Caffeine Extraction, Enzymatic Activity and Gene Expression of Caffeine Synthase from Plant Cell Suspensions
Summary
This protocol describes an efficient methodology for the extraction and quantification of caffeine in cell suspensions of C. arabica L. and an experimental process for evaluating the enzymatic activity of caffeine synthase with the expression level of the gene that encodes this enzyme.
Abstract
Caffeine (1,3,7-trimethylxanthine) is a purine alkaloid present in popular drinks such as coffee and tea. This secondary metabolite is regarded as a chemical defense because it has antimicrobial activity and is considered a natural insecticide. Caffeine can also produce negative allelopathic effects that prevent the growth of surrounding plants. In addition, people around the world consume caffeine for its analgesic and stimulatory effects. Due to interest in the technological applications of caffeine, research on the biosynthetic pathway of this compound has grown. These studies have primarily focused on understanding the biochemical and molecular mechanisms that regulate the biosynthesis of caffeine. In vitro tissue culture has become a useful system for studying this biosynthetic pathway. This article will describe a step-by-step protocol for the quantification of caffeine and for measuring the transcript levels of the gene (CCS1) encoding caffeine synthase (CS) in cell suspensions of C. arabica L. as well as its activity.
Introduction
Caffeine is a secondary metabolite that is biosynthesized by plants of the genus Coffea1. This alkaloid belongs to the methylxanthine family and is regarded as a chemical plant defense because it can act against the adverse effects of pathogens and herbivores2,3. In addition, this metabolite is responsible for the stimulating properties of the coffee drink, which is commonly consumed worldwide4,5. Due to its properties, several research groups are interested in studying the biosynthetic pathway and catabolism of caffeine6,7. Currently, plant in vitro cell/tissue cultures serve as an alternative for evaluating caffeine accumulation under various biotic and abiotic strategies8,9.
Caffeine biosynthesis involves the hydrolytic release of 7-methylxanthine from the corresponding ribose nucleoside followed by ordered N-methylations at positions 3 and 1. A specific S-adenosyl methionine (SAM)-dependent N-methyltransferase (NMT) catalyzes the methylation at position 7, whereas theobromine synthase (TS) and CS are involved in the 3- and 1-methylations, respectively, producing theobromine and caffeine. The study of genes encoding distinct NMTs has allowed understanding the mechanism that regulates caffeine production10,11. CS, which has N-methyltransferase activity, catalyzes the last two steps of the biosynthetic pathway of caffeine11. In coffee tree seedlings, it has been shown that light radiation can increase CS activity, which results in an increase in caffeine biosynthesis. Recently, we showed that the maintenance of cell suspensions of C. arabica L. under light irradiation is the optimal condition for evaluating the effects that produce abiotic stress factors impacting the biosynthetic pathway of caffeine8. The information obtained in these studies may have applications in metabolic engineering and systems biology for maximizing the study of the caffeine biosynthetic pathway in such in vitro systems.
Given the advantages of obtaining a suitable model for the study of caffeine biosynthesis, we optimized the extraction conditions for caffeine on cell suspensions of C. arabica L. It was also possible to develop a useful protocol for studying the enzymatic activity as well as methodological steps for assessing the level of gene transcripts of Coffea caffeine synthase 1 (CCS1) encoding this enzyme. Herein, we report a protocol to extract and quantify caffeine in C. arabica cell suspensions by thin-layer chromatography and densitometry (TLC-densitometry).
Protocol
1. Caffeine Extraction in Cell Suspensions of C. arabica L.
- Use C. arabica cell suspensions9. Maintain the suspensions by biweekly subcultures in Murashige and Skoog medium at pH 4.3 with constant 100 rpm shaking at 25 °C under continuous light (8.3 W/m2).
- Harvest the cells under vacuum filtration using 11 µm pore filter paper and a Buchner funnel.
- Register the fresh weight of the collected cells using a scale, wrap them in aluminum foil, freeze them in liquid nitrogen and keep them at -80 °C until analysis.
- Lyophilize the frozen cellular material for 72 h.
- Register the weight of the lyophilized cells in order to estimate the dry weight yield.
- Store the powdered material in reusable polyethylene bags (9 cm x 7.5 cm). Macerate the material to a fine powder and store in a desiccator until use.
- Measure 0.5 g of dry cellular matter on the analytical scale and add it to a glass flask (25 mL).
- Add 10 mL of acetone to the lyophilized cells and mix in a vortex mixer for 30 s for homogenization. Then, seal the flask with aluminum foil.
- Mix at 100 rpm using a rotator at room temperature (25 °C) for 5 h.
- Transfer all material to a 15 mL conical tube and centrifuge at 13,000 x g for 10 min. Transfer the liquid phase to a new tube and reduce to it dryness at room temperature in the fume hood.
- Resuspend the sample with 25 µL of acetone.
2. Conditions for the Assessment of Caffeine by Thin Layer Chromatography (TLC)-Densitometry
- Apply 1 µL of the previously resuspended caffeine extract to the stationary phase.
Note: Silica gel plates (F254) are used as the stationary phase. Before applying the samples, the plates were developed with 10 mL of chloroform-methanol [9:1 v/v] in a chromatography chamber (14 cm x 12 cm x 9.5 cm), and the plates were dried at room temperature. The characteristics of the chromatographic plate where the samples are applied are shown in Figure 1. The chamber was saturated for 30 min with solvent before developing the plate. TLC plates should handle using gloves to avoid contamination. - Develop the TLC in the chromatography chamber with 10 mL of the mobile phase cyclohexane-acetone [40:50 v/v].
Note: This mixture allows the separation of caffeine at a retention factor (Rf) of 0.34.- Remove the TLC plate from the chamber and let it dry completely at room temperature.
- Visualize caffeine bands in short wavelength ultraviolet light (UV, 254 nm). Use a compact UV lamp. Additionally, for this step, use goggles with UV protection.
- Quantify caffeine levels by densitometry at 273 nm. The instrument is controlled with chromatography software.
3. Ribonucleic Acid (RNA) Isolation
- Take the cell suspensions from step 1.1 and vacuum filtrate with filter paper (11 µm pore size).
- Collect the cellular material from the filter paper, weigh out 0.5 g of cellular material on an analytical balance and pack it in aluminum foil. Freeze the sample with liquid N2.
- Macerate the cell sample in a porcelain mortar with liquid nitrogen and add 1 mL of RNA isolation reagent until homogenized.
Note: Sterilize the porcelain mortars in a muffle furnace at 300 °C for 6 h. RNA isolation reagent contains phenol, guanidine isothiocyanate and other components.- Transfer 500 µL of the sample to a sterile microcentrifuge tube (1.5 mL). Add 300 µL of chloroform-isoamyl alcohol (24:1) and 300 μL of equilibrated phenol (pH 8.0).
- Mix the sample with a vortex mixer and then centrifuge it at 20,000 x g for 15 min at 4 °C.
- Transfer 300 μL of the upper phase to a microcentrifuge tube. Add 200 μL of isopropanol. Incubate the tube for 1 h at -20 °C.
- Centrifuge the tube at 12,000 x g for 10 min at 4 °C, and then decant the liquid phase.
- Add 1 mL of ethanol (70%) to form a pellet in the tube. Centrifuge at 12,000 x g for 10 min and decant. Repeat this step twice.
- Dry the sample for 1.5 h at room temperature (25 °C). Resuspend the RNA extract with 25 μL of diethyl pyrocarbonate (DEPC)-treated water.
Note: For the RNA extraction, use water treated with 0.1% DEPC v/v.
4. Incubation of RNA Transcript with Deoxyribonuclease (DNase)
- Into a sterile microcentrifuge tube, add 2 µg of total RNA extract. Add 1 μL of 10x reaction buffer (100 mM Tris-HCl, 25 mM MgCl2, 1 mM CaCl2). Add 1 μL of 1 U/µL DNase.
- Bring the reaction to a final volume of 10 μL with DEPC-treated water. Mix the sample and centrifuge briefly at 2,000 x g for 1 min.
- Incubate the sample at 37 °C for 30 min.
- Stop the reaction with the addition of 1 μL of 50 mM disodium ethylenediaminetetraacetate dihydrate (Na2EDTA) and incubate the sample at 65 °C for 10 min.
- Analyze the integrity of the RNA by agarose gel electrophoresis.
- Prepare a standard 1% agarose gel and stain it with 1 µL of 3x intercalating nucleic acid stain solution.
- Apply 500 ng of RNA to the agarose gel and run the gel.
- Visualize the integrity of the RNA using a gel photo documentation system.
- Use RNA as the template for cDNA synthesis by reverse transcriptase.
5. Complementary Deoxyribonucleic Acid (cDNA) Synthesis
- Add 2.5 μg of total RNA to a microcentrifuge tube. Add 1 μL of oligo-deoxythymine (dT) primer and fill the tube to a 13 μL final volume with nuclease-free water. Mix the sample and then centrifuge a 2,000 x g for 1 min.
- Incubate the sample at 65 °C for 5 min and then at 4 °C for 2 min.
- Add 4 μL of 5x reaction buffer used for reverse transcriptase, 2 μL of deoxynucleotide triphosphates (dNTPs) mix (10 mM of each dNTP) and 1 μL of reverse transcriptase (200 U/µL). Mix gently and centrifuge at 2,000 x g for 1 min.
- Incubate the sample at 45 °C for 50 min and then at 70 °C for 10 min.
- Quantify the cDNA concentration using a UV spectrophotometer.
- Use the cDNA as the template for the polymerase chain reaction (PCR).
Note: The reaction buffer was used as 10x (step 4.1.1) and 5x times concentrated (step 5.3), respectively.
6. Real-Time Quantitative PCR (qPCR) to Amplify the Gene CCS1
- Add 7.5 μL of Taq DNA polymerase 2x (0.1 U/mL) and 0.1 μM of 5-carboxy-X-rhodamine (ROX) to a PCR microcentrifuge tube.
Note: The hot start Taq DNA polymerase 2x was used as a solution 2x concentrated.- Add 5 μL of nuclease-free water.
- Add 0.75 μL each of 10 μM forward and reverse primer to amplify the gene CCS1 (AB086414) (forward: 5’-CCTGTATCCTGCGATGAACA-3’ and reverse: 5’-AACACTATCATAGAAGCCTTTG-3’). Use primers for tubulin as the house-keeping gene in a separate reaction (forward: 5’-GTGCCCAACTGGGTTCAA-3’ and reverse: 5’-CCTTCCTCCATACCTTCACC-3’)9.
- Add 600 ng of cDNA template.
- Perform the amplification reaction at 50 °C for 2 min and 95 °C for 5 min. Incubate the reaction in the real-time PCR system for 40 cycles of 95 °C for 30 s, 60 °C for 30 s and 72 °C for 30 s.
- Incubate the sample at 50 °C for 30 s and 20 °C for 10 s.
- Analyze the data with the PCR software.
- Calculate the relative expression by the 2-ΔΔCT method described by Livak and Schmittgen (2001)12.
Note: Analyze a control reaction that contains all components except the DNA template (no template control, NTC) to discard contaminated reagents or amplification primer-dimers.
7. Total Protein Extract of Cell Suspensions of C. arabica L.
- Filter cell suspensions under vacuum with a medium-pore filter paper.
- Weigh 1 g of cellular material and pack it in aluminum foil.
- Freeze the sample with liquid nitrogen. Macerate the sample in a sterilized porcelain mortar to obtain a fine powder.
- Transfer the sample to a glass vial and add 2.5 mL of extraction buffer (400 mM tris(hydroxymethyl)aminomethane hydrochloride (Tris-HCl), at pH 8.0, containing 10 mM β-mercaptoethanol, 5 mM Na2EDTA and 0.5% (w/v) sodium ascorbate.
- Mix the sample in a vortex mixer for 2 min.
Note: It is important that the sample is kept on ice to prevent protein denaturation.
- Mix the sample in a vortex mixer for 2 min.
- Centrifuge the sample at 20,000 x g for 20 min at 4 °C and transfer 500 µL aliquots into cryogenic vials (2 mL). Store samples at -72 °C until use.
- Measure the protein concentration of the sample at 562 nm in a spectrophotometer using bicinchoninic acid assay with bovine serum albumin as the standard.
8. Enzymatic Assay for Caffeine Synthase
- Prepare a reaction mixture in a microcentrifuge tube containing 200 µM SAM, 4.07 kBq of methyl [3H]-SAM, 200 µM theobromine, 200 µM MgCl2, and 5 µM caffeine. Increase the volume to 200 µL with 100 mM Tris-HCl at a pH of 8.0.
- Keep the microcentrifuge tube on ice and add a volume of the soluble fraction containing 7-9 mg of protein. Then, mix by vortex and incubate for 30 min at 30 °C in a thermostatic bath/circulator. For a batch of several samples, incubate each sample in duplicate in 1 min periods to give enough time to stop the reactions for all samples in an exact time period of 30 min.
- Add 1 mL of chloroform to stop the reaction and then shake the sample with a vortex mixer. Shake the sample using a vortex mixer for 3 min to remove the radioactive caffeine in the solvent.
- Centrifuge samples at 11,000 x g for 5 min and carefully recover a volume of 900 µL. Then, transfer the samples to scintillation vials.
- Evaporate the chloroform to complete dryness in the hood at room temperature at 25 °C. Add 5 mL of scintillation fluid to the vial and mix the sample.
- Analyze the radioactivity incorporated into the caffeine in a scintillation counter.
Representative Results
Caffeine extracts obtained via the process presented here were analyzed by TLC-densitometry by subjecting the samples to plate chromatography according to the scheme shown in Figure 1. To quantify the levels of caffeine in the cell extracts, a curve with various concentrations of commercial standard for this compound was used (Figure 2A). The pattern of absorbance for caffeine was analyzed in the visible light spectrum (UV-VIS) using a maximum absorption peak at 273 nm (Figure 2B), and the peak separation of caffeine on the TLC plate showed a Rf between 0.34 and 0.39 (Figure 2C).
The activity of caffeine synthase evaluated using this protocol indicated a specific activity for this enzyme of 1.2 pkat (Table 1). This enzyme's activity was similar to that reported by other authors who used purified protein.
The first step before evaluating the CCS1 gene expression was the isolation of high quality RNA. The quality of the extract of the total RNA derived from cell suspensions was evaluated, and the separation of 28S, 18S and 5S rRNA subunits was demonstrated, indicating that the RNA extracted from samples was of high quality (Figure 3A). However, these RNA extracts had to be treated with DNase enzyme to remove genomic DNA contamination (Figure 3B), which interferes with the preparation of cDNA. Subsequently, qPCR was performed using the protocol described above, and the result of the melt curve showed a single amplification product for the caffeine synthase gene (Figure 4).
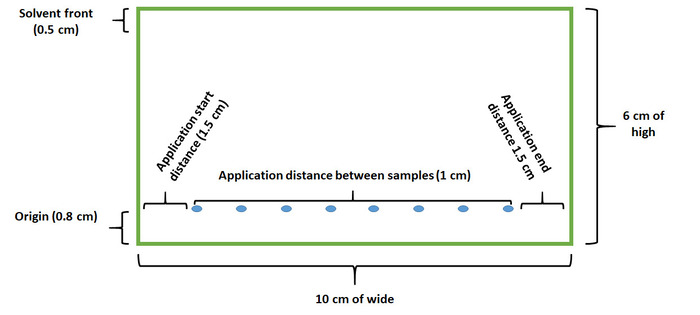
Figure 1. Scheme used for application of the caffeine extract samples on a TLC plate. The scheme shows a silica plate dimension (6 x 10 cm) that is used to apply up to eight samples including caffeine extracts or standards. Consider a distance of 1 cm between samples in the TLC plate to obtain better resolution of the caffeine band separation. Please click here to view a larger version of this figure.
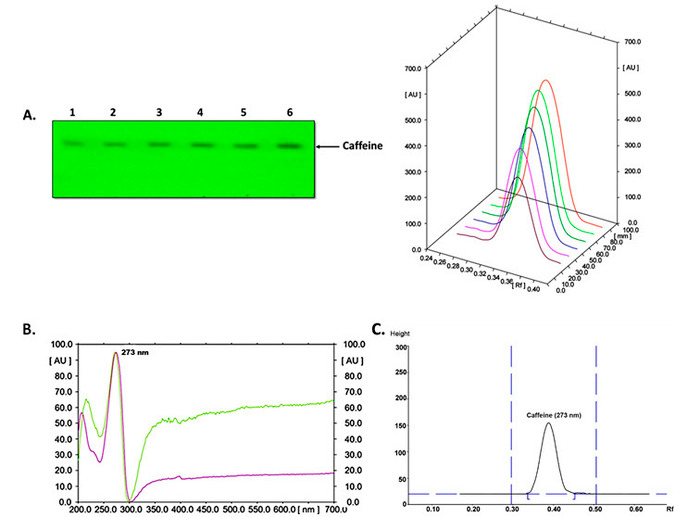
Figure 2. Detection of caffeine by TLC-densitometry in cell extracts of C. arabica L. (A) Separation of various caffeine standard concentrations (lane 1, 0.4; 2, 0.6; 3, 0.8; 4, 1; 5, 1.2; and 6, 1.4 µg) and separation of their peaks in the image on the right. (B) Absorbance patterns of a caffeine standard (purple line) and caffeine extract (green line) at different wavelengths using absorption spectra (ultraviolet and visible). (C) Peak separation of caffeine in the cell extract. Please click here to view a larger version of this figure.
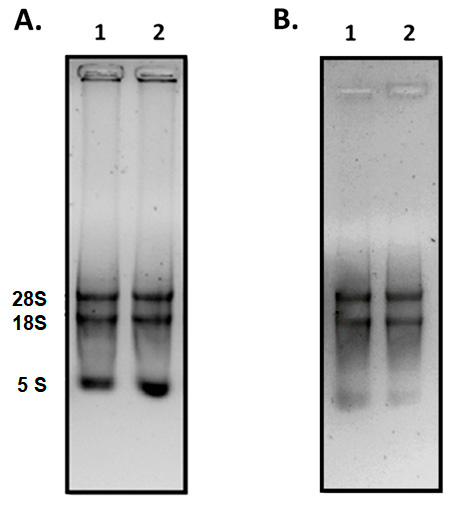
Figure 3. Extraction of the total RNA. The RNA extracts of cellular suspensions of C. arabica L. were analyzed by electrophoresis. (A) RNA extraction, (lanes 1-2 duplicates); (B) RNA samples subjected to treatment with DNase. 1% agarose gels were prepared and stained with 3x intercalating nucleic acid stain solution. Please click here to view a larger version of this figure.
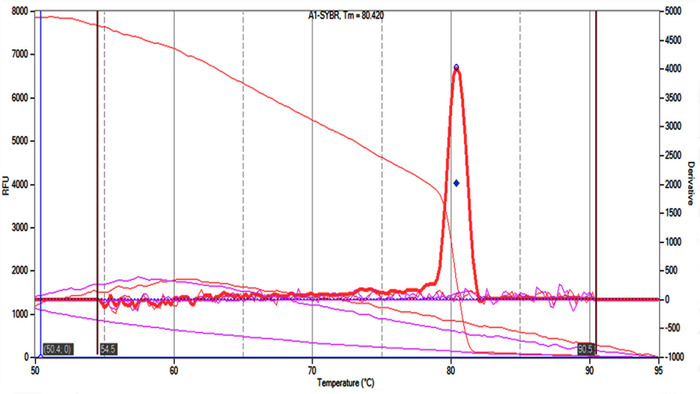
Figure 4. Melting curve produced using qPCR software and RT-qPCR data. The curve marked in red is a single amplified product that corresponds to the caffeine synthase gene. Please click here to view a larger version of this figure.
| Plant | Protein extract | CS specific activity | Reference |
| Coffea | |||
| Cell suspension | Crude | 1.2 pkat | This protocol |
| Endosperm | Purified | 5.7 pkat | 12 |
| Tea | |||
| Leaves | Purified | 11 pkat | 13 |
| Guarana | |||
| Seeds | Purified | 20 fkat | 14 |
Table 1. Comparison of the CS-specific activity in different crops.
Discussion
We present here the optimal conditions for evaluating the caffeine content, CS activity and transcript levels in an in vitro plant tissue culture, such as cell suspensions of C. arabica. Previous reports have confirmed that maintaining cells under light irradiation and in the presence of theobromine in the culture medium are suitable parameters for increasing the level of caffeine, making it possible to evaluate the caffeine separation methods using reversed-phase high-performance liquid chromatography (RP-HPLC). Currently, there are few reports that use a simple and rapid method with economic advantages for studying caffeine biosynthesis in cell suspensions. Here, an alternative assessment showed a separation method for caffeine using TLC-densitometry, which is efficient for the quantitative analysis of caffeine in cell extracts. To evaluate caffeine in the cells, it was not necessary to feed theobromine to the culture cells as reported previously14,15.
Caffeine synthase activity has been evaluated in several plant tissues, including the endosperm, leaves and seeds. In these studies, the methods for the determination of caffeine synthase activity included a step of partial protein purification that suggested that it was necessary to use the purified enzyme16,17. Here, we performed an optimal protocol for the study of the CS enzymatic activity using a soluble protein extract without the need for steps involving the purification of the protein. CS activity in the crude extract of cell suspensions can be measured in units of pkat, as other authors have reported. We evaluated the caffeine levels and enzymatic activity in leaves, and they have similar levels to those in in vitro culture.
Finally, although previous authors have studied the expression of caffeine synthase, several have used differentiated tissue culture6,18. The methodology presented here is useful to obtain an RNA extract and to evaluate the expression levels of the gene CCS1 by RT-qPCR in different in vitro models of plant cell suspensions, such as tea and cocoa.
Offenlegungen
The authors have nothing to disclose.
Acknowledgements
The work of our laboratory was funded by a grant from the Consejo Nacional de Ciencia y Tecnología (CONACyT 219893) to SMTHS. This research was also supported by a fellowship granted to RJPK (No. 37938) by CONACyT and the Sistema Nacional de Investigadores (4422). The authors thank CIATEJ for the use of its installations during the writing of this manuscript. Special thanks are extended to Dr. Víctor Manuel González Mendoza for all recommendations in the molecular biology section and Valentín Mendoza Rodríguez, IFC, UNAM for the facilities during the filming of this article.
Materials
| Murashige & Skoog Basal salt mixture | PhytoTechnology Laboratories | M524 | Packge Size: 50 L Reagent (mg/L) Ammonium Nitrate (1650) Boric acid (6.2) Calcium chloride, anhydrous (322.2) Cobalt Chloride•H2O (0.025) Cupric Sulfate•5H2O (0.025) Na2EDTA•2H2O (37.26) Ferrous Sulfate•7H2O (27.8) Magnesium Sulfate, Anhydrous (180.7) Manganese Sulfate•H2O (16.9) Molybdic Acid (Sodium Salt)• 2H2O (0.25) Potassium Iodide (0.83) Potassium Nitrate (1900) Potassium Phosphate, Monobasic (170) Zinc Sulfate•7H2O (8.6) Supplemented with myo-inositol (100) thiamine (10) cysteine (25) sucrose (30000) 2,4-dichlorophenoxyacetic acid (3) 6-benzylamine purine (1) |
| Caffeine | SIGMA | C0750-5G | STANDARD-5g |
| Theobromine | SIGMA | T4500 | 20 g |
| CAMAG TLC Scanner-4 | CAMAG | 27.62 | |
| WinCATS Planar Chromatography Manager software | CAMAG | 1.4.10 | Software |
| Isoamyl alcohol (24:1) | SIGMA | C-0549 | 500 mL |
| Cyclohexane | JALMEX | C4375-13 | 1 L |
| Acetone | J.T. BAKER | 900643 | 4 L |
| Methanol | J.T. BAKER | 9093-03 | 4 L |
| Chloroform | JALMEX | C-4425-15 | 3.5 L |
| TLC silica gel 60 F254 | Merck | 1.05554.0001 | TLC plate |
| β-mercaptoethanol | M6250 | SIGMA | 100 mL |
| (+)-sodium L- ascorbate | A4034 | SIGMA | 100 g |
| Trizma base | SIGMA | T6066 | 1 Kg |
| Hydrochloric acid 36.5-38% | J.T. Baker | 9535-05 | 2.5 L |
| Pierce BCA Protein Assay Kit | Thermo scientific | 232227 | Kit |
| Methyl [3H]-S-adenosyl methionine | Perkin Elmer | NET155 | Specific activity of 15 Ci/mmol |
| Liquid scintillation vials | SIGMA | Z253081 | |
| Thermostatic bath/circulator | Cole Parmer | 60714 | |
| Micro centrifugue tube | Eppendorf | Tube of 1.5 mL | |
| Cryogenic vials | Heathrow Scientific | HS23202A | 2 mL |
| Centrifuge 5804 | Eppendorf | 5804 000925 | |
| Vortex | Thermolyne | LR 5947 | |
| Porcelain mortar | Fisherbrand | FB961B | |
| Filter paper | Whatman | Z274844 | Porosity medium |
| Picofuge | Stratagene | 400550 | 2000 x g |
| Analytical balance | AND | HR-120 | Model HR-120 |
| Scintillation counter | Beckman Coulter | 6500 | |
| Gel photodocumentation system | Bio-Rad | Chemic XRS | Model Chemic XRS |
| Compact UV lamp | UVP | 95002112 | UVGL-25 |
| Scienceware HDPE Buchner funnel | SIGMA | 2419907 | Type 37600 mixer |
| TRIzol reagent | Thermo scientific | 15596-018 | 200 mL |
| ReverdAid Reverse transcriptase | Thermo scientific | #EP0441 | 10000 U |
| Oligo (dT)18 primer | Thermo scientific | #S0131 | 100 µM |
| DNase I, RNase-free | Thermo scientific | #EN0525 | 1000 U |
| Magnesium chloride | Thermo scientific | EN0525 | 1.25 mL |
| Ethylenediaminetetraacetic acid | Thermo scientific | EN0525 | 1 mL |
| dNTP mix | Thermo scientific | R0191 | R0191 |
| SYBR Green qPCR Master Mix (2X) | Thermo scientific | K0251 | For 200 reactions of 25 µL |
| PikoReal | Thermo scientific | 2.2 | Software |
| Phenol, pH 8.0, equilibrated, Molecular Biology Grade, Ultrapure | USB | J75829 | 100 mL |
| Isopropyl alcohol | Karal | 2040 | 1 L |
| Ethyl alcohol | SIGMA | 64175 | 1 L |
| Diethyl pyrocarbonate | SIGMA | D5758 | 100 mL |
| Lab Rotator | LW Scientific | Mod. LW210 |
Referenzen
- Ferruzzi, M. G. The influence of beverage composition on delivery of phenolic compounds from coffee and tea. Physiolgy & Behavior. 100 (1), 33-41 (2010).
- Majhenič, L., Škerget, M., Knez, &. #. 3. 8. 1. ;. Antioxidant and antimicrobial activity of guarana seed extracts. Food Chemistry. 104 (3), 1258-1268 (2007).
- Sledz, W., Los, E., Paczek, A., Rischka, J., Motyka, A., Zoledowska, S., Lojkowska, E. Antibacterial activity of caffeine against plant pathogenic bacteria. Acta Biochimica Polonica. 62 (3), 605-612 (2015).
- Lipton, R. B., Diener, H. C., Robbins, M. S., Garas, S. Y., Patel, K. Caffeine in the management of patients with headache. Journal of Headache and Pain. 18 (1), 1-11 (2017).
- De Mejia, E. G., Ramirez-Mares, M. V. Impact of caffeine and coffee on our health. Trends in Endocrinology & Metabolism. 25 (10), 489-492 (2014).
- Uefuji, H., Tatsumi, Y., Morimoto, M., Kaothien-Nakayama, P., Ogita, S., Sano, H. Caffeine production in tobacco plants by simultaneous expression of three coffee N-methyltrasferases and its potential as a pest repellant. Plant Molecular Biology. 59 (2), 221-227 (2005).
- Denoeud, F., Carretero-Paulet, L., Dereeper, A., Droc, G., Guyot, R., Pietrella, M., Aury, J. M. The coffee genome provides insight into the convergent evolution of caffeine biosynthesis. Science. 345 (6201), 1181-1184 (2014).
- Kurata, H., Matsumura, S., Furusaki, S. Light irradiation causes physiological and metabolic changes for purine alkaloid production by a Coffea arabica cell suspension culture. Plant Science. 123 (1-2), 197-203 (1997).
- Pech-Kú, R., Muñoz-Sánchez, J. A., Monforte-González, M., Vázquez-Flota, F., Rodas-Junco, B. A., González-Mendoza, V. M., Hernández-Sotomayor, S. T. Relationship between aluminum stress and caffeine biosynthesis in suspension cells of Coffea arabica L. Journal of Inorganic Biochemistry. 181, 177-182 (2018).
- Huang, R., O’Donnell, A. J., Barboline, J. J., Barkman, T. J. Convergent evolution of caffeine in plants by co-option of exapted ancestral enzymes. Proceedings of the National Academy of Sciences. 113 (38), 10613-10618 (2016).
- Mizuno, K., Okuda, A., Kato, M., Yoneyama, N., Tanaka, H., Ashihara, H., Fujimura, T. Isolation of a new dual-functional caffeine synthase gene encoding an enzyme for the conversion of 7-methylxanthine to caffeine from coffee (Coffea arabica L.). FEBS letters. 534 (1-3), 75-81 (2003).
- Kato, M., Mizuno, K., Fujimura, T., Iwama, M., Irie, M., Crozier, A., Ashihara, H. Purification and characterization of caffeine synthase from tea leaves. Plant Physiology. 120 (2), 579-586 (1999).
- Livak, K. J., Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Method. 25 (4), 402-408 (2001).
- Kurata, H., Achioku, T., Furusaki, S. The light/dark cycle operation with an hour-scale period enhances caffeine production by Coffea arabica cells. Enzyme and Microbial Technology. 23 (7-8), 518-523 (1998).
- Sartor, R. M., Mazzafera, P. Caffeine formation by suspension cultures of Coffea dewevrei. Brazilian Archives of Biology Technology. 43 (1), 1-9 (2000).
- Koshiro, Y., Zheng, X. Q., Wang, M. L., Nagai, C., Ashihara, H. Changes in content and biosynthetic activity of caffeine and trigonelline during growth and ripening of Coffea arabica and Coffea canephora fruits. Plant Science. 171 (2), 242-250 (2006).
- Schimpl, F. C., Kiyota, E., Mayer, J. L. S., de Carvalho Gonçalves, J. F., da Silva, J. F., Mazzafera, P. Molecular and biochemical characterization of caffeine synthase and purine alkaloid concentration in guarana fruit. Phytochemistry. 105, 25-36 (2014).
- Perrois, C., Strickler, S. R., Mathieu, G., Lepelley, M., Bedon, L., Michaux, S., Privat, I. Differential regulation of caffeine metabolism in Coffea arabica (Arabica) and Coffea canephora (Robusta). Planta. 241 (1), 179-191 (2015).

