Hydrogel Arrays Enable Increased Throughput for Screening Effects of Matrix Components and Therapeutics in 3D Tumor Models
Summary
The present protocol describes an experimental platform to assess the effects of mechanical and biochemical cues on chemotherapeutic responses of patient-derived glioblastoma cells in 3D matrix-mimetic cultures using a custom-made UV illumination device facilitating high-throughput photocrosslinking of hydrogels with tunable mechanical features.
Abstract
Cell-matrix interactions mediate complex physiological processes through biochemical, mechanical, and geometrical cues, influencing pathological changes and therapeutic responses. Accounting for matrix effects earlier in the drug development pipeline is expected to increase the likelihood of clinical success of novel therapeutics. Biomaterial-based strategies recapitulating specific tissue microenvironments in 3D cell culture exist but integrating these with the 2D culture methods primarily used for drug screening has been challenging. Thus, the protocol presented here details the development of methods for 3D culture within miniaturized biomaterial matrices in a multi-well plate format to facilitate integration with existing drug screening pipelines and conventional assays for cell viability. Since the matrix features critical for preserving clinically relevant phenotypes in cultured cells are expected to be highly tissue- and disease-specific, combinatorial screening of matrix parameters will be necessary to identify appropriate conditions for specific applications. The methods described here use a miniaturized culture format to assess cancer cell responses to orthogonal variation of matrix mechanics and ligand presentation. Specifically, this study demonstrates the use of this platform to investigate the effects of matrix parameters on the responses of patient-derived glioblastoma (GBM) cells to chemotherapy.
Introduction
The expected cost of developing a new drug has steadily risen over the past decade, with over $1 billion in current estimates1. Part of this expense is the high failure rate of drugs entering clinical trials. Approximately 12% of drug candidates ultimately earn approval from the United States (US) Food & Drug Administration (FDA) in 2019. Many drugs fail in Phase I due to unanticipated toxicity2, while others that pass safety trials may fail due to a lack of efficacy3. This attrition due to non-efficacy can partly be explained by the fact that cancer models used during drug development are notoriously non-predictive of clinical efficacy4.
Functional disparities between in vitro and in vivo models may be attributed to removing cancer cells from their native microenvironment, including non-tumor cells and the physical ECM5,6. Commonly, research groups use commercially available culture matrices, such as Matrigel (a proteinaceous basement membrane matrix derived from mouse sarcomas) to provide cultured tumor cells with a 3D matrix microenvironment. Compared to 2D culture, 3D culture in membrane matrix has improved the clinical relevance of in vitro results7,8. However, culture biomaterials from decellularized tissues, including the membrane matrix, typically exhibit batch-to-batch variability that may compromise reproducibility9. Furthermore, matrices derived from tumors with different tissue origins from those studied may not provide the appropriate physiological cues10. Finally, cancers with high degrees of intratumoral heterogeneity have microenvironmental features that vary on a submicron-size scale and which the membrane matrix cannot be tuned to recapitulate11.
Glioblastoma (GBM), a uniformly lethal brain tumor with a median survival time of approximately 15 months, is a cancer for which treatment development has been particularly difficult12,13. The current standard of care for GBM consists of primary tumor resection, followed by radiotherapy, and then chemotherapy using temozolomide (TMZ)14. Yet, more than half of clinical GBM tumors exhibit treatment resistance through various mechanisms15,16,17. Predicting the efficacy of a treatment regimen for an individual patient is extremely difficult. Standard preclinical models used to predict individual outcomes consist of patient-derived tumor cells xenografted orthotopically into immunocompromised mice. While patient-derived xenografts can recapitulate many aspects of clinical GBM tumors and are valuable for preclinical models18, they are inherently expensive, low throughput, time-consuming, and involve ethical concerns19. Cultures of patient-derived cells, on 2D plastic surfaces or as spheroids, mostly avoid these issues. While patient-derived cells preserve genetic aberrations, their cultures in 2D or as suspended spheroids have been largely poor representations of patient-derived xenografts in rodents and original patient tumors20. Previously, we, and others, have shown that GBM cells cultured in a 3D ECM that mimics the mechanical and biochemical properties of brain tissue can preserve drug resistance phenotypes10,21,22,23.
Interactions between hyaluronic acid (HA), a polysaccharide abundant in the brain ECM and overexpressed in GBM tumors, and its CD44 receptor modulate the acquisition of drug resistance in vitro21,24,25,26,27. For example, the inclusion of HA within soft, 3D cultures increased the ability of patient-derived GBM cells to acquire therapeutic resistance. This mechano-responsivity was dependent on HA binding to CD44 receptors on GBM cells21. Additionally, integrin binding to RGD-bearing peptides, incorporated into 3D culture matrices, amplified CD44-mediated chemoresistance in a stiffness-dependent manner21. Beyond HA, the expression of several ECM proteins, many containing RGD regions, vary between normal brain and GBM tumors28. For example, one study reported that 28 distinct ECM proteins were upregulated in GBM tumors29. Within this complex tumor matrix microenvironment, cancer cells integrate mechanical and biochemical cues to yield a particular resistance phenotype, which depends on relatively small differences (e.g., less than an order of magnitude) in Young's modulus or density of integrin-binding peptides28,29,30.
The present protocol characterizes how tumor cells interpret unique combinations of matrix cues and identify complex, patient-specific matrix microenvironments that promote treatment resistance (Figure 1A). A photochemical method for generating miniaturized, precisely tuned matrices for 3D culture provides a large, orthogonal variable space. A custom-built array of LEDs, run by a microcontroller, was incorporated to photocrosslink hydrogels within a 384-well plate format to increase automation and reproducibility. Exposure intensity was varied across well to alter micro-mechanical properties of resulting hydrogels, as assessed using atomic force microscopy (AFM). While this manuscript does not focus on constructing the illumination array itself, a circuit diagram (Figure 1B) and parts list (Table of Materials) are provided as aids for device reproduction.
This report demonstrates the rapid generation of an array of GBM cells cultured in unique, 3D microenvironments in which Young's modulus (four levels across a single order of magnitude) and integrin-binding peptide content (derived from four different ECM proteins) were varied orthogonally. The approach was then used to investigate the relative contributions of hydrogel mechanics and ECM-specific integrin engagement on the viability and proliferation of patient-derived GBM cells as they acquire resistance to temozolomide (TMZ) chemotherapy.
Protocol
Patient-derived GBM cell lines (GS122 and GS304) were provided by Professor David Nathanson (our collaborator), who developed these lines under a protocol approved by the UCLA Institutional Review Board (IRB# 10-000655). Cells were provided de-identified so that the cell lines could not be linked back to the individual patients.
1. Preparation of hydrogel solution
- Prepare HEPES-buffered solution by dissolving HEPES powder at 20 mM in Hank's balanced salt solution (HBSS). Adjust pH to 7 following full solvation.
- In the HEPES-buffered solution, dissolve thiolated HA (700 kDa nominal molecular weight, see Table of Materials), prepared following the previous report31, so that 6%-8% of carboxylic acid residues on each glucuronic acid are modified with a thiol, at a concentration of 10 mg/mL in buffer solution.
NOTE: An amber vial is recommended to prevent thiol oxidation by ambient light.- Stir using a magnetic stir plate (<1,000 rpm) at room temperature until fully dissolved, typically around 45 min.
- While HA is dissolving, prepare separate solutions of (1) 100 mg/mL of 8-arm-PEG-Norbornene (20 kDa), (2) 100 mg/mL of 4-arm-PEG-Thiol (20 kDa), (3) 4 mM of cysteine or cysteine-containing peptide (e.g., GCGYGRGDSPG), and (4) 4 mg/mL of LAP in microcentrifuge tubes (see Table of Materials).
- Prepare each of these four solutions in the HEPES-buffered solution prepared in step 1.1. Vortex the solutions to ensure full dissolution of each reagent prior to performing step 4.
NOTE: If testing multiple different peptides, each must contain a cysteine or other source of thiol moiety for this conjugation chemistry. - Prepare solutions (4 mM available thiol) of all peptides to be tethered within a single hydrogel at this point.
NOTE: Peptide sequences and ECM proteins from which they were derived and used in this study are listed in Table 1. N-acetyl cysteine (see Table of Materials), to which cells do not bind, can be substituted for a bioactive, thiol-containing peptide to titrate the concentration of an adhesive peptide or act as a negative control31.
- Prepare each of these four solutions in the HEPES-buffered solution prepared in step 1.1. Vortex the solutions to ensure full dissolution of each reagent prior to performing step 4.
- Mix the individual solutions of HA, PEG-Norbornene, PEG-thiol, and cysteine/thiol-containing peptides (see Table of Materials) to achieve the final concentrations for the final hydrogel matrices listed in Table 2. Stir (<1,000 rpm) on a magnetic stir plate for at least 30 min to mix fully.
NOTE: HA solutions are highly viscous and best handled using a positive displacement pipette (see Table of Materials). If a positive displacement pipette is unavailable, viscous solutions can also be dispensed with a standard micropipette by slowly pipetting using wide-orifice tips.
2. Illumination and photocrosslinking of hydrogels via an LED array
CAUTION: Wear UV protective eyewear and cover the illumination field with UV-absorbing material.
NOTE: The LED array described in this protocol consists of six sets of eight LEDs placed in series, as illustrated by the provided circuit diagram (Figure 1A). Each set of LEDs can be independently powered, which allows for up to six different irradiances per run. Supplementary File 1 contains screenshots corresponding to the following directions for further guidance.
- Download the Illumination Device.zip file from the Supplementary Coding Files. This directory contains the following files: Arduino.zip (Supplementary Coding File 1), Drivers.zip (Supplementary Coding File 2), GUI.zip (Supplementary Coding File 3), and Holder.zip (Supplementary Coding File 4).
NOTE: 3D Print the top and bottom portions for holding the circuit board in place (see Supplementary Coding Files for details). - Download and install the microcontroller software (see Table of Materials).
- Download and install the GUI software (see Table of Materials). Refer to Supplementary File 1 for software operating instructions.
- Open Processing and install the controlIP5 library via clicking on Sketch > Import Library > Add Library. Then, search for controlIP5 in libraries and click on Install. Perform this for the very first time.
- Power the illumination device (see Table of Materials) using the 36 Volt power supply and connect it to a PC using a micro-USB cable.
NOTE: Some devices will not install drivers automatically for various Arduino nano boards. One set of drivers is provided in the device zip file. - Open the Arduino.ino file, located in the Adruino.zip folder, using Arduino IDE.
- Compile the Arduino.ino file by clicking on the Checkmark button. Upload the compiled code by clicking on the Arrow button.
- Open the GUI.pde file, located in the GUI.zip folder, using Processing.
- Click on Run in the processing program to launch the graphical user interface for controlling the illumination device.
- In the graphical user interface window, click on Intensity for the column containing hydrogel precursor solution to be crosslinked and input the desired intensity. Click on the Time box and input desired time. For the solution provided in Table 2, this will be 15 s.
NOTE: End-users need to calibrate digital intensity values to irradiance using a radiometer. Examples of typical intensities are provided in Figure 2A. - Align the samples with the illumination device (Figure 2B) with every other LED in a single column of the silicone molds (see Table of Materials) or 384-well plate. Click on Finish to begin illumination. Repeat this process as necessary for illumination of multiple slides or other wells of a 384-well plate.
NOTE: The holder is designed such that the 384-well plate sits flush with one corner of the inner chamber during illumination.- Following illumination, when placed in one corner, move the well plate to the next corner and repeat. To illuminate wells on the other half of the plate, lift the plate out of the holder and rotate 180°.
- Generate hydrogels with varying mechanics for mechanical characterization following the steps below.
- Clean the glass slides and silicone molds using tape to remove debris. Adhere the silicone molds to the glass slide, press down to ensure a good seal, and displace any air bubbles.
- Pipette 80 µL of hydrogel precursor solution, as prepared in step 1.4, into each silicone mold on the glass slide.
- Place the glass slide onto the illumination device aligned with every other LED in a single column. Expose the hydrogel precursors to UV light for 15 s, as described in step 2, to photocrosslink.
- Once illumination has stopped, retrieve the slides, and loosen the gels from the molds by tracing the inner circumference of the mold with a fine tip (10 µL pipette tip, 30 G needle, etc.). Remove silicone molds with tweezers/forceps.
- Move crosslinked hydrogels into individual wells of a 12-well plate by wetting a spatula and gently pushing them off the glass slide. Fill each well with 2 mL of DPBS (see Table of Materials) prior to adding the hydrogel. Swell the gels in DPBS solution for at least 12 h (typically overnight) at room temperature (for the next day's mechanical characterization).
3. Atomic Force Microscopy (AFM) measurements
- Turn on the atomic force microscope (AFM) according to the manufacturer's instructions (see Table of Materials). This protocol provides brief instructions for using the instrument and the related software.
- Install the AFM probe (see Table of Materials).
NOTE: For the present study, a triangular silicon nitride cantilever with a nominal spring constant of 0.01 N/m was modified with a spherical 2.5 µm silicon dioxide particle. - Following installation, align the laser to the apex of the triangular probe, and then adjust mirror and laser deflection to maximize signal sum (typically between 1.5-2.2 Volts).
- Immerse the probe in DPBS and wait for up to 15 min to obtain thermal equilibrium. Click on the Calibration button and select Contact-Dependent calibration. Click on the Collect Thermal Tuning button, and following data collection, select the peak around 3 kHz for calibration.
NOTE: Slight adjustment of the mirror and laser deflectors may be necessary following immersion into a liquid due to refractive index changes. - Approach the surface of a Petri dish (plastic) by setting the Approach Parameters auf Constant Velocity, a target height of 7.5 µm, and an approach speed of 15 µm/s. Enable Baseline Measurement Per Run for Approach so that the approach runs continuously and does not stop early due to drift in the deflector.
- Upon approach, set acquisition parameters for force mapping to 4 nN turnarounds, 2 µm indentation distance, 1 µm/s velocity, and 0 s contact time. Press the Start button to begin collecting a force curve on the plastic surface (e.g., a well plate).
- Return to the calibration window and select the portion of the force curve corresponding to contact and indentation of the plastic. Accept the calculated sensitivity and stiffness values for the probe to complete calibration.
- Following calibration, raise the AFM probe and place the hydrogel sample for interrogation. Approach hydrogel following the settings provided in step 5.
NOTE: During the approach procedure toward the hydrogel surface, the unit may mistakenly trigger the approached state. To verify the actual approach, obtain a force curve as in step 4.6. Repeat the approach procedure if the resulting curve does not show contact and resulting indentation. - When the surface approach is successful, switch to the Force Mapping mode and set acquisition parameters to a map of 8 x 8 size with 40 µm length per axis. Obtain force maps in various regions to assess the uniformity of stiffness measurements.
- Interpret force curves using the software program JPK SPM Data Processing through a Hertz/Sneddon model fit (Equations 1 and 2, see Table 3 for the definition of all the variables) with the spherical geometry selected32,33,34.
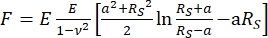 Equation 132
Equation 132
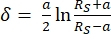 Equation 232
Equation 232
- Interpret force curves using the software program JPK SPM Data Processing through a Hertz/Sneddon model fit (Equations 1 and 2, see Table 3 for the definition of all the variables) with the spherical geometry selected32,33,34.
4. Setting up and drug treatment of 3D, matrix-embedded cultures
- Prepare desired cells as a single cell solution.
NOTE: Different cell types may require different passaging methods. A typical protocol for passaging a suspension culture of GBM spheroids from a T-75 flask is reported in reference31. - Collect GBM spheroids (roughly 150 µm in diameter) from a T-75 flask suspension culture into a 15 mL conical tube. Rinse the culture flask with 5 mL of DPBS to remove any residual cells and media and add this volume to the conical tube.
- Centrifuge the conical tube containing cells at 200 x g for 5 min at room temperature. Following centrifugation, remove the supernatant with a 5 mL serological pipette, taking care not to disturb the cell pellet, and resuspend in 5 mL of DPBS.
- Centrifuge at 200 x g for 5 min at room temperature to wash cells. Aspirate the supernatant with a 5 mL serological pipette, taking care not to disturb the cell pellet, and then resuspend cells in 2 mL of cell dissociation reagent (see Table of Materials).
- Incubate at room temperature for 10-15 min. Add 3 mL of complete medium (see Table of Materials) and gently pipette 3-5 times to break down the spheroids to a single cell suspension31.
- Centrifuge the single-cell suspension at 400 x g (single-cell suspensions may be spun faster for pellet formation) for 5 min to pellet cells at room temperature. Aspirate the supernatant with a 5 mL serological pipette, taking care not to disturb the cell pellet. Resuspend cells in 1 mL of complete medium.
NOTE: If the cells remain in clumps, rather than as single cells in suspension, following passaging, cells can be passed through a 40 µm cell strainer to achieve a single cell suspension. - Remove a portion of the cells for counting using a hemocytometer. Dilute this portion two-fold with trypan blue, which permeates cells with compromised viability. Count only the live, colorless cells. Typically, a T-75 seeded at 800,000 cells per flask yields 2-3 million cells after a week in culture.
- Determine the number of cells necessary for encapsulation. Transfer a volume of media containing the total number of cells needed into a sterile 1.7 mL microcentrifuge tube. Spin down at 400 x g for 5 min at room temperature.
NOTE: For example, a minimum of 2.5 million cells resuspended in 1 mL of gel volume is needed to encapsulate cells at 2.5 million cells/mL. A gel volume of 1 mL allows users to dispense 100 gel drops, where each gel drop is of 10 µL volume. Preparing an extra ~20% volume of cells suspended in hydrogel solution is recommended to account for loss during pipette transfer. Thus, one would prepare 3 million cells and 1.2 mL of hydrogel precursor solution in this example. A minimum density of 500 thousand cells/mL is recommended. - Aspirate the supernatant with a micropipette, taking care not to disturb the cell pellet. Resuspend the cell pellet in the hydrogel precursor solution, as prepared in step 1.4, mixing well by pipetting up and down with a 1,000 µL micropipette 4-5 times.
- Load the cells into a repeat pipettor (see Table of Materials) set to dispense 10 µL. To avoid bubbles and uneven dispensing, prime the repeat pipettor by dispensing an additional 1-2 times into a waste container.
- In each well of a 384-well plate, dispense 10 µL of cells suspended in hydrogel solution from the repeat pipettor. Using the LED array, illuminate each well containing cells (step 2) for 15 s with intensities (example results in Figure 2A utilized intensities of 1.14, 1.55, 2.15, 2.74 mW/cm2) to achieve the desired mechanical properties.
NOTE: It is suggested to start with five replicates per experimental condition and scale up or down depending on the desired throughput and variance of the endpoint assay. - Add 40 µL of complete media to each well containing the cells. Add 50 µL of DPBS to non-experimental, dry wells surrounding the gels to minimize losses due to evaporation.
- For GBM cells, add 40 µL of the media-containing drug (e.g., TMZ, see Table of Materials) to achieve the final desired concentration (10 µM-100 µM in dimethylsulfoxide (DMSO) or vehicle (DMSO), accordingly, starting 3 days after encapsulation.
5. CCK8 proliferation assay
- Add 10 µL of CCK8 reagent (see Table of Materials) to each well containing the cells.
NOTE: If performing this assay for the first time, include negative control wells such as media only or cell-free hydrogel in media. - Incubate for 1-4 h according to the manufacturer's instructions.
NOTE: This time may vary as a function of cell type and density, and thus incubation times need to be tested for each application so that absorbance values fall within a linear range, a requirement for applying Beer's Law35. - Read absorbances at 450 nm for all wells following incubation.
- Calculate the average absorbance at 450 nm obtained in step 3 for the vehicle condition for each group. Divide each drug-treated well by the average of the vehicle control per group.
- Calculate confidence intervals by generating bootstrap distributions (N = 10,000) through the percentile method36.
NOTE: Generally, one may utilize 95% confidence intervals and interpret conditions whose confidence intervals do not cross over 1 to be significant and warrant further investigation. Setting confidence intervals to 95% is congruent with setting a significance cutoff of p = 0.05. For the data shown in the results, there is utility in distinguishing conditions that either promote or inhibit matrix-mediated drug resistance, requiring a two-side analysis.
Representative Results
AFM measurements confirmed precise control of hydrogel mechanics as a function of UV irradiance (mW/cm2) during photo-crosslinking using a custom-built, Arduino-controlled LED array (Figure 2A). The hydrogel formulation used in this protocol can be found in Table 2. The spacing of the LEDs on the provided template matches the spacing for every other well of a 384-well plate, allowing for the formation of gels inside the plate (Figure 2B). AFM interrogation of micron-scale regions at the surfaces of single hydrogels showed that hydrogels with softer average Young's moduli also had smaller ranges of moduli than stiffer hydrogels (Figure 2C–E).
Cell seeding densities that maximize viability should be determined empirically for each cell type. This study demonstrates that 3D cultures of GS122 cells seeded at densities of 2,500,000 cells/mL exhibited substantially higher viabilities when assessed after 7 days in culture compared to those seeded at densities of 500,000 cells/mL (Figure 3A). Furthermore, GS122 and GS304 cells were used as models for culturing patient-derived GBM cells to investigate the dependence of chemotherapy response on the stiffness and biochemical composition of the matrix microenvironment (Figure 3B–D). Cell viability was assessed through the CCK8 assay after treatment with TMZ for 4 days leading to a total culture time of 7 days by scaling OD450 measurement by a corresponding vehicle control and generating 95% confidence intervals by bootstrapping (N = 10,000) with the percentile method36. With these distributions, conditions in which confidence intervals did not overlap with a value of 1 (dashed line) were considered significant. Compared to more commonly used statistical methods such as t-tests or ANOVA, estimation of confidence intervals, using bootstrapping to estimate distributions that would be present for larger sample sizes, is preferred for screening assays whose goal is to identify a smaller subset of conditions for further investigation. One additional benefit of this method is that conditions with a smaller spread in a confidence interval can be prioritized over other conditions with a similar mean value but a higher spread in the confidence interval. This study indicated that GS122 cells gained survival benefits from microenvironmental interactions (Figure 3B). This survival benefit was significant in the 0.8, 1, and 4 kPa conditions but not in the 8 kPa condition for GS122 cells. GS304 cells were insensitive to both stiffness and TMZ treatment.
The effect of the biochemical composition of the matrix microenvironment was then examined by varying the inclusion of ECM-derived, integrin-binding peptides (Table 1) known to be upregulated in the GBM tumor microenvironment at two stiffnesses, 0.8 kPa, and 8 kPa. Again, GS304 cells received no significant survival benefit from matrix inclusion and were insensitive to TMZ. However, GS122 cells showed survival gains in the 8 kPa condition when osteopontin-derived peptides were included in the matrix, while the incorporation of integrin-binding sialoprotein (IBSP)- or tenascin-C-derived peptides provided minimal survival benefits, such as culture in matrices with the general RGD peptide (Figure 3C). In contrast, no peptides conferred survival gains in the 0.8 kPa culture condition (Figure 3D). Together, the results suggest intrinsic differences in both matrix and drug responses between the two patient-derived cell lines evaluated.
3D hydrogel cultures can be visualized using standard light microscopy to assess how cell morphology and invasive behaviors are affected by culture conditions in a cell-line-dependent manner (Figure 4). Both GS122 and GS304 cells spread when cultured in soft or stiff hydrogel matrices including RGD-containing peptides (Figure 4A). While peptides affected cell spreading, the ability of a cell to spread did not necessarily predict the ability of the culture to acquire TMZ resistance. For example, GS122 cells show a similar lack of spreading in both 0.8 and 8 kPa with osteopontin; however, GS122 only showed enhanced resistance to TMZ in the 8 kPa condition (Figure 4B). Finally, this miniaturized, 3D culture platform can be used to culture human cells from other tumor types, including viable organoids of terminally differentiated, neuroendocrine prostate cancer cells (Figure 4C).
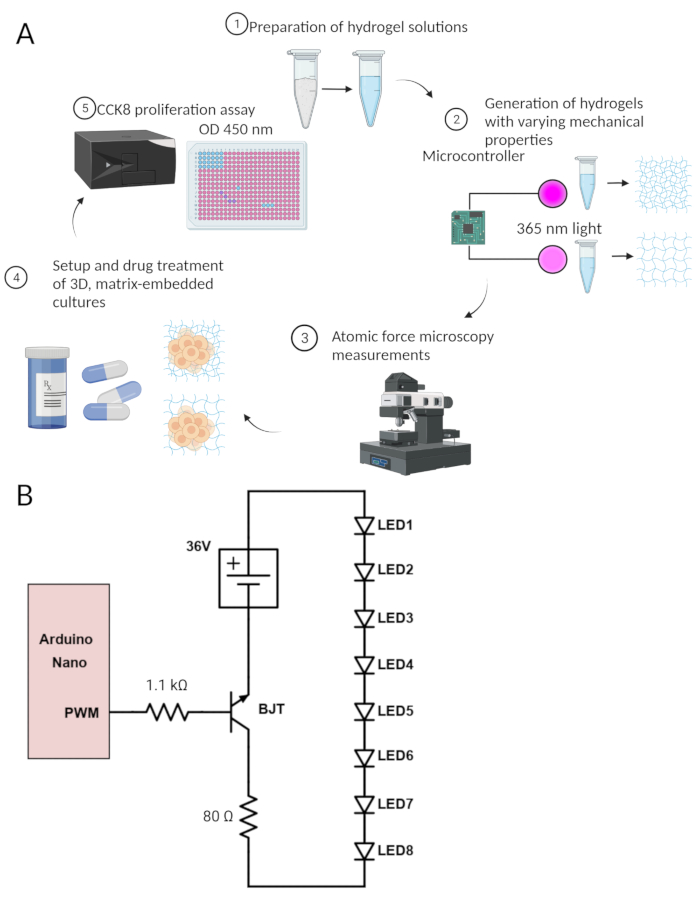
Figure 1: Cartoon depiction of the protocol. (A) Cartoon depiction of the process for 3D culture generation and monitoring. (1) HA-based hydrogel solutions are prepared. (2) Hydrogel solutions are then crosslinked with variable intensity through LEDs controlled by an Arduino microcontroller. (3) Resulting hydrogel mechanics are assessed by AFM to verify the difference in gel mechanics. (4) Solutions matching the formulation from Step 1 are then used to encapsulate patient-derived GBM cells and treated with the drug. (5) Following 7 days, cell viability is read out via CCK8 colorimetric assay. (B) Circuit diagram for custom LED illumination array used in this protocol. The individual components are listed in the Table of Materials. Please click here to view a larger version of this figure.
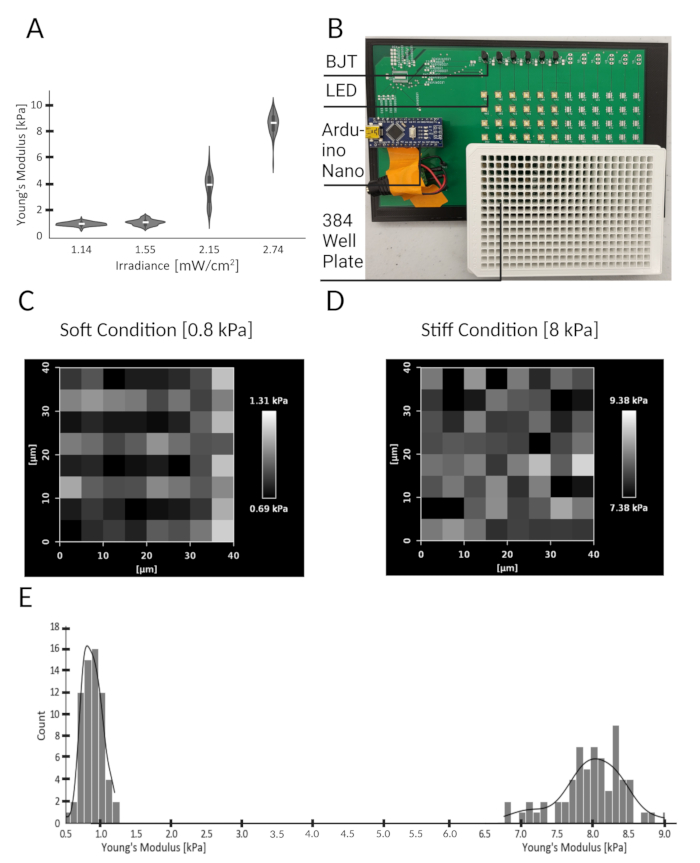
Figure 2: Hydrogels fabricated with varying stiffness using tunable LEDs to modify irradiance. (A) Violin plots show the calculated Young's Modulus from force curves generated by AFM across three surface regions, spanning 40 µm x 40 µm, of individual hydrogels. Young's modulus of each hydrogel is shown as a function of UV irradiance during photocrosslinking. Horizontal white lines indicate the median for each experimental group. (B) LED array with spacing matching the pitch of multi-well plates (384 wells). (C) Heat map showing the regional variation of Young's modulus (mean = 0.8 kPa) for a typical gel crosslinked by exposure to 1.55 mW/cm2 for 15 s. (D) Heat map showing the regional variation of Young's modulus (mean = 8 kPa) for a typical gel crosslinked upon exposure to 2.74 mW/cm2 for 15 s. (E) Histogram illustrating the range of Young's modulus measurements across the surface of hydrogels shown in C and D. Please click here to view a larger version of this figure.
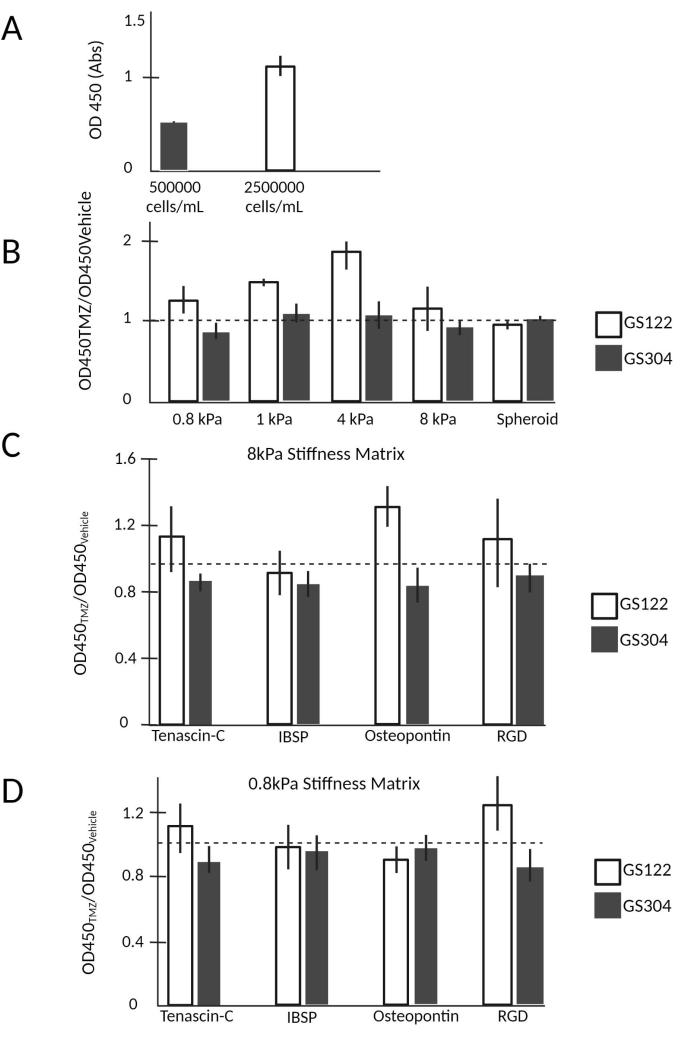
Figure 3: Orthogonal presentation of stiffness and integrin-binding peptide reveals intrinsic biological differences between GBM cell lines. (A) Typical absorbance values for GS122 cells encapsulated in a 10 µL hydrogel at a density of 500,000 or 2,500,00 cells per mL. (B) Drug response data for GS122 and GS304 cell lines are visualized by normalizing the OD450 value of the drug-treated wells (N = 5) by the average of the vehicle-treated wells (N = 5). Viability in the context of drug treatment was observed to vary nonlinearly for hydrogel stiffness, demonstrating variation between cell lines. (C,D) Drug response data for GS122 and GS304 cell lines when the type of integrin-binding peptide was included and matrix stiffness was varied orthogonally. All error bars represent the 95% confidence intervals obtained from bootstrapping each condition by N = 10,000. The dashed line (y-axis = 1) corresponds to the case where OD450 for the treatment conditions equals the vehicle. All experimental values were obtained after 7 days in culture; TMZ was added 3 days after initial encapsulation for drug studies. Please click here to view a larger version of this figure.
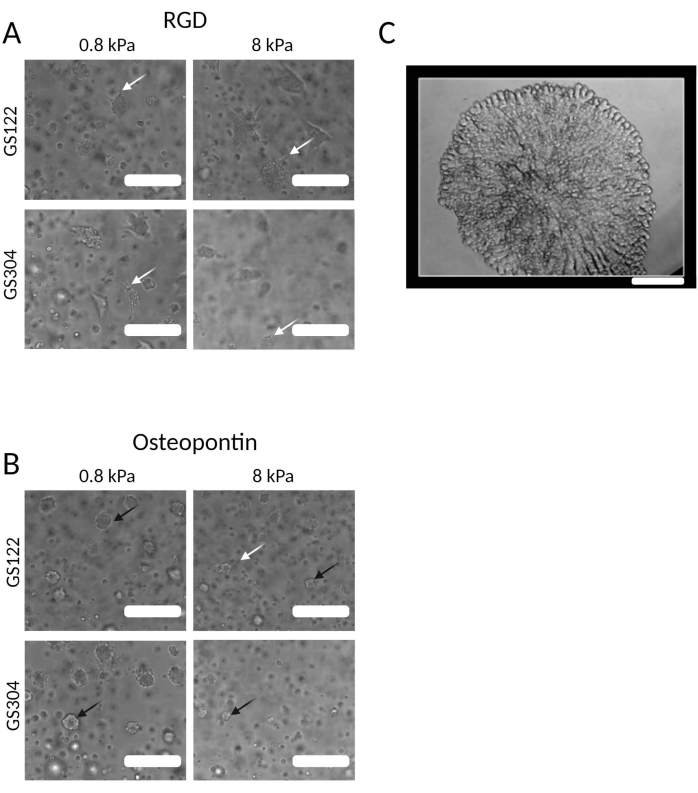
Figure 4: Morphological differences between cells encapsulated in different HA-based hydrogel environments. (A) Phase-contrast images of GS122 and GS304, when cultured in a hydrogel (0.8 and 8 kPa), displayed an RGD motif. White arrows indicate cells with spread morphologies. (B) Phase images of GS122 and GS304, when cultured in a hydrogel (0.8 and 8kPa), displayed a peptide derived from osteopontin. White arrows indicate cells with spread morphologies. Black arrows indicate cells with rounded morphologies. After 7 days in culture, images were taken, and TMZ was added 3 days after initial encapsulation. (C) Phase image of a terminally differentiated neuroendocrine prostate organoid. Scale bars = 200 µm (for A,B); 100 µm (for C). Please click here to view a larger version of this figure.
| Protein | Peptide Sequence |
| RGD | GCGYGRGDSPG |
| Tenascin-C | GCGYGRSTDLPGLKAATHYTITIRGV |
| Integrin-binding sialoprotein (IBSP) | GCGYGGGGNGEPRGDTYRAY |
| Osteopontin | GCGYGTVDVPDGRGDSLAYG |
Table 1: ECM proteins and derived peptide sequences.
| Reagent | Initial Concentration | Volume (µL) | Final Concentration |
| HA-SH Solution | 10 mg/mL | 2300 | 5 mg/mL |
| PEG-SH | 100 mg/mL | 503 | Varies per experiment |
| PEG-Norbornene | 100 mg/mL | 443 | Varies per experiment |
| Peptide | 4 µM | 288 | 0.250 µM |
| LAP | 4 mg/mL | 288 | 0.25 mg/mL |
| HEPES-HBSS | N/A | 798 |
Table 2: Typical final formulation components for hydrogel.
| Variable | Parameter |
| F | Force |
| E | Young’s Modulus |
| ν | Poissons’s Ratio |
| δ | Indentation (vertical tip position) |
| a | Radius of contact circle |
| RS | Radius of Sphere |
Table 3: Variables and corresponding parameters for AFM calculations.
Supplementary File 1: Guidance regarding the use of the LED Array for Illumination and photocrosslinking of hydrogels. Please click here to download this File.
Supplementary Coding File 1: Arduino file for LED array microcontroller (Arduino.zip). Please click here to download this File.
Supplementary Coding File 2: Third-party drivers for Arduino nano (Drivers.zip). Please click here to download this File.
Supplementary Coding File 3: Processing file for GUI control of LED array microcontroller (GUI.zip). Please click here to download this File.
Supplementary Coding File 4: File for 3D printing external holder for LED array microcontroller (Holder.zip). Please click here to download this File.
Discussion
The current work presents methods to generate 3D, miniaturized cultures within HA-based while simultaneously altering matrix stiffness and peptides available for integrin engagement. This technique enables the systematic study of how matrix parameters affect cellular phenotypes (e.g., the viability of cancer cells exposed to chemotherapy) with increased throughput. Previous approaches, including that presented herein, have tuned hydrogel stiffness by varying the percent total polymer in the final formulation, where stiffer hydrogels have a higher polymer content21,31. However, this approach necessitates preparing a unique hydrogel formulation for each stiffness desired, a process that intrinsically lowers throughput. Here, hydrogel stiffness is tuned by varying UV irradiance during crosslinking so that hydrogels of multiple stiffnesses can be obtained from a single precursor solution. Future practitioners who may not have the opportunity to construct the custom illuminator described here can easily substitute a commercially available UV spot-curing device and adjust illumination as different sectors of a well plate are cured. The downside to using a commercial spot-curing device is decreased throughput compared to the custom illuminator. A limitation of the current methods is that unique solutions must still be prepared for each peptide condition. Similar methods have been used previously by other groups to produce stiffness-gradient-containing hydrogels37,38. However, this study demonstrates how photocrosslinking can enable the miniaturization of experimental samples and reduce the complexity of experimental setups. Overall, these improvements will allow researchers to increase experimental throughput when conducting 3D cultures in matrix-mimetic biomaterials and have utility across several fields in biomedical science beyond neuro-oncology.
The representative results demonstrate how this technique can be used to set up miniaturized, 3D cultures of patient-derived GBM cells in defined matrices, in which available integrin-binding peptides and stiffnesses are varied, within a standard 384-well plate appropriate for several standard assays. This method presents representative results to screen how matrix parameters affect responses to TMZ chemotherapy in GBM cells derived from two unique patients, denoted as GS122 and GS304 cells. Of the two patient-derived cell lines evaluated here, the response of GS122 cells to TMZ treatment was sensitive to the matrix microenvironment, while that of GS304 cells was insensitive. Regarding matrix stiffness, GS122 cells cultured in 3D hydrogels with a 4 kPa micro-compressive modulus maximized TMZ resistance, under which condition treated cells increased in number more than untreated cells over a 7-day experimental course. Mechanical properties had larger effects on TMZ resistance than the integrin-binding peptides included. Thus, it is expected that the impact of varying peptide concentrations, alone and in combination, will enable the discovery of matrix conditions promoting drug resistance in the future. The insensitivity of GS304 cells to their surrounding matrix may indicate that the matrix is more influential on cells, like GS122 cells, that are originally sensitive to TMZ yet acquire resistance during treatment. In contrast, the extent to which matrix cues affect cells that are already treatment-resistant at the beginning of the experiment is unclear, as with GS304 cells.
Results from this study deviated somewhat from previously reported results. The softest HA-based hydrogels evaluated (1 kPa bulk Young's modulus, mimicking the stiffness of native brain) promoted maximal TMZ resistance GBM cells derived from four tumors from different patients than that evaluated here21. There are several possible reasons for this discrepancy. First, an expanded range of hydrogel stiffnesses was interrogated in the current study, which compared hydrogels across a range of 0.8 kPa to 8 kPa micro-compressive moduli, while previous studies compared only hydrogels with 1 kPa and 2 kPa Young's moduli. Additionally, in the previous studies, mechanical characterization was done at the bulk scale using linear mechanical compression to estimate Young's modulus, whereas, in this study, AFM was used to measure a micro-compressive modulus. For researchers seeking to implement the methods present here who do not require micro-scale mechanics measurements, bulk scale moduli, using rheometry or linear mechanical testing, are perfectly acceptable substitutes for AFM. Notably, micro-mechanical analyses revealed the heterogeneous moduli across the surface of a single gel. Comparable AFM measurements on the hydrogels formulated in previous studies have not been made, but it is expected that the variance of stiffnesses presented within a single hydrogel to have been greater than those in the current study. Hydrogels in the previous studies were generated using a Michael-type addition crosslinking reliant on kinetic mixing at 37 degrees C10,21,24. In contrast, photocrosslinking permits thorough mixing of hydrogel precursors prior to light exposure, which improves homogeneity within the 3D hydrogel and, in turn, reduces the variability of cell responses. Finally, GBM tumors exhibit notoriously heterogeneous and unpredictable behavior39 and, thus, it is reasonable to expect that cells derived from individual tumors would likewise have unique properties. This lack of consistency across patient samples motivates the need for a high-throughput platform for elucidating patient-specific tumor characteristics.
Common pitfalls when performing the miniaturized hydrogel photocrosslinking procedure include incomplete mixing of hydrogel precursors, resulting in poor reproducibility, and spontaneous gelation of the HA solution while mixing. These issues can be mitigated by stirring the HA-thiol for a minimum of 45 min and the complete precursor solution for at least an additional 30 min while closely monitoring the pH of hydrogel precursor solutions to ensure it remains below 7 to prevent thiol oxidation and formation of disulfide to crosslinks. In contrast to crosslinking methods using a kinetic Michael-type addition mechanism10,21, the photocrosslinking method used in this protocol lowers the probability of spontaneous gelation when all reagents are combined21. Quality control checkpoints are highly recommended, such as measuring HA thiolation percentage31 and making extra hydrogels for parallel mechanical testing to each batch of 3D cultures. Finally, seeding densities for 3D encapsulation in hydrogels need to be identified for each cell type or line used. Generally, the results show that a minimum concentration of 1 million cells/mL is sufficient for the 3D culture of most cells, which form spheroids, including patient-derived GBM cells, human embryonic stem cells (H9), and human-induced pluripotent stem cells (data not shown). The inclusion of ROCK inhibitor treatment prior to encapsulation (step 4.1) when using particularly sensitive cells is also recommended40.
In the context of developing new treatment approaches for GBM, the protocol presented here provides methods for functionally screening drug responses of patient-derived GBM cells within a physiologically relevant microenvironment. A rich repository of genetic, epigenetic, and clinical mRNA expression data for GBM (and other cancers) is publicly available thanks to the efforts of The Cancer Genome Atlas (TCGA) and others15. Used in conjunction with these large datasets, it is expected that data generated from functional screens of miniaturized, 3D cultures can reveal new correlations, improving the prediction of clinical outcomes in individual patients. For example, subpopulations of patient tumors may be identified for which some treatment, like matrix-disrupting compounds such as cilengitide, may improve clinical outcomes41. In addition to drug response, this culture platform enables assessments of tumor cell invasion using well-plate compatible, high-content imagers. The flexibility of this platform to incorporate many different peptides, alone or in combination, while orthogonally varying stiffnesses may help identify matrix features driving GBM tumor aggression.
Beyond GBM, these methods can be adapted to investigate the effects of matrix parameters on other cell types in the context of other diseases, tissue development, and normal tissue function. In the future, it will be straightforward to increase the throughput of these methods further, as they have been specifically designed to be performed in the context of multi-well plates to facilitate adoption into existing workflows for drug discovery by utilizing commercially available automated liquid handlers and high-content imagers. Facile integration with existing infrastructure and increased automation, which decreases the technical skills required to produce and maintain cell-laden, 3D hydrogel cultures, will significantly lower barriers to adopting this method. While the work here specifically presents cultures within HA-based hydrogels, it is expected that this method can be easily translated to other commonly used photocrosslinkable materials for 3D cell culture, such as methacrylated gelatin, and that the methods reported here will provide a helpful guideline for additional applications.
Offenlegungen
The authors have nothing to disclose.
Acknowledgements
The authors would like to specifically acknowledge Carolyn Kim, Amelia Lao, Ryan Stoutamore, and Itay Solomon for their contributions to earlier iterations of the photogelation scheme. Cell lines GS122 and GS304 were generously provided by David Nathanson. All figures were created with BioRender.com. UCLA core facilities, the Molecular Screening Shared Resources, and the Nano and Pico Characterization Laboratory were instrumental to the work. Chen Chia-Chun was supported by the UCLA Eli and Edythe Broad Center of Regenerative Medicine and Stem Cell Research Training Program. Grigor Varuzhanyan was supported by a Tumor Cell Biology Training Program NIH Grant (T32 CA 009056).
Materials
| 1.1 kOhm resistors, 6 W | Digikey | 35601k1ft | |
| 1.7 mL microcentrifuge tube | Genesse Scientific | 21-108 | |
| 15 mL conical tube | Fisher Scientific | 14-959-70C | |
| 365 nm LED | Digikey | ltpl-c034uvh365 | |
| 384 well plate | Bio Greiner One | 781090 | |
| 40 µm cell strainer | MTC bio | C4040 | |
| 4-Armed thiol terminated polyethlene glycol (20 kDa) | Laysan Bio | 4arm-PEG-SH-20K-1g | |
| 6 NPN BJTs | Digikey | 2n5550ta | |
| 80 Ohm resistors, 0.125 W | Digikey | erjj-6enf80r6v | |
| 8-Armed norbornene terminated polyethylene glycol (20 kDa) | Jenkem Technology | A7025-1 | |
| Accutase | Innovative Cell Technologies | AT104500 | cell dissociation reagent |
| AFM Probes | Novascan | 0.01 N/m Nominal spring constant, 2.5 µm SiO2 particle | |
| Arduino IDE | Arduino | 1.8.19 | |
| Arduino Nano | Makerfire | Mini Nano V3.0 ATmega328P Microcontroller Board | |
| bFGF | Peprotech | 100-18B | 20 ng/mL |
| CCK8 | Abcam | ab228554 | |
| Centrifuge | Thermoscientific | sorvall legend xtr | |
| CP100ST | Gilson | F148415 | Pipette tips for positive displacement pipette |
| Cubis Semi-Micro Balance | Sartorius | MSA225S100DI | |
| DMEM – F12 (50-50) | Life Technologies | 11330057 | 1x |
| DMSO | Fisher Scientific | BP231-100 | |
| DPBS Ca (-) Mg (-) | Genesse Scientific | 25-508 | |
| EGF | Peprotech | AF100-15 | 50 ng/mL |
| Ethanol, Anhydrous | Fisher Scientific | A405P | Add DI water to dilute to 70% |
| Fisherbrand Class B Amber Glass threaded vials | Fisher Scientific | 03-339-23C | |
| Fisherbrand Weighing Paper | Fisher Scientific | 09-898-12B | |
| G21 Supplement | Gemini Bio | 400-160 | 50x |
| Hanks Balanced Salt Solution | Thermo Fisher Scientific | 14175095 | |
| HCl, ACS, 12M | Sigma Aldrich | S25838A | Add DI water to dilute to 1 M |
| Heparin sodium salt from porcine intestinal mucosa | Sigma Aldrich | H3149-100Ku | 25 µg/mL |
| HEPES | Sigma Aldrich | H7006-100G | |
| Hot Air Gun | Wagner | HT1000 | |
| Integrin-binding sialoprotein (IBSP) peptide | Genscript | Custom Order | GCGYGGGGNGEPRGDTYRAY |
| Lithium phenyl-2,4,6 trimethylbenzoylphosphinate (LAP) , >95% | Sigma Aldrich | 900889-1G | |
| Magnetic stir plate | Thermo Scientific | SP194715 | |
| Microcentrifuge | Thermo Scientific | Sorvall legend micro 21R | |
| Microman E single Channel Pipettor | Gilson | FD10004 | Positive displacement pipette |
| Micropipette Tips | Various Manufacturs | Various sizes | |
| mLine micropipette | Sartorious | ||
| N-acetyl Cysteine | Sigma Aldrich | A7250-10G | |
| Nanowizard 4 | Bruker | AFM microscope | |
| NaOH | Fisher Scientific | ss255-1 | Add DI water to dilute to 1 M |
| Normoicin | Invivogen | ant-nr-1 | 500x |
| Osteopontin Peptide | Genscript | Custom Order | GCGYGTVDVPDGRGDSLAYG |
| Pipet Aid | Drummond | 4000102 | |
| Plain Microscope Slides | Globe Scientific | 1301 | |
| Press-To-Seal silicone Isolator, 12-4.5mm diam x 2mm deep | Grace Bio Labs | 664201-A | Cut so that 8 individual molds are made from a single sheet |
| Processing | Processing | 3.5.4 | |
| Repeater M4 | Eppendorf | 4982000322 | |
| Repeater Pipette Tips | Sartorious | 30089430 | 1 mL sizes |
| RGD Peptide | Genscript | GCGYGRGDSPG | |
| Scoth Tape | |||
| Serological Pipettes | Genesse Scientific | 12-102,12-104 | 5,10 mL Pipettes |
| Solder Paste | Digikey | 315-NC191LT15T5-ND | |
| Solder Wire | |||
| Straight dissecting forceps | VWR Scientific | 82027-408 | |
| Synergy H1 Plate Reader | Biotek | ||
| T-75 Cell Culture Treated Flask | Genesee Scientific | 25-209 | |
| Temozolomide | Sigma Aldrich | T2577 | Typically used from 10 µM to 100 µM |
| Tenascin-C Peptide | Genscript | GCGYGRSTDLPGLKAATHYTITIR GV |
|
| Thiolated Hyaluronic Acid (700 kDa), 6-8% modified | Lifecore Biomedical | HA700K5 | |
| VWR Spinbar, Flea Micro | VWR | 58948-375 |
Referenzen
- Scannell, J. W., Blanckley, A., Boldon, H., Warrington, B. Diagnosing the decline in pharmaceutical R&D efficiency. Nature Reviews Drug Discovery. 11 (3), 191-200 (2012).
- Waring, M. J., et al. An analysis of the attrition of drug candidates from four major pharmaceutical companies. Nature Reviews Drug Discovery. 14 (7), 475-486 (2015).
- Khozin, S., Liu, K., Jarow, J. P., Pazdur, R. Why do oncology drugs fail to gain US regulatory approval. Nature Reviews Drug Discovery. 14 (7), 450-451 (2015).
- Booth, B., Ma, P., Glassman, R. Oncology’s trials. Market indicators. Nature Reviews Drug Discovery. 2 (8), 609-610 (2003).
- Da Ros, M., et al. Glioblastoma chemoresistance: The double play by microenvironment and blood-brain barrier. International Journal of Molecular Sciences. 19 (10), 2879 (2018).
- Broekman, M. L., et al. Multidimensional communication in the microenvirons of glioblastoma. Nature Reviews Neurology. 14 (8), 482-495 (2018).
- Grundy, T. J., et al. Differential response of patient-derived primary glioblastoma cells to environmental stiffness. Scientific Reports. 6 (1), 1-10 (2016).
- Gomez-Roman, N., Stevenson, K., Gilmour, L., Hamilton, G., Chalmers, A. J. A novel 3D human glioblastoma cell culture system for modeling drug and radiation responses. Neuro-Oncology. 19 (2), 229-241 (2017).
- Simoni, R. D., et al. Basement membrane complexes with biological activity. Biochemie. 25 (2), 312-318 (2002).
- Xiao, W., et al. Brain-mimetic 3D culture platforms allow investigation of cooperative effects of extracellular matrix features on therapeutic resistance in glioblastoma. Krebsforschung. 78 (5), 1358-1370 (2018).
- Aisenbrey, E. A., Murphy, W. L. Synthetic alternatives to Matrigel. Nature Reviews Materials. 5 (7), 539-551 (2020).
- Spinelli, C., et al. Molecular subtypes and differentiation programmes of glioma stem cells as determinants of extracellular vesicle profiles and endothelial cell-stimulating activities. Journal of Extracellular Vesicles. 7 (1), 1490144 (2018).
- Ostrom, Q. T., Cioffi, G., Waite, K., Kruchko, C., Barnholtz-Sloan, J. S. CBTRUS statistical report: Primary brain and other central nervous system tumors diagnosed in the United States in 2014-2018. Neuro-Oncology. 23, (2021).
- Stupp, R., et al. Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. New England Journal of Medicine. 352 (10), 987-996 (2005).
- Brennan, C. W., et al. The somatic genomic landscape of glioblastoma. Cell. 155 (2), 462-477 (2013).
- Tomczak, K., Czerwińska, P., Wiznerowicz, M. The Cancer Genome Atlas (TCGA): An immeasurable source of knowledge. Contemporary oncology. 19, 68-77 (2015).
- Lee, S. Y. Temozolomide resistance in glioblastoma multiforme. Genes and Diseases. 3 (3), 198-210 (2016).
- Joo, K. M., et al. Patient-specific orthotopic glioblastoma xenograft models recapitulate the histopathology and biology of human glioblastomas in situ. Cell Reports. 3 (1), 260-273 (2013).
- Levy, N. The use of animal as models: Ethical considerations. International Journal of Stroke. 7 (5), 440-442 (2012).
- Phon, B. W. S., Kamarudin, M. N. A., Bhuvanendran, S., Radhakrishnan, A. K. Transitioning preclinical glioblastoma models to clinical settings with biomarkers identified in 3D cell-based models: A systematic scoping review. Biomedicine & Pharmacotherapy. 145, 112396 (2022).
- Xiao, W., et al. Bioengineered scaffolds for 3D culture demonstrate extracellular matrix-mediated mechanisms of chemotherapy resistance in glioblastoma. Matrix Biology. 85-86, 128-146 (2020).
- Brancato, V., Oliveira, J. M., Correlo, V. M., Reis, R. L., Kundu, S. C. Could 3D models of cancer enhance drug screening. Biomaterials. 232, 119744 (2020).
- Xu, X., Farach-Carson, M. C., Jia, X. Three-dimensional in vitro tumor models for cancer research and drug evaluation. Biotechnology Advances. 32 (7), 1256-1268 (2014).
- Xiao, W., Ehsanipour, A., Sohrabi, A., Seidlits, S. K. Hyaluronic-acid based hydrogels for 3-dimensional culture of patient-derived Glioblastoma Cells. Journal of Visualized Experiments: JoVE. (138), e58176 (2018).
- Preston, M. Digestion products of the PH20 hyaluronidase inhibit remyelination. Annals of Neurology. 73 (2), 266-280 (2013).
- Kim, Y., Kumar, S. CD44-mediated adhesion to hyaluronic acid contributes to mechanosensing and invasive motility. Molecular Cancer Research. 12 (10), 1416-1429 (2014).
- Pibuel, M. A., Poodts, D., Díaz, M., Hajos, S. E., Lompardía, S. L. The scrambled story between hyaluronan and glioblastoma. The Journal of Biological Chemistry. 296, 100549 (2021).
- Xiao, W., Sohrabi, A., Seidlits, S. K. Integrating the glioblastoma microenvironment into engineered experimental models. Future Science OA. 3 (3), (2017).
- Trombetta-Lima, M., et al. Extracellular matrix proteome remodeling in human glioblastoma and medulloblastoma. Journal of Proteome Research. 20 (10), 4693-4707 (2021).
- Schregel, K., et al. Characterization of glioblastoma in an orthotopic mouse model with magnetic resonance elastography. NMR in Biomedicine. 31 (10), 3840 (2018).
- Xiao, W., Ehsanipour, A., Sohrabi, A., Seidlits, S. K. Hyaluronic-acid based hydrogels for 3-dimensional culture of patient-derived glioblastoma cells. Journal of Visualized Experiments: JoVE. (138), e58176 (2018).
- Guz, N., Dokukin, M., Kalaparthi, V., Sokolov, I. If cell mechanics can be described by elastic modulus: Study of different models and probes used in indentation experiments. Biophysical Journal. 107 (3), 564-575 (2014).
- Sneddon, I. N. The relation between load and penetration in the axisymmetric boussinesq problem for a punch of arbitrary profile. International Journal of Engineering Science. 3 (1), 47-57 (1965).
- Soofi, S. S., Last, J. A., Liliensiek, S. J., Nealey, P. F., Murphy, C. J. The elastic modulus of MatrigelTM as determined by atomic force microscopy. Journal of Structural Biology. 167 (3), 216-219 (2009).
- Mayerhöfer, T. G., Popp, J. Beer’s law – Why absorbance depends (almost) linearly on concentration. Chemphyschem: A European Journal of Chemical Physics and Physical Chemistry. 20 (4), 511-515 (2019).
- Puth, M. T., Neuhäuser, M., Ruxton, G. D. On the variety of methods for calculating confidence intervals by bootstrapping. Journal of Animal Ecology. 84 (4), 892-897 (2015).
- Lavrentieva, A. Gradient hydrogels. Advances in Biochemical Engineering/Biotechnology. 178, 227-251 (2020).
- Zhu, D., Trinh, P., Li, J., Grant, G. A., Yang, F. Gradient hydrogels for screening stiffness effects on patient-derived glioblastoma xenograft cellfates in 3D. Journal of Biomedical Materials Research. Part A. 109 (6), 1027-1035 (2021).
- da Hora, C. C., Schweiger, M. W., Wurdinger, T., Tannous, B. A. Patient-derived glioma models: From patients to dish to animals. Cells. 8 (10), 1177 (2019).
- Li, W., et al. Characterization and transplantation of enteric neural crest cells from human induced pluripotent stem cells. Molecular Psychiatry. 23 (3), 499-508 (2018).
- Scaringi, C., Minniti, G., Caporello, P., Enrici, R. M. Integrin inhibitor cilengitide for the treatment of glioblastoma: A brief overview of current clinical results. Anticancer Research. 32 (10), 4213-4224 (2012).

