CorrelationCalculator and Filigree: Tools for Data-Driven Network Analysis of Metabolomics Data
Summary
We present CorrelationCalculator and Filigree, two tools for data-driven network construction and analysis of metabolomics data. CorrelationCalculator supports building a single interaction network of metabolites based on expression data, while Filigree allows building a differential network, followed by network clustering and enrichment analysis.
Abstract
A significant challenge in the analysis of omics data is extracting actionable biological knowledge. Metabolomics is no exception. The general problem of relating changes in levels of individual metabolites to specific biological processes is compounded by the large number of unknown metabolites present in untargeted liquid chromatography-mass spectrometry (LC-MS) studies. Further, secondary metabolism and lipid metabolism are poorly represented in existing pathway databases. To overcome these limitations, our group has developed several tools for data-driven network construction and analysis. These include CorrelationCalculator and Filigree. Both tools allow users to build partial correlation-based networks from experimental metabolomics data when the number of metabolites exceeds the number of samples. CorrelationCalculator supports the construction of a single network, while Filigree allows building a differential network utilizing data from two groups of samples, followed by network clustering and enrichment analysis. We will describe the utility and application of both tools for the analysis of real-life metabolomics data.
Introduction
In the last decade, metabolomics has emerged as an omics science due to advances in analytical technologies such as Gas Chromatography-Mass Spectrometry (GC-MS) and Liquid Chromatography-Mass Spectrometry (LC-MS). These techniques allow simultaneous measurement of hundreds to thousands of small molecule metabolites, creating complex multidimensional datasets. Metabolomics experiments can be performed in targeted or untargeted modes. Targeted metabolomics experiments measure specific classes of metabolites. They are usually hypothesis-driven, while untargeted approaches attempt to measure as many metabolites as possible and are hypothesis-generating in nature. Targeted assays usually include internal standards and thus allow for absolute quantification of metabolites of interest. In contrast, untargeted assays allow relative quantification and include many unknown metabolites1.
Analysis of metabolomics data is a multi-step process that leverages many specialized software tools1. It can be divided into the following three major steps: (1) data processing and quality control, (2) statistical analysis, and (3) biological data interpretation. The tools described here are designed to enable the latter step of the analysis.
An intuitive and popular way to interpret metabolomics data is to map the experimental measurements onto metabolic pathways. Numerous tools have been designed to achieve this2,3,4,5, including Metscape, developed by our group6. Pathway mapping is often combined with enrichment analysis, which helps identify the most significant pathways7,8. These techniques first gained prominence in the analysis of gene expression data and have been successfully applied for the analysis of proteomics and epigenomics data9,10,11,12,13. However, the analysis of metabolomics data presents a number of challenges for knowledge-based approaches. First, in addition to the endogenous metabolites, metabolomics assays measure exogenous compounds, including those that come from nutrition and other environmental sources. These compounds, as well as metabolites produced by bacteria, cannot be mapped onto human or metabolic pathways of other eukaryotic organisms. Further, pathway coverage of secondary metabolism and lipid metabolism currently does not allow high-resolution mapping at the level that would easily support the biological interpretation of the data14,15.
Data-driven network analysis techniques can help overcome these challenges. For example, correlation-based networks can help derive relationships among both known and unknown metabolites and facilitate the annotation of the unknowns16. While computing Pearson's correlation coefficients is the most straightforward approach to establishing the linear relationships between metabolites, the disadvantage is that it captures both direct and indirect associations17,18,19. An alternative is to compute partial correlation coefficients that can distinguish between direct and indirect associations. Gaussian graphical modeling (GGM) can be used to estimate partial correlation networks. However, GGM requires that the sample size and the number of features be comparable. This condition is rarely met in untargeted LC-MS data that contains measurements for thousands of metabolic features. Regularization techniques can be utilized to overcome this limitation. Graphical lasso (Glasso) and nodewise regression are popular methods for regularized estimation of the partial correlation network16,20.
The first of the bioinformatics tools presented here, CorrelationCalculator16, is based on the debiased sparse partial correlation (DSPC) algorithm. DSPC relies on de-sparsified graphical lasso modeling. The underlying assumption of the algorithm is that the number of connections among the metabolites is considerably smaller than the number of samples, i.e., the partial correlation network of metabolites is sparse. This assumption allows DSPC to discover the connectivity among large numbers of metabolites using fewer samples, leveraging regularized regression techniques. Further, using a debiasing step for the regularized regression estimates, it obtains sampling distributions for the edge parameters that can be used to construct confidence intervals and test hypotheses of interest (e.g., presence/absence of a single or a group of edges). The presence or absence of an edge in the partial correlation network can thus be formally tested using the computed p-values.
CorrelationCalculator proved to be very useful for single-group analysis16; however, the objective of many metabolomics experiments is the differential analysis of two or more conditions. While CorrelationCalculator can be employed on each of the groups separately to generate partial correlation networks for each condition, this approach limits the number of samples that can be used for network generation. Since a sufficiently large sample size is one of the biggest considerations in data-driven analysis, methods that can leverage all available samples in the data to construct networks are highly desirable. This approach is implemented in the second tool presented here, called Filigree21. Filigree relies on the previously published Differential Network Enrichment Analysis (DNEA) algorithm22. Table 1 shows the applications and the workflow of both tools.
| Number of experimental conditions (k) | k = 1 | k = 2 |
| Software tool | CorrelationCalculator | Filigree |
| Input data | • Metabolites x Samples data matrix | • Metabolites x Samples data matrix • Experimental groups |
| Workflow • Pretreatment • Network estimation • Network clustering • Enrichment analysis |
• Log transformation; autoscaling • DSPC • Via external apps • No |
• Log transformation; autoscaling • Joint network estimation • Consensus clustering • NetGSA |
| Data visualization | Via external app, e.g., Cytoscape | Via external app, e.g., Cytoscape |
| Testing metabolic modules for the association with outcome of interest (optional) | Via external apps | Via external apps |
Table 1: The scope of application and the workflow of CorrelationCalculator and Filigree.
Protocol
1. CorrelationCalculator
- Download a sample comma-delimited input file containing a list of metabolites with experimental measurements at http://metscape.med.umich.edu/kora_data_240.csv.
- Double-click on the downloaded sample file to open it.
- Ensure that the file contains labels for both the samples and the metabolites.
- Since samples are in rows, confirm that the first column is the sample names and the first row is the metabolite names.
- Download the CorrelationCalculator Java application (http://metscape.med.umich.edu/calculator.html). Double-click on the downloaded .jar file to launch the application.
- On the Input tab, click the Browse button to upload the input file.
- Under Specify File Format, use the dropdown arrow to select the appropriate input file format. Select Samples in Rows (Supplementary Figure 1).
- Move to the Data Normalization tab by clicking the Next >> button at the bottom right of the window.
- Under Select Method(s), check the box next to Log2-Transform Data. Check the box next to Autoscale Data.
- Under Normalize Data, click the Run button.
NOTE: Once the normalization is complete, click the View Normalized Data button, located under Normalize Data, and review the updated dataset (Supplementary Figure 2). - Under Normalize Data, click the Save button and save the new data file.
- Move to the Data Analysis tab by clicking the Next >> button at the bottom right of the window.
- Under Calculate Pearson's Correlation, click Run. Determine the best Pearson's Correlation range for the data.
- Click the View Histogram button. Review the frequency of the maximal Pearson's correlation scores per feature.
- Click the View Heatmap button. Review the representation of Pearson's correlation matrix.
- Under Filter by Pearson's Correlations, leave the default numbers to filter by a range of 0.00 to 1.00
NOTE: Slide the small blue arrow at the right end from 1 and the small blue arrow at the left from 0 to change the filter. Entering specific numbers in the text boxes is also an option. - Under Select Partial Correlation Method, select the desired method, DSPC Method.
NOTE: If the number of metabolites is smaller than the number of samples in the dataset, only the DSPC method can be used. - Under Calculate Partial Correlations, click the Run button (Supplementary Figure 3).
- Click the View CSV File and view the results. Click the Save button and save the results.
- Click the View in MetScape button to launch an interactive correlation network.
See Karnovsky, A. et al.6 for more information on using MetScape.
NOTE: MetScape is a Cytoscape application that allows for the creation and exploration of correlation networks.
2. Filigree
- Download a sample comma-delimited input file containing metabolite measurements at http://metscape.med.umich.edu/T1D_primaryMetabolites_noIS_log_scaled_sorted.csv.
- Double-click on the downloaded sample file to open it.
- Ensure the file contains sample names in column 1 and group assignations in column 2. Confirm that the remaining columns contain metabolites/lipids.
- Ensure that each row represents a sample.
NOTE: The metabolite measurements should be log-transformed and auto-scaled unless performing feature aggregation, in which case the measurements should only be log-transformed.
- Download the Filigree Java application (http://metscape.med.umich.edu/filigree.html).
NOTE: A detailed user manual is available at http://metscape.ncibi.org/v0.1.2Filigree_UserManual.pdf. - Double-click on the downloaded .jar file to launch the application.
- On the Data tab, click the Browse button to upload the input file.
- Under Specify Columns/Rows, click on the dropdown arrow next to Sample ID to select the corresponding column/row name from the input file. Select Sample.
- Under Specify Columns/Rows, click on the dropdown arrow next to "Group" to select the corresponding column/row from the input file. Select Group.
- Under Specify Sample Groups, click on the dropdown arrows next to each Group to select the corresponding group column from the input file. For Group 1, select Diabetic. For Group 2, select Non-diabetic.
- Under Feature Grouping, check the box next to the desired method, Calculate Feature Groups.
- Click the View Heatmaps button. View the heatmap and determine a desired percent reduction.
- Use the Feature Reduction slider to select the desired percent reduction of features. Slide the small circle until the percent reduction shows a feature-to-sample ratio of 1.25 (Supplementary Figure 4).
- Move to the Analysis tab by clicking the Next >> button at the bottom right of the window.
- Under Select Output Directory, click the Browse button and select the desired directory location for storing the generated output files.
- Click the Run Analysis button located at the bottom left of the window. The progress bars update for each analysis component (Supplementary Figure 5). Click the OK button on the popup window displaying the message Analysis Completed Successfully.
- On the Analysis tab, click the Browse Networks button to open the interactive Filigree subnetworks in a browser tab.
- Click on the Subnetwork 1 link under the Subnetwork Name column.
- Explore the interactive subnetwork using the various buttons. Click the + button and zoom in on the part of the network. Click the – button and zoom out (Supplementary Figure 6).
- Click on a Group Node and drag it to reposition it within the subnetwork.
NOTE: Node color represents up/down-regulation, and color opacity represents higher/lower fold change. The edge color represents the differential status between groups. - Click the Expand Features button at the top right of the page to expand all the group nodes. Review the specific compounds that make up the group nodes.
- Click the Collapse Features button at the top right of the page to collapse the recently expanded group nodes.
- Click the By Sample Group button at the top right of the page to change the view from a single subnetwork to multiple subnetworks split by a group. Explore and compare the groups using this view of the subnetworks (Supplementary Figure 7).
- Click the All Samples button to go back to the single subnetwork view.
- View the next subnetwork by clicking the Next button at the top right of the page.
- Repeat steps 2.19-2.23 for each subnetwork.
- Click the Differential Network Enrichment Analysis Results link at the top middle of the window to return to the summary table view listing all the subnetworks.
NOTE: Import the edge and/or node output files in a different software tool, such as Cytoscape23, to create additional network visualizations.
3. Additional considerations
- For Mac computers running Big Sur (OSX 11.2) or later, approve the tool in Apple Menu > System Preferences > Security & Privacy > General and select Allow at the bottom of the tab.
- In addition, allow Filigree access to the files in Apple Menu > System Preferences > Security & Privacy > Privacy by selecting Files and Folders in the menu on the left and then selecting Filigree in the menu on the right.
Representative Results
To illustrate the use of CorrelationCalculator, we constructed a partial correlation network using a subset of the metabolomics data from the KORA population study described in Krumsiek et al.24. The dataset contained 151 metabolites and 240 samples. Figure 1 shows the resulting partial correlation network that was visualized in Cytoscape. The network contains 148 nodes and 272 edges. The color of the nodes represents metabolites that belong to different chemical classes, while the edges represent the adjusted p-value of the partial correlation coefficients (adjusted p-value < 0.05) . Notably, despite not using any prior information CorrelationCalculator was able to group together chemically related metabolites. For example, phosphatidylcholines and lysophosphatidylcholines are closely connected in the network. Visualizing metabolite changes in the context of this type of network can facilitate hypothesis generation, help plan future experiments and enable manuscript preparation. To illustrate a potential workflow utilizing a partial correlation metabolite network, we performed consensus network clustering as described in Ma et al.22, resulting in the identification of 9 subnetworks or metabolic modules. These modules had a good agreement with the chemical classes, i.e., metabolites belonging to the same chemical class tended to be part of the same metabolic module. The user can access the clustering tool clusterNet at https://github.com/Karnovsky-Lab/clusterNet.
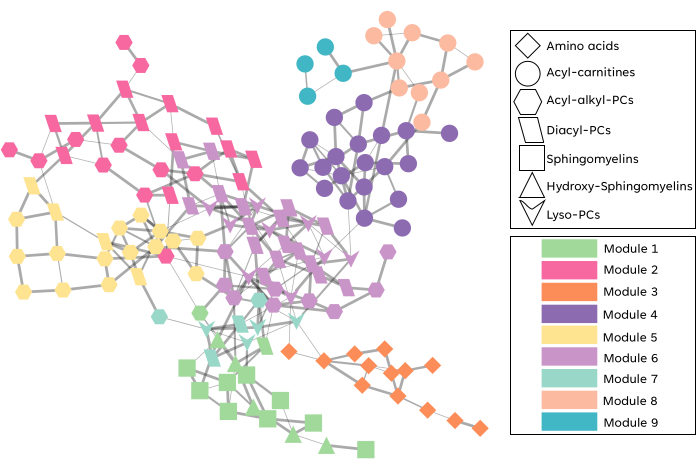
Figure 1: Representative example of a CorrelationCalculator network. The network was constructed from a subset of the KORA population study metabolomics data24 consisting of 151 metabolites across 240 subjects. The nodes represent metabolites, and the edges connecting them are weighted by the adjusted p-value of partial correlation coefficients (adjusted p-value < 0.05). The shape of the nodes represents different metabolic classes, and the color represents metabolic modules obtained by clustering the network using the consensus clustering method. Please click here to view a larger version of this figure.
We illustrate the application of Filigree by analyzing a dataset from a mouse model of type I diabetes (T1D)25,26. Plasma metabolite measurements from T1D and non-diabetic (NOD) mice were used to generate a differential partial correlation network (Figure 2). Notably, we observe a higher degree of network connectivity in the non-diabetic group. The next steps of the analysis identified twelve metabolic modules, nine of which were significantly different between T1D and non-diabetic mice (FDR < 0.05). We refer the reader to the original publication for further insights into biological conclusions that can be drawn from this analysis21.
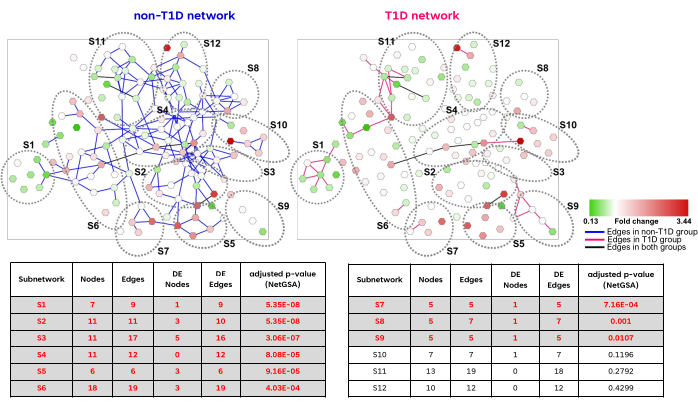
Figure 2: Representative example of a Filigree network. The differential network was constructed utilizing the levels of 163 metabolites from 71 mice (30 T1D and 41 non-T1D)25,26. Differential edges between T1D and non-T1D groups are indicated in pink and blue, respectively. The nodes are colored based on the fold change. The table shows the enrichment results produced by Filigree. Nine out of the twelve identified subnetworks were significantly different between T1D and non-T1D (adjusted p-value < 0.05). Please click here to view a larger version of this figure.
Supplementary Figure 1: CorrCalc_InputTab. Screenshot of the Correlation Calculator's Input tab. Please click here to download this File.
Supplementary Figure 2: CorrCalc_DataNormTab. Screenshot of the Correlation Calculator's Data Normalization tab. Log-2 Transform Data and Autoscale Data are checked. Please click here to download this File.
Supplementary Figure 3: CorrCalc_DataAnalTab. Screenshot of the Correlation Calculator's Data Analysis tab showing filtering to Pearson's Correlation of 0-0.8. In addition, the DSPC Method has been selected. Please click here to download this File.
Supplementary Figure 4: Filigree_DataTab. Screenshot of Filigree's Data tab. Columns, rows, and groups have been specified. The Calculate Feature Groups method has been selected with a feature reduction of 1.25 feature-to-sample ratio. Please click here to download this File.
Supplementary Figure 5: Filigree_AnalysisTab. Screenshot of Filigree's Analysis tab showing the progress of the different analysis components. Please click here to download this File.
Supplementary Figure 6: Filigree_Subnetwork1. A subnetwork generated from Filigree. Node color represents up/down-regulation, and color opacity represents higher/lower fold change. The edge color represents the differential status between groups. Please click here to download this File.
Supplementary Figure 7: Filigree_Subnetwork_SampleGroup. Subnetwork separated by group. The left network represents diabetic samples, and the right network represents non-diabetic samples. Node color represents expression level proportional to the group mean. Please click here to download this File.
Discussion
Partial correlation-based network analysis methods implemented in CorrelationCalculator and Filigree help overcome some of the limitations of knowledge-based metabolic pathway analyses, especially for the datasets with a high prevalence of unknown metabolites and limited coverage of metabolic pathways (e.g., lipidomics data). These tools have been widely used by the research community to analyze a broad range of metabolomics and lipidomics data14,22,27,28,29,30. For example, CorrelationCalculator has been used to analyze the data from many biological systems ranging from microbiome and plants to human disease31,32,33,34. Here we illustrate how data-driven network analysis, enabled by our tools, can be combined with clustering and regression analysis to pinpoint the metabolic modules associated with the phenotype of interest.
Partial correlation networks generated using CorrelationCalculator and Filigree can be clustered using graph clustering algorithms to produce metabolic modules. These modules tend to comprise metabolites that are chemically or functionally related to each other. Such modules are very useful not just from a visualization perspective but also from a biological relevance standpoint. Studying the relationships between metabolic modules and phenotypic outcomes of interest (e.g., survival outcome) can provide more statistical power and generate additional biological insights compared to testing individual metabolites.
Metabolic modules identified through network clustering approaches can also be used in enrichment analysis. Filigree uses metabolic modules identified through consensus clustering instead of predefined biological pathways. Although partial correlation-based metabolic modules are not identical to pathways, they consistently group chemically and biochemically similar metabolites (e.g., amino acids, acylcarnitines, lipids of the same class, etc.). Filigree further tests the significance of these modules using NetGSA algorithm22,35. In addition to differential nodes, NetGSA accounts for disease-specific differences in network structure.
One of the issues to consider when using CorrelationCalculator and Filigree for the analysis of 'real life' metabolomics and lipidomics data is the relationship between the number of metabolites vs. the number of samples in a given experiment. While large-scale epidemiological studies involving thousands of samples are becoming more common, the sample size in the majority of metabolomics experiments remains modest. This is particularly true for mechanistic studies involving systems where low biological variation is expected (i.e., cell lines or genetically homogeneous animal models). The statistical algorithms implemented in both tools can be applied in situations when the number of metabolites exceeds the number of samples, but the increase in that ratio leads to more sparse networks.
Another important consideration for the application of the tools described here concerns the analysis of untargeted metabolomics data that are known to contain a large number of redundant or degenerate features36, which may include isotopes, chemical adducts, in-source fragments, and contaminants. Since many degenerate features originate from the same metabolite, they tend to have a high degree of correlation. Partial correlation-based analysis of such data may require careful annotation and removal of degenerate features.
In conclusion, the tools presented here provide a viable alternative to knowledge-based pathways analysis tools for the interpretation of metabolomics data.
Disclosures
The authors have nothing to disclose.
Acknowledgements
This work was supported by NIH 1U01CA235487 grant.
Materials
| CorrelationCalculator | JAVA | http://metscape.med.umich.edu/calculator.html | |
| clusterNet | https://github.com/Karnovsky-Lab/clusterNet | ||
| Cytoscape | Cytoscape | https://cytoscape.org/ | |
| Filigree | JAVA | http://metscape.med.umich.edu/filigree.html | |
| MetScape | Cytoscape | https://apps.cytoscape.org/apps/metscape | Cytoscape application that allows for the creation and exploration of correlation networks. |
References
- Sas, K. M., Karnovsky, A., Michailidis, G., Pennathur, S. Metabolomics and diabetes: analytical and computational approaches. Diabetes. 64 (3), 718-732 (2015).
- Cottret, L., et al. MetExplore: Collaborative edition and exploration of metabolic networks. Nucleic Acids Research. 46 (W1), W495-W502 (2018).
- Garcia-Alcalde, F., Garcia-Lopez, F., Dopazo, J., Conesa, A. Paintomics: A web based tool for the joint visualization of transcriptomics and metabolomics data. Bioinformatics. 27 (1), 137-139 (2011).
- Kuo, T. C., Tian, T. F., Tseng, Y. J. 3Omics: A web-based systems biology tool for analysis, integration and visualization of human transcriptomic, proteomic and metabolomic data. BMC Systems Biology. 7, 64 (2013).
- Paley, S. M., Karp, P. D. The pathway tools cellular overview diagram and Omics Viewer. Nucleic Acids Research. 34 (13), 3771-3778 (2006).
- Karnovsky, A., et al. Metscape 2 bioinformatics tool for the analysis and visualization of metabolomics and gene expression data. Bioinformatics. 28 (3), 373-380 (2012).
- Chong, J., Xia, J. Using MetaboAnalyst 4.0 for metabolomics data analysis, interpretation, and integration with other omics data. Methods in Molecular Biology. 2104, 337-360 (2020).
- Lopez-Ibanez, J., Pazos, F., Chagoyen, M. MBROLE 2.0-functional enrichment of chemical compounds. Nucleic Acids Research. 44 (W1), W201-W204 (2016).
- Cavalcante, R. G., et al. Broad-Enrich: Functional interpretation of large sets of broad genomic regions. Bioinformatics. 30 (17), i393-i400 (2014).
- Huang, D. W., et al. DAVID bioinformatics resources: Expanded annotation database and novel algorithms to better extract biology from large gene lists. Nucleic Acids Research. 35 (Web Server issue), W169-W175 (2007).
- Lee, P. H., O’Dushlaine, C., Thomas, B., Purcell, S. M. INRICH: interval-based enrichment analysis for genome-wide association studies. Bioinformatics. 28 (13), 1797-1799 (2012).
- Segre, A. V., Groop, L., Mootha, V. K., Daly, M. J., Altshuler, D. Common inherited variation in mitochondrial genes is not enriched for associations with type 2 diabetes or related glycemic traits. PLoS Genetics. 6 (8), e1001058 (2010).
- Subramanian, A., et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proceedings of the National Academy of Sciences of the United States of America. 102 (43), 15545-15550 (2005).
- Afshinnia, F., et al. Lipidomic signature of progression of chronic kidney disease in the chronic renal insufficiency cohort. Kidney International Reports. 1 (4), 256-268 (2016).
- Barupal, D. K., et al. MetaMapp: Mapping and visualizing metabolomic data by integrating information from biochemical pathways and chemical and mass spectral similarity. BMC Bioinformatics. 13, 99 (2012).
- Basu, S., et al. Sparse network modeling and Metscape-based visualization methods for the analysis of large-scale metabolomics data. Bioinformatics. 33 (10), 1545-1553 (2017).
- Krumsiek, J., Suhre, K., Illig, T., Adamski, J., Theis, F. J. Gaussian graphical modeling reconstructs pathway reactions from high-throughput metabolomics data. BMC Systems Biology. 5, 21 (2011).
- Camacho, D., de la Fuente, A., Mendes, P. The origin of correlations in metabolomics data. Metabolomics. 1 (1), 53-63 (2005).
- Steuer, R., Kurths, J., Fiehn, O., Weckwerth, W. Observing and interpreting correlations in metabolomic networks. Bioinformatics. 19 (8), 1019-1026 (2003).
- Bühlmann, P., Van De Geer, S. . Statistics for High-Dimensional Data: Methods, Theory and Applications. , (2011).
- Iyer, G. R., et al. Application of differential network enrichment analysis for deciphering metabolic alterations. Metabolites. 10 (12), 479 (2020).
- Ma, J., et al. Differential network enrichment analysis reveals novel lipid pathways in chronic kidney disease. Bioinformatics. 35 (18), 3441-3452 (2019).
- Shannon, P., et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Reserach. 13 (11), 2498-2504 (2003).
- Krumsiek, J., et al. Mining the unknown: a systems approach to metabolite identification combining genetic and metabolic information. PLoS Genetics. 8 (10), e1003005 (2012).
- Fahrmann, J., et al. Systemic alterations in the metabolome of diabetic NOD mice delineate increased oxidative stress accompanied by reduced inflammation and hypertriglyceremia. American Journal of Physiology. Endocrinology and Metabolism. 308 (11), E978-E989 (2015).
- Grapov, D., et al. Diabetes associated metabolomic perturbations in NOD mice. Metabolomics. 11 (2), 425-437 (2015).
- Jin, Y., Bai, S., Huang, Z., You, L., Zhang, T. Technology characteristics and flavor changes of traditional green wheat product nian zhuan in Northern China. Frontiers in Nutrition. 9, 996337 (2022).
- Lin, Y. S., et al. Probing folate-responsive and stage-sensitive metabolomics and transcriptional co-expression network markers to predict prognosis of non-small cell lung cancer patients. Nutrients. 15 (1), 3 (2022).
- Pan, C., et al. Metabolomics study identified bile acids as potential biomarkers for gastric cancer: A case control study. Frontiers in Endocrinology (Lausanne). 13, 1039786 (2022).
- Pancoro, A., Karima, E., Apriyanto, A., Effendi, Y. (1)H NMR metabolomics analysis of oil palm stem tissue infected by Ganoderma boninense based on field severity Indices. Scientific Reports. 12 (1), 21087 (2022).
- Chele, K. H., et al. A global metabolic map defines the effects of a Si-based biostimulant on tomato plants under normal and saline conditions. Metabolites. 11 (12), 820 (2021).
- Hubert, J., et al. The effect of residual pesticide application on microbiomes of the storage mite Tyrophagus putrescentiae. Microbial Ecology. 85 (4), 1527-1540 (2023).
- Li, K., et al. Metabolomic and exposomic biomarkers of risk of future neurodevelopmental delay in human milk. Pediatric Research. 93 (6), 1710-1720 (2023).
- Marino, C., et al. The metabolomic profile in amyotrophic lateral sclerosis changes according to the progression of the disease: An exploratory study. Metabolites. 12 (9), 837 (2022).
- Ma, J., Shojaie, A., Michailidis, G. Network-based pathway enrichment analysis with incomplete network information. Bioinformatics. 32 (20), 3165-3174 (2016).
- Mahieu, N. G., Patti, G. J. Systems-level annotation of a metabolomics data set reduces 25000 features to fewer than 1000 unique metabolites. Analytical Chemistry. 89 (19), 10397-10406 (2017).

