Global Identification of Co-Translational Interaction Networks by Selective Ribosome Profiling
Summary
Co-translational interactions play a crucial role in nascent-chain modifications, targeting, folding, and assembly pathways. Here, we describe Selective Ribosome Profiling, a method for in vivo, direct analysis of these interactions in the model eukaryote Saccharomyces cerevisiae.
Abstract
In recent years, it has become evident that ribosomes not only decode our mRNA but also guide the emergence of the polypeptide chain into the crowded cellular environment. Ribosomes provide the platform for spatially and kinetically controlled binding of membrane-targeting factors, modifying enzymes, and folding chaperones. Even the assembly into high-order oligomeric complexes, as well as protein-protein network formation steps, were recently discovered to be coordinated with synthesis.
Here, we describe Selective Ribosome Profiling, a method developed to capture co-translational interactions in vivo. We will detail the various affinity purification steps required for capturing ribosome-nascent-chain complexes together with co-translational interactors, as well as the mRNA extraction, size exclusion, reverse transcription, deep-sequencing, and big-data analysis steps, required to decipher co-translational interactions in near-codon resolution.
Introduction
Selective Ribosome Profiling (SeRP) is the only method, to date, that captures and characterizes co-translational interactions, in vivo, in a direct manner1,2,3,4,5,6. SeRP enables global profiling of interactions of any factor with translating ribosomes in near codon resolution2,7.
The method relies on flash freezing of growing cells and preserving active translation. Cell lysates are then treated with RNase I to digest all mRNA in the cell except ribosome-protected mRNA fragments termed "ribosome footprints". The sample is then split into two parts; one part is directly used for the isolation of all the cellular ribosomal footprints, representing all ongoing translation in the cell. The second part is used for the affinity-purification of the specific subset of ribosomes associated with a factor of interest, for example: modifying enzymes, translocation factors, folding chaperones, and complex-assembly interactions. The affinity-purified ribosomal footprints are collectively termed the interactome. Then, the ribosome-protected mRNAs are extracted and used for cDNA library generation, followed by deep sequencing.
Comparative analysis of the total translatome and interactome samples allows for the identification of all orfs which associate with the factor of interest, as well as characterization of each orf interaction profile. This profile reports the precise engagement onset and termination sequences from which one can infer the decoded codons and the respective residues of the emerging polypeptide chain, as well as on the ribosome speed variations during the interaction7,8. Figure 1 depicts the protocol as a schematic.
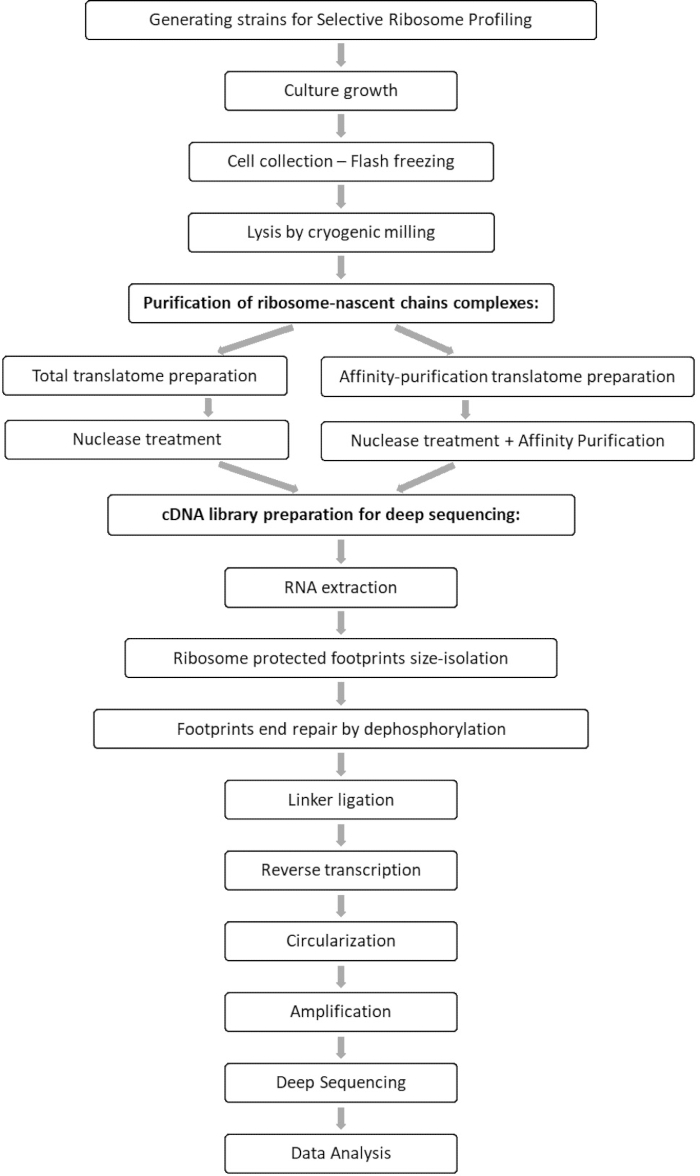
Figure 1: An overview of the SeRP protocol. This protocol can be performed in its entirety within 7-10 days. Please click here to view a larger version of this figure.
Protocol
1. Generating strains for Selective Ribosome Profiling
NOTE: Selective Ribosome Profiling (SeRP) is a method that relies on affinity purification of factors of interest, to assess their mode of interaction with ribosomes-nascent chain complexes. Homologous recombination9, as well as CRISPR/Cas910 based methods are utilized to fuse various factors of interest with tags for affinity purifications. Such tags are GFP, for GFP-trap affinity purifications, TAP-tag for IgG-Sepharose beads purifications as well as AVI-Tag purified by avidin or streptavidin, to list a few successful examples from recent years.
- Perform growth or functional assays to validate that tagging did not impact proteins function. N` versus C` terminal tagging should be evaluated.
NOTE: The ribosomes (rRNA), as well as many ribosome-binding domains in various factors, are highly charged, making highly charged tags (such as polyhistidine) unpreferable to use, since it can lead to false discovery or altered binding mode.
2. Culture growth
- Cultivate the constructed yeast cultures (based on the strain BY4741), containing the desired tagged proteins, in either liquid yeast-extract-peptone-dextrose (YPD)-rich medium, or in synthetic dextrose (SD) minimal medium (1.7 g/L yeast nitrogen base with ammonium sulfate or 1.7 g/L yeast nitrogen base without ammonium sulfate with 1 g/L monosodium glutamic acid, 2% glucose and supplemented with a complete or appropriate mixture of amino acids).
- Grow 250-500 mL of cell culture to a 0.5 OD600 (mid-log), at 30 °C, in an appropriate medium.
3. Cell collection and lysis
- Rapidly collect cells by vacuum filtration on a 0.45 µm nitrocellulose blotting membrane with a glass filtering system (glass filter holder with 1 L glass funnel, vacuum base and cap, stainless steel screen, gasket and spring clamp, 90 mm; ground joint flask 1 L).
- Flash freeze the collected cells, by scraping the pelleted cells with a spatula and immediately immersing them in a liquid nitrogen-filled 50 mL tube.
STOPPING POINT: The cells may be stored at -80 °C for up to 3-4 weeks. - Perform cell lysis by cryogenic grinding in a mixer mill: twice for 2 min at 30 Hz, with 1 mL of the lysis buffer (see Table 1). Chill in liquid nitrogen between millings.
| Reagent | Amount per sample (µL) | Final concentration |
| 10 mg/mL CHX (cycloheximide) | 220 | 0.5 mg/mL |
| 1M Tris-HCl pH 8.0 | 88 | 20 mM |
| 3M KCl | 205.7 | 140 mM |
| 1M MgCl2 | 26.4 | 6 mM |
| 1M PMSF | 4.4 | 1 mM |
| NP-40 | 4.4 | 0.10% |
| Protease inhibitor | 2 tablets | |
| DNase I | 8.8 | 0.02 U/mL |
| Final volume | 4,400 |
Table 1: Recipe for the lysis buffer master mix.
NOTE: Lysis buffer can be altered to contain more protease inhibitors (such as bestatin, leupeptin, aprotinin, etc.) in case the protein of interest is very unstable, but it is important to avoid EDTA in order to maintain the ribosome's small and large subunits assembled during the following steps. For similar reasons, always maintain at least 6 mM MgCl2 in the buffer solution.
CAUTION: HCl is highly corrosive and PMSF is toxic. Wear gloves and handle with care.
- Centrifuge for 2 min at 30,000 x g, 4 °C to clear the lysate and collect supernatant.
4. Purification of ribosome-nascent-chains complexes for SeRP
- For each experiment, divide the supernatant into two parts; each in a different microcentrifuge tube: total RNA sample (~200 µL) and immunopurification (IP) sample (~700 µL) translatome samples.
- Processing the total RNA sample
- Digest total RNA sample using 10 U of RNase I for 25 min at 4 °C; rotate at 30 RPM with a rotating mix rack.
NOTE: Digestion conditions can be calibrated using polysome profiling to ensure no over- or under-digestion of monosomes peak. - Prepare the sucrose cushion master mix as described in Table 2.
- Digest total RNA sample using 10 U of RNase I for 25 min at 4 °C; rotate at 30 RPM with a rotating mix rack.
| Reagent | Amount per sample (µL) | Final concentration |
| 50% Sucrose | 200 | 25% |
| 1M Tris-HCl pH 8.0 | 8 | 20 mM |
| 3M KCl | 18.7 | 140 mM |
| 1M MgCl2 | 4 | 10 mM |
| 100 mg/mL CHX | 0.4 | 0.1 mg/mL |
| Protease inhibitor | 1 tablet | |
| Final volume | 400 |
Table 2: Recipe for sucrose cushion master mix.
- Load the sample onto 400 µL of the sucrose cushion and centrifuge in a TLA120-rotor for 90 min at 245,000 x g and at 4 °C.
- Remove the supernatant quickly with a vacuum pump and overlay pellets with a 150 µL lysis buffer. Resuspend the pellets by shaking for 1 h at 4 °C and at 300 RPM.
- Resuspend the residual pellet by pipetting and transfer to a new 1.5 mL tube.
NOTE: 100-200 µg of total RNA is usually sufficient for ribosome profiling of the total translatome. One can add rRNA depletion step in order to reduce rRNA contamination, which is the most prevalent contaminant of ribosome-nascent chains complexes affinity purifications11 (see discussion for further details).
- Processing the Immunopurification sample
- Wash 100-400 µL of the affinity-binding matrix (1:1 antibody-conjugated beads in 70% EtOH) per sample with 3 x 1 mL lysis buffer (without DNase I and protease inhibitors); resuspend the affinity matrix in the lysis buffer, and then rotate at 30 RPM with a rotating mix rack at 4 °C for 5 min. Precipitate by centrifugation for 30 s at 3,000 x g, 4 °C. Discard the upper liquid. Repeat three times.
- Digest immunopurification samples using 10 U per A260 nm unit of RNase I, together with affinity-binding matrix (for example, 100-400 µL of GFP-TRAP per sample).
- Rotate for 25 min at 30 RPM with a rotating mix rack to bind the protein to the affinity matrix, at 4 °C.
- Prepare the wash buffer master mix as detailed in Table 3.
| Reagent | Amount per sample (µL) | Final concentration |
| 10 mg/mL CHX | 50 | 0.1 mg/mL |
| 1M Tris-HCl pH 8.0 | 100 | 20 mM |
| 3M KCl | 233 | 140 mM |
| 1M MgCl2 | 50 | 10 mM |
| 1M PMSF | 5 | 1 mM |
| NP-40 | 0.5 | 0.01% |
| Protease inhibitor | 2 tablets | |
| 50% Glycerol | 1,000 | 10% |
| Final volume | 5,000 |
Table 3: Recipe for the wash buffer master mix.
- Wash the affinity-binding matrix three times with 1 mL of wash buffer, each time for ~1 min, rotating in the mix rack at 30 RPM, at 4 °C.
- Precipitate by centrifugation at 3,000 x g for 30 s at 4 °C. Discard the upper liquid.
- Wash twice more in 1 mL wash buffer, each time for 5 min, rotating in the mix rack at 30 RPM, at 4 °C.
- Precipitate by centrifugation for 30 s at 3,000 x g and 4 °C.
- Use 50 µL of beads for protein elution with the same amount of 2x sample buffer. Use the rest of the beads for RNA extraction.
- Centrifuge for 30 s at 3,000 x g, 4 °C to pellet the beads and discard the upper liquid.
- Freeze in liquid nitrogen and store at -80 °C. Use these samples for subsequent RNA extraction.
STOPPING POINT: Samples can be stored at -80 °C overnight or longer. This can be a stopping point. - Assess the success of the affinity purification step by western blot or Coomassie staining with aliquots (~10% by volume, after mixing) of each step. Always use mock IP on a non-tagged WT strain as a control for non-specific binding to the affinity matrix.
NOTE: High non-specific background can be overcome by additional washing steps with increasing salt/detergent concentrations. Transient interaction can be stabilized by various cross-linking agents treatment, for example, paraformaldehyde (PFA) treatment of living cells – adding 0.4%-1% PFA to the growth media for 2-5 min, followed by glycine (0.3 M) quenching for 3 min, is highly recommended.
CAUTION: Paraformaldehyde is a suspected carcinogen. Since paraformaldehyde evaporates quickly and is corrosive, work in a chemical safety hood and wear two layers of gloves.
5. cDNA library preparation for deep sequencing
- RNA extraction
NOTE: Work with RNase-free non-stick 1.5 mL tubes to prevent possible RNA or DNA depletion.- Thaw the samples from steps 4.2.5 and 4.3.12 on ice and resuspend samples with 10 mM Tris-HCl pH 7.0 to a final volume of 700 µL.
CAUTION: Acid-phenol and chloroform are volatile and harmful. Work in a chemical safety hood. - Add 40 µL of 20% SDS to 0.7 mL Total RNA or IP elutions. Close and invert a few times. Protein precipitation should turn the samples white.
- Add 0.75 mL of pre-warmed acid-phenol:chloroform to samples. Seal the tubes tightly and shake in a thermal mixer at 1,400 RPM for 5 min and at 65 °C. Chill samples on ice for 5 min.
- Centrifuge the tube from step 5.1.3. at 20,000 x g for 2 min. Transfer the top aqueous layer to a fresh tube and add to it 0.7 mL of acid-phenol:chloroform.
- Incubate for 5 min at room temperature, occasionally vortexing. Centrifuge for 2 min at 20,000 x g. Transfer the top aqueous layer to a fresh tube and add to it 0.6 mL chloroform and vortex.
- Centrifuge for 1 min at 20,000 x g. Transfer the top aqueous layer to a fresh tube.
- Precipitate nucleic acids by adding 78 µL of 3 M NaOAc, pH 5.5, 2 µL of GlycoBlue, and 0.75 mL of isopropanol. Vortex thoroughly for 5 min. Incubate for at least 1 h at -80 °C or 16 h at -20 °C.
- Centrifuge for 30 min at 20,000 x g and at 4 °C and discard the supernatant. Wash the pellets with ice-cold 0.75 mL of 80% ethanol. Invert the tubes for a thorough wash. Centrifuge at 20,000 x g for 5 min at 4 °C, and then discard the supernatant.
- Spin down at 450 x g, 4 °C for 20 s and remove the remaining ethanol and discard the liquids. Dry the pellet with an open lid for 5 min at 65 °C. Resuspend the samples as follows: for IP resuspend the sample in 10 µL of 10 mM Tris-HCl, pH 7.0. For Total translatome analysis, resuspend the sample in 20 µL of 10 mM Tris-HCl, pH 7.0.
STOPPING POINT: RNA can be stored at -80 °C for months.
- Thaw the samples from steps 4.2.5 and 4.3.12 on ice and resuspend samples with 10 mM Tris-HCl pH 7.0 to a final volume of 700 µL.
- Quantify total RNA concentration by fluorometry
NOTE: All the following materials and surfaces should be RNase-free while preparing the cDNA library for next-generation sequencing. While handling RNA samples, wear gloves.- Dilute 1 µL of acid phenol-extracted total RNA in 9 µL of 10 mM Tris-HCl, pH 7.0. Quantify using a fluorometer, as instructed on the manufacturer's website.
- Dilute the samples containing 50 µg of RNA with 10 µL of 10 mM Tris-HCl, pH 7.0.
NOTE: Do not measure IP samples, use everything for the next step.
- Gel-purify ribosome protected footprint fragments
- Set a 15% TBE-urea polyacrylamide gel and submerge in 1x TBE running buffer. Run for 30 min at 200 V prior to sample loading. To each sample, add 20 µL of 2x TBE-urea sample buffer.
NOTE: Expected band size is around 25-35 nt. - Thaw a 10 bp DNA ladder and denature samples (not ladder) at 80 °C for 2 min, and then chill on ice. Load each sample onto every other lane. Run the gel for 50-70 min at 200 V.
- Dilute 6 µL of SYBR Gold (10,000 x concentrate) in 60 mL of 1x TBE buffer and stain while shaking in light-protected boxes for 15-20 min. While staining the gel, prepare a sterile scalpel and 0.5 mL gel-breaker tubes in labeled 1.5 mL tubes.
- Excise the desired bands with a sterile scalpel (use a fresh one or clean well between samples) and place each gel piece in a gel-breaker tube.
- Take an image of the gel to make sure no sample residue is left in the gel.
- Centrifuge the tubes containing cut slices at 20,000 x g for 5 min at 4 °C and transfer the remaining gel pieces from the gel-breaker tube to the 1.5 mL tube.
- Add 0.5 mL of 10 mM Tris, pH 7.0. Shake in a thermal mixer at 1,400 RPM for 10 min at 70 °C.
- Transfer to a cellulose acetate column with a wide bore pipette tip, and centrifuge at 20,000 x g for 3 min at 4 °C.
- Transfer flow-through to a new 1.5 mL tube and chill on ice.
- In order to precipitate the nucleic acids, add: 550 µL of IPA, 55 µL of 3 M NaOAc, and 2 µL of GlycoBlue and vortex to mix thoroughly. Place the samples at -80 °C for at least 1 h.
- Centrifuge for at least 1 h at 20,000 x g and 4 °C and discard the supernatant. Wash the pellets with 0.75 mL of ice-cold 80% ethanol. Invert the tubes for a thorough wash until the pellets separate from the bottom. Centrifuge again at 20,000 x g for 5 min at 4 °C and discard the supernatant.
- Spin down at 450 x g, 4 °C for 20 s and remove the remaining ethanol. Dry the pellets with an open lid for 5 min at 65 °C.
- Add 15 µL of 10 mM Tris, pH 7.0 and resuspend the pellets thoroughly. Spin down at 450 x g, 4 °C for 20 s and transfer the sample to a new 1.5 mL tube.
STOPPING POINT: Purified RNA can be stored at -80 °C for a few months.
- Set a 15% TBE-urea polyacrylamide gel and submerge in 1x TBE running buffer. Run for 30 min at 200 V prior to sample loading. To each sample, add 20 µL of 2x TBE-urea sample buffer.
- Dephosphorylation
- Use 3 µL of the following mix for each sample: Add 1 µL of RNase inhibitor into 2 µL of 10x T4 polynucleotide kinase reaction buffer without ATP. Add 2 µL of T4 polynucleotide kinase to each sample. Pipette gently to mix well and incubate at 37 °C for 2 h, without shaking.
- To inactivate the enzyme, incubate the sample at 75 °C for 10 min and spin down at 450 x g, 4 °C, for 20 s. Add 0.5 mL of 10 mM Tris, pH 7.0.
- To precipitate nucleic acid, add 2 µL of GlycoBlue, 550 µL of IPA, and 55 µL of 3 M NaOAc.
- Vortex to mix thoroughly and chill the samples at -80 °C for at least 1 h.
STOPPING POINT: Samples can be stored at -80 °C overnight or longer. - Repeat steps 5.3.11-5.3.13.
STOPPING POINT: Dephosphorylated RNA can be stored at -80 °C for months.
- Quantification using a Bioanalyzer
- Make a 1:4 dilution of each RNA sample by mixing 1 µL of sample and 4 µL of DEPC-treated water.
CAUTION: DEPC is a carcinogen. Wear gloves and work carefully. - Run a Bioanalyzer Small RNA Chip/TapeStation. Follow the manufacturer's protocol.
NOTE: Expected ribosome-protected RNA fragment size is around 28-30 nt.
- Make a 1:4 dilution of each RNA sample by mixing 1 µL of sample and 4 µL of DEPC-treated water.
- Ligate 3' end with Linker-1
- Dilute 5 pmol of small RNA fragments to 10 µL with 10 mM Tris, pH 7.0. Denature samples at 80 °C for 2 min and chill on ice.
- Prepare the master mix as detailed in Table 4 and use 29 µL per sample.
| Reagent | Amount per sample (µL) | Final concentration |
| 50% sterile-filtered PEG 8000 | 16 | 20% |
| DMSO | 4 | 10% |
| 10× T4 RNA ligase 2 buffer | 4 | 1x |
| SUPERase-In RNase Inhibitor | 2 | 2 U |
| 10 mM adenylated linker 3-L1 | 0.1 | 25 µM |
| DEPC-treated water | 2.9 | |
| Final volume | 29 |
Table 4: Recipe for 3' end ligation master mix.
- Add 1 µL of T4 RNA ligase 2 and pipette gently to mix well. Incubate at 23 °C for 2 h.
- In order to precipitate the nucleic acids, add: 550 µL of IPA, 500 µL of 10 mM Tris, pH 7.0, 55 µL of 3 M NaOAc, and 2 µL of GlycoBlue. Vortex to mix thoroughly, and place the samples at -80 °C for 1 h at least.
STOPPING POINT: Store the samples at -80 °C overnight or longer. - Repeat steps 5.3.2-5.3.12.
- Resuspend the pellet in 6 µL of 10 mM Tris, pH 7.0. Spin down at 450 x g, 4 °C for 20 s, and transfer the sample to a new 1.5 mL tube.
STOPPING POINT: Samples can be stored at -80 °C for months.
- Gel purification of 3' linked footprints
- Set a 10% TBE-urea polyacrylamide gel and submerge in 1x TBE running buffer. Run for 30 min at 200 V prior to sample loading. To each sample, add 6 µL of 2x TBE-urea sample buffer.
NOTE: Expected band size is around 71-73 nt. - Repeat steps 5.3.2-5.3.13.
STOPPING POINT: Samples can be stored at -80 °C for months.
- Set a 10% TBE-urea polyacrylamide gel and submerge in 1x TBE running buffer. Run for 30 min at 200 V prior to sample loading. To each sample, add 6 µL of 2x TBE-urea sample buffer.
- Reverse transcribe 3ʹ linked footprint fragments to generate ssDNA
- Prepare a master mix as detailed in Table 5 and use 3 µL per sample.
| Reagent | Amount per sample (µL) | Final concentration |
| 10 mM dNTPs | 1 | 0.5 mM |
| 25 µM Linker L(rt) | 0.5 | 625 nM |
| DEPC-treated water | 1.5 | |
| Final volume | 3 |
Table 5: Recipe for the reverse transcription buffer master mix prior to nucleic acids' denaturation.
- Vortex and spin down the sample.
- Incubate samples at 65 °C for 5 min.
- Chill samples on ice.
- Prepare a master mix as detailed in Table 6 and use 6 µL per sample. Vortex and spin down the sample. Add 1 µL of Superscript III to each sample and pipette gently to mix well and incubate for 30 min at 50 °C.
| Reagent | Amount per sample (µL) | Final concentration |
| 5× FS buffer | 4 | 1x |
| SUPERase-In RNase Inhibitor | 1 | 2 U |
| DTT 0.1 M | 1 | 5 mM |
| Final volume | 6 |
Table 6: Recipe for the reverse transcription buffer master mix after nucleic acids' denaturation.
- Add 2.3 µL of 1 N NaOH, which hydrolyzes RNA and quenches the reverse transcription.
CAUTION: NaOH is highly corrosive. Wear gloves and eye protection. - Incubate for 15 min at 95 °C, until the sample turns pink.
- Set a 10% TBE-urea polyacrylamide gel and submerge in 1x TBE running buffer. Run for 30 min at 200 V prior to sample loading. To each sample, add 23 µL of 2x TBE-urea sample buffer.
NOTE: The expected DNA band size is 115-117 nt. - Repeat steps 5.3.2-5.3.12.
- Resuspend the pellet in 15 µL of 10 mM Tris, pH 8.0. Spin down at 450 x g, 4 °C for 20 s and transfer the sample to a new 1.5 mL tube.
STOPPING POINT: Samples can be stored at -80 °C for months.
- ssDNA circularization
- Prepare the following master mix and load 4 µL per sample, as detailed in Table 7.
| Reagent | Amount per sample (µL) | Final concentration |
| 10× CircLigase II buffer | 2 | 1x |
| 5 M Betaine (optional) | 1 | 0.25 M |
| 50 mM MnCl2 | 1 | 2.5 mM |
| Final volume | 4 |
Table 7: Recipe for ssDNA circularization master mix.
- Add 1 µL of CircLigase II ssDNA ligase to each sample and incubate for 1 h at 60 °C.
NOTE: The efficiency of this step can be increased by adding 1 µL of CircLigase II ssDNA ligase to each sample after 1 h incubation. - Inactivate the enzyme by incubating at 80 °C for 10 min.
- Chill on ice and continue to PCR amplification or store at -80 °C.
STOPPING POINT: Samples can be stored at -80 °C for years.
- PCR amplification
- Prepare the following PCR master mix and load 82 µL per sample, as detailed in Table 8.
| Reagent | Amount per sample (µL) | Final concentration |
| DEPC-treated water | 61.6 | |
| 5× Phusion HF reaction buffer | 17.6 | 1x |
| 10 mM dNTPs | 1.8 | 200 µM |
| 100 µM PCR forward primer | 0.2 | 225 nM |
| HF Phusion polymerase | 0.8 | 1.6 U |
| Final volume | 82 |
Table 8: Recipe for PCR amplification master mix.
- To each tube containing master mix, add 5 µL of circularized DNA.
NOTE: Store the rest of the circularized DNA samples at -80 °C. - Add a different 1 µL of 20 µM PCR reverse barcode primer to each sample (see Table 9) and vortex to mix thoroughly.
- Aliquot each tube into four separate PCR tubes, each will be used for a different number of PCR cycles.
- Run a PCR reaction according to the following program, as detailed in Table 10.
| Cycle | Denature (98 °C) | Anneal (60 °C) | Extend (72 °C) |
| 1 | 30 s | ||
| 2-16 | 10 s | 10 s | 5 s |
Table 10: PCR program for PCR reaction.
- After cycles 8, 9, 10, and 11 remove PCR tubes (for IP samples, cycles range from 9 to 15) as a first attempt. After each cycle, pause the program, take one aliquot out and put it on ice, and then swiftly resume the program.
NOTE: The number of cycles should be adjusted based on the quantity of circularized DNA in each reaction. Refer to Figure 4 for an example and further clarification. - To each 17 µL reaction, add 3.5 µL of 6x DNA loading dye.
- Thaw a 10 bp DNA ladder.
- For size separation by gel-electrophoresis, submerge 8% TBE polyacrylamide in 1x TBE running buffer and load the samples of each different cycle number into adjacent wells and run the gel for 50 min at 180 V.
- Dilute 6 µL of SYBR Gold (10,000 x concentrate) in 60 mL of 1x TBE buffer and stain while shaking in light-protected boxes for 15-20 min.
- While staining the gel, prepare a sterile scalpel and 0.5 mL gel-breaker tubes in labeled 1.5 mL tubes.
- Take an image of the stained nucleic acids.
- Cut the desired band with an expected band size of 174-176 bp with the sterile scalpel and place the gel slice in the prepared 0.5 mL gel-breaker tube (clean thoroughly in between samples and use RNase inactivating agent, or switch to a new blade).
- Centrifuge the tubes for 5 min at 20,000 x g and 4 °C, and then transfer the remaining gel pieces from the 0.5 mL gel-breaker tube to the 1.5 mL tube.
- Add 500 µL of 10 mM Tris, pH 8.0 and shake in a thermal mixer at 1,400 RPM for 10 min and 70 °C.
- Transfer the dissolved gel to a cellulose acetate column with a wide bore pipette tip.
- Centrifuge the column for 3 min at 20,000 x g and 4 °C and transfer the flow-through to a new 1.5 mL tube and chill on ice.
- To precipitate the nucleic acid, add: 550 µL of IPA, 32 µL of 5 M NaCl, 1 µL of 0.5 M EDTA, and 2 µL of GlycoBlue and vortex to mix thoroughly.
- Keep the samples for at least 1 h at -80 °C, or -20 °C overnight.
STOPPING POINT: Samples can be stored at -80 °C overnight or longer. - Repeat steps 5.3.11-5.3.12.
- Resuspend in 11 µL of 10 mM Tris, pH 8.0. Spin down at 450 x g, 4 °C for 20 s and transfer the sample to a new 1.5 mL tube.
STOPPING POINT: Samples can be stored at -80 °C for years.
- Quantify size distribution by Bioanalyzer
- Make a 1:4 dilution of each sample by mixing 1 µL of sample with 4 µL of DEPC-treated water.
- Run the Bioanalyzer Small RNA Chip. Follow the manufacturer's protocol.
NOTE: The expected length is 175 ± 5 bp.
- Quantify DNA concentration by fluorometer
- Perform a dsDNA high-sensitivity concentration check with a fluorometer according to the manufacturer's recommendations.
- Multiplex and sequence samples according to the Illumina recommendations (Index Adapters Pooling Guide12).
6. Data analysis
- Perform analysis as detailed in the supplementary file.
Representative Results
As illustrated in the flow chart of this protocol (Figure 1), cells were grown to log phase, and then collected swiftly by filtration and lysed by cryogenic grinding. The lysate was then divided into two: one for total ribosome-protected mRNA footprints and the other for selected ribosome-protected mRNA footprints, on which we performed affinity purification to pull-down the tagged protein-ribosome-nascent chains complexes. We ensured tagged protein expression and the success of the pull-down by western blot analysis, as can be seen in Figure 2. We validated the isolation of ribosome-protected footprints, which are typically 20-45 nt long by small RNA electrophoresis (2100 BioAnalyzer system), allowing for 5-10 nt shift in size detection, according to the system manual (Figure 3). Then, we generated a cDNA library for deep sequencing and big-data analysis. While generating the cDNA library, note that under-cycling can lead to low yield (as can be seen in lane 2 in Figure 3), but re-amplification is possible in order to recover the generated library. Over-cycling may occur when PCR primers are depleted but the reaction continues. When dNTPs are still present, the reaction proceeds, generating longer PCR artifacts with chimeric sequences due to PCR products priming themselves13 (as can be seen in lanes 3-4 in Figure 3, indicated by the visible smear). If the dNTPs' concentration also becomes limiting, products indicating the presence of heteroduplexes composed of only partially homologous library fragments can appear. Figure 4 acts as a reference, with lane 2 representing optimal amplification, and lane 3 an acceptable amplification. Samples from lanes 4 and 5 (cycles 10 and 11) should not be used due to the possibility of introducing PCR duplicates and artifacts. The generated library was further validated by high sensitivity DNA electrophoresis (the same BioAnalyzer system was used) for exact size distribution and quantification (Figure 5). After 3' end linker ligation, reverse transcription and PCR amplification, a cDNA length distribution as such is expected, with a sharp peak around 175 nt.
We trimmed and removed adapters and barcodes from the sequenced library, and only the reads between 20 and 45 nt were selected for further analysis. Figure 6 shows the resulting length distribution. The reads were divided into different groups of: coding sequences, introns, and intergenic sequences (Figure 7), and further classified as shown in Figure 8.
Final analysis for detection and characterization of co-translational interactions was performed based on the enrichment of ribosome-protected mRNA fragments, producing the graphs in Figure 9. We compared the normalized ribosome occupancy (at each nucleotide along each orf) of the total translatome to its corresponding selected translatome (nascent-interactome). Per nucleotide comparison eliminates translation rates artifacts. Reproducibility between biological replicates was evaluated by Pearson correlation (threshold > 0.6). We present Selective Ribosome Profiles, analyzing co-translational interactions of Vma2p with three proteins: the ribosome-associated chaperone Ssb1p, Pfk2p (Phosphofructokinase) and Fas1p (Fatty acid synthase), with each protein C` terminally tagged by GFP. We performed the protocol in biological replicates. Figure 9 A, D and G shows the experimental scheme of each affinity purification. We next show the ribosome occupancy of total translatomes compared to Ssb1 interactomes along the Vma2p orf, encoding for a subunit of the vacuolar H+-ATPase (Figure 9 B, E and H). Finally, we performed ratio-based ribosome-enrichment profiling (IP/Total) at each ribosome position in [codon/aa] along the orf (Figure 9 C, F and I). Comparing the co-translational interactions of these three proteins with Vma2p, which is being synthesized by the ribosome, revealed that Ssb1 chaperone engages the nascent Vma2p at four different regions along the orf, as we identified four significant enrichment peaks by SeRP. Differently, Pfk2p shows only one significant enrichment peak, as identified by SeRP, in a different position compared to the co-translational chaperone Ssb1. Analysis of Fas1 co-translational interactions with nascent Vma2p did not detect any significant enrichment. Thus, the comparison of these ratio-based enrichment ribosome profiles demonstrates this protocol`s power in detection and characterization of various co-translational interactions in near codon-resolution.
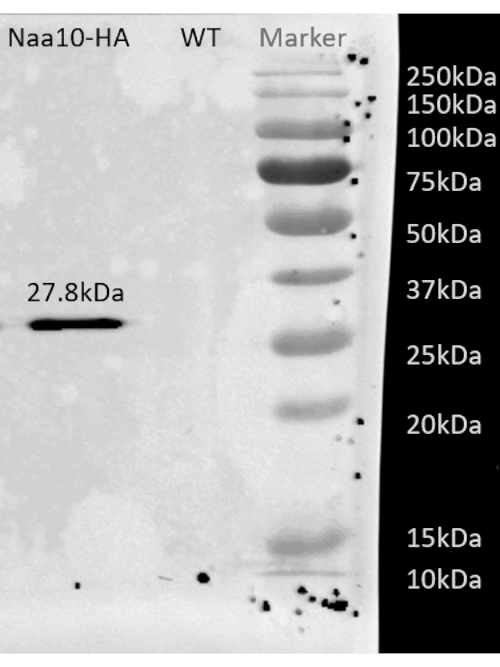
Figure 2: Representative western blot result after affinity purification of BY4741 strain with HA-tagged Naa10. Representative western blot result after affinity purification of BY4741 strain with HA-tagged Naa10 showing a band around 27.8 kDa, while the wild-type, as a negative control, shows no band. Please click here to view a larger version of this figure.
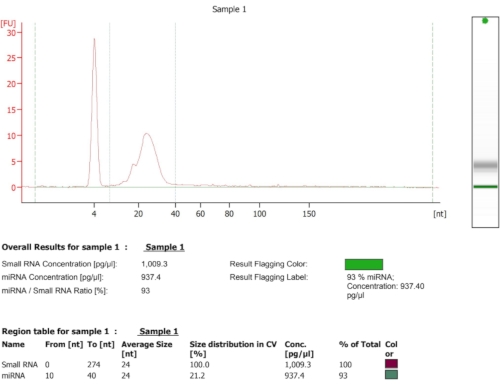
Figure 3: Representative BioAnalyzer result after footprint isolation and RNA extraction with acid-phenol:chloroform, and an average size of 25 nt. Please click here to view a larger version of this figure.
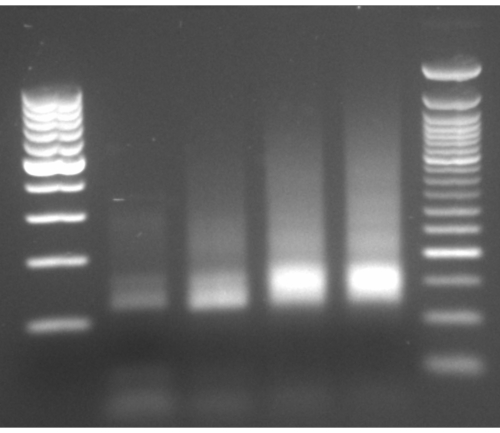
Figure 4: Representative gel-electrophoresis of PCR amplification. Representative gel-electrophoresis of PCR amplification with lanes 2-5 loaded with PCR products from cycles 8-11, and ladders on both sides. Please click here to view a larger version of this figure.
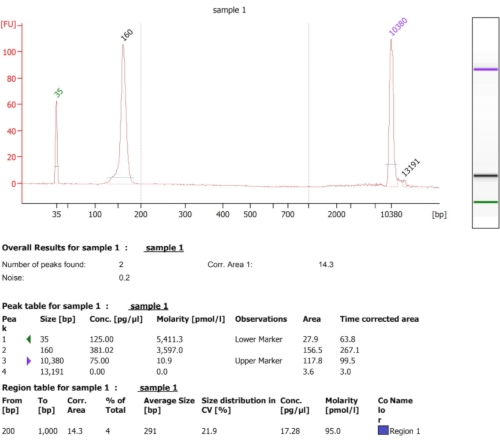
Figure 5: Representative BioAnalyzer result obtained following the creation of a cDNA library. Please click here to view a larger version of this figure.
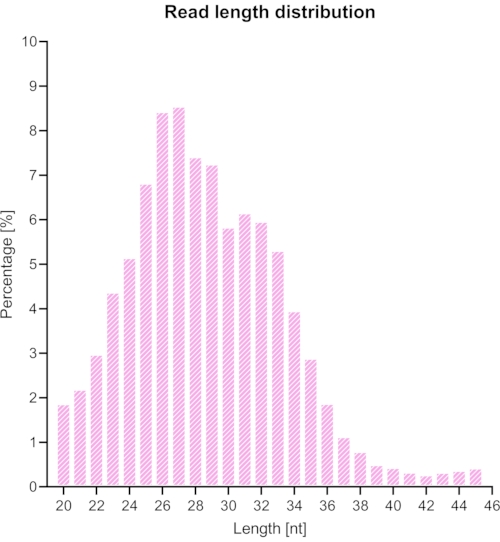
Figure 6: Expected length distribution of reads after the removal of the adaptors with Cutadapt (removing reads shorter than 20 or longer than 45). Please click here to view a larger version of this figure.
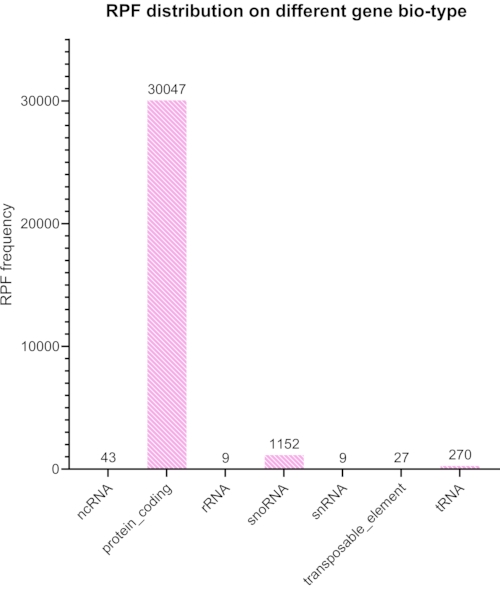
Figure 7: Expected alignment success percentage after removing non-coding RNA reads with Bowtie2 and using TopHat to align the remaining reads to different organisms. The sample was taken from S. cerevisiae (a mutated variant of BY4741). Please click here to view a larger version of this figure.
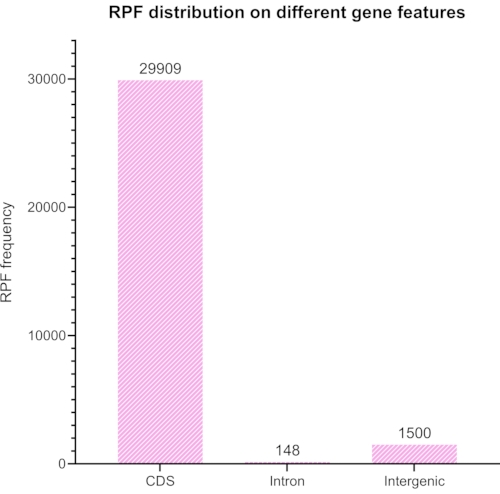
Figure 8: A graph generated with RiboToolkit representing expected coding versus non-coding ratio of aligned reads after using Bowtie2 to remove rRNA elements in the reads. Please click here to view a larger version of this figure.
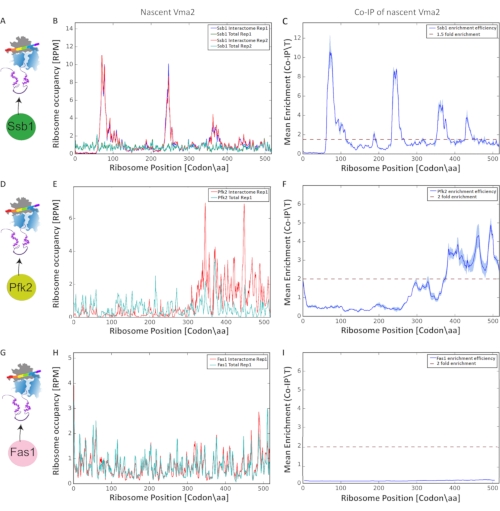
Figure 9: Co-translational interactions of three different proteins: Ssb1p, Pfk2p, and Fas1p with Vma2p, which is being synthesized by the ribosome, analyzed by SeRP. All y axes are shown in reads per million (RPM) reads.(A, D, G) Experimental scheme of SeRP of Ssb1p, Pfk2p, and Fas1p C` terminally tagged by GFP, respectively. (B, E, H) Ribosome occupancy along the orf of total translatomes compared to Ssb1, Pfk2p, and Fas1p interactomes, respectively (in biological replicates). (C, F, I) Mean enrichment of Ssb1p, Pfk2p, and Fas1p (IP/Total ratio) at each ribosome position in [codon/aa] along the orf, respectively. Variation between biological replicates is indicated by the shaded area. Please click here to view a larger version of this figure.
Table 9: 3' Linker and primer sequences. 3' Linker L1: Linker 3-L1 with 5ʹ adenylation and 3ʹ dideoxy-cytidine unique molecular identifiers ('NN…') (RNase-free HPLC purification; Reverse transcription linker: reverse transcription (L(rt)) with 5ʹ phosphorylated, unique molecular identifiers (RNase-free HPLC purification); PCR forward primer: PCRf; HPLC purified. Please click here to download this Table.
Supplementary File. Please click here to download this File.
Discussion
Here, the protocol details the Selective Ribosome Profiling approach for capturing co-translational interactions in near codon resolution. As the ribosome rises as a hub for coordinating the nascent-chain emergence into the crowded cytoplasm, this is a crucial method to identify and characterize the various co-translational interactions required to ensure a functional proteome, as well as for studying various diseases. To date, SeRP is the only method that can capture and characterize these interactions, in a direct manner, in vivo14,15,16.
The first and most critical step is cell collection and lysis. It is imperative to capture, within seconds, ongoing translation, by flash-freezing followed by lysis in a frozen state. Cells collection must be done with haste in order to avoid ribosomal runoff as well as inducing stress translational responses, which can occur rapidly. The second critical step is the affinity purification step. It is imperative to reduce background binding by stringent washing while making sure the co-translational interactions are maintained, which can be facilitated by in vivo cross-linking. As this protocol is based on the highly sensitive NGS (Next Generation Sequencing) high background in the first steps can be amplified in the following cDNA library preparation steps, leading to low signal-to-noise ratios.
Nuclease treatment, to digest all non-protected mRNAs should be evaluated by polysome profiling17 together with careful evaluation of the isolated ribosomal footprints size distribution (as detailed above) to avoid over or under RNA digestion. Calibration of Nuclease concentration and digestion times can facilitate accurate footprint recovery, as over-digestion can lead to ribosomal rRNA digestion, leading to loss of ribosome-protected footprints. It is important to note that under-digestion can also lead to lower discovery rates of ribosome-protected footprints, as the cDNA library preparation steps, as well as data analysis steps described here discard long, uncharacteristic reads.
While rRNA depletion not always constitutes a critical step and is not mandatory, it has some advantages such as cleaner samples and, therefore, a higher rate of genome-mapped reads. On the other hand, there is the possibility of biases, as many rRNA depletion protocols might also cause depletion of the desired ribosome-protected fragments. One should also take into consideration the costs of the rRNA depletion kits. rRNA depletion can be performed after the ribosome-foot print isolation step or after the cDNA circularization step.
cDNA library preparation steps as described here, have been optimized for low mRNA input, as the affinity and ribosome purification steps highly reduce the mRNA input amount, as compared to RNA-seq expression studies. Upscaling the initial amount of cell cultures can greatly facilitate cDNA library generation. Alternatively, any cDNA library protocol of choice can fit with the affinity purification and footprint isolation steps described here. It is important to note that the Nuclease treatment generating the ribosomal footprints requires the resulting mRNA ends repair (cDNA library preparation, Dephosphorylation step) to allow following linker ligation steps in the cDNA protocol described here on in your protocol of choice.
During sequencing, it is important to differentiate SeRP from RNA-Seq, as the generated libraries' heterogeneity greatly varies, depending on the affinity tagged factors. Molecular chaperones and targeting factors are often more promiscuous in binding, interacting with hundreds or thousands of substrates during translation, leading to highly diverse cDNA libraries. However, highly specific interactors, such as co-translational complex assembly interactors can often lead to the generation of much less diverse cDNA libraries. Spike in of diverse and non-diverse libraries on the same lane can greatly improve sequencing and following data analysis results.
Another unique feature of SeRP is its ability to capture local variations in ribosome occupancies along the orf allowing for the discovery of ribosomal shifts in translation rate associated with each set of interactions. It is therefore imperative to compare ribosome occupancies in each codon along the orf to correctly identify enrichment. Utilizing orfs averages can lead to loss of transient interactions or false discovery.
Correct use of the SeRP method opens many co-translational pathways to direct analysis, discovering novel mechanistic features as well as novel ribosome-associated factors, revolutionizing the protein-biosynthesis field.
Offenlegungen
The authors have nothing to disclose.
Acknowledgements
We would like to thank all the lab members for fruitful discussions and Muhammad Makhzumy for the critical reading of the manuscript. This work was funded by ISF (Israeli Science Foundation) grant 2106/20.
Materials
| 3'-Phosphorylated 28 nt RNA control oligonucleotide | IDT | custom order | RNase free HPLC purification; 5'-AUGUAGUCGGAGUCGAGGCGC GACGCGA/3Phos/-3' |
| Absolute ethanol | VWR | 20821 | |
| Acid phenol–chloroform | Ambion | AM9722 | |
| Antibody: mouse monclonal anti-HA | Merck | 11583816001 | 12CA5 |
| Aprotinin | Roth | A162.3 | |
| ATP* | NEB | P0756S | 10 mM |
| Bacto agar | BD | 214030 | |
| Bacto peptone | BD | 211820 | |
| Bacto tryptone | BD | 211699 | |
| Bacto yeast extract | BD | 212720 | |
| Bestatin hydrochloride | Roth | 2937.2 | |
| Chloroform | Merck | 102445 | |
| CircLigase II ssDNA Ligase* | Epicentre | CL9025K | 100 U/μL |
| Colloidal Coomassie staining solution | Roth | 4829 | |
| cOmplete, EDTA-free protease inhibitor cocktail tablets | Roche Diagnostics | 29384100 | |
| Cycloheximide | Biological Industries | A0879 | |
| DEPC treated and sterile filtered water* | Sigma | 95284 | |
| D-Glucose anhydrous | Merck | G5767-500G | |
| Diethylpyrocarbonate | Roth | K028 | |
| Dimethylsulfoxide* | Sigma-Aldrich | 276855 | |
| DNA ladder, 10 bp O'RangeRuler* | Thermo Fisher Scientific | SM1313 | |
| DNA loading dye* | Thermo Fisher Scientific | R0631 | 6× |
| DNase I, recombinant | Roche | 4716728001 | RNAse free |
| dNTP solution set* | NEB | N0446S | |
| EDTA* | Roth | 8043 | |
| Glycerol | VWR | 24388.260. | |
| Glycine solution | Sigma-Aldrich | 67419-1ML-F | 1 M |
| GlycoBlue | Ambion | AM9516 | 15 mg/mL |
| HEPES | Roth | HN78.3 | |
| HF Phusion polymerase* | NEB | M0530L | |
| HK from S. cerevisiae | Sigma-Aldrich | H6380-1.5KU | |
| Hydrochloric acid | AppliChem | A1305 | |
| Isopropanol | Sigma-Aldrich | 33539 | |
| Isopropyl β-D-1-thiogalactopyranoside | Roth | CN08 | |
| Kanamycin | Roth | T832.4 | |
| KCl | Roth | 6781.1 | |
| KH2PO4 | Roth | 3904.1 | |
| Leupeptin | Roth | CN33.4 | |
| Linker L(rt) | IDT | custom order | |
| Liquid nitrogen | |||
| MgCl2 | Roth | KK36.3 | |
| Na2HPO4 | Roth | P030.2 | |
| Na2HPO4·2H2O | Roth | T879.3 | |
| NaCl* | Invitrogen | AM97606 | 5 M |
| NaH2PO4·H2O | Roth | K300.2 | |
| NHS-activated Sepharose 4 fast-flow beads | GE Life Sciences | 17090601 | |
| Nonidet P 40 substitute | Sigma | 74385 | |
| Pepstatin A | Roth | 2936.2 | |
| Phenylmethyl sulfonyl fluoride | Roth | 6367 | |
| Precast gels | Bio-Rad | 5671034 | 10% and 12% |
| RNase I | Ambion | AM2294 | |
| SDS, 20% | Ambion | AM9820 | RNase free |
| Sodium acetate* | Ambion | AM9740 | 3 M, pH 5.5 |
| Sodium azide | Merck | S8032-100G | |
| Sodium chloride | Roth | 9265 | |
| Sodium hydroxide* | Sigma | S2770 | 1 N |
| Sucrose | Sigma-Aldrich | 16104 | |
| SUPERase-In RNase Inhibitor | Ambion | AM2694 | |
| Superscript III Reverse Transciptase* | Invitrogen | 18080-044 | |
| SYBR Gold* | Invitrogen | S11494 | |
| T4 polynucleotide kinase* | NEB | M0201L | |
| T4 RNA ligase 2* | NEB | M0242L | |
| TBE polyacrylamide gel* | Novex | EC6215BOX | 8% |
| TBE–urea polyacrylamide gel* | Novex | EC68752BOX | 10% |
| TBE–urea polyacrylamide gel* | Novex | EC6885BOX | 15% |
| TBE–urea sample buffer* | Novex | LC6876 | 2× |
| Tris | Roth | 4855 | |
| Tris* | Ambion | AM9851 | 1 M, pH 7.0 |
| Tris* | Ambion | AM9856 | 1 M, pH 8.0 |
| UltraPure 10× TBE buffer* | Invitrogen | 15581-044 | |
| * – for library preparation | |||
| gasket and spring clamp , 90 mm, | Millipore | XX1009020 | |
| ground joint flask 1 L , | Millipore | XX1504705 |
Referenzen
- Oh, E., et al. Selective ribosome profiling reveals the cotranslational chaperone action of trigger factor in vivo. Cell. 147 (6), 1295-1308 (2011).
- Shiber, A., et al. Cotranslational assembly of protein complexes in eukaryotes revealed by ribosome profiling. Nature. 561 (7722), 268-272 (2018).
- Becker, A. H., Oh, E., Weissman, J. S., Kramer, G., Bukau, B. Selective ribosome profiling as a tool for studying the interaction of chaperones and targeting factors with nascent polypeptide chains and ribosomes. Nature Protocols. 8 (11), 2212-2239 (2013).
- Galmozzi, C. V., Merker, D., Friedrich, U. A., Döring, K., Kramer, G. Selective ribosome profiling to study interactions of translating ribosomes in yeast. Nature Protocols. , (2019).
- Knorr, A. G., et al. Ribosome-NatA architecture reveals that rRNA expansion segments coordinate N-terminal acetylation. Nature Structural and Molecular Biology. 26 (1), 35-39 (2019).
- Matsuo, Y., Inada, T. The ribosome collision sensor Hel2 functions as preventive quality control in the secretory pathway. Cell Reports. 34 (12), (2021).
- Döring, K., et al. Profiling Ssb-Nascent chain interactions reveals principles of Hsp70-assisted folding. Cell. , (2017).
- Chartron, J. W., Hunt, K. C. L., Frydman, J. Cotranslational signal-independent SRP preloading during membrane targeting. Nature. 536 (7615), 224-228 (2016).
- Janke, C., et al. A versatile toolbox for PCR-based tagging of yeast genes: New fluorescent proteins, more markers and promoter substitution cassettes. Yeast. 21 (11), 947-962 (2004).
- Levi, O., Arava, Y. Expanding the CRISPR/Cas9 Toolbox for Gene Engineering in S. cerevisiae. Current Microbiology. 77 (3), 468-478 (2020).
- Giannoukos, G., et al. Efficient and robust RNA-seq process for cultured bacteria and complex community transcriptomes. Genome Biology. 13 (3), 23 (2012).
- . Illumina Index Adapters – Pooling Guide Available from: https://support.illumina.com/content/dam/illumina-support/documents/documentation/chemistry_documentation/experiment-design/index-adapters-pooling-guide-1000000041074-05.pdf (2019)
- Kanagawa, T. Bias and artifacts in multitemplate polymerase chain reactions (PCR). Journal of Bioscience and Bioengineering. 96 (4), 317-323 (2003).
- Bertolini, M., et al. Interactions between nascent proteins translated by adjacent ribosomes drive homomer assembly. Science. 371 (6524), (2021).
- Kramer, G., Shiber, A., Bukau, B. Mechanisms of cotranslational maturation of newly synthesized proteins. Annual Review of Biochemistry. 88, 337-364 (2019).
- Joazeiro, C. A. P. Mechanisms and functions of ribosome-associated protein quality control. Nature Reviews Molecular Cell Biology. 20 (6), 368-383 (2019).
- Beaupere, C., Chen, R. B., Pelosi, W., Labunskyy, V. M. Genome-wide quantification of translation in budding yeast by ribosome profiling. Journal of Visualized Experiments: JoVE. (130), e56820 (2017).

