Evaluation of Caspase Activation to Assess Innate Immune Cell Death
Summary
This protocol describes a comprehensive method for assessing caspase activation (caspase-1, caspase-3, caspase-7, caspase-8, caspase-9, and caspase-11) in response to both in vitro and in vivo (in mice) models of infection, sterile insults, and cancer to determine the initiation of cell death pathways, such as pyroptosis, apoptosis, necroptosis, and PANoptosis.
Abstract
Innate immunity provides the critical first line of defense in response to pathogens and sterile insults. A key mechanistic component of this response is the initiation of innate immune programmed cell death (PCD) to eliminate infected or damaged cells and propagate immune responses. However, excess PCD is associated with inflammation and pathology. Therefore, understanding the activation and regulation of PCD is a central aspect of characterizing innate immune responses and identifying new therapeutic targets across the disease spectrum.
This protocol provides methods for characterizing innate immune PCD activation by monitoring caspases, a family of cysteine-dependent proteases that are often associated with diverse PCD pathways, including apoptosis, pyroptosis, necroptosis, and PANoptosis. Initial reports characterized caspase-2, caspase-8, caspase-9, and caspase-10 as initiator caspases and caspase-3, caspase-6, and caspase-7 as effector caspases in apoptosis, while later studies found the inflammatory caspases, caspase-1, caspase-4, caspase-5, and caspase-11, drive pyroptosis. It is now known that there is extensive crosstalk between the caspases and other innate immune and cell death molecules across the previously defined PCD pathways, identifying a key knowledge gap in the mechanistic understanding of innate immunity and PCD and leading to the characterization of PANoptosis. PANoptosis is a unique innate immune inflammatory PCD pathway regulated by PANoptosome complexes, which integrate components, including caspases, from other cell death pathways.
Here, methods for assessing the activation of caspases in response to various stimuli are provided. These methods allow for the characterization of PCD pathways both in vitro and in vivo, as activated caspases undergo proteolytic cleavage that can be visualized by western blotting using optimal antibodies and blotting conditions. A protocol and western blotting workflow have been established that allow for the assessment of the activation of multiple caspases from the same cellular population, providing a comprehensive characterization of the PCD processes. This method can be applied across research areas in development, homeostasis, infection, inflammation, and cancer to evaluate PCD pathways throughout cellular processes in health and disease.
Introduction
The innate immune system acts as the first line of defense during infection and in response to sterile stimuli, such as tissue injury and alterations in homeostasis. Innate immune sensors on the cell surface and in the cytoplasm respond to pathogen- or damage-associated molecular patterns (PAMPs or DAMPs, respectively) to trigger inflammatory signaling pathways and cellular responses. One of the key processes of the innate immune response is the induction of cell death to remove infected or damaged cells and drive further innate and adaptive immune responses. Programmed cell death (PCD) is a highly conserved process across species, highlighting its evolutionary importance as an innate immune mechanism.
There are several innate immune PCD pathways that can be activated in all cell types. Caspases are a key family of highly conserved, intracellular, cysteine-dependent proteases that are critical across many PCD pathways, including the traditionally non-inflammatory apoptosis pathway, as well as inflammatory PCD pathways such as pyroptosis, necroptosis, and PANoptosis1,2,3,4,5. There are 11 human and 10 murine caspases that are well defined, as well as pseudo-caspases that may be functional, and most are constitutively expressed as inactive monomeric or dimeric pro-caspases that require cleavage for activation6,7. Caspases also contain important domains for the recruitment and formation of multiprotein complexes. These include the caspase activation and recruitment domain (CARD), which can be found in caspase-1, caspase-2, caspase-4, caspase-5, caspase-9, and caspase-11, or the death effector domain (DED), which is found in caspase-8 and caspase-10. Through both their proteolytic activity and their ability to form multiprotein complexes, caspases are critical drivers of innate immune PCD.
The role of caspases in innate immune PCD was first identified in apoptosis, where the initiator caspases, caspase-2, caspase-8, caspase-9, and caspase-10, activate the executioner caspases, caspase-3, caspase-6, and caspase-7, to drive cell death8,9,10,11,12. Initiator caspases can be activated by diverse signaling cascades; the extrinsic pathway activates caspase-8 through extracellular ligand-induced death receptor signaling, and the intrinsic pathway activates caspase-9 through the disruption of mitochondrial integrity13. Activated initiator caspases cleave the linker separating the large and small catalytic subunits of executioner caspases to produce their active forms. The executioner caspases then cleave their substrates to disassemble the cell, resulting in DNA degradation, membrane blebbing, nuclear fragmentation, and the release of apoptotic bodies14,15. This process typically ends in a non-lytic and non-inflammatory form of cell death when coupled with the immediate clearance of the dying cells by efferocytosis16. However, defects in efferocytosis or a lack of phagocytic cells can lead to the accumulation of apoptotic cells, which then undergo lytic and inflammatory cell death17,18.
The inflammatory caspases, including caspase-1 (human and mouse), caspase-4 and caspase-5 (human), and caspase-11 (mouse), have been discovered to be activated during a form of inflammatory innate immune PCD (III-PCD) called pyroptosis. Caspase-1 activation is associated with the formation of inflammasomes, which are multiprotein complexes containing a cytosolic innate immune sensor, an adaptor molecule (apoptosis-associated speck-like protein containing a CARD [ASC]), and caspase-1. The formation of this complex allows caspase-1 to undergo proximity-mediated autoproteolysis to release its active form, which can cleave target substrates including the pro-inflammatory cytokines interleukin (IL)-1β and IL-18 and the pore-forming molecule gasdermin D (GSDMD)19,20,21,22,23. Caspase-11, caspase-4, and caspase-5 can also activate GSDMD without the upstream formation of the inflammasome after sensing PAMPs such as lipopolysaccharide (LPS)19,20. These caspases undergo dimerization followed by oligomerization and self-cleavage for activation upon binding to cytosolic LPS, which leads to non-canonical inflammasome activation24,25,26 and caspase-1 activation in a cell-intrinsic manner to induce IL-1β and IL-18 maturation20. The maturation and release of these pro-inflammatory cytokines characterize these caspases as "inflammatory." Additionally, the apoptotic caspase-8 has been found to localize to the inflammasome, providing a link between apoptotic and pyroptotic processes. Studies have found that the apoptotic caspase-8 is also critical for regulating another form of PCD called necroptosis. The loss of caspase-8 results in spontaneous receptor-interacting serine-threonine kinase 3 (RIPK3)-mediated mixed lineage kinase domain-like pseudokinase (MLKL) activation to drive the III-PCD pathway of necroptosis27,28,29,30,31,32,33,34,35.
While caspases have historically been classified as "apoptotic" or "inflammatory" based on the type of cell death they initiate, growing evidence suggests there is extensive crosstalk between the innate immune PCD pathways through caspases3,4. For instance, the inflammatory caspase-1 from inflammasomes cleaves the apoptotic caspase-7 at its canonical activation site34. Caspase-1 activation can also lead to the cleavage of apoptotic substrates such as poly(ADP-ribose) polymerase 1 (PARP1)36. In cells lacking GSDMD, caspase-1 can also cleave caspase-337,38. Additionally, the canonically apoptotic caspase-3 can cleave gasdermin E (GSDME) to induce PCD17,18 and also processes GSDMD into an inactive form40. Furthermore, caspase-8 recruitment to the inflammasome complex has been observed39,40,41,42,43,44,45, and caspase-8 is a key regulator of canonical and noncanonical inflammasome activation39. There are also overlapping and redundant roles for caspase-8 and caspase-1 in many inflammatory conditions, and innate immune PCD characterized by the activation of pyroptotic, apoptotic, and necroptotic components occurs across the disease spectrum39,46,47,48,49,50.
Based on this crosstalk between inflammatory and apoptotic caspases, a key gap in the mechanistic understanding of innate immunity and PCD was identified, leading to the discovery of PANoptosis. PANoptosis is a unique form of III-PCD that is activated in response to pathogens, PAMPs, DAMPs, and alterations in homeostasis and is regulated by PANoptosomes – multifaceted macromolecular complexes that integrate components from other cell death pathways44,50,51,52,53,54,55. The totality of the biological effects in PANoptosis cannot be individually accounted for by pyroptosis, apoptosis, or necroptosis alone3,4,35,36,39,46,47,48, as PANoptosis is characterized by the activation of multiple caspases, including caspase-1, caspase-11, caspase-8, caspase-9, caspase-3, and/or caspase-7, depending on the context44,48,49,50,51,52,53,54,56,57,58,59,60,61,62. PANoptosis has been increasingly implicated in infectious and inflammatory diseases, as well as in cancers and cancer therapies3,4,35,36,39,44,46,47,48,49,50,51,52,53,54,56
,57,58,59,60,61,62,63,64,65,66.
Given the essential role of caspases across cell death pathways, including in apoptosis, pyroptosis, necroptosis, and PANoptosis, it is important to develop techniques to characterize their activation and understand the full complexity of the PCD pathways. The protocol here details a method to stimulate cells and measure the subsequent activation of caspases (Figure 1). This method leverages the proteolytic cleavage of caspases, which is generally required for their activation, as a means to study them. Through western blotting, the protein sizes can be determined, allowing for the clear visualization and differentiation of inactive pro-caspases and their activated, cleaved forms.
The major advantages of this protocol are 1) its ability to assess the activation of multiple caspases in parallel from a single population of endogenous cells to more accurately determine PCD activation and 2) the use of relatively simple lab techniques that do not require extensive training or expensive equipment. Previous protocols have used western blotting, fluorescent reporters, or antibody staining to monitor caspase activation in culture supernatants, cell and tissue lysates, whole cells via microscopy, and in vivo67,68,69,70,71, but these techniques generally only monitor one or two caspases in a sample. Furthermore, while synthetic peptide substrates containing caspase cleavage sites that fluoresce upon cleavage have been used to monitor caspase activation in cell or tissue lysates69, these substrates can often be cleaved by more than one caspase, making it difficult to determine the specific activation of individual caspases in this system. Additionally, the use of western blotting rather than the use of fluorescent reporters or other tag-based methods allows researchers to use endogenous cells rather than creating specific cell lines with reporter genes. There are multiple advantages to using endogenous cells, including the fact that many immortalized cell lines are deficient in key cell death molecules72,73, which could affect the results. Additionally, using endogenous cells allows for the evaluation of diverse cell types, such as macrophages, epithelial cells, and endothelial cells, rather than a single lineage. Western blotting is also a relatively simple and cost-effective technique that can be carried out in labs around the world without the need for large, expensive equipment or complicated setups.
This protocol is widely applicable across biology to understand both the cell death-dependent and cell death-independent functions of caspases, including their scaffolding roles and functions in other inflammatory signaling pathways74. Applying this method allows for a unified approach in the study of innate immune PCD pathways and inflammatory signaling across diseases and conditions, and this protocol can be used to identify critical regulatory processes and mechanistic connections that will inform the development of future therapeutic strategies.
Protocol
The animal use and procedures were approved by the St. Jude Children's Research Hospital Committee on the Use and Care of Animals.
1. Preparing the solutions
- Prepare L929-conditioned media.
- Plate 1 × 106 L929 cells (see Table of Materials) in a 182 cm2 tissue culture flask containing 50 mL of L929 culture media (see Table 1 for the preparation of the media).
- Grow the cells in a humidified incubator at 37 °C with 5% CO2.
- After 7 days, collect the supernatant, and filter using a 0.45 µm filter. Prepare 50 mL aliquots (store frozen aliquots at −80 °C for up to 1 year).
- Prepare 500 mL of bone marrow-derived macrophage (BMDM) culture media (Table 1).
- Prepare 500 mL of BMDM stimulation media (Table 1).
- Prepare 500 mL of media for infection (Table 1).
- Prepare 100 mL of 1 M Tris buffer (Table 1).
- Prepare 4x sodium dodecyl sulfate (SDS) buffer (Table 1).
- Prepare 1 mL of 1 M 1,4-dithiothreitol (DTT, Table 1).
- Prepare 40 mL of caspase lysis buffer (Table 1).
- Prepare 100 mL of 1.5 M Tris buffer (Table 1).
- Prepare 100 mL of a 10% (wt/vol) SDS solution (Table 1).
- Prepare 50 mL of a 2x radioimmunoprecipitation assay (RIPA) buffer (Table 1).
- Prepare a 5 mg/mL LPS solution (Table 1).
- Prepare a 0.5 M ATP solution (Table 1).
- Prepare the western blotting solutions.
- Prepare 1 L of 5x running buffer stock (Table 1).
- Prepare 1 L of 10x transfer buffer stock (Table 1).
- Prepare 1 L of Tris-buffered saline with Tween 20 (TBST, Table 1).
- Prepare 100 mL of a 5% (wt/vol) skim milk blocking solution (Table 1).
- Prepare the primary antibody solutions.
- Prepare 10 mL of caspase-1 primary antibody (Table 1).
- Prepare 10 mL of caspase-11 primary antibody (Table 1).
- Prepare 10 mL of caspase-3 primary antibody (Table 1).
- Prepare 10 mL of caspase-7 primary antibody (Table 1).
- Prepare 10 mL of caspase-8 primary antibody (Table 1).
- Prepare 10 mL of caspase-9 primary antibody (Table 1).
- Prepare 10 mL of HRP-conjugated β-actin primary antibody (Table 1).
- Prepare the secondary antibody solutions.
- Prepare 10 mL of anti-rabbit secondary antibody (Table 1).
- Prepare 10 mL of anti-mouse secondary antibody (Table 1).
- Prepare 10 mL of anti-rat secondary antibody (Table 1).
2. Isolating bone marrow-derived macrophages
NOTE: For this protocol, 6-10-week-old wild-type mice with intact PCD pathways or mutant mice with the PCD regulators, effectors, or molecules of interest deleted or altered can be used.
- Euthanize a mouse in a CO2 chamber with a flow rate that displaces 10%-30% of the cage volume per min for 2-3 min. Then, perform a secondary euthanasia method, such as cervical dislocation. Follow all additional facility-, institution-, and government-specific guidelines and regulations wherever applicable.
- Dissect the mouse to retrieve the hind leg bones.
- Pin the mouse on its back so the abdomen is exposed. Spray with 70% (vol/vol) ethanol to sterilize the hind legs and abdomen.
CAUTION: Ethanol is flammable. Keep it away from open flames. - Use scissors to make an incision at the midline of the abdomen; continue cutting toward the legs to make the femurs visible.
- Take the right rear leg and pull the skin away from the body toward the midline. Detach the leg from the body by severing the adductor muscles toward the midline; then, cut the leg between the hip joint and the spine. Next, cut the paw off distal to the ankle, and remove excess tissue from the bone by peeling off the skin and using slightly opened scissors to strip off the calf tissue.
- Place the leg on a 70% (vol/vol) ethanol-soaked towel and dissect the tibia and the femur.
- Clasp the tibia and femur each with a separate set of forceps. Gently press the tibia against the knee joint's natural direction; this will cause the tibia to break at the knee.
- Use the forceps and dissecting scissors if needed to remove any remaining hanging tissue. Save the tibia for later use by placing it on the ethanol-soaked towel.
- Collect the femur in the same way, by snapping off the knee.
- Repeat step 3 and 4 above for the left leg to remove it from the body and dissect the tibia and femur.
- Spray the bones with 70% (vol/vol) ethanol.
- Clean off the bones by placing them on a clean 70% (vol/vol) ethanol-soaked towel, squeezing the fleshy part in the towel, and rubbing the towel against the bone to remove excess tissue.
- Pin the mouse on its back so the abdomen is exposed. Spray with 70% (vol/vol) ethanol to sterilize the hind legs and abdomen.
- Once both femurs and both tibias are clean, spray all four bones with 70% (vol/vol) ethanol. Collect the bones in a sterile Petri dish and rinse them with 10 mL of BMDM culture media by gently swishing the media in the dish.
- Fill a 10 mL syringe with 10 mL of fresh BMDM culture media and attach a 25 G needle.
- Pick up the tibia using forceps; then, cut the ankle joint at a ~45° angle.
- Flush the bone marrow from the tibia.
- Hold the tibia over a 50 mL tube, with the narrow end of the tibia pointing down. Dispense the media over the tibia from the filled syringe.
- Insert the needle (gently at first) at the top end of the marrow and dispense the media.
- Remove the needle and then insert again. Use short, high-pressure pushes to dispense the media and move the needle into the marrow.
- Once the media begins to flow out from the bottom of the bone, use short, high-pressure pushes to continue to flush the cells out. Monitor the color of the bone during this process, and when the bone is white, discard it.
- Do this for both tibias.
- Repeat steps 5 and 6 above for the femurs, cutting at the hip joint as the tibia was cut at the ankle joint. Use the same 50 mL tube to collect the BMDM culture media as the marrow is flushed.
- Once all four bones have been flushed, aspirate the marrow and media from the 50 mL tube up and down 3x through an 18 G needle on a 10 mL syringe, rinsing the sides of the tube each time to disperse the marrow.
- Adjust the final volume in the 50 mL tube to 30 mL with BMDM culture media, and ensure the cells are thoroughly suspended.
- Use a 70 µm cell strainer to filter the BMDM culture media from the 50 mL tube.
- Plate the resulting cell suspension, which contains the bone marrow progenitor cells, into three 150 mm tissue culture dishes by adding 10 mL (or ~20 × 106 cells) to each. Then, add an additional 10 mL of BMDM culture media to each dish. Incubate in a humidified incubator at 37 °C.
3. Differentiating the BMDMs and plating for the experiments
- Incubate the plated bone marrow progenitor cells at 37 °C for 3 days. Then, remove each dish, and add an additional 5-8 mL of BMDM culture media (Table 1). Return to the incubator at 37 °C.
- On day 5 after the initial plating, remove each dish, and add an additional 5 mL of BMDM culture media. Return to the incubator at 37 °C.
- On day 6, remove each dish, and discard the media. Then, add 10 mL of cold (stored at 4 °C) PBS to wash once. Discard the 10 mL of cold PBS wash. Then, add 10 mL of fresh, cold PBS to each dish, and incubate each dish on ice for 5 min.
- Using a cell scraper, gently scrape the cells from all three dishes into one 50 mL tube. Gently spin down the cells at 270 × g at 4 °C for 5 min; then, discard the supernatant.
- Add 20 mL of BMDM culture media, pipette up and down to resuspend the pellet, and count the cells.
NOTE: It is expected that each mouse will yield approximately 60 × 106-100 × 106 cells. - Plan out the 12-well plate layout for the desired in vitro cell death/inflammation stimulation assay. Plan to plate 1 × 106 cells per well. For each planned stimulation, include at least three biological replicates, and plate one set of wells to be harvested in caspase lysis buffer and a second set of wells to be harvested in RIPA buffer.
- Plate 1 × 106 cells in 1 mL of BMDM culture media per well in 12-well plates. Culture overnight in a humidified incubator at 37 °C before proceeding to the cell death/inflammation evaluation. After overnight incubation, remove the media, and add 1 mL of warm (37 °C) PBS to each well to wash the cells.
- Remove the PBS wash, and add 500 µL of BMDM stimulation media with antibiotics (if performing non-bacterial stimulations) or BMDM stimulation media without antibiotics (if performing bacterial stimulations) (Table 1). Incubate for 2 h before moving to step 4 for in vitro stimulation/infection.
4. Stimulating or infecting the cells
CAUTION: The agents included in this protocol are potentially pathogenic and should be handled with the appropriate precautions in a biosafety level 2 (BSL2) facility with approval from the relevant institutional and governmental authorities.
- Stimulate the BMDMs to activate cell death with the trigger of interest.
NOTE: For the purposes of this protocol, influenza A virus (IAV), herpes simplex virus 1 (HSV1), Francisella novicida, and LPS + ATP are used for illustration, but other triggers can be used.- Example stimulation 1: Infect with IAV (A/Puerto Rico/8/34, H1N1 [PR8]) (constructed as per Hoffmann et al.75; the viral titer for the multiplicity of infection [MOI] determination is calculated by a plaque assay in MDCK cells):
- Calculate the volume of virus needed for infection at a multiplicity of infection (MOI) of 20 plaque-forming units (PFU) using equation (1) and equation (2):

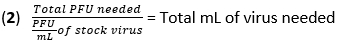

- Remove the media from the BMDMs and wash the cells once with 500 µL of PBS. Add 450 µL of IAV (20 MOI) in high-glucose DMEM without heat-inactivated (HI)-FBS to each well. Incubate the plates at 37 °C for 1 h in a humidified incubator to allow absorption.
- Remove the plates and add 50 µL of HI-FBS. Return the plates to the incubator at 37 °C. Incubate for a total of 12 h.
- Calculate the volume of virus needed for infection at a multiplicity of infection (MOI) of 20 plaque-forming units (PFU) using equation (1) and equation (2):
- Example stimulation 2: Infect with HSV1 (HF strain; cultured as described previously44; the viral titer for MOI determination is calculated by a plaque assay in Vero cells):
- Calculate the volume of virus needed for infection at an MOI of 10 PFU using equation (1) and equation (2) from step 1 in the IAV infection section above.

- Remove the media from the BMDMs. Add 450 µL of HSV1 (MOI 10) in high-glucose DMEM without HI-FBS to each well. Incubate the plates at 37 °C for 1 h in a humidified incubator to allow absorption.
- Remove the plates, and add 50 µL of HI-FBS. Return the plates to the incubator at 37 °C. Incubate for a total of 12 h.
- Calculate the volume of virus needed for infection at an MOI of 10 PFU using equation (1) and equation (2) from step 1 in the IAV infection section above.
- Example stimulation 3: Infect with F. novicida (U112 strain; cultured as described previously44 under aerobic conditions at 37 °C in BBL Trypticase Soy Broth supplemented with 0.2% L-cysteine overnight. Then, subculture the bacteria at a ratio of 1:10 at 37 °C for another 4 h in fresh medium before taking the optical density (OD) at 600 nm using the fresh medium as a blank. An OD value of 1 equates to 1 × 109 colony-forming units (CFU) per mL.)
- Calculate the volume of F. novicida needed for infection at an MOI of 50 CFU using equation (3) and equation (4):

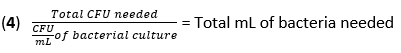
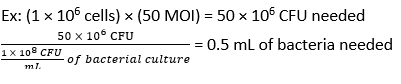
- Remove the media from the BMDMs. Add 500 µL of F. novicida (MOI 50) in BMDM stimulation media without antibiotics to each well. Incubate the plates at 37 °C for 4 h in a humidified incubator to allow absorption.
- Wash the cells three times with warmed (37 °C) PBS and add 500 µL of BMDM stimulation media containing 50 µg/mL gentamycin. Return the plates to the incubator at 37 °C. Incubate overnight (16 h).
- Calculate the volume of F. novicida needed for infection at an MOI of 50 CFU using equation (3) and equation (4):
- Example stimulation 4: Stimulate with LPS + ATP.
- Remove the media from the BMDMs. Add 500 µL of BMDM stimulation media with antibiotics (Table 1) containing 100 ng/mL LPS to each well. Incubate the plates at 37 °C for 3.5 h in a humidified incubator.
- Add 5 µL of 0.5 M ATP stock solution (Table 1) to each well. Return the plates to the incubator at 37 °C. Incubate for 30 min.
- Example stimulation 1: Infect with IAV (A/Puerto Rico/8/34, H1N1 [PR8]) (constructed as per Hoffmann et al.75; the viral titer for the multiplicity of infection [MOI] determination is calculated by a plaque assay in MDCK cells):
5. Collecting the combined supernatant and protein lysate to be used for caspase western blots
- After 4 h, 12 h, or 16 h of incubation (the specific timing is dependent on the trigger used), remove the plate from the incubator.
- Remove 150 µL of the supernatant; discard or save this for other supernatant analyses (e.g., enzyme-linked immunosorbent assay [ELISA]). Do not remove the remaining supernatant.
- Create the protein collection solution by combining 50 µL of caspase lysis buffer + 100 µL of 4x SDS buffer per well (Table 1). Then, add 150 µL of the mix to each well.
- For each well, pipette the mixture up and down to collect the lysed cells and supernatant. While pipetting, also scrape the bottom of the well with the pipette tip to disrupt the cells. After scraping and pipetting, collect the protein lysate into labeled 1.5 mL tubes.
- Use a heat block to heat all the tubes to 100 °C for 12 min.
- Pellet any insoluble components by centrifuging at 14,500 × g for 30 s at room temperature.
NOTE: This is a pause point – the protein from the combined supernatant and protein lysates can either be used immediately or stored at −20 °C for up to 2 months or at −80 °C for up to 6 months until ready to use.
6. Collecting the protein lysate to be used for caspase western blots
- After 4 h, 12 h, or 16 h of incubation (the specific timing is dependent on the trigger used), remove the plate from the incubator. Remove all the supernatant; discard or save this for other supernatant analyses (e.g., ELISA).
- Create the 1x RIPA buffer by diluting the 2x RIPA stock solution (Table 1) in an equal volume of deionized water. Then, add one phosphatase inhibitor tablet and one protease inhibitor tablet, and allow them to dissolve. Add 150 µL of 1x RIPA buffer and 50 µL of 4x SDS to each well.
- For each well, pipette the mixture up and down to collect the lysed cells. While pipetting, also scrape the bottom of the well with the pipette tip to disrupt the cells. After scraping and pipetting, collect the protein lysate into labeled 1.5 mL tubes.
- Use a heat block to heat all the tubes to 100 °C for 12 min.
- Pellet any insoluble components by centrifuging at 14,500 × g for 30 s at room temperature.
NOTE: This is a pause point – the protein lysates can either be used immediately or stored at −20 °C or −80 °C until ready to use.
7. Performing western blotting using the lysates collected from the BMDMs following the steps above or from tissue homogenates
NOTE: If using tissue, it can be homogenized by hand or through a power-driven tissue homogenizer. The protocol by Simpson76 provides a detailed description of tissue homogenization.
- Prepare 1x running buffer: Combine 200 mL of the 5x running buffer stock (Table 1) and 800 mL of deionized water. Make this 1x running buffer just before each experiment.
- Prepare the electrophoresis apparatus with a 12% (wt/vol) polyacrylamide gel with 10 wells. Fill the electrophoresis apparatus with 1x running buffer. Then, remove the gel comb.
NOTE: To analyze caspase-1, caspase-11, caspase-3, caspase-7, caspase-8, and caspase-9 for each sample, six gels will be needed. - For caspase-1, caspase-3, caspase-7, and caspase-8 blots, plan to use 30 µL of the combined supernatant and protein lysate in caspase lysis buffer or the tissue homogenate. For caspase-11 and caspase-9 blots, plan to use 20 µL of the protein lysate in RIPA buffer or the tissue homogenate. If the samples have been stored at −20 °C or −80 °C, thaw them on ice first.
- For all the samples, heat to 100 °C for 5 min, and centrifuge at 14,500 × g for 30 s at 4 °C before loading. Then, slowly load 20 or 30 µL of the sample into each lane. Avoid having any sample overflow into the other lanes. To evaluate all six caspases at once, use the same procedure to load the appropriate samples into each of the six gels.
- Connect the electrophoresis apparatus to the power source. Then, set the power to 80 V for 20 min to begin the gel run, and then adjust the power to 130 V for 45-60 min.
- Watch the dye front carefully, and turn the power off when the dye front is at the bottom of the gel but has not yet been pushed out of the gel.
- While the gel is running, prepare 1x transfer buffer by combining 700 mL of deionized water, 100 mL of the 10x transfer buffer stock (Table 1), and 200 mL of methanol. Make the 1x solution fresh each time.
NOTE: Use caution as methanol is flammable. Perform the transfer buffer preparation away from open flames. - Remove the gel from the electrophoresis apparatus gently by using the gel releaser.
- Set up one transfer stack for each gel.
- Activate a PVDF membrane by soaking it in methanol for 1 min.
- Pre-wet two pieces of filter paper, the gel, and the PVDF membrane in transfer buffer for 5 min. Keep the PVDF membrane and gel in separate containers during this 5 min incubation.
- Assemble the transfer stack on the semi-dry system. Beginning on the bottom platinum anode side, place one piece of filter paper, the PVDF membrane, the gel, and finally one piece of filter paper. Gently roll out or press out air bubbles between the layers, and close the top of the system. Ensure that the safety cover is secured before proceeding further.
- Connect to the power source. Set the power to 25 V for 45 min.
- After the transfer is complete, disassemble the transfer stack, and collect the membrane; place it in a square Petri dish (incubation tray).
- Perform membrane blocking by adding 15 mL of a 5% (wt/vol) skim milk solution (Table 1). Incubate the membrane on a rocking shaker at 50 rpm to 70 rpm at room temperature for 1 h.
NOTE: This is a pause point – the membrane can either be removed after 1 h or stored in blocking solution at 4 °C overnight. - After 1 h or overnight incubation, remove the blocking solution. Add 10 mL of the diluted antibody solution (anti-caspase-1 antibody, anti-caspase-11 antibody, combined anti-caspase-3 and anti-cleaved caspase-3 antibody, combined anti-caspase-7 and anti-cleaved caspase-7 antibody, combined anti-caspase-8 and anti-cleaved caspase-8 antibody, or anti-caspase-9 antibody) (Table 1). Place on a rocking shaker at 50 rpm to 70 rpm to incubate at room temperature for 2 h or at 4 °C overnight (16 h).
- Collect the antibody solution (reuse up to 3x or discard it), and wash by adding 15 mL of TBST (Table 1) to the membrane on a rocking shaker at 50 rpm to 70 rpm at room temperature for 10 min. Discard the TBST.
- Repeat the wash with 15 mL of TBST following step 14 a total of 3x.
- Add 10 mL of the diluted secondary HRP-conjugated antibody solution (anti-rabbit for blots stained with primary antibodies against caspase-3, caspase-7, caspase-8, or caspase-9; anti-mouse for blots stained with primary antibody against caspase-1; anti-rat for blots stained with primary antibody against caspase-11) (Table 1). Incubate on a rocking shaker at 50 rpm to 70 rpm at room temperature for 1 h.
- Remove the antibody solution, and wash by adding 15 mL of TBST to the membrane on a rocking shaker at 50 rpm to 70 rpm at room temperature for 10 min. Discard the TBST.
- Repeat the wash with 15 mL of TBST following step 17 a total of 3x.
- Add 10 mL of the high-sensitivity HRP substrate to the membrane. Let it sit at room temperature in the dark for 1 min.
- Remove the membrane from the substrate. Proceed directly to the imaging, using a chemiluminescence imager with the accessory white trans tray inserted in the lower position. Expose the membrane using the auto exposure mode (generally ~1-2 min of exposure time).
- Using the membrane from the caspase-9 or caspase-11 blotting (i.e., a membrane with RIPA lysate samples), add 10 mL of stripping buffer, and incubate on a rocking shaker at 50 rpm to 70 rpm at room temperature for 5 min.
- Discard the stripping buffer, and wash by adding 15 mL of TBST to the membrane on a rocking shaker at 50 rpm to 70 rpm at room temperature for 10 min. Discard the TBST.
- Repeat the wash with 15 mL of TBST following step 22 a total of 3x.
- Perform membrane blocking by adding 15 mL of a 5% (wt/vol) skim milk solution. Incubate the membrane on a rocking shaker at 50 rpm to 70 rpm at room temperature for 1 h.
NOTE: This is a pause point – the membrane can either be removed after 1 h or stored in blocking solution at 4 °C overnight. - After 1 h or overnight incubation, remove the blocking solution. Add 10 mL of the diluted anti-β-actin (HRP-conjugated) antibody solution. Place on a rocking shaker at 50 rpm to 70 rpm to incubate at room temperature for 1.5 h.
- Remove the antibody solution and wash by adding 15 mL of TBST to the membrane on a rocking shaker at 50 rpm to 70 rpm at room temperature for 10 min. Discard the TBST.
- Repeat the wash with 15 mL of TBST following step 26 a total of 3x.
- Add 10 mL of the standard-sensitivity HRP substrate to the membrane. Let it sit at room temperature in the dark for 1 min.
- Proceed directly to the imaging, using a chemiluminescence imager with the accessory white trans tray inserted in the lower position. Expose the membrane using the auto exposure mode (generally <1 min of exposure time).
Representative Results
PANoptosis has been observed in response to numerous bacterial, viral, and fungal infections and other inflammatory stimuli, as well as in cancer cells44,48,49,50,51,52,53,54,56,57,58,60,61,62. In these cases, the activation of multiple caspases has been reported. Using the example stimulations in this protocol that are known to induce PANoptosis, namely IAV, HSV1, and F. novicida44,48,50,51,53,57, it is expected that cleavage may be observed as a surrogate for the activation of multiple caspases (Figure 2A-C). Additionally, LPS + ATP is included as a control; this combination is a canonical NLRP3 inflammasome trigger that is expected to induce caspase-1 activation. The activation of caspase-1 can be visualized in response to each of these triggers by the cleavage of the p45 pro-form to the p20 active form (Figure 2D). The p20 form can then cleave GSDMD and initiate cell death19.
Similarly, the activation of the apoptotic initiator caspase, caspase-8, is observed, as denoted by the presence of the p18 cleaved form (Figure 2A–D). The weak activation of caspase-9 can also be observed in some cases with these stimuli (Figure 2A–D). Downstream of these initiators, the effector caspases caspase-3 and caspase-7 are activated; this activation is observed as the formation of the p17/19 cleaved form of caspase-3 and the p20 cleaved form of caspase-7 (Figure 2A–D). To date, the activation of caspase-11 has not been widely evaluated in PANoptosis, but its activation has been observed in some contexts54. Caspase-11 remains an important inflammatory caspase, and its activation can be observed by the formation of its p26 cleaved form. While some low-level caspase-11 activation is seen, robust activation is not observed in response to IAV, HSV1, or F. novicida infection or LPS + ATP stimulation (Figure 2A–D).
Cells lacking upstream PANoptosis sensors do not undergo cell death in response to PANoptosis-inducing stimuli. For example, IAV infection induces the formation of the ZBP1-PANoptosome48,51,53,57, and Zbp1-/- cells are protected from cell death during IAV infection (Figure 3A). Similarly, HSV1 and F. novicida infections induce the formation of the AIM2-PANoptosome44, and Aim2-/- cells fail to undergo robust cell death in response to these infections (Figure 3B,C). In cells that are deficient in the upstream sensor necessary for PANoptosome formation, the activation of caspases is significantly reduced or even eliminated, as can be seen by the reduction in the intensity of the cleaved bands in the western blots (Figure 2A–C). Similarly, cells lacking upstream inflammasome sensors are protected from cell death in response to their respective stimuli, and Nlrp3-/- cells do not undergo cell death in response to LPS + ATP (Figure 3D). These cells also show reduced caspase activation (Figure 2D).
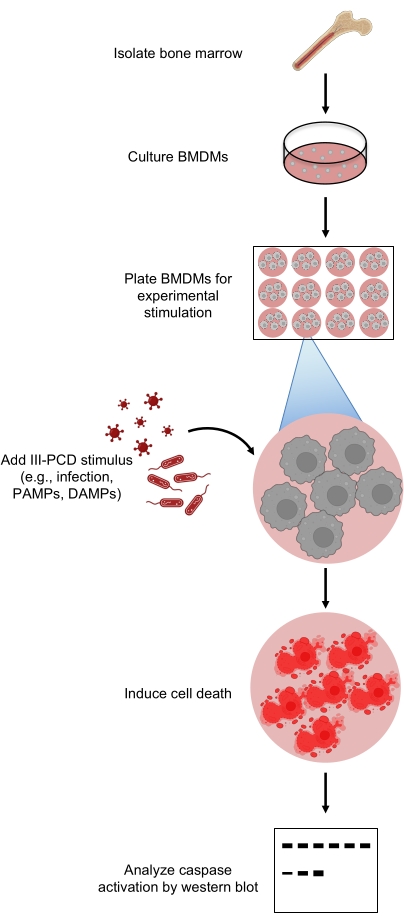
Figure 1: Overview of the experimental procedure. Bone marrow is isolated and differentiated into bone marrow-derived macrophages. These cells are then stimulated with a trigger that activates innate immune signaling and cell death, such as infection or an inflammatory stimulus. After the cells activate and begin to undergo PCD, the lysate is collected and analyzed by western blotting to monitor caspase cleavage to their active forms. Abbreviations: BMDMs = bone marrow-derived macrophages; DAMPs = damage-associated molecular patterns; III-PCD = inflammatory innate immune programmed cell death; PAMPs = pathogen-associated molecular patterns; PCD = programmed cell death. Please click here to view a larger version of this figure.
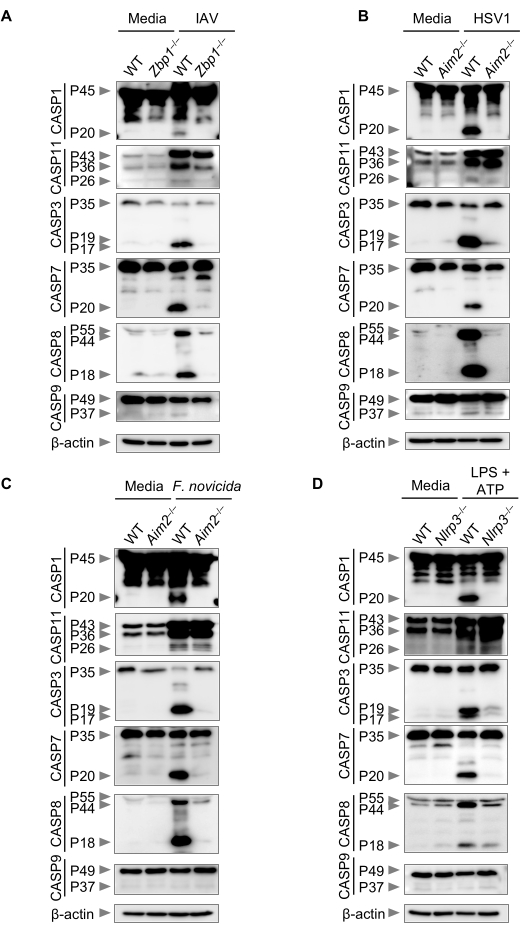
Figure 2: Activation of caspases in response to stimuli. Caspase activation from cell culture lysates can be assessed by evaluating the size of the products run on an electrophoretic gel. (A–D) Immunoblot analysis of pro- (P45) and activated (P20) CASP1, pro- (P43) and cleaved (P36 and P26) CASP11, pro- (P35) and cleaved (P17/P19) CASP3, pro- (P35) and cleaved (P20) CASP7, pro- (P55) and cleaved (P44 and P18) CASP8, and pro- (P49) and cleaved (P37) CASP9 in WT and Zbp1−/−, WT and Aim2−/−, or WT and Nlrp3−/− BMDMs after (A) IAV infection, (B) HSV1 infection, (C) Francisella novicida infection, or (D) LPS + ATP stimulation. Images are representative of three independent experiments. Abbreviations: BMDMs = bone marrow-derived macrophages; CASP1 = caspase-1; CASP3 = caspase-3; CASP7 = caspase-7; CASP8 = caspase-8; CASP9 = caspase-9; CASP11 = caspase-11; HSV1 = herpes simplex virus 1; IAV = influenza A virus; LPS = lipopolysaccharide; WT = wild-type. Please click here to view a larger version of this figure.
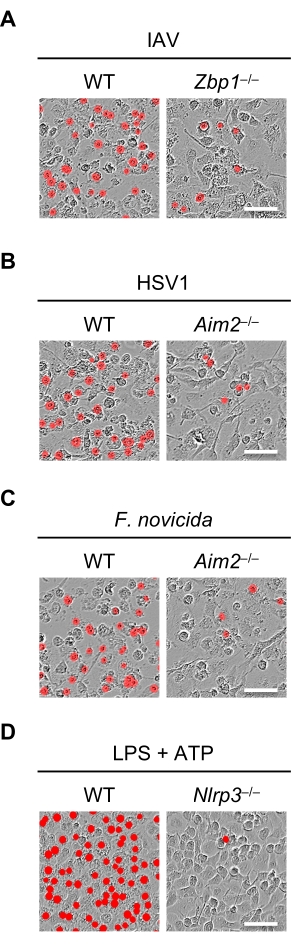
Figure 3: Inhibition of cell death when caspase activation does not occur. (A–D) Representative images of cell death in WT and Zbp1−/−, WT and Aim2−/−, or WT and Nlrp3−/− BMDMs after (A) IAV infection, (B) HSV1 infection, (C) Francisella novicida infection, or (D) LPS + ATP stimulation. The red mask denotes dead cells. Scale bars = 50 µm. Images are representative of three independent experiments. Abbreviations: BMDMs = bone marrow-derived macrophages; HSV1 = herpes simplex virus 1; IAV = influenza A virus; LPS = lipopolysaccharide; WT = wild-type. Please click here to view a larger version of this figure.
Discussion
Monitoring caspase cleavage and activation provides one of the most comprehensive pictures of innate immune PCD activation as part of the innate immune response. The protocol described here demonstrates a strategy to monitor caspase activation in response to IAV, HSV1, and F. novicida infections and the sterile trigger LPS + ATP, but numerous other stimuli can induce PCD and could be used in this method, as has been shown in several publications44,48,49,50,51,52,53,54,56,57,58,59,60,61,62. This procedure can also be used to characterize the innate immune PCD in response to new triggers, where the mechanisms of PCD remain unknown. Using a combination of wild-type and genetically deficient cells and comparing the activation of different caspases between these populations can provide new insights into the sensors and regulators involved in PCD in response to a given stimulus.
Several technical points should be considered when performing this method. First, as this method involves infecting or otherwise stimulating cells to induce cell death, it is important to use sterile techniques for the isolation of the cells and for the subsequent stimulations. Contamination can significantly affect the results, leading to caspase activation even in unstimulated conditions. To test for this, it is best to always include an unstimulated control when collecting the samples and loading gels for the caspase blots. Additionally, when plating the cells, it is important to ensure that the cells are counted accurately and that clumping does not occur. If the cells are clumped, inconsistent numbers of cells will be deposited in the wells, and the MOI and the amount of protein per well will be affected. When performing the western blots, it is good practice to compare the banding pattern of the uncleaved, pro-form of the caspase, as well as to monitor the cleaved form to ensure that consistent amounts of protein were collected and loaded across the wells. However, when there is a high level of caspase activation in one sample, this can cause a distortion in the signal, leading to what appears to be a reduction in the pro-form in the other samples, as we show with caspase-8 in HSV1 and F. novicida infections in this example (Figure 2B,C). This likely occurs due to uneven antibody binding caused by the presence of too much protein in a particular spot on the membrane, or because of imaging artifacts where the substrate becomes oversaturated in a single sample. Adjusting the antibody dilutions or imaging settings can help overcome this issue. Additionally, when establishing an infection or stimulation for the first time, using a time course can be beneficial to identify when the activation of each caspase occurs. Without a time course, the data may be misleading, as the individual time point selected could be too early or too late, causing caspase activation to be missed and inappropriate conclusions to be made about its involvement. Similarly, testing different MOIs for infectious stimuli can also be beneficial to identify the specific conditions under which caspase(s) are activated.
The electrophoresis step also requires precision. When preparing to load the samples onto the gel, the tubes should always be centrifuged first, and only the supernatant should be loaded. Insoluble protein can interfere with the running of the gel and the subsequent analysis of the data. The combined supernatant and protein lysate should be used for the detection of caspase-1 rather than a pure cell lysate; this will greatly improve the sensitivity of detection. Caspase-3, caspase-7, and caspase-8 can also be detected well in the combined supernatant and protein lysate. However, for best results when detecting caspase-11 and caspase-9, the protein lysate collected in the RIPA buffer is sufficient. As the relative abundance of each caspase within the lysate can be low, as much sample as possible should be loaded onto the gel. This requires slow and steady pipetting to avoid overflowing into the neighboring wells. If poor separation is observed between the pro- and cleaved forms of the caspases, the gel run time can be extended, or a lower percentage of acrylamide can be used to optimize the separation.
Finally, the antibodies listed in this protocol have been optimized for use with mouse BMDMs, and several groups have rigorously tested them using genetically deficient cells as controls to confirm there are no non-specific bands at the molecular weights of interest. While other antibodies may also work, the ones listed here are optimal. Additionally, many of these antibodies also work well in tissue lysates, and other cell populations can also be collected and stimulated in vitro for analysis. Using cell lines or primary cells from humans or other species is also possible but will require an additional assessment of the antibodies for optimization. Human anti-caspase-1 (1:1,000) and human anti-caspase-8 (1:1,000; see Table of Materials) have been successfully used for such assessments, and the same anti-caspase-3, anti-cleaved caspase-3, anti-caspase-7, and anti-cleaved caspase-7 antibodies included in this protocol can be used for both mouse and human cells44,56,62.
The importance of understanding PCD pathways, including pyroptosis, apoptosis, necroptosis, and PANoptosis, as an integral component of the innate immune response is growing, as crosstalk between PCD pathways is increasingly being characterized, and cell death molecules are being identified as therapeutic targets across the disease spectrum3,4. Continuing to identify the innate immune sensors that induce PCD as well as III-PCD and characterize the timing and interplay of caspase activation will improve the ability to therapeutically modulate innate immune PCD pathways and improve patient outcomes.
Offenlegungen
The authors have nothing to disclose.
Acknowledgements
We thank members of the Kanneganti lab for their comments and suggestions, and we thank J. Gullett, PhD, for scientific editing support. Work in our lab is supported by National Institutes of Health (NIH) grants AI101935, AI124346, AI160179, AR056296, and CA253095 (to T.-D.K.) and by the American Lebanese Syrian Associated Charities (to T.-D.K.). The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Materials
| 0.45 μm filter | Millipore | SCHVU05RE | |
| 10 mL syringe | BD Biosciences | 309604 | |
| 12% polyacrylamide gel with 10 wells | Bio-Rad | 4561043 | |
| 12-well plate | Corning | 07-200-82 | |
| 18 G needle | BD Biosciences | 305195 | |
| 25 G needle | BD Biosciences | 305122 | |
| 50 mL tube | Fisher Scientific | 50-809-218 | |
| 70 μm cell strainer | Corning | 431751 | |
| 150 mm tissue culture dishes | Corning | 430597 | |
| 182-cm2 tissue culture flask | Genesee Scientific | 25-211 | |
| Accessory white trans tray | Cytiva | 29-0834-18 | |
| Anti–caspase-1 antibody | AdipoGen | AG-20B-0042-C100 | |
| Anti–caspase-11 antibody | Novus Biologicals | NB120-10454 | |
| Anti–caspase-3 antibody | Cell Signaling Technology | 9662 | |
| Anti–caspase-7 antibody | Cell Signaling Technology | 9492 | |
| Anti–caspase-8 antibody | Cell Signaling Technology | 4927 | |
| Anti–caspase-9 antibody | Cell Signaling Technology | 9504 | |
| Anti–cleaved caspase-3 antibody | Cell Signaling Technology | 9661 | |
| Anti–cleaved caspase-7 antibody | Cell Signaling Technology | 9491 | |
| Anti–cleaved caspase-8 antibody | Cell Signaling Technology | 8592 | |
| Anti-mouse HRP-conjugated secondary antibody | Jackson ImmunoResearch Laboratories | 315-035-047 | |
| Anti-rabbit HRP-conjugated secondary antibody | Jackson ImmunoResearch Laboratories | 111-035-047 | |
| Anti-rat HRP-conjugated secondary antibody | Jackson ImmunoResearch Laboratories | 112-035-003 | |
| Anti–β-Actin antibody (C4) HRP | Santa Cruz | sc-47778 HRP | |
| ATP | InvivoGen | tlrl-atpl | |
| BBL Trypticase Soy Broth | BD Biosciences | 211768 | |
| Bead bath | Chemglass Life Sciences | CLS-4598-009 | |
| Biophotometer D30 | Eppendorf | 6133000010 | |
| BME | Sigma | M6250 | |
| Bromophenol blue | Sigma | BO126 | |
| Cell scrapers | CellTreat Scientific Products | 229315 | |
| Chemiluminescence imager (Amersham 600) | Cytiva | 29083461 | |
| CO2 chamber | VetEquip | 901703 | |
| Cuvettes | Fisher Scientific | 14-955-129 | |
| Dissecting scissors | Thermo Fisher Scientific | 221S | |
| DMEM | Thermo Fisher Scientific | 11995-073 | |
| DTT | Sigma | 43815 | |
| Eelectrophoresis apparatus | Bio-Rad | 1658004 | |
| Ethanol | Pharmco | 111000200 | |
| Fetal bovine serum | Biowest | S1620 | |
| Filter paper | Bio-Rad | 1703965 | |
| Forceps | Fisher Scientific | 22-327379 | |
| Francisella novicida (U112 strain) | BEI Resources | NR-13 | |
| Gel releaser | Bio-Rad | 1653320 | |
| Gentamycin | Gibco | 15750060 | |
| Glycerol | Sigma | G7893 | |
| Glycine | Sigma | G8898 | |
| HCl | Sigma | H9892 | |
| Heat block | Fisher Scientific | 23-043-160 | |
| Herpes simplex virus 1 (HF strain) | ATCC | VR-260 | |
| High glucose DMEM | Sigma | D6171 | |
| Human anti–caspase-1 antibody | R&D Systems | MAB6215 | |
| Human anti–caspase-8 antibody | Enzo | ALX-804-242 | |
| Humidified incubator | Thermo Fisher Scientific | 51026282 | |
| Image analysis software | ImageJ | v1.53a | |
| IMDM | Thermo Fisher Scientific | 12440-053 | |
| Influenza A virus (A/Puerto Rico/8/34, H1N1 [PR8]) | constructed per Hoffmann et al. | ||
| L929 cells | ATCC | CCL-1 | cell line for creating L929-conditioned media |
| L-cysteine | Thermo Fisher Scientific | BP376-100 | |
| Luminata Forte Western HRP substrate | Millipore | WBLUF0500 | standard-sensitivity HRP substrate |
| MDCK cells | ATCC | CCL-34 | cell line for determining IAV viral titer |
| Methanol | Sigma | 322415 | |
| Microcentrifuge | Thermo Fisher Scientific | 75002401 | |
| Non-essential amino acids | Gibco | 11140050 | |
| Nonfat dried milk powder | Kroger | ||
| NP-40 solution | Sigma | 492016 | |
| PBS | Thermo Fisher Scientific | 10010023 | |
| Penicillin and streptomycin | Sigma | P4333 | |
| Petri dish | Fisher Scientific | 07-202-011 | |
| PhosSTOP | Roche | PHOSS-RO | |
| Power source | Bio-Rad | 164-5052 | |
| Protease inhibitor tablet | Sigma | S8820 | |
| PVDF membrane | Millipore | IPVH00010 | |
| Rocking shaker | Labnet | S2035-E | |
| SDS | Sigma | L3771 | |
| Sodium chloride | Sigma | S9888 | |
| Sodium deoxycholate | Sigma | 30970 | |
| Sodium hydroxide | Sigma | 72068 | |
| Sodium pyruvate | Gibco | 11360-070 | |
| Square Petri dish | Fisher Scientific | FB0875711A | |
| Stripping buffer | Thermo Fisher Scientific | 21059 | |
| Super Signal Femto HRP substrate | Thermo Fisher Scientific | 34580 | high-sensitivity HRP substrate |
| Tabletop centrifuge | Thermo Fisher Scientific | 75004524 | |
| Trans-Blot semi-dry system | Bio-Rad | 170-3940 | |
| Tris | Sigma | TRIS-RO | |
| Tween 20 | Sigma | P1379 | |
| Ultrapure lipopolysaccharide (LPS) from E. coli 0111:B4 | InvivoGen | tlrl-3pelps | |
| Vero cells | ATCC | CCL-81 | cell line for determining HSV1 viral titer |
Referenzen
- Alnemri, E. S., et al. Human ICE/CED-3 protease nomenclature. Cell. 87 (2), 171 (1996).
- Man, S. M., Kanneganti, T. D. Converging roles of caspases in inflammasome activation, cell death and innate immunity. Nature Reviews Immunology. 16 (1), 7-21 (2016).
- Gullett, J. M., Tweedell, R. E., Kanneganti, T. D. It’s all in the PAN: Crosstalk, plasticity, redundancies, switches, and interconnectedness encompassed by PANoptosis underlying the totality of cell death-associated biological effects. Cells. 11 (9), 1495 (2022).
- Pandian, N., Kanneganti, T. D. PANoptosis: A unique innate immune inflammatory cell death modality. Journal of Immunology. 209 (9), 1625-1633 (2022).
- Galluzzi, L., et al. Molecular mechanisms of cell death recommendations of the Nomenclature Committee on Cell Death 2018. Cell Death & Differentiation. 25 (3), 486-541 (2018).
- Shi, Y. Caspase activation: Revisiting the induced proximity model. Cell. 117 (7), 855-858 (2004).
- Galluzzi, L., Lopez-Soto, A., Kumar, S., Kroemer, G. Caspases connect cell-death signaling to organismal homeostasis. Immunity. 44 (2), 221-231 (2016).
- Fernandes-Alnemri, T., Litwack, G., Alnemri, E. S. CPP32, a novel human apoptotic protein with homology to Caenorhabditis elegans cell death protein Ced-3 and mammalian interleukin-1 beta-converting enzyme. Journal of Biological Chemistry. 269 (49), 30761-30764 (1994).
- Tewari, M., et al. Yama/CPP32 beta, a mammalian homolog of CED-3, is a CrmA-inhibitable protease that cleaves the death substrate poly(ADP-ribose) polymerase. Cell. 81 (5), 801-809 (1995).
- Nicholson, D. W., et al. Identification and inhibition of the ICE/CED-3 protease necessary for mammalian apoptosis. Nature. 376 (6535), 37-43 (1995).
- Stennicke, H. R., et al. Pro-caspase-3 is a major physiologic target of caspase-8. Journal of Biological Chemistry. 273 (42), 27084-27090 (1998).
- Twiddy, D., Cohen, G. M., Macfarlane, M., Cain, K. Caspase-7 is directly activated by the approximately 700-kDa apoptosome complex and is released as a stable XIAP-caspase-7 approximately 200-kDa complex. Journal of Biological Chemistry. 281 (7), 3876-3888 (2006).
- Kesavardhana, S., Malireddi, R. K. S., Kanneganti, T. D. Caspases in cell death, inflammation, and pyroptosis. Annual Reviews of Immunology. 38, 567-595 (2020).
- Kerr, J. F., Wyllie, A. H., Currie, A. R. Apoptosis: a basic biological phenomenon with wide-ranging implications in tissue kinetics. British Journal of Cancer. 26 (4), 239-257 (1972).
- Taylor, R. C., Cullen, S. P., Martin, S. J. Apoptosis: Controlled demolition at the cellular level. Nature Reviews Molecular Cell Biology. 9 (3), 231-241 (2008).
- Morioka, S., Maueröder, C., Ravichandran, K. S. Living on the edge: Efferocytosis at the interface of homeostasis and pathology. Immunity. 50 (5), 1149-1162 (2019).
- Wang, Y., et al. Chemotherapy drugs induce pyroptosis through caspase-3 cleavage of a gasdermin. Nature. 547 (7661), 99-103 (2017).
- Rogers, C., et al. Cleavage of DFNA5 by caspase-3 during apoptosis mediates progression to secondary necrotic/pyroptotic cell death. Nature Communications. 8, 14128 (2017).
- Kayagaki, N., et al. Caspase-11 cleaves gasdermin D for non-canonical inflammasome signalling. Nature. 526 (7575), 666-671 (2015).
- Shi, J., Gao, W., Shao, F. Pyroptosis: Gasdermin-mediated programmed necrotic cell death. Trends in Biochemical Sciences. 42 (4), 245-254 (2017).
- Sborgi, L., et al. GSDMD membrane pore formation constitutes the mechanism of pyroptotic cell death. EMBO Journal. 35 (16), 1766-1778 (2016).
- Aglietti, R. A., et al. GsdmD p30 elicited by caspase-11 during pyroptosis forms pores in membranes. Proceedings of the National Academy of Sciences of the United States of America. 113 (28), 7858-7863 (2016).
- Hagar, J. A., Powell, D. A., Aachoui, Y., Ernst, R. K., Miao, E. A. Cytoplasmic LPS activates caspase-11: implications in TLR4-independent endotoxic shock. Science. 341 (6151), 1250-1253 (2013).
- Kayagaki, N., et al. Noncanonical inflammasome activation by intracellular LPS independent of TLR4. Science. 341 (6151), 1246-1249 (2013).
- Shi, J., et al. Inflammatory caspases are innate immune receptors for intracellular LPS. Nature. 514 (7521), 187-192 (2014).
- Lamkanfi, M., et al. Targeted peptidecentric proteomics reveals caspase-7 as a substrate of the caspase-1 inflammasomes. Molecular & Cellular Proteomics. 7 (12), 2350-2363 (2008).
- Kalai, M., et al. Tipping the balance between necrosis and apoptosis in human and murine cells treated with interferon and dsRNA. Cell Death & Differentiation. 9 (9), 981-994 (2002).
- Li, C., et al. Development of atopic dermatitis-like skin disease from the chronic loss of epidermal caspase-8. Proceedings of the National Academy of Sciences of the United States of America. 107 (51), 22249-22254 (2010).
- Kovalenko, A., et al. Caspase-8 deficiency in epidermal keratinocytes triggers an inflammatory skin disease. Journal of Experimental Medicine. 206 (10), 2161-2177 (2009).
- Kang, T. B., et al. Caspase-8 serves both apoptotic and nonapoptotic roles. Journal of Immunology. 173 (5), 2976-2984 (2004).
- Oberst, A., et al. Catalytic activity of the caspase-8-FLIP(L) complex inhibits RIPK3-dependent necrosis. Nature. 471 (7338), 363-367 (2011).
- Kaiser, W. J., et al. RIP3 mediates the embryonic lethality of caspase-8-deficient mice. Nature. 471 (7338), 368-372 (2011).
- Zhang, H., et al. Functional complementation between FADD and RIP1 in embryos and lymphocytes. Nature. 471 (7338), 373-376 (2011).
- Shi, J., et al. Cleavage of GSDMD by inflammatory caspases determines pyroptotic cell death. Nature. 526 (7575), 660-665 (2015).
- Hitomi, J., et al. Identification of a molecular signaling network that regulates a cellular necrotic cell death pathway. Cell. 135 (7), 1311-1323 (2008).
- Malireddi, R. K., Ippagunta, S., Lamkanfi, M., Kanneganti, T. D. Cutting edge: Proteolytic inactivation of poly(ADP-ribose) polymerase 1 by the Nlrp3 and Nlrc4 inflammasomes. Journal of Immunology. 185 (6), 3127-3130 (2010).
- Tsuchiya, K., et al. Caspase-1 initiates apoptosis in the absence of gasdermin D. Nature Communications. 10 (1), (2019).
- Taabazuing, C. Y., Okondo, M. C., Bachovchin, D. A. Pyroptosis and apoptosis pathways engage in bidirectional crosstalk in monocytes and macrophages. Cell Chemical Biology. 24 (4), 507-514 (2017).
- Gurung, P., et al. FADD and caspase-8 mediate priming and activation of the canonical and noncanonical Nlrp3 inflammasomes. Journal of Immunology. 192 (4), 1835-1846 (2014).
- Man, S. M., et al. Inflammasome activation causes dual recruitment of NLRC4 and NLRP3 to the same macromolecular complex. Proceedings of the National Academy of Sciences of the United States of America. 111 (20), 7403-7408 (2014).
- Man, S. M., et al. Salmonella infection induces recruitment of Caspase-8 to the inflammasome to modulate IL-1beta production. Journal of Immunology. 191 (10), 5239-5246 (2013).
- Van Opdenbosch, N., et al. Caspase-1 engagement and TLR-induced c-FLIP expression suppress ASC/caspase-8-dependent apoptosis by inflammasome sensors NLRP1b and NLRC4. Cell Reports. 21 (12), 3427-3444 (2017).
- Pierini, R., et al. AIM2/ASC triggers caspase-8-dependent apoptosis in Francisella-infected caspase-1-deficient macrophages. Cell Death & Differentiation. 19 (10), 1709-1721 (2012).
- Lee, S., et al. AIM2 forms a complex with pyrin and ZBP1 to drive PANoptosis and host defence. Nature. 597 (7876), 415-419 (2021).
- Sagulenko, V., et al. AIM2 and NLRP3 inflammasomes activate both apoptotic and pyroptotic death pathways via ASC. Cell Death & Differentiation. 20 (9), 1149-1160 (2013).
- Lukens, J. R., et al. Dietary modulation of the microbiome affects autoinflammatory disease. Nature. 516 (7530), 246-249 (2014).
- Gurung, P., Burton, A., Kanneganti, T. D. NLRP3 inflammasome plays a redundant role with caspase 8 to promote IL-1beta-mediated osteomyelitis. Proceedings of the National Academy of Sciences of the United States of America. 113 (16), 4452-4457 (2016).
- Kuriakose, T., et al. ZBP1/DAI is an innate sensor of influenza virus triggering the NLRP3 inflammasome and programmed cell death pathways. Science Immunology. 1 (2), (2016).
- Malireddi, R. K. S., et al. TAK1 restricts spontaneous NLRP3 activation and cell death to control myeloid proliferation. Journal of Experimental Medicine. 215 (4), 1023-1034 (2018).
- Malireddi, R. K. S., et al. Innate immune priming in the absence of TAK1 drives RIPK1 kinase activity-independent pyroptosis, apoptosis, necroptosis, and inflammatory disease. Journal of Experimental Medicine. 217 (3), (2020).
- Christgen, S., et al. Identification of the PANoptosome: A molecular platform triggering pyroptosis, apoptosis, and necroptosis (PANoptosis). Frontiers in Cellular and Infection Microbiology. 10, 237 (2020).
- Malireddi, R. K. S., et al. RIPK1 distinctly regulates Yersinia-induced inflammatory cell death, PANoptosis. Immunohorizons. 4 (12), 789-796 (2020).
- Zheng, M., Karki, R., Vogel, P., Kanneganti, T. D. Caspase-6 is a key regulator of innate immunity, inflammasome activation, and host defense. Cell. 181 (3), 674-687 (2020).
- Karki, R., et al. ADAR1 restricts ZBP1-mediated immune response and PANoptosis to promote tumorigenesis. Cell Reports. 37 (3), 109858 (2021).
- Wang, Y., et al. Single cell analysis of PANoptosome cell death complexes through an expansion microscopy method. Cellular and Molecular Life Sciences. 79 (10), 531 (2022).
- Malireddi, R. K. S., et al. Inflammatory cell death, PANoptosis, mediated by cytokines in diverse cancer lineages inhibits tumor growth. Immunohorizons. 5 (7), 568-580 (2021).
- Kesavardhana, S., et al. The Zα2 domain of ZBP1 is a molecular switch regulating influenza-induced PANoptosis and perinatal lethality during development. Journal of Biological Chemistry. 295 (24), 8325-8330 (2020).
- Banoth, B., et al. ZBP1 promotes fungi-induced inflammasome activation and pyroptosis, apoptosis, and necroptosis (PANoptosis). Journal of Biological Chemistry. 295 (52), 18276-18283 (2020).
- Karki, R., et al. Interferon regulatory factor 1 regulates PANoptosis to prevent colorectal cancer. JCI Insight. 5 (12), 136720 (2020).
- Zheng, M., et al. Impaired NLRP3 inflammasome activation/pyroptosis leads to robust inflammatory cell death via caspase-8/RIPK3 during coronavirus infection. Journal of Biological Chemistry. 295 (41), 14040-14052 (2020).
- Karki, R., et al. Synergism of TNF-α and IFN-γ triggers inflammatory cell death, tissue damage, and mortality in SARS-CoV-2 infection and cytokine shock syndromes. Cell. 184 (1), 149-168 (2021).
- Karki, R., et al. ZBP1-dependent inflammatory cell death, PANoptosis, and cytokine storm disrupt IFN therapeutic efficacy during coronavirus infection. Science Immunology. 7 (74), (2022).
- Jiang, W., Deng, Z., Dai, X., Zhao, W. PANoptosis: A new insight into oral infectious diseases. Frontiers in Immunology. 12, 789610 (2021).
- Chi, D., et al. Real-time induction of macrophage apoptosis, pyroptosis, and necroptosis by Enterococcus faecalis OG1RF and two root canal isolated strains. Frontiers in Cellular and Infection Microbiology. 11, 720147 (2021).
- Lin, J. F., et al. Phosphorylated NFS1 weakens oxaliplatin-based chemosensitivity of colorectal cancer by preventing PANoptosis. Signal Transduction and Targeted Therapy. 7 (1), 54 (2022).
- Song, M., et al. Self-assembled polymeric nanocarrier-mediated co-delivery of metformin and doxorubicin for melanoma therapy. Drug Delivery. 28 (1), 594-606 (2021).
- Boucher, D., Chan, A., Ross, C., Schroder, K. Quantifying caspase-1 activity in murine macrophages. Methods in Molecular Biology. 1725, 163-176 (2018).
- Boucher, D., Duclos, C., Denault, J. B. General in vitro caspase assay procedures. Methods in Molecular Biology. 1133, 3-39 (2014).
- Kaushal, V., Herzog, C., Haun, R. S., Kaushal, G. P. Caspase protocols in mice. Methods in Molecular Biology. 1133, 141-154 (2014).
- Swacha, P., Gekara, N. O., Erttmann, S. F. Biochemical and microscopic analysis of inflammasome complex formation. Methods in Enzymology. 625, 287-298 (2019).
- Talley, S., et al. A caspase-1 biosensor to monitor the progression of inflammation in vivo. Journal of Immunology. 203 (9), 2497-2507 (2019).
- Pelegrin, P., Barroso-Gutierrez, C., Surprenant, A. P2X7 receptor differentially couples to distinct release pathways for IL-1beta in mouse macrophage. Journal of Immunology. 180 (11), 7147-7157 (2008).
- Yu, J. W., et al. Cryopyrin and pyrin activate caspase-1, but not NF-kappaB, via ASC oligomerization. Cell Death & Differentiation. 13 (2), 236-249 (2006).
- Henry, C. M., Martin, S. J. Caspase-8 acts in a non-enzymatic role as a scaffold for assembly of a pro-inflammatory "FADDosome" complex upon TRAIL stimulation. Molecular Cell. 65 (4), 715-729 (2017).
- Hoffmann, E., Neumann, G., Kawaoka, Y., Hobom, G., Webster, R. G. A DNA transfection system for generation of influenza A virus from eight plasmids. Proceedings of the National Academy of Sciences of the United States of America. 97 (11), 6108-6113 (2000).
- Simpson, R. J. Homogenization of mammalian tissue. Cold Spring Harbor Protocols. (7), (2010).

