Preparation of Human Tissues Embedded in Optimal Cutting Temperature Compound for Mass Spectrometry Analysis
Summary
Sphingolipids are bioactive metabolites with well-established roles in human disease. Characterizing alterations in tissues with mass-spectrometry can reveal roles in disease etiology or identify therapeutic targets. However, the OCT-compound used for cryopreservation in biorepositories interferes with mass-spectrometry. We outline methods to analyze sphingolipids in human tissues embedded in OCT with LC-ESI-MS/MS.
Abstract
Sphingolipids are cellular components that have well-established roles in human metabolism and disease. Mass spectrometry can be used to determine whether sphingolipids are altered in a disease and investigate whether sphingolipids can be targeted clinically. However, properly powered prospective studies that acquire tissues directly from the surgical suite can be time consuming, and technically, logistically, and administratively challenging. In contrast, retrospective studies can take advantage of cryopreserved human specimens already available, usually in large numbers, at tissue biorepositories. Other advantages of procuring tissues from biorepositories include access to information associated with the tissue specimens including histology, pathology, and in some instances clinicopathological variables, all of which can be used to examine correlations with lipidomics data. However, technical limitations related to the incompatibility of optimal cutting temperature compound (OCT) used in the cryopreservation and mass spectrometry is a technical barrier for the analysis of lipids. However, we have previously shown that OCT can be easily removed from human biorepository specimens through cycles of washes and centrifugation without altering their sphingolipid content. We have also previously established that sphingolipids in human tissues cryopreserved in OCT are stable for up to 16 years. In this report, we outline the steps and workflow to analyze sphingolipids in human tissue specimens that are embedded in OCT, including washing tissues, weighing tissues for data normalization, the extraction of lipids, preparation of samples for analysis by liquid chromatography electrospray ionization tandem mass spectrometry (LC-ESI-MS/MS), mass spectrometry data integration, data normalization, and data analysis.
Introduction
Sphingolipids are bioactive metabolites known for their roles in human metabolism and disease1,2. They regulate complex cellular processes such as cell migration, cell survival and death, cell movement, vesicular trafficking, cellular invasion and metastasis, angiogenesis, and the production of cytokines1,2,3,4,5,6,7,8,9. Defects in the regulation of sphingolipid metabolism contribute to the initiation and progression of cancers, determine how aggressive cancers are, and how cancers respond to and develop resistance to therapy3,10. Therefore, because of these broad impacts on the etiology of disease, analytical methods that can precisely establish disease-specific sphingolipid alterations are important tools. Mass spectrometry (MS) is the most accurate and reliable method to analyze sphingolipid alterations.
Human specimens that can be used for the analysis of sphingolipid alterations can be obtained prospectively from the surgical suite or retrospectively from tissue biorepositories. Fresh tissues from surgery are advantageous because they can be directly analyzed by MS or other analytical methods. However, acquiring tissues prospectively has administrative, technical, and logistical hurdles, and collecting sufficient specimens to reach statistical power can be challenging. Obtaining tissues from biorepositories is advantageous because they can be acquired retrospectively, in large numbers, and biorepositories confirm histology and pathology, use standard operating procedures to cryogenically preserve and store tissues, and can provide clinicopathological data that can be used for correlation analyses. However, to preserve molecular and structural features, biorepositories may cryopreserve tissues by embedding them in optimal cutting temperature (OCT) compound, which we have shown interferes with data normalization assays and the quantification of sphingolipids by liquid chromatography electrospray ionization tandem mass spectrometry (LC-ESI-MS/MS)11. It has also been shown that polyvinyl alcohol and polyethylene glycol, the primary components in OCT compound, result in ion suppression in other MS analysis platforms12,13,14,15. Therefore, OCT compound must be removed from tissues prior to sphingolipidomic analysis by MS.
In a previous report, we have validated a protocol for the removal of OCT compound from human specimens for LC-ESI-MS/MS analysis11 and the methodology used for weighing tissues for data normalization11. Here, we detail the steps of the sphingolipidomic OCT compound-removal protocol (sOCTrP) and show representative data from human lung adenocarcinoma tumors and normal adjacent uninvolved tissues.
Protocol
De-identified human lung tissues were obtained from the Virginia Commonwealth University (VCU) Tissue and Data Acquisition and Analysis Core under an internal review board (IRB) approved protocol (#HM2471). The use of mice for research and harvesting of mice tissues was approved by the VCU institutional animal care and use committee (IACUC).
1. Preparation of materials
NOTE: These steps should be performed a day prior to the tissue washing.
- Pre-label and weigh 1.5 mL polypropylene centrifuge tubes with concise but clear identifier codes for each sample that will be processed the next day. Use a chemically resistant permanent marker (see Table of Materials) that will not fade under the repeated tube handling.
- Open an electronic spreadsheet and label consecutive columns with the following column headers: tube identifier (ID), tube alone, tube w/tissue, and total tissue. Enter the tube identifiers that will be processed into the column labeled tube ID.
- Pre-calibrate and zero an analytical scale. Place a clean 10 mL Erlenmeyer flask in the center of the balance plate (the flask will act as a tube holder). Close the weighing chamber door and tare the scale with the flask.
- Weigh 1.5 mL centrifuge tubes. Carefully place the tube in the flask and close the weighing chamber door. Allow the weight to stabilize and record the weight in the column labeled tube alone.
NOTE: Avoid moving the 10 mL Erlenmeyer flask when weighing tubes. If the flask is moved, re-center the flask and re-tare the balance. Place the closed tubes in a rack and store until needed. - Pre-label 13 x 100 mm screw-top glass tubes and caps with the identifier codes and fill them with 2 mL of LC-MS grade methanol. Place them in a tube rack, cap the tubes, and store them in a refrigerator (4 °C) until use.
NOTE: Use a bottle top solvent dispenser (see Table of Materials). If one is not available, use glass pipettes to avoid the use of plastics when dispensing organic solvents. - Pre-label 13 x 100 mm glass test-tubes. Cover and store the test tubes at room temperature.
- Pre-label autoinjector vials with identifier codes. Cover and store the autoinjector vials at room temperature until use.
- Prepare tissue-drying wicks.
- Begin by cleaning surgical scissors by immersing them in LC-MS grade methanol for 5 min, and then wipe them with a clean laboratory grade tissue.
- Using powder free gloves, twirl the end of a laboratory grade tissue (see Table of Materials) until a finely tapered 30-40 mm wick is formed.
- Using methanol-cleaned scissors, cut the wick on the thick end. Place wicks in a clean carboard box. Make double the number of wicks than will be needed to process all the samples.
- Prepare shortened 1 mL pipette tips. Using methanol-cleaned surgical scissors, cut 4-5 mm off the tip of a 1 mL pipette tip, and place the tip back in the tip holder. Prepare twice as many tips as samples to be processed.
NOTE: These will be used in the retrieval of washed tissues from tubes. Do not touch the end of the tip. - Pre-chill molecular biology grade phosphate buffered saline (PBS) to 4 °C (0.5 L is sufficient). This will be used to retrieve specimens. For each sample that will be processed, pre-chill (4 °C) 60 mL of ultra-pure deionized water.
- Prepare mass spectrometry internal standards. Dissolve the following lipids in ethanol:methanol:water (7:2:1; v/v/v) at a concentration of 25 nmol/mL: d17:1-sphingosine, (2S,3R,4E)-2-aminoheptadec-4-ene-1,3-diol; d17:1-sphingosine-1-phosphate, heptadecasphing-4-enine-1-phosphate; C12-Ceramide (d18:1/C12:0), N-(dodecanoyl)-sphing-4-enine; C12-sphingomyelin (d18:1/C12:0), N-(dodecanoyl)-sphing-4-enine-1-phosphocholine; C12-glucosylceramide (d18:1/C12:0), N-(dodecanoyl)-1-β-glucosyl-sphing-4-eine; C12-lactosylceramide (d18:1/C12:0), N-(dodecanoyl)-1-ß-lactosyl-sphing-4-ene.
2. Tissue washing
NOTE: Completing all steps in section 1 one day prior allows a skilled researcher to process up to 40 samples per day, and novices ~10-15 samples per day. Begin early in the morning and do not stop until all steps in sections 2.1-3.8 are completed.
- On the day that the tissues will be processed, prepare the biosafety cabinet work area by equipping it with a vortexer, a fully charged serological pipettor, and an ice tray. For all steps in this section, wear appropriate PPE, including a lab coat, a double layer of gloves, and protective eyewear. CAUTION: Human tissues and fluids should be considered and treated as biohazardous materials.
- Pre-chill (4 °C) a swinging bucket centrifuge, any adapters needed for holding 15 mL conical centrifuge tubes, and centrifuge biocontainment lids. Pre-chill (4 °C) a fixed angle centrifuge capable of holding 1.5 mL centrifuge tubes.
- If tissues are not aliquoted into 15 mL polypropylene centrifuge tubes, use a methanol cleaned spatula (clean for each new tissue sample) to transfer them to pre-labeled 15 mL polypropylene tubes. Label the 15 mL caps also.
- Place a tube rack capable of holding 15 mL centrifuge tubes in an ice tray and fill the tray with ice. Thaw tissues that will be processed that day by placing them in the rack on ice.
NOTE: A minimum of 3-4 mg of tissue should be requested from the biorepository for processing as it has been previously shown that tissue washing and weighing protocols are accurate and reliable at these weights11.
- Place a tube rack capable of holding 15 mL centrifuge tubes in an ice tray and fill the tray with ice. Thaw tissues that will be processed that day by placing them in the rack on ice.
- Move the ice tray with the samples into the biosafety cabinet.
- Perform three sOCTrP cycles11 as discussed below (i.e., perform steps 2.5.1-2.5.5 three times).
- Add 10 mL of ice-cold ultra-pure deionized water to each tube and cap them tightly. Allow the tubes to incubate for 10 min.
- Vigorously vortex each tube for 10-20 s or until all the tissue deposited on the walls of the tube, or a tissue pellet, is completely resuspended.
NOTE: In sOCTrP cycles 2-3, it is critical that the pellet be resuspended so that any OCT in the pellet is diluted into the wash solution. - Transfer the tubes to the pre-chilled (4 °C) swinging bucket centrifuge. Centrifuge samples at 4,000-5,000 x g for 10 min at 4 °C.
NOTE: Avoid the generation of and exposure to biohazardous aerosols generated during centrifugation by sealing the centrifuge carriers with a biocontainment lid (see Table of Materials). - Retrieve samples from the centrifuge and transfer to the ice tray. Place the tray in a biosafety cabinet and uncap the tubes. Save the caps.
- Carefully aspirate the wash solution. Avoid aspirating the tissue material in the pellet by leaving ~0.5 mL of the supernatant in the tube. Cap the tubes and avoid cross-contamination by matching labeled caps with tubes.
NOTE: The wash solution (the supernatant) in step 2.5.5 is a biohazardous material; handle and dispose of it following biosafety guidelines appropriate for specimens of human origin.
3. Tissue weighing (for data normalization)
- After the last sOCTrP wash cycle, carefully aspirate most of the wash solution but avoid disturbing the pellet. Leave ~0.5 mL of the wash solution in the tube.
- Using the shortened pipette tips (prepared in step 1.9), take-up 500 µL of ice-cold PBS and add it to the tube.
- Using the same tip, gently resuspend the pellet and retrieve all tissue. Avoid turbulent pipetting to decrease tissue loss by its adherence to the tube and pipette tip walls.
- Add the resuspended tissue to its corresponding pre-labeled and pre-weighed 1.5 mL centrifuge tube and place the tube on ice. Repeat for all tissues in the data set.
- Place the 1.5 mL tubes into a pre-chilled centrifuge and pellet the tissues at 7,000 x g for 5-7 min and 4 °C.
- Retrieve tubes and carefully aspirate as much of the PBS as possible without disturbing the pellet. Place the tube back on ice.
- Centrifuge tubes for an additional 3 min at 7,000 x g to ensure tissues are well pelleted. Transfer the tubes to ice.
NOTE: The additional centrifugation step is important to prevent tissue loss when removing all wash solution and wicking. If processing more than five samples, repeat the 3 min, 7,000 x g centrifugation step after every fifth sample is processed to ensure tissues remain pelleted. - Using the wicks prepared in step 1.8, gently remove any remaining wash solution. Use a clean wick for each sample. It is acceptable to gently dab the tissue with the wick, which will help remove most of the wash solution but avoid losing any tissue by adherence to the wick. Close the tube and place it on ice.
- Weigh each tube in the recently calibrated, tared, and zeroed analytical balance.
- Place each tube in the glass Erlenmeyer flask and close the chamber door. When the weight has stabilized, record it in the electronic spreadsheet column labeled tube w/tissue.
- After weighing, place the tube back on ice.
- Add 300 µL of ice-cold PBS. Using a shortened pipette tip (step 1.9), carefully retrieve all tissue and deposit in a pre-labeled screw-top glass tube (prepared in step 1.5) containing 2 mL of ice-cold LC-MS grade methanol (add the tissue to the methanol, not to the side of the tube).
- Calculate the amount of tissue that was washed by subtracting the tube alone from the tube w/tissue value. Record this number in the total tissue column. The total tissue value will be used to normalize mass spectrometry data in step 6.12.
- Store the samples at -80 °C until ready to perform steps 4.1-4.15.
NOTE: This is a good stopping point, and the samples can be stored safely for a couple of weeks at -80 °C. For other tissue normalization methods, such as protein concentration or lipid phosphate content, refer to Rohrbach et al.11.
4. Lipid extraction
NOTE: It is important to begin these steps in the morning, particularly when many samples will be processed. Before starting, pre-heat a water bath or incubator to 48 °C.
- Retrieve the samples prepared in step 3.8 and the MS lipid standards (prepared in step 1.11) from the freezer. Allow them to equilibrate to room temperature for 20 min.
- Vortex and immerse the internal standard solution in an ultrasonic water bath (see Table of Materials) for 2-3 min or until the MS lipid standards are fully dissolved prior to dispensing into samples. This will avoid errors in data normalization.
- Using a repeating pipette, add 250 pmol (10 µL from the 25 nmol/mL solution prepared in step 1.11) of internal mass spectrometry standards to each tube. Re-cap the tubes lightly.
- To facilitate homogenization, adjust the methanol volume to 2 mL. Only use LC-MS/MS grade methanol.
- Working one tube at a time, use a homogenizer to triturate the tissue until no clumps are visible (typically 5-20 s). Move the homogenizer tip in a circular motion and gently press against the tube wall to assist trituration. Re-cap the tube.
NOTE: Ensuring all tissues are finely triturated will maximize lipid extraction. Do not add chloroform to samples prior to homogenization as the disposable homogenizer tips are made of plastic that will dissolve in solutions containing chloroform. - Immerse the tube at a 45° angle in an ultrasonic water bath for 5-10 s.
- If no tissue clumps form upon immersing in the ultrasonic bath, cap it, and move on to the next tube.
- If tissue clumps form when the tube is immersed in the ultrasonic bath, re-homogenize, and target the clumps.
- Repeat this process (homogenization/immersion in an ultrasonic bath) iteratively until all large tissue clumps are broken-up.
- In a fume-hood, add LC-MS grade chloroform and methanol to each tube (use a bottle top dispenser) to achieve a ratio of 2:1:0.1 methanol:chloroform:water. Using clean caps, seal the tubes tightly and leave on the benchtop until the end of the day.
- For tissues weighing 50 mg or less, use a total extraction volume of 4 mL, and adjust using this ratio (50 mg:4 mL extraction solution) for larger specimens.
- Before leaving that evening, place the tightly capped tubes in a 48 °C water bath or incubator and incubate them overnight.
NOTE: It is critical that the tubes be tightly capped so that the solvent ratios do not change due to evaporation. - The next morning, retrieve the samples from the 48 °C water bath and pellet any solvent-insoluble debris by centrifugation at 4,000-5,000 x g at 4 °C for 20 min.
- Working in a fume-hood, carefully decant the supernatant into the pre-labeled 13 x 100 mm borosilicate glass test-tubes. Tilt the 13 x 100 mm tube at a 30-45° angle to facilitate decanting.
NOTE: If processing more than 12 samples, do not allow the centrifuged tubes to sit for extended periods of time as the pellets will soften and insoluble debris will be transferred when performing step 4.10. To avoid this, only take 12 tubes out of the centrifuge at a time and continue centrifuging the other tubes until you are ready to decant them. - Transfer the tubes to a vacuum concentrator capable of holding 13 x 100 mm tubes. Evaporate all solvent with an initial 1 h heating step (40 °C) and run under the maximum vacuum.
- Remove tubes from the vacuum concentrator and add 500 µL of LC-MS grade methanol (use a bottle top dispenser) to each tube.
- Resuspend the dried lipid extracts by vortexing for 5-10 s, followed by immersing tubes at a 45° angle in an ultrasonic water bath for 2 min while rotating the tube. Re-vortex for 5 s and immerse in ultrasonic water bath for an additional minute.
NOTE: This is a critical step to ensure the maximal recovery of extracted lipids. Do not move ahead until all the dried material on the wall of the tube has been resuspended. - Place the 13 x 100 mm tubes in a pre-chilled (4 °C) centrifuge and remove any insoluble debris by centrifugation at 4,000-5,000 x g at 4 °C for 20-30 min.
- Carefully decant the clarified supernatant into a pre-labeled autoinjector vial. Tilting the autoinjector vial at a 30-45° angle will facilitate decanting. Ensure that the entire contents of the 13 x 100 mm tube are decanted into the autoloader vial.
- Cap the tubes tightly and store tubes at -80 °C until analysis by LC-ESI-MS/MS.
5. LC-ESI-MS/MS analysis
NOTE: The following procedure uses a binary pump system coupled to an autoinjector, degasser, and LC-MS/MS system operating in a triple-quadrupole mode. The column temperature was maintained using a column oven. See Table of Materials for details on the equipment used.
- Prepare mobile phase A1 (CH3OH:H2O:HCOOH, 58:41:1, v:v:v, with 5 mM ammonium formate) and mobile phase B1 (CH3OH:HCOOH, 99:1, v:v, with 5 mM ammonium formate). Use only LC-MS grade solvents, water, and reagents.
- For each set of samples, prepare four Blank autoloader vials containing 500 µL of LC-MS grade methanol.
- For each set of samples prepare two Internal Standard (label as IS) autoloader vials by diluting 10 µL (250 pmol total each standard) of the internal standard solution prepared in step 1.11 in 500 µL of LC-MS grade methanol.
- Set the column oven temperature to 60 °C, and the ion source temperature to 500 °C.
- For the MS/MS analysis, utilize the triple-quadrupole mode of the spectrometer. Set the first quadrupole (Q1) to pass precursor ions, while setting the third quadrupole (Q3) to pass molecularly distinctive product ions (or a scan across multiple m/z in Q1 or Q3) using N2 to collisionally induce dissociations in quadrupole 2 (Q2), which is offset from Q1 by 30-120 eV.
- Set the multiple reaction monitoring pairs (MRM) detection window at 120 s with a target scan time of 1.0 s. Refer to Table 1 for a listing of MRM pairs, ionization energies, and collision energies.
- Use the following parameters to resolve sphingolipids in the LC step: using a flow rate of 0.7 mL/min, equilibrate a 2.1 (i.d) x 50 mm C18 reverse phase column for 0.5 min with a solvent mixture of 95% mobile phase A1 and 5% mobile phase B1.
- After sample injection, maintain the mobile phase A1/B1 ratio at 95/5 for 2.25 min, followed by a linear gradient to 100% B1 over 1.5 min, which is then held at 100% B1 for 5.5 min, followed by a 0.5 min gradient return to 95/5 A1/B1.
- After the run, re-equilibrate the column with a mixture of 95/5 A1/B1 for 0.5 min.
- Use the following settings for the sample autoloader: rinsing volume, 500 µL; needle stroke, 52 mm (this should be adjusted depending on vial size and volume in each vial); rinsing speed, 35 µL/s; sampling speed, 5.0 µL/s; rinse dip time, 2 s; rinse mode, before and after aspiration; rinse time,2 s.
- Place samples prepared in step 4.15 onto an autoloader tray in the following order: Blank; IS; Blank; Samples prepared in step 4.15; Blank, IS, Blank.
- Analyze 5-10 µL of each sample by LC-ESI-MS/MS. Analyze the same volume for all samples in the data set.
6. Data processing, integration, and normalization
NOTE: Although the following steps outline a procedure for specific software (see Table of Materials), similar procedures on equivalent LC-MS instruments and software can be used to integrate MRM pairs for the analyzed lipid classes and corresponding internal standards.
- Open the quantitation software. In the Quantitate tab double click on the Quantitation Wizard. In the pop-up window, in the left-most workspace, navigate to the correct data set and left click on it. The available samples will be shown in the middle workspace.
- Highlight the samples to be analyzed and then click on the right-handed arrow to add samples to the Selected Samples workspace. Click on Next to navigate to the settings and query screen where the default settings are typically used.
- Hit Next to navigate to the integration method selection screen. Select an appropriate integration method containing MRM transition pairs for lipids to be analyzed. Refer to the Table 1 for MRM transition pairs. Hit Finish.
NOTE: Retention times will vary depending on instrumentation and individual settings and must therefore be verified for each individual setup. - In the navigation bar, click on the Peak Review pane so that MRM pairs can be inspected. To facilitate review, right-click on the Peak Review pane, change the Automatic Zooming auf 100% of largest peak, and a Zoom Window of 1.00 – 2.00 minutes.
- Right-click on the Quantitation Display window and select Analyte and the desired sphingolipid to be examined. Verify that there are no peaks in the Blank samples and that the correct peak has been integrated in the IS samples.
- Begin integrating unknown samples. Working one sphingolipid class at a time, inspect the retention time for the MRM pair of the internal standard (i.e., C12:0 ceramide) and ensure that the software has integrated the correct peak.
- Continue onto the analytes in the same lipid class (i.e., C14:0 ceramide, C16:0-ceramide, etc.), beginning with the shortest chain-length lipid in the class. Verify for each chain-length lipid that the software routine has integrated the correct MRM pair peak.
- When all the analytes have been integrated, create an electronic spreadsheet for each sphingolipid species analyzed (e.g., ceramides, sphingomyelins, etc.). Label column headers to match the order of the analytes and chain-lengths in the LC-MS/MS quantitation software. Also include a column header for sample ID and sample normalization weights determined in step 3.9.
NOTE: As previously described11, other common normalization measures include total lipid phosphate content or protein amount. - Export or copy the peak areas for all analytes and internal standards to the electronic spreadsheet.
- Normalize data by dividing the integrated peak area of each chain-length species for a given sphingolipid class by the peak area of its corresponding internal standard. For instance, C14:0 ceramide, C16:0 ceramide, C18:1 ceramide, etc., should each be divided by the C12:0 ceramide internal standard peak area.
- Convert peak area to moles of lipid recovered by multiplying the normalized peak area (calculated in step 6.10) with the amount of internal standard, in picomoles, that was added to the sample in step 4.3. In this protocol, this amount is 250 pmol.
- To obtain the relative amount of lipid in each sample, divide the picomoles of each lipid chain-length by the tissue weight determined in step 3.9. This will give units of picomoles of lipid per mg of tissue.
Representative Results
In this protocol, we describe in detail a method to remove OCT from cryo-preserved human tissues and weigh the tissues for analysis by LC-ESI-MS/MS. The materials required for this procedure are listed in Table of Materials. Shown in Figure 1 are results of a typical experiment where 10 human lung adenocarcinoma tumors and 10 normal adjacent tissues were washed to remove OCT and analyzed by LC-ESI-MS/MS. Importantly, as we have previously shown11, there are various chain-length species of ceramides (Figure 1A) and monohexosyl-ceramides (Figure 1B) that are significantly elevated in lung adenocarcinoma tumors as compared to normal adjacent uninvolved tissues.
In control experiments to assess the effect of OCT on the lipid extraction steps outlined in section 4, 200 mg of mice livers (C57BL6; males; 14-16 weeks old) were resuspended in 2 mL of phosphate buffered saline and homogenized until finely triturated. The resulting fine liver suspension was then extensively sonicated with a probe sonicator (three rounds of 1 min sonication on ice, 1 min rest on ice, 60% power; microtip). From this solution, six 300 µL liver-homogenate replicates were prepared. To three replicates, 200 mg of OCT was added (average amount found in biorepository specimens11), and 200 µL of water was added to the other three. The samples were then processed in parallel following the steps outlined in section 4. Importantly, following centrifugation of the single-phase Bligh-Dryer solution, samples containing OCT were cloudy whereas the control samples containing water were clear (Figure 2A). The cloudiness in the single-phase lipid extraction solution persisted even after increased centrifugation and extensive sonication (not shown). After the solvent was evaporated, samples containing OCT had large pellets (Figure 2B), which could not be reconstituted in methanol even under vigorous vortexing and extensive sonication. Samples containing OCT also showed a large loss of signal for internal standards of all species analyzed (Figure 3A–G). We have also previously shown that samples containing OCT showed loss of signal for many of the sphingolipids analyzed11.
Typically, it is expected that the sphingolipid standards will elute sequentially as shown in Figure 4 in the following order: d17:1 sphingosine, d17:1 sphingosine-1-phosphate, C12:0 lactosyl-ceramide, C12:0 monohexosyl-ceramide, C12:0 sphingomyelin, and C12:0 ceramide. For each of these sphingolipid species, using the gradient and instrumentation settings described in section 5 and previously11, there will be approximately a +0.15 min shift in the retention time for every 2-carbon increase in chain-length. For example, as shown in Figure 5, C14:0-ceramide (Figure 5B) will elute approximately +0.15 min later than C12:0-ceramide (Figure 5A). A double bond in the fatty acid often results in a -0.15 shift in retention time, so C16:0-ceramide (Figure 5C) will have a retention time similar to C18:1-ceramide (Figure 5D). However, longer chain-length lipids (i.e., C24:0, C26:0) may have larger retention time shifts (Figure 5I,K, respectively).
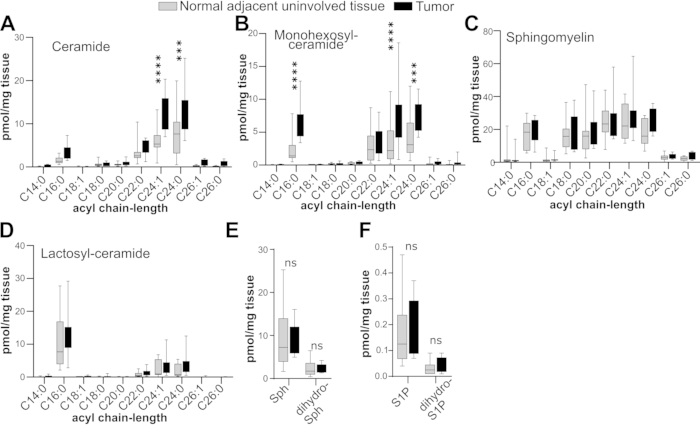
Figure 1: Sphingolipid profiles of lung adenocarcinomas and adjacent uninvolved lung tissues. Tissues were cryogenically stored in OCT, thawed, processed with three sOCTrP cycles, weighed, and analyzed by LC-ESI-MS/MS. Levels are in pmol per mg of tissue. The indicated acyl chain-length species of ceramide (A), monohexosylceramide (B), sphingomyelin (C), lactosylceramide (D), and sphingosine (E; Sph), sphingosine-1-phopshate (F; S1P), dihydro-Sph (E), and dihydro-S1P (F) were quantified. n = 10 tumors and n = 10 adjacent uninvolved tissues. Box plots show medians (black line) and whiskers of min to max. ns, not significant; ***, p≤0.005; ****, p ≤ 0.0005. Please click here to view a larger version of this figure.
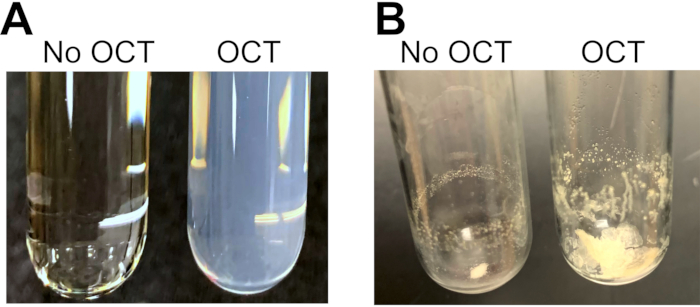

Figure 2: Representative images of extracted samples with and without OCT. Mouse livers were homogenized and sonicated, split into six identical samples: 200 mg of OCT was added to three samples and water to the other three. (A) After methanol and chloroform were added to achieve a ratio of 2:1:0.1 methanol:chloroform:water (v:v:v), overnight incubation at 48 °C, followed by centrifugation, the supernatants of samples that contained OCT were cloudy and could not be clarified by sonication, vortexing, or centrifugation. (B) Following solvent evaporation, a large pellet was recovered from samples containing OCT that could not be fully reconstituted in 0.5 mL of methanol after extensive sonication and vortexing. Please click here to view a larger version of this figure.
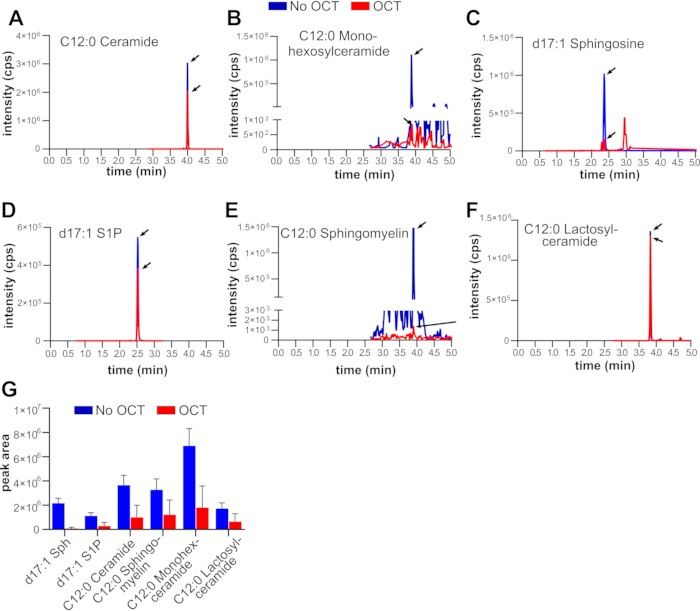
Figure 3: LC-ESI-MS/MS quantification of sphingolipid standards in the presence or absence of OCT. Mouse livers were homogenized, sonicated, and split into six identical samples. To three out of the six samples, 200 mg of OCT was added, while to the other three, water was added. After addition of internal standards, methanol, and chloroform, samples were processed identically and extracted lipids were analyzed by LC-ESI-MS/MS. Shown are multiple reaction monitoring (MRM) pairs of the indicated lipids (A–F), and corresponding integrated peak areas (G). Abbreviations: Sph, sphingosine; S1P, sphingosine-1-phosphate. Black arrows point to the correct MRM pair for the indicated sphingolipid. Please click here to view a larger version of this figure.
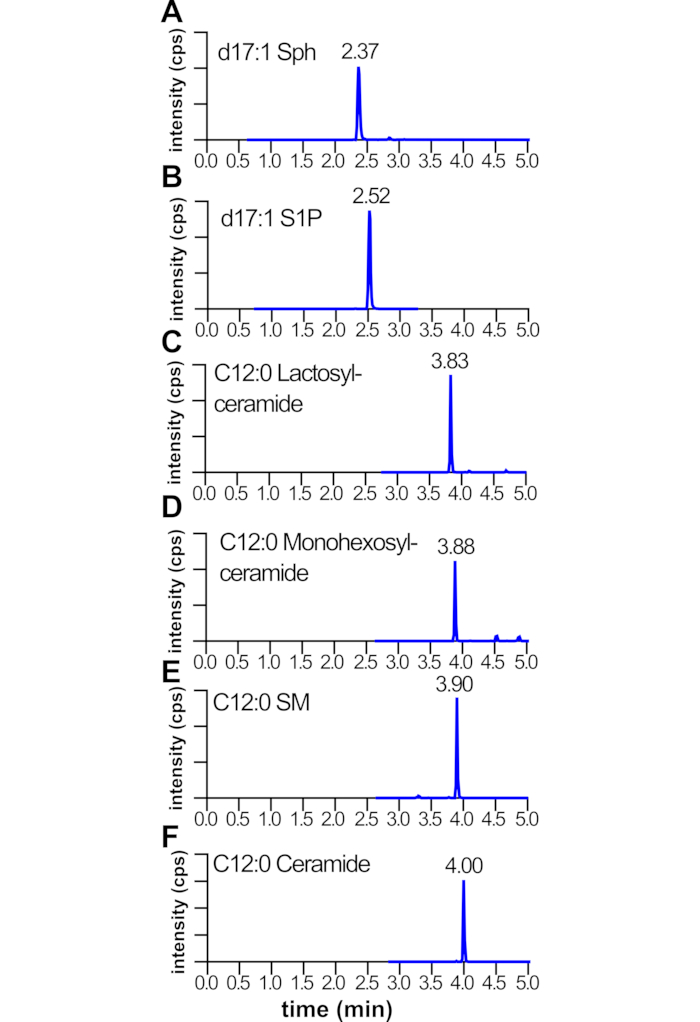
Figure 4: Relative retention times for LC-ESI-MS/MS multiple reaction monitoring pairs of sphingolipid standards. Mouse livers were homogenized and sonicated; internal standards, methanol, and chloroform added; and lipids extracted and analyzed by LC-ESI-MS/MS. Shown are retention times of multiple reaction monitoring pairs of the indicated lipids (A–F). Note the relative order in which lipids elute during the liquid chromatography gradient. Abbreviations: Sph, sphingosine; S1P, sphingosine-1-phosphate; SM, sphingomyelin. The X-axis is MRM pair retention time in minutes. The Y-axis is signal intensity for the MRM pairs. Please click here to view a larger version of this figure.
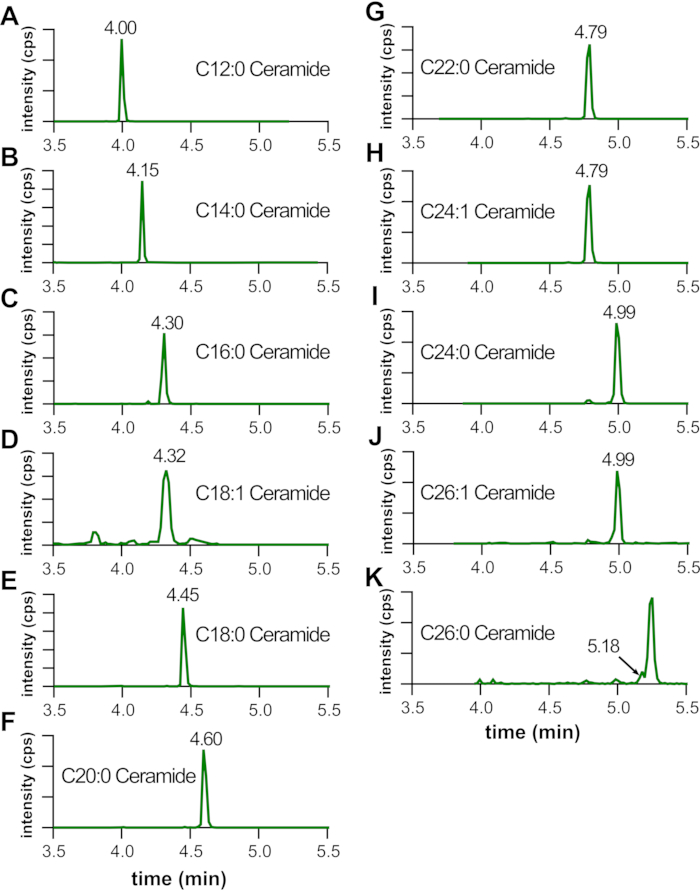
Figure 5: Acyl chain-length dependent shift in retention time of LC-ESI-MS/MS MRM pairs. Mouse livers were homogenized and sonicated; internal standards, methanol, and chloroform added; and lipids extracted and analyzed by LC-ESI-MS/MS. Shown are retention times and MRM pairs of the indicated lipids. Note the shift in retention time with increasing acyl-chain length for various lipids (A–K), which typically corresponds to a +0.15 min retention time per 2-carbon increase. A double bond will result in a -0.15 min retention time shift (D,H,J). However, for longer chain lipids, the relative increase in retention time may be longer (G–K). The X-axis is MRM pair retention time in minutes. The Y-axis is signal intensity for the MRM pairs. Please click here to view a larger version of this figure.
Table 1: MRM pairs, ionization parameters, and collision energies for LC-ESI–MS/MS analysis. DP, de-clustering potential; CE, collision energy. Please click here to download this Table.
Discussion
OCT is a common long-term cryo-preservation agent used in biorepositories. However, OCT can result in ion suppression when tissues are analyzed by various mass spectrometry platforms12,13,14,15, or result in loss of signal when samples are analyzed by LC-ESI-MS/MS11. OCT in cryopreserved tissues can also interfere with tissue normalization methods such as weighing and protein quantification assays such as Bradford and BCA11. Here, we present a detailed protocol to remove OCT from human tissues and subsequently be analyzed by mass spectrometry. The protocol may be modified to use other tissue sources that are not lung, or from mice tissues as we have previously described11. In addition, other mass spectrometry data normalization strategies can be used, including those we have previously described11, as well as other validated methods.
An important limitation of this protocol is that it is not adequate for the analysis of tissues that have been previously fixed with formaldehyde or another fixing agent. It is only adequate for tissues that have not been fixed and have been cryopreserved in OCT. In addition, this method is not adequate for processing tissues that weigh less than 1-2 mg as there is always some tissue loss during the sOCTrP steps and transfers between tubes. It will be difficult to weigh such small tissues using a standard analytical scale, and any left-over PBS not removed during the wicking stage will contribute greatly to the experimental error. Below are other important considerations when washing tissues to remove OCT with this method.
To ensure the smooth execution of this protocol, particularly if many samples will be processed, steps outlined in section 1 should be performed ahead of time, including pre-labeling and pre-weighing 1.5 mL tubes, pre-labeling 13 x 100 mm screw cap tubes and caps, 13 x 100 mm culture tubes, and autoinjector vials, preparing all wicks, pre-shortening 1 mL pipette tips, and pre-chilling wash solutions. This will prevent time-consuming procedures having to be performed on the day that tissues will be washed, which can lead to delays. In general, we find that if many of these steps are performed the day prior, an experienced researcher can wash 30-40 specimens in a normal workday. However, because the first suggested stopping point is after all tissues have been washed, weighed, and placed in methanol and stored at -80 °C, not performing 1.5 mL pre-weighing and labeling ahead of time will impact efficiency and extend the length of time required to wash 30-40 tissues.
Tissue weighing is one of the most important steps, as errors or carelessness will impact data quality as they will carry over into data normalization. It is critical that the analytical balance used for weighing be calibrated, properly zeroed, and tared. Accordingly, when using wicks, removing as much of the wash solution as possible is important, especially for tissues weighing less than 5 mg. At these tissue weights, leftover wash solution will make weighing inaccurate by a large percentage and carry over as an error in data normalization. When removing wash solution, we find that tissues can be gently touched with the wick for several seconds to remove as much wash solution as possible without leading to significant losses of tissue by adhesion to the wick. However, each tissue type being washed should be tested for adhesion to wicks, and these steps in the protocol adjusted accordingly.
To facilitate the tissue washing workflow, it is recommended that tissue shavings be requested from the biorepository in 15 mL polypropylene tubes. This will reduce the handling of tissues prior to washing.
When washing tissues, if a gel-like material is observed at the bottom of a tube after the third sOCTrP cycle, this is indicative that not all OCT has been removed and more sOCTrP cycles are needed. Keep track of how many cycles are needed, and for consistency, perform the same number of sOCTrP cycles in all specimens in the data set. We have previously evaluated up to 9 sOCTrP cycles and found no effect on depletion of sphingolipids in tissues11.
When using wicks to remove wash solution, gently dabbing the tissue will ensure that all wash solution is removed. This increases the accuracy of tissue weight estimation. However, great care must be taken with very small tissues as these can stick to the wick and result in the loss of specimen. It is helpful to use several wicks for the same tube to remove as much solution as possible. To prevent cross-sample contamination, always use a new, and clean, wick for each sample. When making wicks, use laboratory grade tissue (see Table of Materials) as we previously tested these and found to contain a negligible amount of contaminating sphingolipids11. Wicks from other materials can be used if they are tested as described11.
When retrieving specimens from 1.5 mL tubes after weighing, it is critical that all tissue be recovered. Failing to do so will result in errors in data normalization. If more PBS is needed to recover all the tissue, add the equivalent amount of PBS used to all other tubes to avoid differences in extraction efficiency that might result from samples containing different amounts of PBS. Adjust chloroform and methanol volumes to account for any increase in water volume to maintain a 2:1:0.1 methanol:chloroform:water ratio.
When thawing specimens, it is important that this step be done on ice to avoid the degradation of labile lipids such as sphingosine-1-phosphate. Fully thawing samples until the OCT is in a liquid state will also facilitate its removal as it will more easily be dissolved in the cold washing solution rather than remaining as a pellet during early wash steps.
Occasionally, especially when washing lung tissues, there will be fragments that will not pellet and will float on top of the wash solution after centrifugation. With care, immersing the aspirating tip beneath the floating tissues to remove the wash solution can prevent loss during aspiration.
Do not add chloroform to samples prior to homogenization as the disposable homogenizer tips are made of plastic and will dissolve in solutions containing chloroform. This can significantly impact the LC-MS/MS quantification of lipids in the samples. Therefore, in general, the use of plastics that are not resistant to solvents should be avoided when dispensing chloroform and methanol. These solvents are also toxic and should be handled in an appropriate fume hood. Wear appropriate PPE when handling solvent such as methanol and chloroform.
Human tissues should be processed in an appropriate biosafety cabinet (e.g., Class II). Users should be trained in the handling of biohazardous materials and fluids of human origin and follow safety protocols such as wearing appropriate personal protective equipment and decontaminating any surface or equipment that comes in contact with specimens. Users should thoroughly disinfect (see Table of Materials for suggested decontaminant) any surface or equipment (centrifuges, centrifuge carriers and adapters, pipettors, vortexers, fume hood surfaces, balances, glassware, ice trays, computer keyboards, computer mouse, etc.) used during the wash procedure. Lab members should not use equipment or hoods used for tissue washing until they have been thoroughly disinfected. Avoid the use of bleach on steel surfaces and electronics.
When processing many samples, use caution to prevent cross contamination using clean caps in re-capping steps during the lipid extraction steps. Caps can also be re-used if they are labeled. However, in practice, we find that markers stand out poorly on the black caps, and adhesive labels fall off tubes during the overnight 48 °C incubation.
Sphingolipids are demonstrated important players in the etiology of disease and human cancers. Therefore, there is great interest in precisely defining the sphingolipid metabolism alterations in these diseases as they may reveal therapeutic targets. However, many of the tissues that are easily accessible to researchers are cryopreserved in OCT, which is not compatible with mass spectrometry analysis. Thus, the detailed protocol described here will expand the repertoire of tissues adequate for quantitative sphingolipidomic analysis, and thus help increase our understanding of the biology of lipid metabolism alterations in human disease and cancer.
Offenlegungen
The authors have nothing to disclose.
Acknowledgements
Services and support of the research project were provided by the VCU Massey Cancer Center Tissue and Data Acquisition and Analysis Core and the VCU Lipidomics and Metabolomics Core, which are supported in part with funding from NIH-NCI Cancer Center Support Grant P30CA016059. This work was supported by National Institutes of Health Grants R21CA232234 (Santiago Lima).
Materials
| 1 mL polypropylene pipette tips | NA | NA | Used to retrieve specimens |
| 1.5 mL polypropylene centrifuge tubes | NA | NA | |
| 10 mL Erlenmeyer flask | VWR | 89091-116 | Used for tube and tissue weighing |
| AB Sciex Analyst 1.6.2 | Sciex | NA | Software to analyze and integrate MS data |
| Ammonium formate | Fisher Scientific | A11550 | For LC mobile phases |
| Analytical scale | NA | NA | Scale that is accurate to 0.1 mg |
| Bottle top dispenser | Sartorius | LH-723071 | Used for dispensing solvents |
| C12-Ceramide (d18:1/C12:0); N-(dodecanoyl)-sphing-4-enine | Avanti Polar Lipids | LM2212 | Internal standard |
| C12-glucosylceramide (d18:1/12:0); N-(dodecanoyl)-1-β-glucosyl-sphing-4-eine | Avanti Polar Lipids | LM2511 | Internal standard |
| C12-lactosylceramide (d18:1/12:0); N-(dodecanoyl)-1-ß-lactosyl-sphing-4-ene | Avanti Polar Lipids | LM2512 | Internal standard |
| C12-Sphingomyelin (d18:1/C12:0), N-(dodecanoyl)-sphing-4-enine-1-phosphocholine | Avanti Polar Lipids | LM2312 | Internal standard |
| CHLOROFORM OMNISOLV 4L | VWR | EM-CX1054-1 | |
| ClickSeal Biocontainment Lids | Thermo Scientific | 75007309 | To prevent biohazard aeresols during centrifugation |
| Conflikt | Decon Labs | 4101 | Decontaminant |
| CTO-20A/20AC Column Oven | Shimadzu | NA | For LC |
| d17:1-Sphingosine; (2S,3R,4E)-2-aminoheptadec-4-ene-1,3-diol | Avanti Polar Lipids | LM2000 | Internal standard |
| d17:1-Sphingosine-1-phosphate; heptadecasphing-4-enine-1-phosphate | Avanti Polar Lipids | LM2144 | Internal standard |
| DGU20A5R degasser | Shimadzu | NA | |
| Disposable Culture Tubes 13x100mm | VWR | 53283-800 | 13×100 mm screw top tubes |
| Heated water bath | NA | NA | For overnight lipid extraction |
| Homogenizer 150 | Fisher Scientific | 15-340-167 | triturate tissues |
| Homogenizer Plastic Disposable Generator Probe | Fisher Scientific | 15-340-177 | for homogenization |
| Kimwipes | Kimtech | 34120 | Laboratory grade tissue used to make wicks |
| Methanol LC-MS Grade 4L | VWR | EM-MX0486-1 | |
| Nexera LC-30 AD binary pump system | Shimadzu | NA | For LC-MS |
| Permanent marker | VWR | 52877-310 | |
| Phenolic Screw Thread Closure, Kimble Chase (caps for disposable culture tubes) | VWR | 89001-502 | 13×100 mm screw top tube caps |
| Phosphate bufffered saline | Thermo Scientific | 10010023 | To retrieve specimens from tubes after washing |
| Repeater pipette | Eppendorf | 4987000118 | To dispense LC-MS internal standards |
| Screw Caps, Blue, Red PTFE/White Silicone | VWR | 89239-020 | Autoinjector vial caps |
| Screw Thread Glass Vials with ID Patch | VWR | 46610-724 | Autoinjector vials |
| SIL-30AC autoinjector | Shimadzu | NA | |
| SpeedVac | Thermo Scientific | SPD2030P1220 | For drying solvents |
| Supelco 2.1 (i.d.) x 50 mm Ascentis Express C18 column | Sigma Aldrich | 53822-U | For LC-MS |
| Triple Quad 5500+ LC-MS/MS System | Sciex | NA | For LC-ESI-MS/MS |
| Ultrasonic water bath | Branson | Model 2800 | for homogenization and resuspension of extracted and dried lipids |
| Vortexer | NA | NA | For sOCTrP and resuspending dried lipids |
| VWR Culture Tubes Disposable Borosilicate Glass | VWR | 47729572 | 13×100 glass culture tubes |
| Water Hipersolve Chromanorm LC-MS | VWR | BDH83645.400 |
Referenzen
- Maceyka, M., Spiegel, S. Sphingolipid metabolites in inflammatory disease. Nature. 510, 58-67 (2014).
- Ogretmen, B. Sphingolipid metabolism in cancer signalling and therapy. Nature Reviews. Cancer. 18 (1), 33-50 (2018).
- Hannun, Y. A., Obeid, L. M. Sphingolipids and their metabolism in physiology and disease. Nature Reviews Molecular Cell Biology. 19 (3), 175-191 (2018).
- Hla, T., Dannenberg, A. J. Sphingolipid signaling in metabolic disorders. Cell Metabolism. 16 (4), 420-434 (2012).
- Lima, S., Milstien, S., Spiegel, S. Sphingosine and Sphingosine Kinase 1 involvement in endocytic membrane Trafficking. The Journal of Biological Chemistry. 292 (8), 3074-3088 (2017).
- Lima, S., et al. TP53 is required for BECN1- and ATG5-dependent cell death induced by sphingosine kinase 1 inhibition. Autophagy. , 1-50 (2018).
- Young, M. M., et al. Sphingosine Kinase 1 cooperates with autophagy to maintain endocytic membrane trafficking. Cell Reports. 17 (6), 1532-1545 (2016).
- Young, M. M., Wang, H. G. Sphingolipids as regulators of autophagy and endocytic trafficking. Advances in Cancer Research. 140, 27-60 (2018).
- Shen, H., et al. Coupling between endocytosis and sphingosine kinase 1 recruitment. Nature Cell Biology. 16 (7), 652-662 (2014).
- Morad, S. A. F., Cabot, M. C., Chalfant, C. E., Fisher, P. B. . Advances in Cancer Research. 140, 235-263 (2018).
- Rohrbach, T. D., et al. A simple method for sphingolipid analysis of tissues embedded in optimal cutting temperature compound. Journal of Lipid Research. 61 (6), 953-967 (2020).
- Weston, L. A., Hummon, A. B. Comparative LC-MS/MS analysis of optimal cutting temperature (OCT) compound removal for the study of mammalian proteomes. Analyst. 138 (21), 6380-6384 (2013).
- Holfeld, A., Valdés, A., Malmström, P. -. U., Segersten, U., Lind, S. B. Parallel proteomic workflow for mass spectrometric analysis of tissue samples preserved by different methods. Analytical Chemistry. 90 (9), 5841-5849 (2018).
- Zhang, W., Sakashita, S., Taylor, P., Tsao, M. S., Moran, M. F. Comprehensive proteome analysis of fresh frozen and optimal cutting temperature (OCT) embedded primary non-small cell lung carcinoma by LC-MS/MS. Methods. 81, 50-55 (2015).
- Shah, P., et al. Tissue proteomics using chemical immobilization and mass spectrometry. Analytical Biochemistry. 469, 27-33 (2015).

