Characterization of Multi-subunit Protein Complexes of Human MxA Using Non-denaturing Polyacrylamide Gel-electrophoresis
Summary
This article describes a simple and rapid protocol to evaluate the oligomeric state of the dynamin-like GTPase MxA protein from lysates of human cells using a combination of non-denaturing PAGE with western blot analysis.
Abstract
The formation of oligomeric complexes is a crucial prerequisite for the proper structure and function of many proteins. The interferon-induced antiviral effector protein MxA exerts a broad antiviral activity against many viruses. MxA is a dynamin-like GTPase and has the capacity to form oligomeric structures of higher order. However, whether oligomerization of MxA is required for its antiviral activity is an issue of debate. We describe here a simple protocol to assess the oligomeric state of endogenously or ectopically expressed MxA in the cytoplasmic fraction of human cell lines by non-denaturing polyacrylamide gel electrophoresis (PAGE) in combination with Western blot analysis. A critical step of the protocol is the choice of detergents to prevent aggregation and/or precipitation of proteins particularly associated with cellular membranes such as MxA, without interfering with its enzymatic activity. Another crucial aspect of the protocol is the irreversible protection of the free thiol groups of cysteine residues by iodoacetamide to prevent artificial interactions of the protein. This protocol is suitable for a simple assessment of the oligomeric state of MxA and furthermore allows a direct correlation of the antiviral activity of MxA interface mutants with their respective oligomeric states.
Introduction
The quaternary structure of a protein plays a crucial role in many cellular processes. Signaling pathways, gene expression, and enzyme activation/deactivation all rely on the proper assembly of protein complexes 1-4. This process also known as homo- or hetero-oligomerization is due to irreversible covalent or reversible electrostatic and hydrophobic protein-protein interactions. Oligomerization not only diversifies the different cellular processes without increasing the genome size, but also provides a strategy for proteins to build stable complexes that are more resistant towards denaturation and degradation 5. Defects in oligomerization have an impact on the function of proteins and can lead to the development of diseases. For example, the enzyme phenylalanine hydroxylase forms a tetrameric complex. Some mutations within the protein complex can weaken the tetramer formation and lead to the disease phenylketonuria 6.
The human MxA protein is an interferon (IFN)-induced antiviral effector protein exerting a broad antiviral activity against various RNA as well as DNA viruses 7. It belongs to the superfamily of dynamin-like large GTPases and has the capacity to form large oligomeric structures in vitro 8. Oligomerization has been suggested to protect MxA from rapid degradation 9,10. Despite intense efforts by many research groups, the molecular mechanism of action remains largely elusive and the role of the oligomerization state of MxA for its antiviral function is under debate 9,11,12. In this regard, Gao and coworkers proposed a model where MxA exerts its antiviral activity by interacting with viral nucleoproteins in form of large ring-like oligomeric structures 11. However, more recently, we demonstrated that MxA dimers exhibit antiviral activity and interact with the nucleoprotein of influenza A virus 12. Based on the crystal structure of MxA, Gao and coworkers identified several amino acid residues in the interface regions that are critical for its oligomerization in vitro and its antiviral function 11,13. Therefore, in order to elucidate which oligomeric state of MxA exerts antiviral activity, we sought to establish a simple protocol to rapidly determine the oligmeric state of MxA interface mutants expressed in human cells as well as endogenous MxA expressed after IFNα stimulation.
Although there are many techniques that are commonly used to investigate the interaction between proteins such as the split-Green Fluorescent Protein (split-GFP) complementation assay 14, surface plasmon resonance 15 and Förster resonance energy transfer (FRET) 16, they do not provide information of the exact stoichiometry of an oligomeric protein complex. For investigation of this particular aspect, techniques such as multi-angle light scattering (MALS) 17 and analytical ultracentrifugation 18 are very useful. Usually, the proteins analyzed using these methods are purified proteins. Oligomerization processes may also depend on other cellular factors. If these factors are unknown, the analysis is more difficult. Additionally, some proteins are difficult to express in E. coli and to purify. Therefore, these methods are not the optimal choice to analyze protein oligomerization in the cellular environment. In addition, these techniques require expensive instruments which are not readily available.
Non-denaturing polyacrylamide gel electrophoresis (PAGE), size exclusion chromatography or chemical crosslinking followed by conventional Sodium dodecyl sulfate (SDS)-PAGE are useful tools for the characterization of the formation of oligomers from cell lysates 2,19,20. These methods do not require specialized equipment and can be easily performed in a standard laboratory. We initially evaluated various chemical cross-linking protocols that invariantly led to non-specific aggregation and precipitation of MxA. Therefore, we next tested non-denaturing PAGE protocols. As non-denaturing PAGE excludes the use of SDS, the migration of proteins depends on their native charge. Blue-native PAGE uses coomassie brilliant blue G250 to load proteins with an overall negative charge, similar to SDS, but does not denature the protein 21. Unfortunately, coomassie brilliant blue precipitates in the presence of high salts and divalent cations (e.g. Mg2+) that are often included in lysis buffers. Depending on the buffers used, it may be difficult to analyze the sample without further optimization of steps that could have an effect on the oligomeric protein complex.
Here we present a simple protocol based on a previously published method 22 to determine oligomerization of human MxA protein derived from cellular lysates using non-denaturing PAGE.
Protocol
NOTE: This protocol is based on the previously published non-denaturing PAGE protocol 12. In that study, the oligomeric state of the MxA protein was assessed using either Vero cells overexpressing MxA or IFN-α-stimulated A549 cells expressing endogenous MxA. The protocol described below can be used to analyze the oligomeric state of any protein in addition to MxA. However, further optimization may be required.
1. Preparation of Cell Lysate for Non-denaturing PAGE
NOTE: To analyze the oligomeric state of the human MxA protein in either Vero or A549 cells, 1.0 x 106 cells were harvested. Depending on the cell type or the abundance of the protein to analyze, the cell number should be adjusted. It is also important to protect the lysis buffer from light exposure, as soon as the light-sensitive iodoacetamide is added.
- Seed 0.3 x 106 A549 or Vero cells per well into 6 well-dishes. Keep the cells in 2 ml growth medium per well (see Table 1). Incubate cells overnight in a cell culture incubator (37 °C, 5% CO2).
- Harvest the cells by washing with 1 ml of phosphate buffered saline (PBS) and detach by adding 0.5 ml of 0.25% trypsin-ethylenediaminetetraacetic acid (EDTA) 1x solution for approximately 5 min at room temperature.
- As soon as the cells detach from the dish, add 0.5 ml growth medium and carefully mix by pipetting up and down.
- Transfer the cells of each well into one 2 ml tube and pellet them using a table top centrifuge (5,000 x g, 4 °C, 5 min).
- Carefully remove the supernatant by pipetting without disturbing the cell pellet.
- Wash the cells with 1 ml ice-cold PBS by carefully pipetting the cell suspension up and down.
- Pellet cells in a table top centrifuge (5,000 x g, 4 °C, 5 min).
- Carefully remove the supernatant by pipetting without detaching the cell pellet.
- Resuspend cells in 200 μl ice-cold lysis buffer (see Table 1) by pipetting up and down and put on ice.
- Immediately, protect lysate from light by covering the tubes using tin foil and incubate for 30 min on ice.
NOTE: After incubation for 30 min on ice, it is no longer essential to protect the lysate from light exposure, since the protection of the free thiol groups is irreversible. - Remove cell debris by centrifugation in a pre-chilled table top centrifuge (13,000 x g, 4 °C, 20 min).
- Equilibrate dialysis columns in dialysis buffer (Table 1) in the cold room at 4 °C for 20 min during the centrifugation step. Use a column with a molecular weight cut off of 10,000.
- Attach the columns to a float buoy and put them into a beaker filled with dialysis buffer. To ensure gentle stirring, use a magnetic stirrer. Do not touch the membrane.
NOTE: Dialysis columns can be purchased or prepared from 1.5 ml tubes according to the protocol described by Fiala and coworkers 19.
- Attach the columns to a float buoy and put them into a beaker filled with dialysis buffer. To ensure gentle stirring, use a magnetic stirrer. Do not touch the membrane.
- Remove the columns from the dialysis buffer and the float buoy. Transfer the cleared lysates into the prepared dialysis column by pipetting without touching the membrane. Attach the columns to a float buoy and put them back into the beaker filled with dialysis buffer.
- Dialyze the lysate in a beaker containing ice-cold dialysis buffer (Table 1) for at least 4 hr (or preferably overnight) at 4 °C while carefully stirring using a magnetic stirrer. Use at least 100 ml dialysis buffer for a 200 µl lysate.
- Transfer the dialysed sample into a 1.5 ml tube. Remove precipitates by centrifugation in a table top centrifuge (13,000 x g, 4 °C, 20 min). To prevent the dissociation of the oligomeric protein complexes continue with the protocol (section 2) immediately after dialysis. Do not freeze the prepared lysates.
2. Electrophoresis
NOTE: Electrophoresis was performed as described before with some modifications 22. In the protocol described below, pre-cast gradient gels were used (4-15% gradient). Alternatively, the gels can be prepared in the laboratory. It is very important to exclude any denaturing agent such as SDS to prevent the dissociation of the oligomeric protein complexes. Time of electrophoresis was optimized for the different oligomeric states of the human MxA protein. However, it can vary for other proteins, depending on the size of the oligomeric complex as well as the range of separation that is supposed to be achieved to analyze the complex. Therefore, the optimal time of electrophoresis should be determined empirically. For optimal resolution of the oligomers to be analyzed the current should not exceed 25 mA.
- Assemble the non-denaturing PAGE gel in the gel chamber. Fill the inner and outer chamber with pre-chilled running buffer (Table 1).
- Pre-run the gel with pre-chilled running buffer at 25 mA per gel for 15 min in the cold room at 4 °C.
- Mix 15 µl of the above prepared lysates with 5 µl of 4x sample buffer (Table 1). Do not boil the sample.
- Load 15 µl of sample and a native protein standard of choice on the gel. Run the gel at 25 mA for 4 hr in the cold room at 4 °C.
NOTE: For semi-quantitative analyses, a protein quantification protocol (e.g. a Bradford protein assay 23) can be performed in order to ensure loading of equal amounts of total protein per lane.
3. Western Blot
NOTE: Described below is the protocol of a wet western blot system. Any blotting membrane can be used. Activate polyvinylidene fluoride (PVDF) membranes in 100% methanol before equilibration in blotting buffer. The Semi-dry western blot technique can be used alternatively, but has to be optimized for large oligomeric complexes.
- Disassemble the gel and carefully transfer it into SDS buffer (Table 1).
- Incubate for 10 min at room temperature while gently shaking.
- Prepare 2 sponges, 4 cellulose filter paper sheets and a blotting membrane per gel. Soak them in blotting buffer (Table 1).
- Assemble the sandwich as follows (bottom to top): 1 sponge, 2 cellulose filter paper sheets, membrane, gel, 2 cellulose filter paper sheets, 1 sponge.
- Put the sandwich into the blotting tank. Make sure that the membrane faces the plus pole while the gel faces the minus pole.
- Fill the blotting tank with pre-chilled blotting buffer.
- Blot at 90 mA overnight at 4 °C for best protein transfer results.
- Disassemble the sandwich and visualize the protein standard by incubating the membrane in Ponceau S solution for 5 min at room temperature.
- Destain the membrane by carefully washing off the Ponceau S with deionized water until you can clearly see the bands of the protein standard.
- Mark the bands of the protein standard using a pen.
NOTE: Residual Ponceau S can interfere with the immunostaining. To avoid this, the membrane can be destained further by incubation in 0.1 M NaOH for 1 min and subsequent washing with deionized water. - Block the membrane with Blocking buffer (see Table 1) for at least 1 hr at room temperature or overnight at 4 °C.
- Visualize protein(s) of interest by immunostaining using antibodies directed against the protein to be analyzed.
NOTE: The human MxA protein was visualized using the rabbit polyclonal antibody specific for human Mx1 diluted 1:1,000 in Blocking buffer (Table 1). The antibody solution was incubated overnight at 4 °C. Alternatively, the monoclonal anti-MxA antibody (clone 143) can be used (data not shown) 24.
Representative Results
Using non-denaturing PAGE, we analyzed the oligomeric state of the human wild type MxA, the dimeric interface mutants MxA(R640A) and MxA(L617D) as well as the monomeric interface mutant MxA(M527D) from cell lysates 12. Cells were lysed in a buffer containing 1% octylphenoxypolyethoxyethanol (NP-40) and iodoacetamide to ensure protein solubilization and protection of free thiol groups (see Figure 1). As described before, salt and small metabolites were removed by dialysis 19. Protein separation was carried out by non-denaturing PAGE. To facilitate efficient western blotting, the gel was incubated in SDS buffer before blotting. The MxA proteins were visualized by immunostaining using a rabbit polyclonal antibody directed against MxA. The workflow is described in Figure 2.
To compare the oligomeric state of endogenous human MxA protein from IFN-α stimulated A549 cells, we transfected Vero cells (lacking endogenous MxA) with recombinant wildtype, monomeric and dimeric MxA variants. These recombinant wild type, monomeric and dimeric MxA variants formed stable tetramers, monomers and dimers, respectively, when compared to an unstained native protein marker (Figure 3A). Therefore, we used these recombinant proteins to assess the oligomeric state of endogenous human MxA protein derived from IFN-α stimulated A549 cells. Figure 3B reveals that the size of MxA in lysates of IFN-α-stimulated A549 cells corresponds to a tetramer.
Taken together, we describe a method to determine the oligomeric state of the human MxA protein from cell lysate. Our non-denaturing PAGE approach can also be used to assess the oligomeric state of other oligomeric protein complexes.
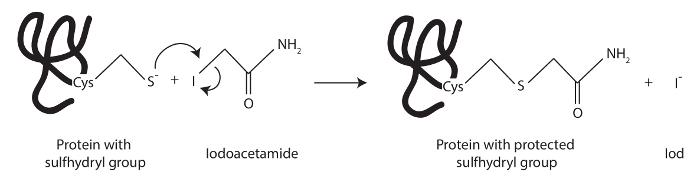
Figure 1: Structure and reaction scheme of iodoacetamide. Iodoacetamide irreversibly protects the thiol group of free cysteins by forming a thioether bond. This stable modification results from the nucleophilic substitution of the iodine with the sulfur atom from the cystein. Please click here to view a larger version of this figure.
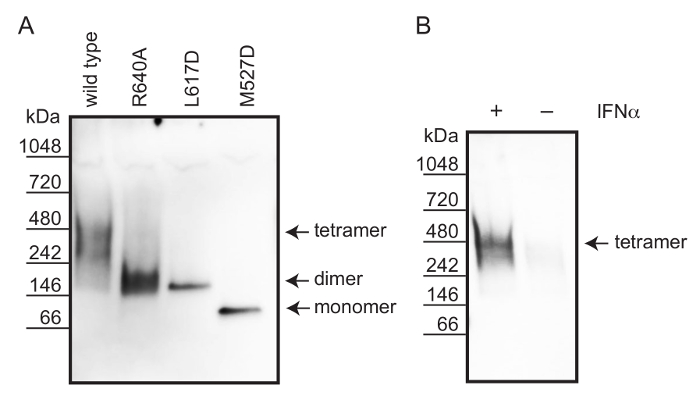
Figure 2: Workflow diagram of non-denaturing PAGE. A systematic representation of the non-denaturing PAGE approach of cell lysates. During cell lysis, detergents solubilize the proteins and thiol groups are protected by iodoacetamide to prevent protein aggregation. Dialysis removes small metabolites and salts that could interfere with non-denaturing PAGE 19. The complex separation is performed under non-denaturing conditions. Detection of the oligomeric complexes is achieved by Western blot followed by immunostaining. Please click here to view a larger version of this figure.
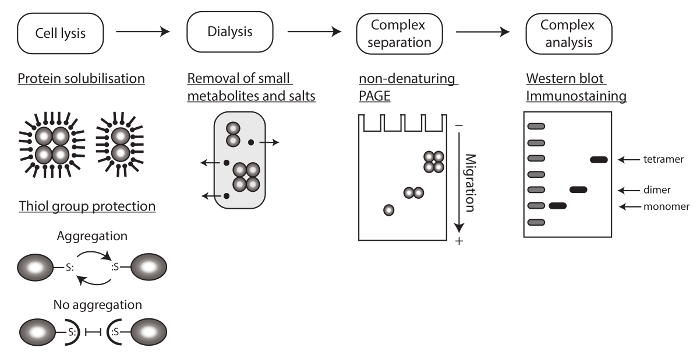
Figure 3: Determination of the oligomeric state of the human MxA protein using non-denaturing PAGE and Western blotting. (A) Recombinant MxA variants ectopically expressed in Vero cells. The complexes of wild type MxA (tetramer) interface mutants MxA(R640A), MxA(L617D) (dimers) and MxA(M527D) migrate at their expected molecular weights, confirming their oligomeric state. (B) A549 cells were stimulated with 1,000 IU per ml of IFN-α to induce MxA expression. The endogenous MxA shows a band that corresponds to the tetrameric form. Please click here to view a larger version of this figure.
| Buffer name | Content | Comments | ||
| Lysis buffer | 20 mM Tris-HCl (pH 7.5), 150 mM NaCl, 5 mM MgCl2, 100 μM iodoacetamide, 50 mM NaF, 1 mM Na3VO4, 1% NP-40, 50 mM β-glycerophosphate, 1 tablet per 50 ml Lysis buffer of EDTA free, protease inhibitor cocktail |
Add iodacetamide, NaF, Na3VO4, octylphenoxypolyethoxyethanol (NP-40), β-glycerophosphate and protease inhibitor cocktail right before cell lysis Iodacetamide is light-sensitive and instable when in solution. Prevent light exposure and dissolve right before use. |
||
| Dialysis buffer | 20 mM Tris-HCl (pH 6.8), 10% glycerol, 0.1% CHAPS, 0.5 mM DTT |
CHAPS can be raplaced by other non-denaturing detergents | ||
| Running buffer | 25 mM Tris-HCl (pH 8.3), 192 mM glycine, 0.1% CHAPS, 0.5 mM DTT |
At pH 8.3, most proteins are negatively charged. However, for basic proteins, an acidic pH should be used. Otherwise, the proteins will run into the opposite direction and will be lost. CHAPS can be replaced by other non-denaturing detergents |
||
| Sample buffer | 310 mM Tris-HCl (pH6.8), 0.05% bromophenol blue, 50% glycerol |
|||
| SDS buffer | 25 mM Tris-HCl (pH 8.3), 192 mM glycine, 0.1% SDS |
|||
| Blotting buffer | 25 mM Tris, 192 mM glycine, 20 % Methanol |
For very large complexes, Methanol can be ommitted | ||
| Blocking buffer | 50 mM Tris-HCl (pH 7.4) 150 mM NaCl 0.05 % Tween 20 5 % Milk powder |
|||
| Growth medium | Dulbecco's modified medium 1x Penicillin/Streptomycin 2 mM Glutamine 10% Fetal Calf Serum |
|||
Table 1: Buffer recipes required for non-denaturing PAGE.
Discussion
Here we describe a simple method that allows the rapid determination of the oligomeric state of proteins expressed in mammalian cells by non-denaturing PAGE followed by Western blot analysis. The major advantage of this approach is that the oligomeric state of a given protein can be determined from whole cell lysates without prior protein purification. This may be important for proteins that oligomerize or exert their function in association with auxiliary factors. In addition, the proteins are still in their native state and if further extracted from the gel, the enzymatic activity or other protein functions can be determined and correlated to the oligomeric state.
A critical aspect of this protocol is the choice of detergents during sample preparation. This is of particular importance for proteins associated with cellular membranes. MxA appears to be primarily associated with membranes of the smooth endoplasmic reticulum 25. For cell lysis the non-ionic detergent NP40 was optimal, preventing the precipitation of MxA. After buffer exchange and removal of low molecular weight impurities of the lysates by dialysis as described previously 19 the presence of 0.1 % 3-[(3-cholamidopropyl)dimethylammonio]-1-propanesulfonate (CHAPS) was required to prevent precipitation of MxA during gel electrophoresis. In addition, CHAPS does not interfere with the enzymatic activity of MxA as determined with purified recombinant MxA protein expressed in E. coli 12. It is of great importance that the detergent does not denature the proteins or disrupts protein-protein interactions during lysis. Non-denaturing detergents such as NP40, Octoxinol 9, digitonin and CHAPS are suitable for solubilization. The choice of the detergent and its concentration should be determined empirically. The detergent might also influence downstream experiments e.g. for certain assays the detergent needs to be removed which is easily achieved with CHAPS by dialysis, but not with Octoxinol 9 26.
Since the described method analyzes proteins derived from cell lysates under non-denaturing conditions, no prior purification of the proteins is required. This is an advantage since purification of recombinant proteins sometimes requires buffer optimizations by the addition of high salts and other additives to prevent the protein from aggregation or precipitation. However, these additives and the high salt concentrations might have an impact on the oligomerization of the protein that might not necessarily resemble its natural state. Especially in the case of the human MxA protein, the salt concentration and the presence of nucleotides play a crucial role in the formation of higher oligomeric states 27. In cell lysates, proteins are more easily stabilized since the cellular stabilization factors are still present. Therefore it is possible to analyze protein complexes under more cell physiological conditions (e.g. physiological salt concentrations, and pH). This protocol should also be applicable for other proteins forming oligomers, for example for the structurally related MxB or dynamin 8.
Another important aspect of the protocol is the protection of free thiol groups to prevent the formation of artificial disulfide bridges of cytoplasmic proteins during lysis (Figure 1). Initial experiments showed that addition of 1,4-dithiothreitol (DTT) or β-mercaptethanol is not sufficient to prevent the formation of artificial disulfide bonds during sample preparation. Both reducing agents protect the thiol group reversibly from forming disulfide bridges. This reversible protection might not be sufficient to protect all thiol groups permanently. In case of the MxA protein, this leads to irreversible aggregation of the protein. However, addition of iodoacetamide that irreversibly protects the free thiol groups of cysteines greatly reduced aggregation of the MxA protein Furthermore, iodoacetamide treatment of lysates prepared from MxA expressing mammalian cells had no influence on the GTPase activity of immunoprecipitated MxA when compared to MxA from lysates treated with DTT (data not shown).
Other crucial considerations for the exact determination of the number of protomers in an oligomer are the choice of the polyacrylamide concentration range as well as the protein molecular weight reference. The range of the polyacrylamide concentration should be first established to allow maximal separation of the bands at the expected molecular masses of oligomers. Non-denaturing PAGE does not contain any SDS. Lacking SDS, the charge, molecular mass and the shape of the protein determines its electrophoretic mobility. Therefore, the choice of the protein molecular weight marker is crucial. Ideally, a molecular weight reference would be a recombinant purified form of the protein of interest with known oligomeric states. Since this is not always available, a non-denatured or native protein marker should be used. Since MxA has been shown to associate with other cellular proteins such as UAP56 or viral nucleoproteins of Thogotovirus, La Crosse virus or influenza virus 12,28-30, we also tested by immunostaining using specific antibodies whether UAP56 or influenza A nucleoprotein would co-separate with MxA on the non-denaturing PA gels. However we found no evidence for the formation of MxA hetero-oligomers. This probably is due to the fact that MxA-UAP56 and MxA-nucleoprotein interactions are of low affinity 12,24. Moreover, the fraction of MxA protein associating with UAP56 or viral nucleoproteins might be very low and hence difficult to detect by this method.
Further, it is important to take into consideration the pKa of the protein to be analyzed. In the described non-denaturing PAGE protocol, the proteins are separated by electrophoresis at pH 8.3. Most proteins are negatively charged at this pH. However, basic proteins exhibit a positive net charge at pH 8.3 and therefore will run into the opposite direction. As a consequence, it is important, for the analysis of basic proteins to adjust the pH of the running buffer to ensure an overall negative charge of the protein.
Taken together we present here a protocol for the rapid assessment of the oligomeric state of proteins expressed in mammalian cells without the need of its prior purification.
Offenlegungen
The authors have nothing to disclose.
Acknowledgements
This work was funded by a Grant from the Swiss National Science foundation (Grant nr. 31003A_143834) to JP.
Materials
| Slide-A-Lyzer MINI Dialysis Units, 10K MWCO, 0.5 mL | Thermo Fisher Scientific | 69570 | Pre-equilibrate in dialysis buffer ( if Glycerol removal is desired) Can be self-made according to Fiala et al. 2011 |
| 4–15% Mini-PROTEAN TGX Precast Protein Gels, 10-well, | Bio-Rad | 456-1083 | Pre-run in running buffer to adjust buffer system |
| cOmplete, Mini, EDTA-free | Roche | 11836170001 | use 1 tablet per 50 ml |
| PBS, pH 7.4 bottle a 500ml Gibco | Thermo Fisher Scientific | 14190-094 | |
| Ponceau S solution | Sigma-Aldrich | P7170 | toxic! wear gloves and protect eyes! |
| NativeMark Unstained Protein Standard 50ul | Invitrogen | P/N 57030 | load 5 ul/well |
| A549 cells | ATCC | ATCC CCL185 | Grow in growth medium (see Table 1) |
| Vero cells | ATCC | ATCC CCL81 | Grow in growth medium (see Table 1) |
| anti-Mx1 antibody | Novus Biologicals | H00004599_D01P | Use at a 1:1000 dilution |
| ECL Anti-rabbit IgG, Horseradish Peroxidase linked whole antibody (from donkey) | GE-Healthcare | NA934V | Use at a 1:10000 dilution |
| 0.5% Trypsin-EDTA (1x) Life Technologies | Thermo Fisher | 15400-054 | |
| Iodoacetamide 5g | Sigma-Aldrich | I-6125 | stock 100mM |
| Bromphenolblue | Sigma-Aldrich | B0126-25G | |
| DMEM +4.5g/l Gluc,+L-Glut,+Pyruvate life technologies | Thermo Fisher Scientific | 41966-029 | |
| Pen Strep 100 x 100ml life technologies | Thermo Fisher Scientific | 15140 – 130 | |
| Glutamax 100xStock, 100ml life technologies | Thermo Fisher Scientific | 350500-038 | |
| Fetal Bovine Serum, Dialyzed , US Origin 500ml Gibco Lot:42G9552K | Thermo Fisher Scientific | 10270-106 | |
| Cellulose filter paper | Bio-Rad | 1703965 | |
| PVDV blotting membrane | GE-Healthcare | 10600022 | |
| Tris(hydroxymethyl)aminomethane | Biosolve | 0020092391BS | |
| sodium fluoride (NaF) | Sigma Aldrich | S-7920 | |
| NP-40 | Calbiochem | 492015 | |
| cOmplete, Mini, EDTA-free | Roche | 11836170001 | |
| Tween 20 | Calbiochem | 6555204 | |
| CHAPS 10% solution | Amresco | N907 | |
| DL-Dithiothreitol (DTT) | Sigma Aldrich | 43819 | |
| Glycine | Biosolve | 0007132391BS | |
| sodium orthovanadate (Na3VO4) | Sigma Aldrich | 450243 | |
| Glycerol | Sigma Aldrich | G7757 | |
| b-Glycerophospate | Sigma Aldrich | G9422 | |
| Milk powder | Migros/Switzerland | ||
| Methanol | Millipore | 1.06009 | |
| sodium cloride (NaCl) | Sigma Aldrich | 71380 | |
| magnesium chloride (MgCl2) | Amresco | 288 | |
| Sodium dodecyl sulphate (SDS) | Sigma Aldrich | L4509 | |
| sodium hydroxide (NaOH) | Sigma Aldrich | S-8045 |
Referenzen
- Baisamy, L., Jurisch, N., Diviani, D. Leucine zipper-mediated homo-oligomerization regulates the Rho-GEF activity of AKAP-Lbc. J Biol Chem. 280, 15405-15412 (2005).
- Chen, C. P., Posy, S., Ben-Shaul, A., Shapiro, L., Honig, B. H. Specificity of cell-cell adhesion by classical cadherins: Critical role for low-affinity dimerization through beta-strand swapping. Proc Natl Acad Sci U S A. 102, 8531-8536 (2005).
- Jackson-Fisher, A. J., Chitikila, C., Mitra, M., Pugh, B. F. A role for TBP dimerization in preventing unregulated gene expression. Mol Cell. 3, 717-727 (1999).
- Torshin, I. Activating oligomerization as intermediate level of signal transduction: analysis of protein-protein contacts and active sites in several glycolytic enzymes. Front Biosci. 4, 557-570 (1999).
- Goodsell, D. S., Olson, A. J. Structural symmetry and protein function. Annu Rev Biophys Biomol Struct. 29, 105-153 (2000).
- Flydal, M. I., Martinez, A. Phenylalanine hydroxylase: function, structure, and regulation. IUBMB Life. 65, 341-349 (2013).
- Haller, O., Kochs, G. Human MxA protein: an interferon-induced dynamin-like GTPase with broad antiviral activity. J Interferon Cytokine Res. 31, 79-87 (2011).
- Haller, O., Staeheli, P., Schwemmle, M., Kochs, G. Mx GTPases: dynamin-like antiviral machines of innate immunity. Trends Microbiol. 23, 154-163 (2015).
- Di Paolo, C., Hefti, H. P., Meli, M., Landis, H., Pavlovic, J. Intramolecular backfolding of the carboxyl-terminal end of MxA protein is a prerequisite for its oligomerization. J Biol Chem. 274, 32071-32078 (1999).
- Janzen, C., Kochs, G., Haller, O. A monomeric GTPase-negative MxA mutant with antiviral activity. J Virol. 74, 8202-8206 (2000).
- Gao, S., et al. Structure of myxovirus resistance protein a reveals intra- and intermolecular domain interactions required for the antiviral function. Immunity. 35, 514-525 (2011).
- Nigg, P. E., Pavlovic, J. Oligomerization and GTP-binding Requirements of MxA for Viral Target Recognition and Antiviral Activity against Influenza A Virus. J Biol Chem. 290, 29893-29906 (2015).
- Gao, S., et al. Structural basis of oligomerization in the stalk region of dynamin-like MxA. Nature. 465, 502-506 (2010).
- Ghosh, I., Hamilton, A. D., Regan, L. Antiparallel leucine zipper-directed protein reassembly: Application to the green fluorescent protein. J Am Chem Soc. 122, 5658-5659 (2000).
- Patching, S. G. Surface plasmon resonance spectroscopy for characterisation of membrane protein-ligand interactions and its potential for drug discovery. Biochim Biophys Acta. 1838, 43-55 (2014).
- Kenworthy, A. K. Imaging protein-protein interactions using fluorescence resonance energy transfer microscopy. Methods. 24, 289-296 (2001).
- Wyatt, P. J. Light-Scattering and the Absolute Characterization of Macromolecules. Analytica Chimica Acta. 272 (93), 1-40 (1993).
- Howlett, G. J., Minton, A. P., Rivas, G. Analytical ultracentrifugation for the study of protein association and assembly. Curr Opin Chem Biol. 10, 430-436 (2006).
- Fiala, G. J., Schamel, W. W., Blumenthal, B. Blue native polyacrylamide gel electrophoresis (BN-PAGE) for analysis of multiprotein complexes from cellular lysates. J Vis Exp. , (2011).
- Zou, H., Li, Y., Liu, X., Wang, X. An APAF-1.cytochrome c multimeric complex is a functional apoptosome that activates procaspase-9. J Biol Chem. 274, 11549-11556 (1999).
- Schagger, H., Cramer, W. A., von Jagow, G. Analysis of molecular masses and oligomeric states of protein complexes by blue native electrophoresis and isolation of membrane protein complexes by two-dimensional native electrophoresis. Anal Biochem. 217, 220-230 (1994).
- Walker, J. M. Nondenaturing polyacrylamide gel electrophoresis of proteins. Methods Mol Biol. 32, 17-22 (1994).
- Stoscheck, C. M. Quantitation of protein. Methods Enzymol. 182, 50-68 (1990).
- Wisskirchen, C., Ludersdorfer, T. H., Muller, D. A., Moritz, E., Pavlovic, J. Interferon-induced antiviral protein MxA interacts with the cellular RNA helicases UAP56 and URH49. J Biol Chem. 286, 34743-34751 (2011).
- Stertz, S., et al. Interferon-induced, antiviral human MxA protein localizes to a distinct subcompartment of the smooth endoplasmic reticulum. J Interferon Cytokine Res. 26, 650-660 (2006).
- Seddon, A. M., Curnow, P., Booth, P. J. Membrane proteins, lipids and detergents: not just a soap opera. Biochim Biophys Acta. 1666, 105-117 (2004).
- Kochs, G., Haener, M., Aebi, U., Haller, O. Self-assembly of human MxA GTPase into highly ordered dynamin-like oligomers. J Biol Chem. 277, 14172-14176 (2002).
- Kochs, G., Haller, O. GTP-bound human MxA protein interacts with the nucleocapsids of Thogoto virus (Orthomyxoviridae). J Biol Chem. 274, 4370-4376 (1999).
- Reichelt, M., Stertz, S., Krijnse-Locker, J., Haller, O., Kochs, G. Missorting of LaCrosse virus nucleocapsid protein by the interferon-induced MxA GTPase involves smooth ER membranes. Traffic. 5, 772-784 (2004).
- Wisskirchen, C., Ludersdorfer, T. H., Muller, D. A., Moritz, E., Pavlovic, J. The cellular RNA helicase UAP56 is required for prevention of double-stranded RNA formation during influenza A virus infection. J Virol. 85, 8646-8655 (2011).

