Using Human Intestinal Organoids to Understand the Small Intestine Epithelium at the Single Cell Transcriptional Level
Summary
The protocol combines human intestinal organoid technology with single cell transcriptomic analysis to provide significant insight into previously unexplored intestinal biology.
Abstract
Single cell transcriptomics has revolutionized our understanding of the cell biology of the human body. State-of-the-art human small intestinal organoid cultures provide ex vivo model systems that bridge the gap between animal models and clinical studies. The application of single cell transcriptomics to human intestinal organoid (HIO) models is revealing previously unrecognized cell biology, biochemistry, and physiology of the GI tract. The advanced single cell transcriptomics platforms use microfluidic partitioning and barcoding to generate cDNA libraries. These barcoded cDNAs can be easily sequenced by next generation sequencing platforms and used by various visualization tools to generate maps. Here, we describe methods to culture and differentiate human small intestinal HIOs in different formats and procedures for isolating viable cells from these formats that are suitable for use in single-cell transcriptional profiling platforms. These protocols and procedures facilitate the use of small intestinal HIOs to obtain an increased understanding of the cellular response of human intestinal epithelium at the transcriptional level in the context of a variety of different environments.
Introduction
The small intestinal epithelium has two distinct zones: the crypt that houses the intestinal stem cell (ISC) and the villus, which is comprised of differentiated cells of the secretory and absorptive lineages. Adding to this complexity is the regional specificity of the epithelium that provides unique functional properties between the regions of the small intestine. Pioneering work established culture conditions in which both the human small intestinal crypt and villus zones can be generated ex vivo from surgical tissues or tissue biopsies1. These cultures are bridging the gap between animal studies and clinical trials and are revealing previously unrecognized cell biology, biochemistry, and physiology of the gastrointestinal tract. The HIOs are propagated as 3D spherical structures using a media with growth factors that promote stem cell viability and extracellular support matrices. These conditions result in a crypt-like HIO model that consists mostly of progenitors and stem cells. Removal of the growth factors promotes ISC differentiation (villus-like model) and production of mature intestinal epithelial cells (goblet, enteroendocrine, tuft, enterocyte) in appropriate ratios along with cell proliferation and differentiation, polarization, barrier integrity, regional specific features, and appropriate physiological responses2. HIO cultures are genetically stable and can be propagated indefinitely in their crypt-like state. Easy access to the apical surface of these cultures is provided by culture conditions that allow growth in a monolayer format3. HIOs also allow considerations of host individual variability such as genetics, age, sex, ethnicity, and disease status to be included in biological analyses. Analytic and functional assessment tools are identical to those used in approaches centered on transformed cell lines and include a variety of molecular techniques such as flow cytometry, microscopy, transcriptomics, proteomics, and metabolomics.
Single cell transcriptomics is revolutionizing our understanding of the biology and physiology of the small intestine by providing insight into the individual contributions of each cell type to a biological process. Pioneering work using this technology has provided a landscape of the cell types present in the native human intestine4,5,6. Single cell transcriptional profiling platforms allow exploration of the transcriptional landscape of individual cells, allowing cell heterogeneity to come to the forefront of scientific exploration. In some single cell transcriptional profiling platforms, microfluidic partitioning and barcoding are used to generate cDNA libraries from cellular polyadenylated mRNAs obtained from up to 10,000 cells per sample. On this platform, droplets containing single cells, barcoded oligonucleotides, reverse transcription reagents, and oil form a reaction vesicle that results in all cDNAs from a single cell having the same barcode. The barcoded cDNAs can then be efficiently sequenced using next generation sequencing. Data generated can be handled through software and visualization tools, which convert the barcoded sequences into visualization maps and single cell transcriptional profiles. Cell populations can be identified using publicly available databases of human small intestinal epithelium4,5,6. Although many studies have utilized this platform to interrogate murine intestinal organoids at the single cell level, the analysis of single cell transcriptional responses of HIOs has lagged behind7,8,9,10,11,12.
Here, we provide a step-by-step guide to isolating viable cells from small intestinal HIOs for processing on single cell transcriptional profiling platforms (Figure 1). We provide culturing and differentiation guidelines along with media components that have been optimized for epithelial differentiation. We outline cell recovery methods for three different culture formats: three dimensional (3D) and monolayer cultures either on plastic or on membrane cell culture inserts. We provide sample clustering data obtained using open-source software to derive differentially expressed genes in each cluster.
Protocol
The organoid lines used here were obtained from the Texas Medical Center Digestive Disease Center GEMS Core. Briefly, to initially establish organoid lines, donor tissue samples were washed and enzymatically digested to release the intestinal crypts. Crypts were embedded in a basement membrane and cultured in a medium. The Institutional Review Board at Baylor College of Medicine approved the study protocol to obtain tissue samples from which organoid lines were established, and informed consent was obtained from all donors to establish organoid lines from the donated tissue.
1. Passaging of 3D HIOs to expand for differentiation
- Remove and discard medium from wells of organoids grown in 24-well plates (around the solid basement membrane) with a P1000 pipet. Add 300 µL of 0.05% Trypsin-EDTA to the well and mechanically break up organoids by pipetting with a P1000 pipet up and down 5x.
- Incubate plate at 37 °C for 4 min. Add 500 µL of complete media without growth factors (CMGF- medium) with 10% FBS to each well (Table 1). Use a P1000 pipet and pipet CMGF medium and HIO mixture, 10x back and forth in the well. Transfer the entire volume into a 15 mL tube. Combine passage from multiple wells into the same 15 mL tube. Do not combine more than 10 wells per tube.
- Spin down cells at 4 °C using a swing rotor centrifuge at 100 x g for 5 min. Remove all the medium, keeping the pellet at the bottom of the tube. Keep the tube on ice.
- Calculate the amount of basement membrane needed for the number of HIO plated, and resuspend the HIO pellet in 30 µL/well of basement membrane using cold P200 pipet tips. Pipette the HIO basement membrane mixture as a droplet into the center of a 24-well plate using cold P200 pipet tips.
- Transfer the plate to a 37 °C incubator. Let the basement membrane solidify for 5-10 min, then add 500 µL of room temperature complete medium containing WNT, R-spondin, noggin, EGF, and ROCK inhibitor (WRNE+Y; Table 1) to each well and culture in a 37 °C incubator. Refresh culture with WRNE+Y every other day for 7 days.
NOTE: A healthy HIO will be cystic in shape with a shiny budding epithelium (Figure 2). Cells in the middle of the cyst cannot be visualized. An unhealthy HIO will look dull and have a thick epithelium with a solid middle. The most important characteristic is the shiny appearance.
2. HIO differentiation: 3D format
- Use one well of 3D HIOs for 3 wells of 3D differentiated HIOs 1 week after culturing.
- Remove media from wells. Disperse basement membrane and HIOs using a P1000 pipet and add cold 500 µL of CMGF- per well and pipet up and down at least 5x. Spin down HIOs at 4 °C using a swing rotor centrifuge at 100 x g for 5 min. Remove all the medium keeping the pellet at the bottom of the tube.
- Add 30 µL of basement membrane per well of HIOs being plated. Pipette the 30 µL droplet of basement membrane with HIOs into a well of a 24-well plate. Incubate at 37 °C for 10 min. Add 500 µL of WRNE+Y/Diff (50/50) media.
- Switch the medium to differentiation medium after 24 h for 3 additional days, refreshing the medium every other day. Prepare 3 wells per sample to pool to reduce technical variability.
3. HIO differentiation: Monolayer format
- Coat the membrane cell culture inserts or wells of a 96-well plate with 100 µL/well of human placental collagen IV (1mg/ml in 100mM acetic acid) diluted in cold H2O (1:30). Incubate at 37 °C for a minimum of 1.5 h or maximum overnight. For membrane cell culture insert, add sterile H2O in the plate space to humidify the plate; for 96-well plate, add sterile H2O in the unused wells.
- Collect 3D HIOs cultured in WRNE for 7 days by removing the media and recovering the basement membrane and cells by adding 500 µL/well of cold 0.5 mM EDTA (0.5 M EDTA diluted 1:1000 in sterile PBS). Use a P1000 pipet to pipet organoids and solution up and down 5x and transfer to a 15 mL microcentrifuge tube. Do not pool more than 10 wells of organoids per tube. Centrifuge for 5 min at 300 x g in a swinging bucket rotor at 4 °C.
NOTE: To prepare 3 wells on a 96-well plate, use 1 well of 3D HIOs. To prepare 1 membrane cell culture insert, use 1 well of 3D HIOs (seeding density of single cells is approximately 1-2 x 105 cells/well for 96-well plate, 2.5-3 x 105 cells/well for membrane cell culture insert). - Remove 500 µL of media, leaving the pellet at the bottom of the tube. Dissociate the HIOs by adding 500 µL of 0.05% Trypsin/0.5 mM EDTA and incubating for 4-5 min at 37 °C. Vigorously pipette up and down for ~50x using a P1000. Avoid making bubbles by pipetting against the side of the tube.
- Inactivate the trypsin by adding 1 mL of CMGF- containing 10% FBS. Filter out large clumps of undigested cells by wetting a 40 µm cell strainer with 1 mL of CMGF- containing 10% FBS and adding the cells to the top of the cell strainer. Use gravity to allow the cells to pass through the strainer and discard the cell strainer containing cell clumps.
- Collect the cells by centrifugation for 5 min at 400 x g in a swinging bucket rotor at room temperature.
- Remove the liquid from the collagen-coated membrane cell culture insert or from the 96-well plate. Resuspend the HIO pellet in 100 µL/well of WRNE containing 10 µM Y-27632, then pipette into the upper compartment of the membrane cell culture insert or into the 96-well plate. For membrane cell culture insert, add 600 µL of WRNE containing 10 µM Y-27632 into the lower compartment of the well.
- Remove the medium after 24 h from both the upper and lower part of the membrane cell culture insert or from the 96-well plate and replace with differentiation medium for 5 days. Prepare 3 wells per sample to pool to reduce technical variability.
4. Preparation of single cell suspensions from differentiated 3D HIOs for single cell transcriptomics
- Place enzymatic cell digestion reagent at 37 °C in a CO2 incubator to thaw and warm 30 min prior to initiation of 3D HIO digestion. Check the HIOs using a bright field microscope at 10x to ensure cell health.
- Remove the media from the wells and add 500 µL of cold cell recovery solution to the wells. Pipette up and down a few times to liberate the cells from the basement membrane using a P1000 pipette and standard pipette tips. Transfer the cell suspension to a 1.5 mL microcentrifuge tube.
NOTE: The use of replicate wells ensures adequate cell numbers and reduces well-to-well variation. Do not combine wells before step 4.7 below. - Incubate the tubes of cells on ice for 10 min, pipetting up and down every 5 min. Collect the cells by centrifugation for 3 min at 400 x g at 4 °C.
- Remove the liquid from the tube, taking care not to disturb the cell pellet. Resuspend the cells in 500 µL of pre-warmed enzymatic cell digestion reagent. Pipette up and down with a P1000 at least 10x.
- Incubate the HIOs for 30 min at 37 °C in a CO2 incubator. Every 10 min, pipette the entire volume 10x using a P1000 pipette to help dissociate the cells. Avoid introducing air bubbles into the cell suspension.
- Pellet the cells for 3 min at 400 x g at 4 °C. Remove the supernatant, resuspend the cells in 1 mL of differentiation media, and store on ice.
NOTE: From this point on, the HIOs must always be kept on ice. - Pipette the cells up and down 50x rapidly using a P1000 to ensure HIOs are dissociated into single cells. Take care not to introduce air bubbles at this step, as it will cause cell loss.
- Filter the entire volume through a 70 µm tip strainer using a P1000 pipette tip. Use a 1.5 mL microcentrifuge tube to collect the eluate.
NOTE: The filter tip will not become clogged if digestion time is sufficient. Use multiple filter tips if the filter becomes clogged; do not force the volume through the filter. - Repeat step 4.8 with a 40 µm tip strainer. Store samples on ice.
- Count viable single cells on a cell counting chamber using trypan blue exclusion by adding 5 µL of 0.4% trypan blue to 5 µL of cell suspension. Dilute the sample as needed to 1000 cells/µL. If the cell density is less than 1000 cells/µL, pellet cells for 2 min at 400 x g at 4 °C, remove the original media, and resuspend in cell culture media. This protocol typically yields 5 x 105-1 x 106 cells.
NOTE: Preparation of a high-quality suspension (90% viable and 90% single cells) is essential for high quality single-cell data. However, experience indicates viability as low as 75% can be accepted in cases where treatment, such as infection or small molecule application, impact viability. - Prepare DNA libraries for single cell transcriptional profiling platforms according to the manufacturer's protocols. Process the data according to best practice protocols for scRNAseq data analysis13,14.
5. Preparation of single cell suspensions from HIOs differentiated on membrane cell culture insert
- Place enzymatic cell digestion reagent at 37 °C in a CO2 incubator to thaw and warm 30 min prior to initiation of HIO membrane cell culture insert digestion. Divide the pre-warmed enzymatic cell digestion reagent into 200 µL aliquots in 1.5 mL microcentrifuge tubes, 1 tube per membrane cell culture insert. Keep the tubes in the 37 °C incubator until ready for use.
- Prepare PBS-EDTA by adding 5 µL of 0.5 M EDTA to 5 mL of PBS. Aliquot 500 µL of PBS-EDTA into a 24-well plate, 1 well per membrane cell culture insert.
- Wash the cells with PBS-EDTA: transfer the membrane cell culture inserts from wells containing growth media to the wells containing PBS-EDTA. Remove the cell culture media from the apical side and replace it with 100 µL of PBS-EDTA. Take care not to disturb the cell layer.
- Immediately discard the PBS-EDTA added to the apical side of the membrane cell culture inserts. Remove the membrane cell culture insert from the well, blot it by gently touching the edge to a paper towel, and then invert it on the benchtop. Use a sharp scalpel blade to remove the membrane from the insert by cutting around the inner circumference of the membrane. Work quickly to prevent drying.
- Use forceps to transfer the membrane to the pre-warmed 1.5 mL microcentrifuge tube filled with enzymatic cell digestion reagent and ensure the membrane is completely submerged. Repeat for all additional membrane cell culture inserts.
NOTE: Increasing the volume of the enzymatic cell digestion reagent to 500 µL may aid in dissociating dense monolayers. - Dissociate the cells by incubating tubes with membranes at 37 °C in a CO2 incubator. Mix by gently pipetting the entire volume 5x with a 200 µL pipette every 10 min for a total of 30 min. Avoid bubbles and suction of the membrane to preserve viability.
- Transfer a small aliquot of cells (1-2 µL) to a glass slide every 10 min to assess dissociation by qualitatively assessing the percent of single cells. It may take up to 50 min with additional pipetting to achieve 80%-90% single cells, depending on the HIO type. Gently scrape the membrane with a pipette tip during mixing steps to encourage dissociation, particularly for hard to dissociate cells, however scraping may impact viability.
- Combine 3 replicate wells per condition in a single 1.5 mL microcentrifuge tube (600 µL total). Add 400 µL of cell culture media (diff) + 10 µM ROCKi (1 mL total volume). Mix gently by pipetting.
NOTE: The use of replicate wells ensures adequate cell numbers and reduces well-to-well variation. Wells should be pooled at this step. - Filter the entire volume through a 70 µm tip strainer using a 1 mL pipette tip. Collect the flow through in a fresh 1.5 mL microcentrifuge tube.
NOTE: The filter tip will not become clogged if digestion time is sufficient. Use multiple filter tips if the filter becomes clogged; do not force the volume through. - Repeat step 5.9 with a 40 µm tip strainer. Store samples on ice.
- Count viable single cells on a cell counting chamber using trypan blue exclusion by adding 5 µL of 0.4% trypan blue to 5 µL of cell suspension. Dilute sample as needed to 1000 cells/ µL. If the cell density is less than 1000 cells/ µL, spin 2 min at 400 x g at 4 °C to pellet the cells, remove the original media, and resuspend in WRNE+Y at the desired cell density. This protocol typically yields 4 x 105 – 6 x 105 cells.
NOTE: Preparation of high-quality suspension (90% viable and 90% single cells) is essential for high quality single-cell data. However, experience indicates viability as low as 75% can be accepted in cases where treatment, such as infection or small molecule application, impact viability. - Prepare DNA libraries for the single cell transcriptional profiling platforms according to the manufacturer's protocols. Process the data according to best practice protocols for scRNAseq data analysis13,14.
6. Preparation of single cell suspensions from HIOs differentiated as 96-well monolayers
- Place enzymatic cell digestion reagent at 37 °C in a CO2 incubator to thaw and warm 30 min prior to initiation of monolayer HIO digestion.
- Prepare PBS-EDTA by adding 5 µL of 0.5 M EDTA to 5 mL of PBS. Discard the cell culture media from the 96-well plate and replace it with 100 µL of PBS-EDTA. Take care not to disturb the cell layer.
- Dissociate the cells: Discard the PBS-EDTA and add 100 µL of warm enzymatic cell digestion reagent to each well. Place the plate in the 37 °C in a 5% CO2 incubator and mix by gently pipetting the entire volume 5x with a 200 µL pipette tip every 10 min for 20-30 min. Avoid bubbles to preserve viability.
NOTE: Increasing the volume of enzymatic cell digestion reagent to 500 µL may aid in dissociating dense monolayers. - Transfer a small aliquot of cells (1-2 µL) to a glass slide every 10 min to assess dissociation. It may take up to 50 min to achieve 80%-90% single cells, depending on the HIO type. Gently scrape the plate with a pipette tip during mixing steps to encourage dissociation, particularly for hard to dissociate cells; however, scraping may impact viability.
- Combine 3 replicate wells per condition in a single 1.5 mL microcentrifuge tube (300 µL total). Add 700 µL of cell culture media (diff) + 10 uM ROCKi (1 mL total volume). Mix gently by pipetting.
NOTE: The use of replicate grown wells ensures adequate cell numbers and reduces well-to-well variation. - Filter the entire volume through a 70 µm tip strainer using a 1 mL pipette tip. Collect the flow through in a fresh 1.5 mL microcentrifuge tube.
NOTE: The filter tip will not become clogged if enzymatic cell digestion reagent digestion time is sufficient. Use multiple filter tips if the filter becomes clogged; do not force the volume through. - Repeat step 6.6 with a 40 µm tip strainer. Store samples on ice.
- Count viable single cells on a cell counting chamber using trypan blue exclusion by adding 5 µL of 0.4% trypan blue to 5 µL of cell suspension. Dilute as needed to 1000 cells/µL. If the cell density is less than 1000 cells/µL, spin 2 min at 400 x g 4 °C to pellet the cells, remove the original media, and resuspend in cell culture media + ROCKi at the desired cell density. This protocol typically yields 2 x 105 – 6 x 105 cells.
NOTE: Preparation of high-quality suspension (90% viable and 90% single cells) is essential for high quality single-cell data. However, experience indicates viability as low as 75% can be accepted in cases where treatment, such as infection or small molecule application, impact viability. - Prepare DNA libraries according to the single cell transcriptional profiling platform manufacturer's protocols. Process the data according to best practice protocols for scRNAseq data analysis13,14.
Representative Results
Single-cell suspensions were pooled from 2-3 wells of membrane cell culture insert, monolayer, and 3D HIOs to ensure sufficient cell yield and reduce well-to-well variation. Single cell libraries were prepared using reagents specific to the single cell transcriptional profiling platform. and sequenced with paired end reads on a next generation sequencing platform, 30,000 reads/cell. Reads were mapped, counted, and analyzed using analytical tools for single cell genomics. Low-quality cells with more than 20% mitochondrial read, or less than 200 counted genes were excluded from the analysis. To assess the quality of the cells, we plotted the number of genes, UMIs, and percent mitochondrial reads per cell (Figure 3); high-quality cells contain approximately 3500 genes, 15,000 UMIs, and 10% mitochondrial reads. Following filtering, we obtained 4000 to 9500 single cells depending on the HIO growth format. Unsupervised clustering and UMAP dimension reduction identified distinct clusters representing expected cell types, including absorptive enterocytes and secretory cells (Figure 4).
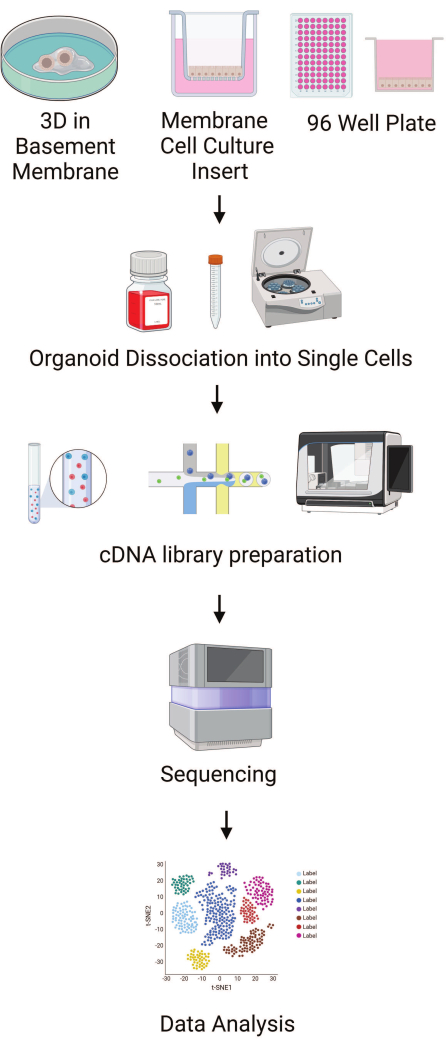
Figure 1: Outline of protocol steps. Schematics outlining the major steps in plating HIOs in different formats to single cell data workflow. This image was created using Biorender. Please click here to view a larger version of this figure.

Figure 2: Bright field images. HIOs ready for processing were imaged using a bright field microscope at 5X magnification. Left, 3D organoids; middle, 96-well plate; right, transwell. Scale bars = 200 µm. Please click here to view a larger version of this figure.
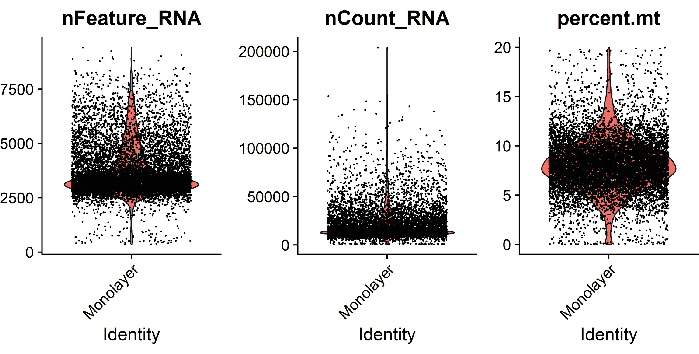
Figure 3: Quality control of data. Example metrics of analyzed single cells isolated from jejunal HIOs grown as monolayers on 96-well plates. Violin plots show the number of genes (nFeature_RNA), UMIs (nCount_RNA), and percent mitochondrial reads (percent.mt) per cell. Please click here to view a larger version of this figure.
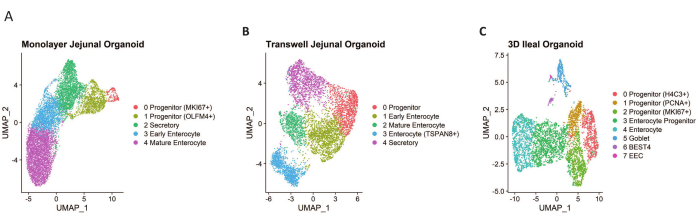
Figure 4: UMAPs of analyzed single cells. (A) A total of 9952 single cells were isolated from differentiated jejunal HIOs grown in 96-well plates. (B) A total of 5623 single cells were isolated from differentiated jejunal HIOs grown on membrane cell culture inserts. (C) A total of 4402 single cells were isolated from differentiated ileal 3D HIOs. Please click here to view a larger version of this figure.
| WRNE Growth Medium | Differentiation medium | ||
| Component | Volume | Final Concentration | WRNE medium without L-WRN conditioned medium, Nicotinamide and SB202190 |
| *Y-27632 (5 mM) optional | 200 µL | 10 µM | |
| A 83-01 (500 µM) | 100 µL | 500 nM | |
| Advanced DMEM/F12 | 45 mL | ||
| B27 | 2 mL | ||
| EGF (50 µg/mL) | 100 µL | 50 ng/mL | |
| Gastrin (10 µM) | 100 µL | 10 nM | |
| GlutaMax (100X) | 1 mL | ||
| HEPES 1 M | 1 mL | ||
| L-WRN Con- Media | 50 mL | ~50% | |
| N2 | 1 mL | ||
| N-Acetylcysteine (500 mM) | 100 µL | 500 µM | |
| Nicotinamide 1 M | 1 mL | 10 mM | |
| SB202190 (10 mM) | 100 µL | 10 µM | |
| Total Volume | 100 mL | ||
| CMGF- Medium | |||
| Advanced DMEM/F12 | 500 mL | ||
| Glutamax | 5 mL | 100X | |
| 1 M Hepes | 5 mL |
Table 1: Composition of various media used.
Discussion
Using single cell genomics platforms, complex biological systems, such as tissue derived HIO cultures that model the intestinal epithelium, can be broken down to yield individual cellular contributions to overall biological response4,5,6. Cellular heterogeneity and rare cell populations can also be identified and interrogated. Cellular input needs to be optimized to maximize output using single cell transcriptomic-based platforms. Here we describe protocols that will yield highly viable and good quality single cell suspensions that will increase the likelihood of obtaining downstream high quality transcriptomic data. Familiarity with the protocol is of utmost importance, and it is recommended that several practice runs be performed before proceeding with library preparation. Reproducible achievement of viable single cell suspensions is suggested prior to using the cells for library preparation. The rapidness at which the protocol is implemented is of utmost importance as speed is directly proportional to viability outcomes. Based on this, limiting the number of samples handled at one time is advised. Interfacing with the entity that will prepare the libraries prior to initiating these protocols for assistance with troubleshooting is recommended.
HIO viability directly influences the DNA library sent for sequencing. Cell viability of the HIO depends on several steps in the preparation of the single cell suspension, including dissociation, centrifugation, buffers, pipetting, and counting accuracy. The goal of preparing the HIOs for the platform is to balance the maintenance of cell viability with sample quality. Critical to obtaining high quality data is the minimization of cellular aggregates, dead cells, non-cellular nucleic acids, and biochemical inhibitors of reverse transcription. Dying cells increase nucleic acid noise and contribute to cell clumping, leading to data and platform failures7,8,9,10,11,12. Filtering with an appropriate filter size can help remove large aggregates and improve data quality.
Several factors need to be considered to minimize HIO cell death when preparing single cell suspensions. Obtaining at least 80% viable cells is necessary, and >90% viability is even more desirable. Samples with high numbers of non-viable cells can be improved using strategies to remove dead cells, such as column-based microbeads, although these require more initial cells and larger sample volumes due to loss during handling. HIOs are particularly sensitive to buffers and will undergo cell death if left too long at RT in certain buffers. Organoid media results in the least amount of cell loss and aggregate formation. Pipetting can also affect HIO viability during dissociation. Regular bore tips can shear organoids, helping to dissociate into single cell suspensions; however, pipetting too rapidly with a regular bore tip can cause HIO cell damage from the shearing forces. Slow pipetting with wide bore tips minimizes stress on the HIO cells and greatly improves viable cell numbers downstream. Centrifugation, washing, and resuspending are other factors that can influence the quality of HIO single cells suspensions. Swinging bucket centrifugation results in gentler pelleting of HIO cells, resulting in lower numbers of dead cells while minimizing cell loss. Finally, counting accuracy to determine concentration is essential to ensure the maximum number of cells reach the platform for library synthesis. Attention to these details will result in the minimization of stress on the HIO cells during the isolation process and ensure the highest probability of data return after processing.
Although single cell transcriptomics is a powerful tool to dissect biological profiles and responses at the single cell level4,5,6, there are several limitations that need to be considered when applying the technology to analyze human intestinal organoid cultures. First, there can be significant transcriptional variation because of technical variability introduced during library preparation and sequencing or due to batch effects resulting from sample preparation. In addition, sequencing can result in coverage bias where there is uneven sequencing depth across genes or cells. This causes challenges when detecting lowly expressed genes or identifying rare cell populations. Second, the identification of cell types can be difficult due to the cellular and transcriptional heterogeneity of the organoid cultures and a less differentiated cellular profile when compared to the native intestinal epithelium. Coupling transcriptional responses to functional phenotypes is also a challenge and requires additional validation using other approaches or assays. Third, this approach does not include consideration of spatial context or temporal dynamics of the cells. The dissociation procedure removes the ability to infer cell-cell interactions from the transcriptional data. It only provides a single snapshot in time of the cellular state rather than a biologic continuum. Finally, single cell transcriptional approaches can be costly and require specialized computational tools and bioinformatics expertise. However, this approach still remains a dominant state-of-the-art tool for understanding cellular heterogeneity and identifying novel cell types.
Offenlegungen
The authors have nothing to disclose.
Acknowledgements
The authors acknowledge U19 AI157984, U01 DK103168, U19 AI144297, P30 DK56338, P01 AI057788, U19 AI116497 grants and NASA Cooperative Agreement Notice/TRISH NNX16AO69A.
Materials
| [Leu15]-Gastrin I | Sigma-Aldrich | G9145 | 10 nM |
| 0.05% Trypsin-EDTA | Invitrogen | 25300054 | |
| 0.4% Trypan blue | Millipore-Sigma | T8154 | |
| 0.5 M EDTA | Corning | 46-034-CI | |
| 1x PBS Ca- Mg- | Corning | 21-040-CM | |
| 24 mm Transwell | Costar | 3412 | |
| 24 well Nunclon delta surface tissue culture dish | Thermo Scientific | 142475 | |
| 40 µm cell strainer | Falcon | 352340 | |
| 40 µm Flowmi tip strainer | SP Bel-Art Labware | H13680-0040 | |
| 70 µm Flowmi tip strainer | SP Bel-Art Labware | H13680-0070 | |
| 96 well plate | Corning | 3595 | |
| A-83-01 | Tocris | 2939 | 500 nM |
| Accutase | StemCell Technologies | 7920 | |
| Advanced DMEM/F12 | Invitrogen | 12634-028 | |
| B27 supplement | Invitrogen | 17504-044 | 1X |
| Chromium Next GEM Single Cell 3’ GEM, Library & Gel Bead Kit v3.1 | 10x Genomics | PN-1000128 | |
| Collagen IV | Sigma-Aldrich | C5533-5MG | 33 µg/mL |
| Corning Cell Recovery Solution | VWR | 354253 | |
| DPBS (Mg2-, Ca2-) | Invitrogen | 14190-136 | 1X |
| GlutaMAX-I | Invitrogen | 35050-061 | 2 mM |
| HEPES 1M | Invitrogen | 15630-080 | 10 mM |
| L-WRN conditioned media | ATCC | CRL-3276 | |
| Matrigel, GFR, phenol free | Corning | 356231 | |
| mouse recombinant EGF | Invitrogen | PMG8043 | 50 ng/mL |
| N2 supplement | Invitrogen | 17502-048 | 1X |
| N-Acetylcysteine | Sigma-Aldrich | A9165-5G | 500 µM |
| Nicotinamide | Sigma-Aldrich | N0636 | 10 mM |
| SB202190 | Sigma-Aldrich | S7067 | 10 µM |
| Transwell | Corning | 3413 | |
| Y27632 | Stem Cell Technologies | 72308 | 10 µM |
Referenzen
- Sato, T., et al. Long-term expansion of epithelial organoids from human colon, adenoma, adenocarcinoma, and Barrett’s epithelium. Gastroenterology. 141 (5), 1762-1772 (2011).
- Blutt, S. E., et al. Gastrointestinal microphysiological systems. Exp Biol Med. 242 (16), 1633-1642 (2017).
- VanDussen, K. L., et al. Development of an enhanced human gastrointestinal epithelial culture system to facilitate patient-based assays. Gut. 64 (6), 911-920 (2015).
- Elmentaite, R., et al. Cells of the human intestinal tract mapped across space and time. Nature. 597 (7875), 250-255 (2021).
- Hickey, J. W., et al. Organization of the human intestine at single-cell resolution. Nature. 619 (7970), 572-584 (2023).
- Burclaff, J., et al. A proximal-to-distal survey of healthy adult human small intestine and colon epithelium by single-cell transcriptomics. Cell Mol Gastroenterol Hepatol. 13 (5), 1554-1589 (2022).
- Triana, S., et al. Single-cell transcriptomics reveals immune response of intestinal cell types to viral infection. Mol Syst Biol. 17 (7), e9833 (2021).
- Triana, S., et al. Single-cell analyses reveal SARS-CoV-2 interference with intrinsic immune response in the human gut. Mol Syst Biol. 17 (4), e10232 (2021).
- Fujii, M., et al. Human intestinal organoids maintain self-renewal capacity and cellular diversity in niche-inspired culture condition. Cell Stem Cell. 23 (6), 787-793 (2021).
- Beumer, J., et al. BMP gradient along the intestinal villus axis controls zonated enterocyte and goblet cell states. Cell Rep. 38 (9), 110438 (2022).
- Beumer, J., et al. High-resolution mRNA and secretome atlas of human enteroendocrine cells. Cell. 182 (4), 1062-1064 (2020).
- Busslinger, G. A., et al. Human gastrointestinal epithelia of the esophagus, stomach, and duodenum resolved at single-cell resolution. Cell Rep. 34 (10), 108819 (2021).
- Luecken, M. D., Theis, F. J. Current best practices in single-cell RNA-seq analysis: a tutorial. Mol Syst Biol. 15 (6), e8746 (2019).
- Heumos, L., et al. Best practices for single-cell analysis across modalities. Nat Rev Genet. 24 (8), 550-572 (2023).

.
