adapted to perform preclinical screening of HDT for TB in cultured macrophages. Compared to CFU enumeration, the main advantage of this technique is that it considerably reduces the experimental labor and time required to quantify intracellular mycobacterial growth/survival. This technique relies on access to an automated culture instrument that can be used to assess intracellular mycobacterial survival in immune cells treated with a broad range of pharmacological reagents targeting cellular functions to boost host immunity.
Protocol
The experiments outlined in this protocol were carried out using the attenuated H37Ra strain of Mtb, which can be handled in a Containment Level 2 laboratory. All manipulations of live mycobacteria were carried out in Class II biological safety cabinet (BSC). Experimental procedures were designed to minimize the generation of aerosols. Eukaryotic cell culture (THP-1 cells) was also carried out in a Class II BSC. The laboratory carried out a risk assessment and ensured that all procedures were carried out in line with institutional and national biological safety regulations. The human monocytic THP-1 cell line was used to perform the method as described (step 1). Cells are differentiated into macrophages following stimulation with phorbol 12-myristate 13-acetate (PMA) before infection with mycobacteria.
1. Cell culture
- Propagate H37Ra seed stock to log phase in Middlebrook 7H9 (MB) broth supplemented with albumin-dextrose-catalase (ADC) enrichment (10%) and 0.05% polysorbate 80. Store H37Ra stock in 1 mL aliquots in a -80 °C freezer for up to 1 year.
- Thaw a 1 mL vial of Mtb-H37Ra and transfer it to a T25 flask with a filter cap containing 9 mL of MB supplemented broth approximately 1 week before the planned experiment. Incubate at 37 °C for 5-7 days in a static incubator.
- Grow THP-1 cells in RPMI-1640 supplemented with non-heat killed 10% fetal calf serum (complete (c)RPMI) in a T75 flask in a CO2 humidified incubator at 5% CO2/37 °C and subculture twice per week to maintain a density of less than 1 x 106 cells/mL.
- Differentiate THP-1 cells into macrophages 3 days before infection by gently pipetting cells several times using a serological pipette in T75 flasks to disperse any clumps and place them in a 50 mL conical tube.
- Centrifuge cells at 300 x g for 10 min at room temperature, decant off the supernatant, and gently resuspend the pellet in 2 mL of RPMI. Perform cell count to estimate cells/mL.
- Seed 2 mL of THP-1 macrophages in 12-well tissue culture plates at a density of 100,000 cells/mL in cRPMI with 100 ng/mL PMA for 72 h. Remove PMA-containing medium from cells and replenish with fresh cRPMI before Mtb infection.
- Set up individual plates for each time point required.
- Seed cells at the same density (100,000 cells/mL) in 2-well glass chamber slides to determine the multiplicity of infection (MOI).
- Place in a 5% CO2 humidified incubator for 3 days at 37 °C.
2. Quantification of Mtb uptake
- Determination of Mtb uptake by macrophages (MOI)
- Set up the Class II biological safety cabinet (BSC) on the day of infection and work on two layers of tissue paper to catch any spills. Set up waste discard containers according to local regulations.
- Remove 6-8 mL of mycobacterial culture from the T25 flask and transfer it into a 15 mL polypropylene tube.
NOTE: Smaller volume tubes can be used for smaller experiments. - Centrifuge the tube in a benchtop centrifuge at room temperature for 10 min at 2890 x g.
- Carefully remove the tube from the centrifuge and transfer it to the biological safety cabinet. Wait 1 min to allow the bacteria to settle.
- Pour off the supernatant into the disinfectant discard container, recap tube, and resuspend the bacteria in the remaining medium by tapping the side of the tube. Wait 1 min to allow the bacteria to settle.
- Add 2 mL of pre-warmed cRPMI, mix gently, and transfer to a 50 mL conical tube.
- Resuspend the mycobacteria very carefully using a 25 G needle and 5 mL syringe. To resuspend, draw up the mycobacteria suspension into the syringe and eject very gently down the sidewall of the tube to minimize aerosol production. Repeat 6-8 times.
NOTE: Exercise utmost caution as this is a high-density culture of mycobacteria. To avoid the risk of needle stick injury, use blunt needles where possible, and Luer lock syringes. - Dispose of the needle and syringe in a sharps container in the BSC.
- Transfer the suspension to a 2 mL microfuge tube (with screw-on cap) and centrifuge at room temperature for 3 min at 100 x g to pellet any remaining clumps. Return the tube to the safety cabinet and wait 1 min to allow the bacteria to settle.
- Transfer the top 1-1.5 mL of the supernatant to a new tube. Discard the original tube in the waste bucket containing disinfectant. Mix well and add various amounts of the mycobacterial suspension (e.g., 5, 25, 50, 150 µL) to the 2-well glass chamber slides and incubate for 3 h in a CO2 incubator at 37 °C.
- Staining for acid fast bacteria (AFB)
NOTE: After 3 h incubation, the macrophages are washed and fixed with paraformaldehyde to inactivate mycobacteria. The slides are then stained using a Modified Auramine O kit (see Table of Materials) to estimate mycobacteria phagocytosed per cell. Due to their waxy cell wall, mycobacteria retain the Auramine dye after an acid-alcohol wash. The macrophage nuclei are then counter-stained with Hoechst. This method allows for the number of phagocytosed bacteria per cell to be counted and is used to determine the multiplicity of infection (MOI) of the macrophages.- Remove the medium from the glass chamber slide after pipetting up and down three times to dislodge bacteria that have not been phagocytosed.
- Wash once with 2 mL of room temperature PBS.
- Store stocks of 4% paraformaldehyde, dissolved in PBS in aliquots at -20 °C for up to 6 months. Thaw an aliquot of 4% paraformaldehyde immediately before use. Dilute to 2% paraformaldehyde with PBS and add 2 mL per well.
- Incubate for 10 min at room temperature. The glass chamber slide can be removed from the safety cabinet at this stage for staining.
- Wash the slide under a gentle stream of tap water.
- Dispense enough Auramine onto the slide to cover the cells using a plastic transfer pipette and incubate for 1 min at room temperature in the dark (cover with aluminum foil).
- Wash excess dye off the slide with tap water and add the decolorizer/quencher for 1 min in the dark.
- Wash off the excess with tap water and incubate for 15 min at room temperature with Hoechst 33342 (10 µg/mL in PBS) in the dark.
- Wash off the Hoechst stain with tap water, remove the chambers, drain excess water from the slide, add a drop of antifade and coverslip, and air dry.
- Examine the slide under the fluorescent microscope using the 100x oil objective. Mycobacteria will fluoresce green under the FITC filter. The nuclei fluoresce blue under the DAPI filter (Figure 1C).
- Determine the MOI by counting the number of mycobacteria phagocytosed per cell and the percentage of cells infected.
- Calculate the volume of mycobacterial suspension needed to achieve the required MOI based on the surface area of a well in the plate; for example, the surface area of the glass chamber slides used in this experiment is 4 cm2. Low MOI (approximately 1-2 bacilli/cell) is desirable for experiments conducted over several days (e.g., 5 days).
- Infection of macrophages
- Mix the mycobacteria suspension well and add the amount needed to the cells on 12-well plates once the volume required to achieve the desired MOI has been determined.
- Incubate at 37 °C for 3 h to allow mycobacteria to be phagocytosed.
- Remove extracellular bacteria by washing the wells with either warm RPMI or HBSS several times.
- Lyse the macrophages in one well (3 h sample) to determine the percentage time to positivity (TTP) of the initial inoculum (3 h sample) as outlined in step 3 below.
- Add fresh cRPMI and the required drug doses or vehicle control to the remaining wells, incubate the plates in the CO2 incubator at 37 °C for the time necessary (dependent on the experimental design but usually at several intervals between 1 to 8 days).
3. Harvesting samples for the liquid culture detection system
NOTE: On the day of infection, extracellular mycobacteria are removed by washing, and intracellular mycobacteria are harvested by lysis of one well of macrophages (3 h sample) to determine the initial amount phagocytosed as a baseline control for infection. At subsequent times both the medium, cell lysate, and washes are combined to measure total mycobacterial growth. Extracellular and intracellular growth can also be assessed separately if desired.
- Harvesting 3 h sample to determine TTP
- Wash off extracellular mycobacteria from all the wells after the initial 3 h of infection as outlined in step 2.3.3. Add 1 mL of fresh media to the 3 h control well to equalize the lysate volume with those of later time points.
NOTE: See step 3.2.7 if extracellular mycobacteria are to be excluded from the analysis.
- Wash off extracellular mycobacteria from all the wells after the initial 3 h of infection as outlined in step 2.3.3. Add 1 mL of fresh media to the 3 h control well to equalize the lysate volume with those of later time points.
- Sample collection
- Warm MB broth and instrument culture bottles to bring them to room temperature.
- Transfer the medium from the 12-well plate to the corresponding labeled conical tubes.
- Add 500 µL of sterile lysis buffer (0.1% Triton x-100 in PBS filtered through a sterile 0.2 µm filter) to each well for 10 min.
- Gently scrape the cells from the well with a sterile scraper and combine with the medium in the appropriate conical tube.
- Wash the well with 0.5 mL of sterile PBS and transfer to the appropriate tube.
- Gently pass each sample through a needle and syringe (25 G) 6-8 times to break up the clumps. Dilute samples 1:10 in MB broth; 100 µL sample + 900 µL MB medium.
- At the required times/days (usually between 1 to 8 days), harvest the remaining samples by following steps 3.2.1-3.2.6 above.
NOTE: Investigators may prefer to exclude extracellular mycobacteria from their analysis, in which case, in step 3.2.2 above, the medium from each well is discarded, and macrophages are washed several times before adding lysis buffer.
- Inoculating and loading instrument culture bottles
NOTE: Details of the liquid culture instrument and related consumables are provided in the Table of Materials.- Sterilize the rubber cap of the instrument culture bottle with tissue paper soaked in 70% alcohol and allow it to air dry.
NOTE: This step needs to be carried out in the BSC. - Prepare bottles by transferring enough Nutrient supplements for all the samples into a conical tube (0.5 mL/bottle). Use a needle and syringe to inject 0.5 mL of Nutrient supplement into the instrument culture bottle.
- Pipette 500 µL of the diluted sample (1:10) into a 1 mL V-bottomed tube.
- Use a needle and syringe to inject the 500 µL of sample into the assigned instrument culture bottle.
- Sterilize the rubber cap of the instrument culture bottle with tissue paper soaked in 70% alcohol and allow it to air dry. Wipe the bottles with tissue paper soaked in 70% alcohol before removal from the BSC.
- Carefully transport bottles from the biosafety cabinet to the instrument for loading.
- Press the loading button on the automated microbial detection system.
- Scan the barcodes on instrument culture bottles and place the bottles into the detection system incubator at 37 °C for up to 42 days. Read and record the time taken to reach positivity from the instrument screen.
NOTE: The barcode allows the instrument to identify the bottle and link reflectance readings with a particular bottle. - Calculate percentage time to positivity (TTP) by comparing the TTP of the initial intracellular inoculum (Day 0) to that of macrophages cultured for the indicated times. A positive change in percentage TTP means mycobacterial growth13.
For example, for day 3:
Percentage change in time to positivity =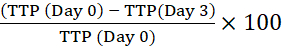
- Sterilize the rubber cap of the instrument culture bottle with tissue paper soaked in 70% alcohol and allow it to air dry.
Representative Results
The automated liquid culture instrument used in this study monitors CO2 levels every 10 min. A color change in the sensor at the bottom of the instrument bottle is measured colorimetrically and expressed as reflectance units. The instrument software then applies detection algorithms to calculate time to positivity (TTP), i.e., the number of days from inoculation until cultures are flagged as positive (Figure 1A). An inverse relationship between TTP and log10CFU (determined by the agar plate method)12 in the initial inoculum is illustrated in Figure 1B. When Mtb growth within macrophages-infected at MOI ranging from 1-2 to 5-10 in the presence or absence of pharmacological inhibitors (Figure 1C) was compared by the automated culture method described or by enumeration of colonies on solid agar, there was a significant correlation between results obtained by both methods (Figure 1D). To display Mtb growth graphically, the percentage change in TTP was calculated according to the equation above by comparing the TTP values for up to 8 days after infection of macrophages to the initial TTP value12. Results demonstrated a similar trend between liquid culture and CFU, showing significant inhibition of Mtb growth in macrophages in the presence of AtRA in solution or the equivalent dose of AtRA encapsulated in PLGA microparticles (MP) (Figure 2A,B).
The efficacy of another HDT, IFNγ, which has been used to treat multi-drug resistant TB14, in combination with the second-line antibiotic linezolid, was also investigated. A dose-response experiment was first carried out in infected THP-1 cells to determine the efficacy of linezolid alone at concentrations ranging from 0.6-5.0 µg/mL (Figure 2C) before the combination of linezolid at a suboptimal dose (1.25 µg/mL) and IFNγ was tested: there was no significant interaction between the drugs (Figure 2D).
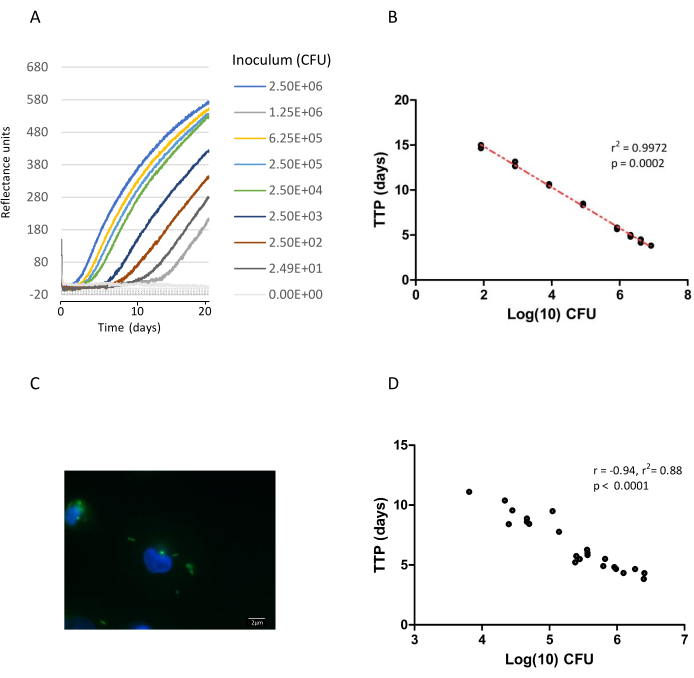
Figure 1: Quantification of Mtb growth by automated liquid culture and on solid medium. Mtb H37Ra was diluted in Middlebrook broth over a range of dilutions (1:2 to 1:100,000). A 300 µL aliquot of each dilution was injected into duplicate instrument culture bottles, incubated on the liquid culture instrument, and their growth was monitored. Simultaneously, an aliquot (10 µL) was spread on MB agar plates containing 0.5% glycerol and 10% OADC (Oleic acid, albumin, dextrose, catalase supplement) in triplicate, for CFU enumeration as previously published12. (A) Data points from the liquid culture system of each dilution are plotted as reflectance units versus time15. To allow for comparisons between samples, the background was normalized to the 9 h reflectance reading for each sample. Growth was monitored for 42 days in all (the first 21 days is shown on graph), time to positivity (TTP) ranged from 3.83 to 11.1 days. (B) TTP from diluted samples was plotted against log10 CFU estimation; each data point represents the values for two replicates from one experiment and describes two separate experiments. The line of best fit and regression coefficient (r2) are shown. (C) Example of AFB staining of intracellular Mtb H37Ra in THP-1 macrophages with auramine (green) which is used to calculate multiplicity of infection, nuclei are counterstained with Hoechst 333258 (blue). Images were generated using an Epifluorescent Microscope with a 100x (numerical aperture [NA], 1.3) oil objective. Scale bar represents 2 µm. (D) Correlation analysis (Pearson) of paired TTP (liquid culture) and CFU estimation of Mtb H37Ra growth in THP-1 macrophage lysates from multiple experiments (n = 23), treated with/without pharmacological reagents and infected at MOI ranging from 1-2 to 5-10 bacilli/infected macrophage. Please click here to view a larger version of this figure.
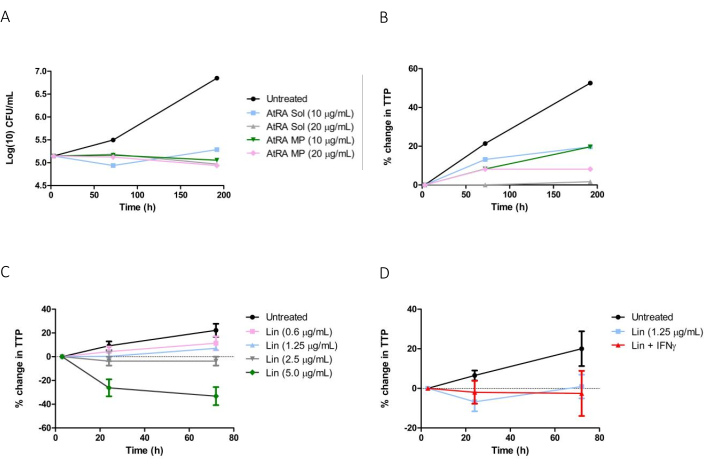
Figure 2: Evaluation of host-directed therapies for TB using an automated liquid culture system. THP-1 macrophages were infected with Mtb H37Ra for 3 h, extracellular bacteria were removed, and cells were treated with candidate host-directed therapies for up to 192 h. (A) CFU estimation of Mtb growth in macrophages treated with AtRA solution (Sol) or 2 µM AtRA microparticles (MP) (10-20 µg/mL). (B) % change in (time to positivity TTP) in macrophages treated with AtRA solution or AtRA PLGA MP. Infected THP1 cells cultured in 0.1% DMSO in cRPMI were designated as untreated controls. (C) Macrophages were treated with increasing concentrations of linezolid solution (0.6 µg/mL to 5 µg/mL), % change in TTP was determined 24 and 72 h post-infection using an automated liquid culture system. (D) Macrophages infected with Mtb H37Ra were treated with linezolid alone (1.25 µg/mL) or linezolid + IFNγ (5 ng/mL) up to 72 h, % change in TTP was calculated. Untreated controls consisted of infected THP1 cells cultured in cRPMI alone for the indicated times. Please click here to view a larger version of this figure.
Discussion
The authors have used the liquid culture method described in this protocol to monitor Mtb growth in monocyte-derived macrophages and alveolar macrophages and THP-1 cells differentiated with PMA11,16,17. This technique can also be modified for use with non-adherent cells12. More recently, the instrument was also used in preclinical studies to evaluate inhaled all-trans retinoic acid (AtRA) as an HDT for TB17. Critical steps in the protocol include (1) controlling for uptake of Mtb by AFB staining to mitigate factors such as differential phagocytosis due to drug pre-treatment and minimize cell death which is less likely to occur at lower MOI, (2) depending on the MOI, cell lysates may need to be diluted to maximize the dynamic range of the assay. Aiming for a TTP of approximately 5-12 days has yielded reproducible results.
In this protocol, attenuated Mtb H37Ra has been used as a surrogate for virulent Mtb, but the method can also be applied to study drug efficacy against virulent laboratory and clinical strains in a Containment Level 3 facility with appropriate modification. Confirmation of these results in a mouse model of virulent TB whereby inhaled AtRA, both in solution and encapsulated in poly(lactic-co-glycolic acid (PLGA) microparticles (MP), significantly improved clearance of Mtb-H37Rv from the lung while reducing pathology, indicates that H37Ra is a helpful surrogate for virulent Mtb in these studies17.
As HDTs are designed to be used as adjuncts to conventional therapeutics, it is essential to evaluate their efficacy in vitro in combination with anti-tubercular antibiotics to analyze potential drug interactions18. To obtain a measurable TTP, antibiotics-particularly those with high intracellular efficacy-must first be titrated to suboptimal concentrations (below the minimum inhibitory concentration) before evaluating HDT/antibiotic combinations in macrophages. In the example shown in this protocol (Figure 2C,D), linezolid alone was first titrated before being evaluated in combination with IFN-γ. In another example, lysates of H37Ra-infected THP-1 macrophages treated with rifampicin at or above 0.6 µg/mL do not reliably turn positive in the liquid culture system17. Therefore, a lower concentration of rifampicin is needed to evaluate any HDT combined with this antibiotic in macrophages.
Due to the tendency of Mtb to form aggregates, it is necessary to break up clumps to provide a single cell suspension to achieve uniform aliquoting of mycobacteria into multiple wells of cultured macrophages in each experiment. Similarly, this disaggregation step is repeated after macrophages have been lysed to provide an accurate estimation of growth. In the present protocol, the mycobacteria are passed through a 25 G needle using a syringe to generate a single-cell suspension. Other dispersal methods include sonication of the mycobacterial suspension in a sonicating water bath or vortexing with glass beads19. However, both of these techniques can alter the composition of the outer capsule of Mtb20,21 and may influence binding to and phagocytosis by macrophages21.
Pharmacological compounds can influence the uptake of mycobacteria if they are incubated with macrophages before or during infection. In addition, when performing experiments with primary human macrophages, there may be variation in phagocytosis of mycobacteria between donors. In these circumstances, using a fixed ratio of mycobacteria: macrophages would lead to differences in uptake during the 3 h infection period. This may lead to erroneous assumptions about the efficacy of compounds. This can be avoided by performing staining for acid fast bacilli (AFB) using the auramine method described here (or another AFB stain) in the presence of each HDT and adjusting the initial inoculum accordingly22. While prone to a certain amount of error due to difficulty distinguishing extracellular from intracellular mycobacteria19 and the likelihood that small clumps are still present, the AFB staining procedure helps to equalize the initial level of intracellular infection for each treatment.
In this protocol, for samples collected at times other than the initial 3 h infection sample, the lysate and supernatant are combined to collect intracellular mycobacteria and those that have been released extracellularly. The reason for this is that, since the macrophages have been washed after the initial 3 h incubation with Mtb, extracellular mycobacteria present are likely to have been released by necrotic macrophages. On the other hand, if investigators prefer to harvest the remaining intracellular mycobacteria and exclude extracellular mycobacteria, this method is easily adapted to do so.
Advantages of automated liquid culture-based instruments compared to the solid medium are simplicity of set up, a more efficient experimental workflow as analysis of serial dilutions is unnecessary, an accurate readout is obtained compared to the more subjective manual counting of colonies, faster time to results (approximately 5-12 days compared to 21 or more for agar plates), and higher sensitivity5. Disadvantages include (1) dependence on researchers having access to a suitable instrument, (2) the purchase of instrument bottles can add significantly to the cost per sample. In addition, (3) detection of growth depends on bacillary replication: sub-populations of non-replicating dormant mycobacteria will not give a signal23,24. However, this drawback also applies to growth on solid media and prolonged incubation in liquid media may restore growth of these sub-populations25,26. Furthermore, (4) using this liquid broth culture method necessitates manual harvesting of lysates and would not be suitable for medium to high throughput workflows required to screen compound libraries. In such cases, automated imaging systems are a more practical option27. However, even in high throughput experiments, it is essential to confirm the antimicrobial activity of hits using a method that measures growth directly28. In the future, the authors plan to use this method to assess the bactericidal effects of autophagy-inducing drugs in combination with conventional anti-tubercular drugs on Mtb survival in macrophages.
Disclosures
The authors have nothing to disclose.
Acknowledgements
This work was funded by Science Foundation Ireland (SFI 08/RFP/BMT1689), the Health Research Board in Ireland [HRA-POR/2012/4 and HRA-POR-2015-1145] and Royal City of Dublin Hospital Trust.
Materials
| IX51 Fluorescent Microscope | Olympus, Japan | N/A | AFB detection and imaging |
| 2 mL microtube, flat bottom, screw cap, sterile | Sarstedt, North Carolina, USA | 72.694.006 | Mtb infection of macropahges |
| 5 mL syringe, Luer lock | BD Biosciences, San Jose, CA, USA | SZR-150-031K | Mtb infection of macropahges/CFU |
| 50 mL tube, sterile | Sarstedt, North Carolina, USA | 62.547.254 | Mtb infection of macropahges |
| all-trans-Retinoic Acid (ATRA) ≥98% (HPLC) | Sigma Aldrich, Missouri, USA | R2625 | Host directed therapy candidate |
| BacT/ALERT 3D Microbial Detection System | Biomerieux ( Hampshire, UK) | 247001 | Broth-based colormetric detection system |
| BACT/ALERT MP BACT/ALERT MP Nutrient Supplement | Biomerieux ( Hampshire, UK) | 414997 | Broth-based colormetric detection assay |
| BACT/ALERT MP culture bottles | Biomerieux ( Hampshire, UK) | 419744 | Broth-based colormetric detection assay |
| BD BBL Middlebrook ADC Enrichment, 20 mL | BD Biosciences, San Jose, CA, USA | M0553 | Mycobacterium liquid culture |
| BD BBL Middlebrook OADC Enrichment, 20mL | BD Biosciences, San Jose, CA, USA | M0678 | Colony Forming Units |
| Cell scraper, 25 cm | Sarstedt, North Carolina, USA | 83.1830 | Harvest of lmacrophage lysates |
| Corning Syringe Filter, 0.2 µm | Corning Incorporated, Germany | 431219 | Sterilization of lysis buffer |
| Cover glass (borosilicate), 24 x 50 mm, #1.5 thickness | VWR International Limited | 631 – 0147 | |
| Cycloheximide, from microbial | Sigma Aldrich, Missouri, USA | C7698 | Colony Forming Units |
| Dako Fluorescent Mounting Medium | Agilent Technologies Ireland Limited | S3023 | Antifade mounting medium |
| Dulbecco’s Phosphate Buffered Saline | Sigma Aldrich, Missouri, USA | D8537 | Mtb infection of macropahges |
| Fetal Bovine Serum, Gibco | Thermo Fisher, Massachusetts, USA | 10270106 | Macrophage cell culture |
| Glycerol, Difco | BD Biosciences, San Jose, CA, USA | 228220 | Colony Forming Units |
| Hoescht 33342 (bisBenzimide H 33342 trihydrochloride) | Sigma Aldrich, Missouri, USA | B2261 | Nuclear stain |
| IFNγ, recombinant human | R&D Systems Inc, Minnesota, USA | 285-IF | Host directed therapy candidate |
| Labtek 2-well chamber slide, sterile, Nunc | Thermo Fisher, Massachusetts, USA | TKT-210-150R | Mtb infection of macropahges |
| L-Asparagine, anhydrous | Sigma Aldrich, Missouri, USA | A4159 | Colony Forming Units |
| Linezolid | Sigma Aldrich, Missouri, USA | PZ0014 | Antibiotic |
| Microlance Hypodermic Needle 25 G | BD Biosciences, San Jose, CA, USA | 300400 | Mtb infection of macropahges/CFU |
| Middlebrook 7H10 Agar Base | BD Biosciences, San Jose, CA, USA | M0303 | Colony Forming Units |
| Middlebrook 7H9 Broth Base | BD Biosciences, San Jose, CA, USA | M0178 | Mycobacterium liquid culture |
| Modified Auramine O Stain and Decolourizer | Scientific Device Laboratory, IL, USA | 345-250 | AFB stain |
| Paraformaldehyde | Sigma Aldrich, Missouri, USA | 158127 | Mtb infection of macropahges |
| Petri dishes, 92 x 16mm (20/bag) | Sarstedt, North Carolina, USA | 82.1473.001 | Colony Forming Units |
| Phorbol 12-myristate 13-acetate (PMA) | Sigma Aldrich, Missouri, USA | P8139 | Macrophage cell culture |
| Polysorbate 80, Difco | BD Biosciences, San Jose, CA, USA | 231181 | Mycobacterium liquid culture |
| RPMI-1640, Gibco | Thermo Fisher, Massachusetts, USA | 52400025 | Macrophage cell culture |
| Sterile Cell Spreader, L-Shaped | Fisherbrand, Thermo Fisher, MA, USA | RB-44103 | Colony Forming Units |
| T25 TC flask, angled neck, filter cap, sterile, Nunc | Thermo Fisher, Massachusetts, USA | 156367 | Mycobacterium liquid culture |
| THP-1 cell line | ATCC, Virginia, USA | ATCC TIB-202 | Macrophage cell culture |
References
- WHO. Global Tuberculosis Report. World Health Organization. , (2020).
- Russell, D. G. Mycobacterium tuberculosis and the intimate discourse of a chronic infection. Immunological Reviews. 240 (1), 252-268 (2011).
- Young, C., Walzl, G., Du Plessis, N. Therapeutic host-directed strategies to improve outcome in tuberculosis. Mucosal Immunology. 13 (2), 190-204 (2020).
- Angeby, K. A., et al. Evaluation of the BacT/ALERT 3D system for recovery and drug susceptibility testing of Mycobacterium tuberculosis. Clinical Microbiology and Infection. 9 (11), 1148-1152 (2003).
- Asmar, S., Drancourt, M. Rapid culture-based diagnosis of pulmonary tuberculosis in developed and developing countries. Frontiers in Microbiology. 6, 1184 (2015).
- Diacon, A. H., et al. Time to liquid culture positivity can substitute for colony counting on agar plates in early bactericidal activity studies of antituberculosis agents. Clinical Microbiology and Infection. 18 (7), 711-717 (2012).
- Diacon, A. H., et al. Time to positivity in liquid culture predicts colony forming unit counts of Mycobacterium tuberculosis in sputum specimens. Tuberculosis. 94 (2), 148-151 (2014).
- O’Sullivan, D. M., et al. Evaluation of liquid culture for quantitation of Mycobacterium tuberculosis in murine models. Vaccine. 25 (49), 8203-8205 (2007).
- Heinrichs, M., et al. Mycobacterium tuberculosis Strains H37ra and H37rv have equivalent minimum inhibitory concentrations to most antituberculosis drugs. International Journal of Mycobacteriology. 7 (2), 156-161 (2018).
- Sorrentino, F., et al. Development of an intracellular screen for new compounds able to inhibit mycobacterium tuberculosis growth in human macrophages. Antimicrobial Agents and Chemotherapy. 60 (1), 640-645 (2016).
- O’Leary, S. M., et al. Cigarette smoking impairs human pulmonary immunity to Mycobacterium tuberculosis. American Journal of Respiratory and Critical Care Medicine. 190 (12), 1430-1436 (2014).
- Ryan, R. C., O’Sullivan, M. P., Keane, J. Mycobacterium tuberculosis infection induces non-apoptotic cell death of human dendritic cells. BMC Microbiology. 11, 237 (2011).
- Keane, J., Remold, H. G., Kornfeld, H. Virulent mycobacterium tuberculosis strains evade apoptosis of infected alveolar macrophages. The Journal of Immunology. 164 (4), 2016-2020 (2000).
- Gao, X. F., Yang, Z. W., Li, J. Adjunctive therapy with interferon-gamma for the treatment of pulmonary tuberculosis: a systematic review. International Journal of Infectious Diseases. 15 (9), 594-600 (2011).
- Thorpe, T. C., et al. BacT/Alert: an automated colorimetric microbial detection system. Journal of Clinical Microbiology. 28 (7), 1608-1612 (1990).
- Gleeson, L. E., et al. Cutting edge: Mycobacterium tuberculosis induces aerobic glycolysis in human alveolar macrophages that is required for control of intracellular bacillary replication. The Journal of Immunology. 196 (6), 2444-2449 (2016).
- O’Connor, G., et al. Inhalable poly(lactic-co-glycolic acid) (PLGA) microparticles encapsulating all-trans-Retinoic acid (ATRA) as a host-directed, adjunctive treatment for Mycobacterium tuberculosis infection. European Journal of Pharmaceutics and Biopharmaceutics. 134, 153-165 (2019).
- O’Connor, G., et al. Sharpening nature’s tools for efficient tuberculosis control: A review of the potential role and development of host-directed therapies and strategies for targeted respiratory delivery. Advanced Drug Delivery Reviews. 102, 33-54 (2016).
- Rodriguez, D. C., Ocampo, M., Salazar, L. M., Patarroyo, M. A. Quantifying intracellular Mycobacterium tuberculosis: An essential issue for in vitro assays. Microbiologyopen. 7 (2), 00588 (2018).
- Ortalo-Magne, A., et al. Molecular composition of the outermost capsular material of the tubercle bacillus. Microbiology. 141, 1609-1620 (1995).
- Stokes, R. W., et al. The glycan-rich outer layer of the cell wall of Mycobacterium tuberculosis acts as an antiphagocytic capsule limiting the association of the bacterium with macrophages. Infection and Immunity. 72 (10), 5676-5686 (2004).
- O’Sullivan, M. P., O’Leary, S., Kelly, D. M., Keane, J. A caspase-independent pathway mediates macrophage cell death in response to Mycobacterium tuberculosis infection. Infection and Immunity. 75 (4), 1984-1993 (2007).
- Cheon, S. H., et al. Bactericidal activity in whole blood as a potential surrogate marker of immunity after vaccination against tuberculosis. Clinical and Diagnostic Laboratory Immunology. 9 (4), 901-907 (2002).
- Nathan, C., Barry, C. E. TB drug development: immunology at the table. Immunological Reviews. 264 (1), 308-318 (2015).
- Bowness, R., et al. The relationship between Mycobacterium tuberculosis MGIT time to positivity and cfu in sputum samples demonstrates changing bacterial phenotypes potentially reflecting the impact of chemotherapy on critical sub-populations. Journal of Antimicrobial Chemotherapy. 70 (2), 448-455 (2015).
- Basu Roy, R., et al. An auto-luminescent fluorescent BCG whole blood assay to enable evaluation of paediatric mycobacterial responses using minimal blood volumes. Frontiers in Pediatrics. 7, 151 (2019).
- Christophe, T., et al. High content screening identifies decaprenyl-phosphoribose 2′ epimerase as a target for intracellular antimycobacterial inhibitors. PLoS Pathogens. 5 (10), 1000645 (2009).
- Franzblau, S. G., et al. Comprehensive analysis of methods used for the evaluation of compounds against Mycobacterium tuberculosis. Tuberculosis. 92 (6), 453-488 (2012).

