An Electroporation Method to Transform Rickettsia spp. with a Fluorescent Protein-Expressing Shuttle Vector in Tick Cell Lines
Summary
Electroporation is a rapid, broadly adopted method for introducing exogenous DNA into the genus Rickettsia. This protocol provides a useful electroporation method for the transformation of obligate intracellular bacteria in the genus Rickettsia.
Abstract
Rickettsioses are caused by a broad range of obligate intracellular bacteria belonging to the genus Rickettsia that can be transmitted to vertebrate hosts through the bite of infected arthropod vectors. To date, emerging or re-emerging epidemic rickettsioses remain a public health risk due to the difficulty in diagnosis, as diagnostic methods are limited and not standardized or universally accessible. Misdiagnosis resulting from a lack of recognition of the signs and symptoms may result in delayed antibiotic treatment and poor health outcomes. A comprehensive understanding of Rickettsia characteristics would ultimately improve clinical diagnosis, assessment, and treatment with improved control and prevention of the disease.
Functional studies of rickettsial genes are crucial for understanding their role in pathogenesis. This paper describes a procedure for the electroporation of the Rickettsia parkeri strain Tate’s Hell with the shuttle vector pRAM18dSFA and the selection of transformed R. parkeri in tick cell culture with antibiotics (spectinomycin and streptomycin). A method is also described for the localization of transformed R. parkeri in tick cells using confocal immunofluorescence microscopy, a useful technique for checking transformation in vector cell lines. Similar approaches are also suitable for the transformation of other rickettsiae.
Introduction
Rickettsioses are caused by a broad range of obligate intracellular bacteria that belong to the genus Rickettsia (family Rickettsiaceae, order Rickettsiales). The genus Rickettsia is classified into four major groups based on phylogenetic characteristics1,2: the spotted fever group (SFG), which contains those rickettsiae that cause the most severe and fatal tick-borne rickettsioses (e.g., Rickettsia rickettsii, the causative agent of Rocky Mountain Spotted Fever), the typhus group (TG, e.g., Rickettsia prowazekii, the agent of epidemic typhus), the transitional group (TRG, e.g., Rickettsia felis, the causative agent of flea-borne spotted fever), and the ancestral group (AG, e.g., Rickettsia bellii).
Among the oldest known vector-borne diseases, rickettsioses are mainly acquired following transmission of the pathogens through the bites of infected arthropod vectors, including ticks, fleas, lice, and mites3,4. Although the discovery of effective antibiotics improved treatment outcomes, emerging and re-emerging epidemic rickettsioses continue to challenge traditional prevention and control strategies. Thus, a comprehensive understanding of rickettsia/host/vector interactions would ultimately establish a strong foundation for developing new approaches to prevent and cure these ancient diseases.
In nature, horizontal gene transfer (HGT) in bacteria occurs through conjugation, transduction, and transformation5. In vitro bacterial transformation utilizes these HGT concepts, although the intracellular nature of rickettsiae presents some challenges. The restricted growth conditions and poorly understood conjugation and transduction systems in different species of rickettsiae have prevented the application of conjugation and transduction methods in rickettsiae6,7,8. Compared with other obligate intracellular bacterial genera (e.g., Chlamydia, Coxiella, Anaplasma, and Ehrlichia), the genus Rickettsia differs with regard to the growth and replication strategies within the cell cytoplasm, which imposes specific challenges to the genetic modification of rickettsiae due to their unique lifestyle features9.
The initial hurdle to overcome when attempting the genetic modification of rickettsiae is to achieve successful transformation. Thus, designing a feasible approach with high transformation efficiency would be extremely valuable for developing genetic tools for rickettsiae. Here, we focus on electroporation, a broadly recognized transformation method that has been used to introduce exogenous DNA successfully into several species of rickettsiae, including Rickettsia prowazekii, Rickettsia typhi, Rickettsia conorii, Rickettsia parkeri, Rickettsia montanensis, Rickettsia bellii, Rickettsia peacockii, and Rickettsia buchneri10,11,12,13,14,15,16.
This paper describes a procedure for the electroporation of the R. parkeri strain Tate's Hell (accession: GCA_000965145.1) with the shuttle vector pRAM18dSFA derived from the Rickettsia amblyommatis strain AaR/SC plasmid pRAM18 engineered to encode mKATE, a far-red fluorescent protein, and aadA, conferring spectinomycin and streptomycin resistance13,15,20. Transformed R. parkeri are viable and stably maintained under antibiotic selection in tick cell lines. In addition, we show that the localization of transformed R. parkeri in live tick cells via confocal microscopy can be used to assess the quality of transformation rates in vector cell lines.
Protocol
1. Propagation and purification of R. parkeri from tick cell culture
NOTE: All cell culture procedures are to be performed in a class II biosafety cabinet.
- Preparing R. parkeri-infected tick cells
- Grow ISE6 cells in 25 cm2 cell culture flasks at 34 °C in 5 mL of L15C300 medium17, supplemented with 5% fetal bovine serum (FBS), 5% tryptose phosphate broth (TPB), and 0.1% lipoprotein concentrate (LPC).
NOTE: ISE6 (Ixodes scapularis embryo-derived cell line) is a tick cell line extensively used in many laboratories and is, thus, an essential model to study tick-pathogen interactions17,18,19. The growth rate of ISE6 cells is slower than that of mammalian cells, with a population doubling time of ≥72 h. For example, ISE6 cells subcultivated from a 100% confluent culture at a ratio of 1:5 will take 3-4 weeks to regain 100% confluency. Healthy ISE6 cells will be generally rounded with numerous adhering filaments17. - Count the ISE6 cells with a hemocytometer under a light microscope. Use at a concentration of ~106 cells/mL (100% confluent).
- Gently rinse the cells from the flask with medium and disperse the cells by pipetting to generate a homogenous cell suspension. Add 20 µL of cell suspension between the hemocytometer and cover glass to count the cells using a hemocytometer.
- Add host cell-free wild-type R. parkeri20 (average number: 5 × 107-10 × 107) to the ISE6 cell cultures in 25 cm2 cell culture flasks. Incubate infected R. parkeri cell cultures at 34 °C in 5 mL of complete medium consisting of L15C300 medium, 10% FBS, 5% TPB, 0.1% LPC, 0.25% sodium bicarbonate (NaHCO3), and 25 mM HEPES until 90%-100% of the cells are infected.
NOTE: The infection rate of R. parkeri in ISE6 cells is determined by Giemsa staining, and the incubation time to reach 90%-100% infection ranges from 5 days to 7 days on average. - Determine the infection rate via Giemsa staining.
- In the biosafety cabinet, resuspend the infected cell cultures and transfer 50 µL of the suspension to a 1.5 mL microcentrifuge tube.
- Dilute the suspension with complete medium (1:5 dilution) and centrifuge 100 µL onto glass microscope slides using a centrifuge at 113 × g for 5 min. To keep live R. parkeri contained, use a centrifuge with a sealed rotor that is designed to be removed from the centrifuge. This enables the transport of the sealed rotor in and out of the hood.
NOTE: As the R. parkeri are still alive before centrifugation, the sample needs to be contained until the rickettsia are killed. Thus, the centrifuge used to spin the R. parkeri culture onto the slide must have a sealed rotor that is designed to be lifted out of the centrifuge. With the rotor in the laminar flow hood, load the samples into the funnel-microscope slide assembly, reseal the rotor, and place the sealed rotor back into the centrifuge. Then, turn on the centrifuge, and the samples are deposited onto the microscope slides. After the run is finished, remove the sealed rotor from the centrifuge and place it in the laminar flow hood. There, open the rotor lid, and unload the funnel-microscope slide assembly inside the hood. Once dry, immerse the glass microscope slides in absolute methanol, which kills all pathogens. The funnels and the metal carriers are submerged in diluted DMQ, which kills R. parkeri. - Air-dry the slides and fix in absolute methanol for 5 min at room temperature (RT).
- Stain the slides with Giemsa stain (4% in Sørenson's buffer, pH 6.6) for 30 min at 37 °C and rinse with water for 5 s.
- Determine the infection percentage (number of cells infected with rickettsiae/100 cells) by light microscopy.
NOTE: Figure 1 shows typical Giemsa stain results.
- Grow ISE6 cells in 25 cm2 cell culture flasks at 34 °C in 5 mL of L15C300 medium17, supplemented with 5% fetal bovine serum (FBS), 5% tryptose phosphate broth (TPB), and 0.1% lipoprotein concentrate (LPC).
- Preparing cell-free R. parkeri
NOTE: When 90%-100% of the cells in the culture are infected, the rickettsiae should be isolated from the ISE6 cells and electroporation should be performed.- Add sterile 60-90 silicon carbide grit to sterile 2 mL microcentrifuge tubes to a volume of approximately 0.2 mL.
- Remove 2 mL of medium from 100% R. parkeri-infected cultures (step 1.1.3) and resuspend the cells in the remaining 3 mL of medium.
- Transfer equal portions of this suspension to tubes prepared in step 1.2.1.
- Vortex each tube at high speed for 30 s and then place the tubes on ice. Let the grit settle within seconds by gravity.
- In a class II biosafety cabinet, have a sterile 5 mL Luer-lock syringe ready with the plunger extended and the end of the plunger secured in a polystyrene base for 15 mL conical tubes.
- Using a sterile, 1 mL barrier pipet tip fitted to a suitable pipettor, remove the supernatant of the vortexed cell lysate from the tube, taking care not to aspirate any grit. Insert the pipet tip into the syringe hub opening and eject the contents into the syringe with gentle pressure. Alternatively, use a sterile, barrier 2 mL Pasteur pipette operated with a rubber bulb.
- Attach a sterilized 2 µm pore size filter onto the syringe hub and filter the R. parkeri culture from step 1.2.6 into sterile 1.5 mL tubes.
- Collect the R. parkeri by centrifugation at 13,600 × g at 4 °C for 5 min, and discard the supernatant.
- Resuspend the pellet in 1.2 mL of ice-cold 300 mM sucrose and centrifuge again at 13,600 × g at 4 °C for 5 min. Repeat the resuspension and centrifugation for a total of two sucrose washes.
- Combine the R. parkeri pellets into one chilled, sterile 1.5 mL microcentrifuge tube in 50 µL of ice-cold 300 mM sucrose per transformation. As one 25 cm2 flask of R. parkeri provides enough rickettsiae for two to three transformations, add 100-150 µL of 300 mM cold sucrose.
NOTE: If desired, the number of rickettsiae can be calculated using a Petroff-Hausser counting chamber. When using an initial inoculation of 5 × 107 -10 × 107 cell-free R. parkeri, it will take 5-7 days on average to reach 90%-100% infection, yielding approximately 5 × 107-10 × 1010 cell-free R. parkeri. - Gently resuspend the pellet by pipetting until it is thoroughly dispersed. Divide the sample into 50 µL portions in chilled, sterile 1.5 mL microcentrifuge tubes.
NOTE: If desired, rickettsial viability may be assessed by flow cytometry following staining with a fluorescent dye 21.
2. Transformation of R. parkeri with the pRAM18dSFA plasmid
- On ice, add 3 µg of endotoxin-free pRAM18dSFA plasmid DNA13,15,20 to each tube containing 50 µL of R. parkeri suspension and stir the mixture with the pipet tip gently but thoroughly.
NOTE: When preparing plasmid DNA, use a purification kit that includes an endotoxin-removal step to yield endotoxin-free plasmid DNA23. We found that a range of plasmid concentrations of between 1 µg and 3 µg per 50 µL of R. parkeri suspension resulted in successful transformations, whereas increasing the amount of plasmid to 10 µg inhibited transformation. - Transfer the above mixture of DNA and R. parkeri into a chilled, sterile 0.1 cm gap electroporation cuvette (gently tap the cuvette until the mixture is evenly distributed) and let it sit on ice for 10-30 min.
- Remove the medium from a 100% confluent culture of ISE6 cells and resuspend the cell layers in 1.5 mL of fresh medium with NaHCO3 and HEPES buffer (described above in step 1.1.3). Use one flask of confluent cells per transformation.
- Transfer the resuspended cells to a sterile 2 mL microcentrifuge tube (one tube per 25 cm2 flask of ISE6 cells).
- Electroporate the R. parkeri/pRAM18dSFA mixture at 1.8 KV, 200 ohms, 25 µF using an electroporator.
- Using a sterile, extended fine-tip transfer pipet, transfer a small amount of the ISE6 cell suspension (step 2.4) into the cuvette and gently pull the liquid up and down to wash out the cuvette. Transfer the mixture into the 2 mL microcentrifuge tube containing the remainder of the ISE6 cell suspension and gently mix the transformation mixture with the cells.
- Centrifuge the transformed samples at 700 × g for 2 min at RT, and then increase the centrifugation speed to 13,600 × g for 1 min more at RT.
NOTE: The two centrifugation speeds promote rickettsial binding with and entry into the cells: 700 × g is used to pull down the ISE6 cells, while 13,600 × g is used to pull down the rickettsiae. - Let samples sit at RT or 34 °C for 15 min to 1 h.
- Resuspend the transformants (two or three) in ISE6 cells with a sterile 2 mL barrier pipet tip fitted to a pipettor and transfer into two or three 25 cm2 cell culture flasks containing 3.5 mL of fresh medium with NaHCO3 and HEPES buffer. Alternatively, use a sterile, barrier 2 mL Pasteur pipette operated with a rubber bulb.
- Rock the flasks to spread the mixture evenly and then incubate at 34 °C.
- After 16-24 h, add 10 µL of spectinomycin (50 mg/mL) and 10 µL of streptomycin (50 mg/mL) to each flask in step 2.10.
3. Observation of the transformed R. parkeri
NOTE: Use an epifluorescence microscope with rhodamine/TRITC filters to observe the flasks prepared in step 2.11 after 3-7 days. Once plaques are evident in the cultures (5-14 days), transformed R. parkeri can be seen that express the red fluorescent protein mKATE, encoded on the pRAM18dSFA plasmid.
- Confocal microscopy
- Resuspend the cell cultures from the step 2.11 flasks and mix 100 µL of these cell cultures with 5 µL of Hoechst 33342 solution for 10-30 min at RT in the dark.
NOTE: Hoechst 33342 is a cell-permeable nuclear counterstain that binds to live tick cell DNA and emits blue fluorescence. - Centrifuge 50 µL of the mixture onto glass microscope slides using a centrifuge at 5 × g for 3 min.
- Add 3 µL of 1x PBS to the spot of cells deposited on the slide, overlay with a cover slip, and view with a confocal microscope (60x objective). Use the following excitation and emission parameters for fluorescent imaging: for 4′,6-diamidino-2-phenylindole (DAPI), excitation at 350 nm, emission at 470 nm; for tetramethylrhodamine (TRITC), excitation at 557 nm, emission at 576 nm.
- Resuspend the cell cultures from the step 2.11 flasks and mix 100 µL of these cell cultures with 5 µL of Hoechst 33342 solution for 10-30 min at RT in the dark.
Representative Results
The morphology of R. parkeri in ISE6 cells under a light microscope after Giemsa staining are shown in Figure 1. In Figure 2, transformed R. parkeri expressing red fluorescence protein in ISE6 cells are shown using confocal microscopy. There is a substantial increase in the infection rate of transformed R. parkeri (red) in ISE6 cells (blue, corresponds to the nuclei) from (A) day 7 to (B) day 10 of incubation.
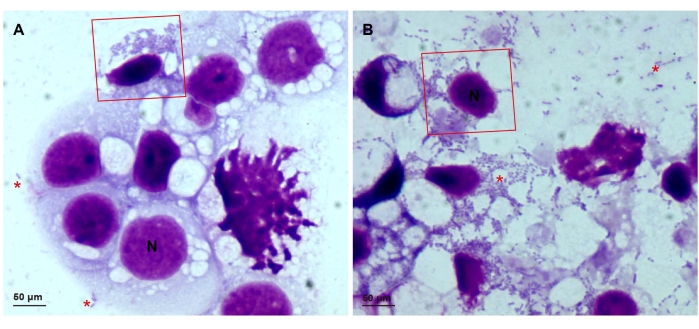
Figure 1: Slides showing Giemsa-stained Rickettsia parkeri in ISE6 cells. ISE6 cells infected with wild-type R. parkeri at (A) low infection and (B) 90%-100% infection rates. The nuclei of the ISE6 cells stain to dark purple with Giemsa; the R. parkeri appear as dark purple rods. The red boxes show intracellular rickettsiae, and the red asterisks indicate extracellular rickettsiae. All images were taken on a light microscope with a 100x objective. Scale bars = 50 µm. Abbreviation: N = nuclei. Please click here to view a larger version of this figure.
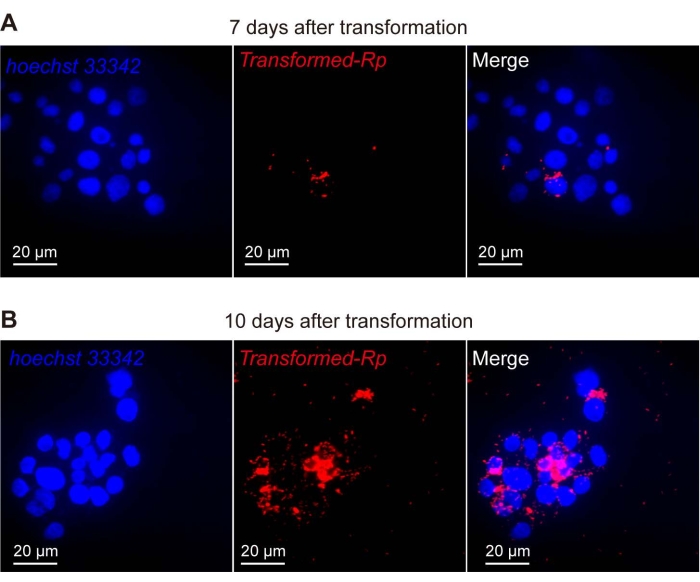
Figure 2: Transformed Rickettsia parkeri expressing red fluorescence protein in ISE6 cells. pRAM18dSFA-transformed R. parkeri (red) in ISE6 cells (blue), stained with Hoechst 33342 and detected by confocal microscopy (A) 7 days and (B) 10 days after transformation. Hoechst 33342 stains the nuclei, and its excitation and emission wavelengths are similar to DAPI. The merged signals are a combination of DAPI and TRITC filter images. All images were taken on a confocal microscope with a 60x objective. Scale bars = 20 µm. Abbreviations: DAPI = 4',6-diamidino-2-phenylindole; TRITC = tetramethylrhodamine. Please click here to view a larger version of this figure.
Discussion
Here, we demonstrate a method for introducing exogenous DNA encoded on the shuttle plasmid pRAM18dSFA into rickettsiae using electroporation. In this procedure, cell-free rickettsiae were purified from host cells, transformed with a rickettsial shuttle vector, and released onto tick cells for infection. Also described is a confocal immunofluorescence procedure to detect red fluorescence protein-expressing R. parkeri in tick cells. Similar methods are applicable to other Rickettsia species and with further modifications might be adaptable to other obligate intracellular bacteria that are capable of maintaining a plasmid.
A successful transformation in this protocol requires three components: exogenous DNA, Rickettsia spp. bacteria, and a suitable type of host cell22. Depending on the specific research goal, these components can be changed. First, the exogenous DNA was introduced via the pRAM18dSFA plasmid, which contains spectinomycin- and streptomycin-selectable markers and expresses mKATE (a far-red fluorescent protein). This plasmid also confers ampicillin resistance for growth in Escherichia coli. As shown in previous studies, it is possible to replace antibiotic-resistance markers and the genes for fluorescent proteins13. Second, ISE6 cells were selected to represent transformed cell lines as this cell line is extensively used in many laboratories and is an essential model for the study of vector-pathogen interactions17,18,19. Other species of tick16 or mammalian cells23,24 can also be used to grow rickettsiae for transformation. Finally, in this protocol, R. parkeri was used, which can be manipulated in a biosafety level 2 (BSL-2) laboratory using a class II biosafety cabinet with proper safety precautions. Other more pathogenic Rickettsia spp. (e.g., R. rickettsii; R. prowazekii) require a BSL-3 setting for manipulation; therefore, a more stringent biosafety standard protocol must be used when working with such agents.
Although this protocol can be used as a standard procedure for rickettsiae transformation, several key technical steps require particular attention. First, a successful purification step must ensure cell-free rickettsiae survival and infectivity9; hence, the most critical aspect of the transformation procedure is to isolate infectious, cell-free rickettsiae. Any residual salt in the purified rickettsiae will result in arcing and transformation failure. Therefore, washing the isolated cell-free rickettsiae with sucrose is required to remove all salts. The cold sucrose solution also protects the rickettsial membrane integrity in the extracellular condition.
When 90%-100% of the cells in a culture are infected, the rickettsiae should be isolated from the ISE6 cells, and electroporation should be performed without delay and without interruptions, as rickettsiae are intracellular organisms and do not survive well outside cells. Although rickettsial cultures can be stored at 4 °C, this type of material should not be used for electroporation. A culture that has been stored at 4 °C can be used to inoculate a fresh layer of ISE6 cells, which can then be used for electroporation once infection has reached 90%-100%.
Second, the electroporation settings, including voltage, resistance, capacitance, and time constant, are specific to different rickettsial species. For example, the field strength required during electroporation is dependent on the size of the rickettsiae and exogenous DNA7. Finally, it is important to maintain the transformed rickettsiae under antibiotic selection to prevent the loss of exogenous plasmids during multiple passages in host cells9. The concentration and type of antibiotic employed are dependent on the Rickettsia species being transformed, as previously described13,15,16,23, and it is important that the antibiotic marker selected should not be one that is applied in clinical treatments.
To follow this protocol, both spectinomycin and streptomycin should be used for selection until all the residual untransformed rickettsiae have been eliminated. Combined, the two antibiotics kill both intracellular and extracellular rickettsiae and reduce the probability of resistant wild-type rickettsiae emerging. In addition, using both of these antibiotics does not affect downstream experiments, such as the ability to use them separately to select for two different plasmids, since the aadA gene confers resistance to both spectinomycin and streptomycin. The two antibiotics used in this protocol do not confer resistance to antibiotics used in the clinical treatment of rickettsial disease.
The ISE6 tick cell line utilized in this study has unique advantages. First, ISE6 cells were isolated from the black-legged tick Ixodes scapularis Say (Acari: Ixodidae), the principal vector for seven human pathogens in the United States. Second, ISE6 is an extensively used tick cell line in many laboratories and has been successfully used to recover many tick-associated bacteria (Rickettsia, Anaplasma, and Ehrlichia) following electroporation. Third, some pathogenic rickettsiae can only be propagated in tick cells but not in mammalian cells17,18,19,24. However, tick cell lines are relatively fragile compared to mammalian cells and have more intensive culture requirements25,26,27,28. In addition, the growth rate of ISE6 cells is significantly slower than that of mammalian cell lines, even if seeded at the same initial density, although slow replication can be advantageous when the rickettsial transformants exhibit reduced growth.
This protocol also provides a method to evaluate the transformation efficiency of rickettsiae in live cells at critical experimental steps, which helps to optimize electroporation settings or in testing the efficiency of various cell lines for recovering transformants. Nevertheless, this protocol has limitations, especially for the quantification of the transformants obtained. Transformation efficiency as an indicator of a successful transformation could be demonstrated in a future study. Rickettsial viability could be assessed using a suitable staining kit and flow cytometry21 to quantify the transformants that are obtained. Two parameters would be used to determine transformation efficiency-the number of live rickettsiae used for transformation and the number of transformed rickettsiae that express mKATE.
In addition, image analysis with fluorescence signals could be used to compare the transformation rates of different plasmids. As the underlying mechanism of rickettsial plasmid maintenance is poorly understood, well-characterized transformation systems for introducing exogenous plasmids can be valuable tools for further investigation. Furthermore, the direct visualization of fluorescent protein-expressing rickettsiae in cells or tissues will improve our understanding of rickettsia/host/vector interactions. This will provide information for designing strategies to control and prevent rickettsioses.
Data availability:
All data underlying the results of this study are publicly available.
Offenlegungen
The authors have nothing to disclose.
Acknowledgements
We thank Timothy J. Kurtti and Benjamin Cull for their insightful discussions and suggestions. This study was financially supported by a grant to U.G.M. from the NIH (2R01AI049424) and a grant to U.G.M. from the Minnesota Agricultural Experiment Station (MIN-17-078).
Materials
| 0.1 cm gap gene pulser electroporation cuvette | Bio-Rad | 1652083 | |
| 2 μm pore size filter | GE Healthcare Life Sciences Whatman | 6783-2520 | |
| 5 mL Luer-lock syringe | BD | 309646 | |
| 60-90 silicon carbide grit | LORTONE, inc | 591-056 | |
| absolute methanol | Fisher Scientific | A457-4 | |
| Bacto tryptose phosphate broth | BD | 260300 | |
| Cytospin centrifuge Cytospin4 | Thermo Fisher Scientific | A78300003 | The rotor is detatchable so the whole rotor can be put into the hood to load infectious samples |
| EndoFree Plasmid Maxi Kit (10) | QIAGEN | 12362 | used to obtain endotoxin-free pRAM18dSFA plasmid |
| extended fine tip transfer pipet | Perfector Scientific | TP03-5301 | |
| fetal bovine serum | Gemini Bio | 900-108 | The FBS batch has to be tested to make sure ISE6 cells will grow well in it. |
| Gene Pulser II electroporator with Pulse Controller PLUS | Bio-Rad | 165-2105 & 165-2110 | |
| hemocytometer | Thermo Fisher Scientific | 267110 | |
| HEPES | Millipore-Sigma | H4034 | |
| ImageJ Fiji | National Institute of Health | raw image editing | |
| KaryoMAX Giemsa stain | Gibco | 2021-10-30 | |
| Leibovitz's L-15 medium | Gibco | 41300039 | |
| lipoprotein concentrate | MP Biomedicals | 191476 | |
| Nikon Diaphot | Nikon | epifluorescence microscope | |
| NucBlue Live ReadyProbes Reagent | Thermo Fisher Scientific | R37605 | |
| Olympus Disc Scanning Unit (DSU) confocal microscope | Olympus | ||
| Petroff-Hausser Counting Chamber | Hausser Scientific | Chamber 3900 | |
| sodium bicarbonate | Millipore-Sigma | S5761 | |
| Vortex | Fisher Vortex Genie 2 | 12-812 |
Referenzen
- Gillespie, J. J., et al. Plasmids and rickettsial evolution: Insight from Rickettsia felis. PLoS One. 2 (3), 266 (2007).
- Murray, G. G., Weinert, L. A., Rhule, E. L., Welch, J. J. The phylogeny of Rickettsia using different evolutionary signatures: How tree-like is bacterial evolution. Systematic Biology. 65 (2), 265-279 (2016).
- Parola, P., Paddock, C. D., Raoult, D. Tick-borne rickettsioses around the world: Emerging diseases challenging old concepts. Clinical Microbiology Reviews. 18 (4), 719-756 (2005).
- Fang, R., Blanton, L. S., Walker, D. H. Rickettsiae as emerging infectious agents. Clinics in Laboratory Medicine. 37 (2), 383-400 (2017).
- Von Wintersdorff, C. J., et al. Dissemination of antimicrobial resistance in microbial ecosystems through horizontal gene transfer. Frontiers in Microbiology. 7, 173 (2016).
- Winkler, H. H. Rickettsia species (as organisms). Annual Review of Microbiology. 44 (1), 131-153 (1990).
- Wood, D. O., Azad, A. F. Genetic manipulation of rickettsiae: A preview. Infection and Immunity. 68 (11), 6091-6093 (2000).
- Perlman, S. J., Hunter, M. S., Zchori-Fein, E. The emerging diversity of Rickettsia. Proceedings of the Royal Society B: Biological Sciences. 273 (1598), 2097-2106 (2006).
- McClure, E. E., et al. Engineering of obligate intracellular bacteria: Progress, challenges and paradigms. Nature Reviews Microbiology. 15 (9), 544-558 (2017).
- Rachek, L. I., Tucker, A. M., Winkler, H. H., Wood, D. O. Transformation of Rickettsia prowazekii to rifampin resistance. Journal of Bacteriology. 180 (8), 2118-2124 (1998).
- Troyer, J. M., Radulovic, S., Azad, A. F. Green fluorescent protein as a marker in Rickettsia typhi transformation. Infection and Immunity. 67 (7), 3308-3311 (1996).
- Kim, H. K., Premaratna, R., Missiakas, D. M., Schneewind, O. Rickettsia conorii O antigen is the target of bactericidal Weil-Felix antibodies. Proceedings of the National Academy of Sciences of the United States of America. 116 (39), 19659-19664 (2019).
- Burkhardt, N. Y., et al. Development of shuttle vectors for transformation of diverse Rickettsia species. PLoS One. 6 (12), 29511 (2011).
- Welch, M. D., Reed, S. C., Lamason, R. L., Serio, A. W. Expression of an epitope-tagged virulence protein in Rickettsia parkeri using transposon insertion. PloS One. 7 (5), 37310 (2012).
- Oliver, J. D., Burkhardt, N. Y., Felsheim, R. F., Kurtti, T. J., Munderloh, U. G. Motility characteristics are altered for Rickettsia bellii transformed to overexpress a heterologous rickA gene. Applied and Environmental Microbiology. 80 (3), 1170-1176 (2014).
- Kurtti, T. J., Burkhardt, N. Y., Heu, C. C., Munderloh, U. G. Fluorescent protein expressing Rickettsia buchneri and Rickettsia peacockii for tracking symbiont-tick cell interactions. Veterinary Sciences. 3 (4), 34 (2016).
- Oliver, J. D., Chávez, A. S., Felsheim, R. F., Kurtti, T. J., Munderloh, U. G. An Ixodes scapularis cell line with a predominantly neuron-like phenotype. Experimental and Applied Acarology. 66 (3), 427-442 (2015).
- Grabowski, J. M., Gulia-Nuss, M., Kuhn, R. J., Hill, C. A. RNAi reveals proteins for metabolism and protein processing associated with Langat virus infection in Ixodes scapularis (black-legged tick) ISE6 cells. Parasites & Vectors. 10 (1), 1-4 (2017).
- Al-Rofaai, A., Bell-Sakyi, L. Tick cell Lines in research on tick control. Frontiers in Physiology. 11, 152 (2020).
- Wang, X. R., et al. Mitochondrion-dependent apoptosis is essential for Rickettsia parkeri infection and replication in vector cells. mSystems. 6 (2), 01209-01220 (2021).
- Berney, M., Hammes, F., Bosshard, F., Weilenmann, H. U., Egli, T. Assessment and interpretation of bacterial viability by using the LIVE/DEAD BacLight Kit in combination with flow cytometry. Applied and Environmental Microbiology. 73 (10), 3283-3290 (2007).
- Riley, S. P., Macaluso, K. R., Martinez, J. J. Electrotransformation and clonal isolation of Rickettsia species. Current Protocols in Microbiology. 39, 1-20 (2015).
- Burkhardt, N. Y., et al. Examination of rickettsial host range for shuttle vectors based on dnaA and parA genes from the pRM plasmid of Rickettsia monacensis. Applied and Environmental Microbiology. 88 (7), 00210-00222 (2022).
- Kurtti, T. J., et al. Rickettsia buchneri sp. nov., a rickettsial endosymbiont of the blacklegged tick Ixodes scapularis. International Journal of Systematic and Evolutionary Microbiology. 65, 965 (2015).
- Munderloh, U. G., Kurtti, T. J. Formulation of medium for tick cell culture. Experimental & Applied Acarology. 7 (3), 219-229 (1989).
- Bell-Sakyi, L. Continuous cell lines from the tick Hyalomma anatolicum anatolicum. The Journal of Parasitology. 77 (6), 1006-1008 (1991).
- Mattila, J. T., Burkhardt, N. Y., Hutcheson, H. J., Munderloh, U. G., Kurtti, T. J. Isolation of cell lines and a rickettsial endosymbiont from the soft tick Carios capensis (Acari: Argasidae: Ornithodorinae). Journal of Medical Entomology. 44 (6), 1091-1101 (2007).
- Bell-Sakyi, L., Růžek, D., Gould, E. A. Cell lines from the soft tick Ornithodoros moubata. Experimental and Applied Acarology. 49 (3), 209-219 (2009).

