Fluorescence Anisotropy to Determine Transcription Factor-DNA Binding Affinity
Abstract
Source: Jung, C. et al., High Sensitivity Measurement of Transcription Factor-DNA Binding Affinities by Competitive Titration Using Fluorescence Microscopy. J. Vis. Exp. (2019).
This video describes high-performance fluorescence anisotropy that helps to monitor the interaction between transcription factors and DNA. The assay monitors the interactions of a fluorophore-labeled DNA molecule with its specific transcription factor by measuring the degree of polarization due to molecular rotation or anisotropy of the fluorophore-labeled DNA.
Protocol
1. Polarization Microscopy
- For widefield laser illumination, focus a 638 nm line of a continuous diode laser (40 mW) on the aperture of multimode optical fiber for beam cleanup. Mount a linear polarizer at the output of the fiber to set the polarization of laser light.
- Block the excitation component of the emitted light with a dichroic mirror (640 nm cut-off) and a bandpass filter (bandpass 700/75).
- Let the fluorescence signal pass through a polarizing beam splitter, which splits the emitted light into its perpendicular and parallel polarized components. Then, focus the non-reflected beam (parallel component) and the reflected beam (perpendicular component) with an achromatic lens of 200 mm focal length on the chip of a back-illuminated EM-CCD camera (Figure 1b and 1c). Use a mirror to adjust the direction of the perpendicular beam toward the lens.
2. Design and Testing of Fluorescent-Labelled Reference DNA Oligomer
- Determine the core sequence of the reference DNA: the method is based on a competitive assay that measures the dissociation constant (KD2) between a transcription factor and unlabeled competitor DNA oligomer that competes for binding with a fluorescently labeled DNA whose affinity to the TF acts as a reference (KD1). The consensus sequence obtained from other sources like DNase footprinting or bacterial 1-hybrid can serve as a starting point.
NOTE: As a rule of thumb, a suitable reference DNA has a 3 to 7-fold decrease in binding affinity to the TF compared to the consensus sequence. - Measure by HiP-FA the KD1s of 2-3 tentative single mutations of the consensus sequence derived in the previous step. Try to mutate positions in consensus sequences that are not too specific to avoid complete loss of binding.
NOTE: It is important that the reference sequence is bound by the transcription factor of interest (we used in this protocol Giant Gt), but not too strongly, so that weaker competitors can outcompete it at high concentrations. - Extend the core motif (8-12 base pairs generally) to a length of 16 base pairs or more by adding a symmetrically flanking sequence at both sides (add side chains for proper binding). If necessary (for longer binding domains, for example), use longer sequences (up to ~50 base pairs in length were tested with the HiP-FA assay).
CAUTION: Be careful not to add bases that are expected to create ectopic binding sites. Use computational tools that predict binding sites from available PWMs to facilitate this process (e.g., PySite). - As labeled reference DNA, order oligomers that are fluorescently labeled on either forward or reverse strand at the 3' or 5' end. Use, for example, Cy5, Bodipy-650, or any other suitable dye at a concentration of 10 µM (100 µM 10x stock) in water, and dilute stepwise as described in step 3.1.
- Prepare 500 mL of 1x binding buffer by adding 33 mM potassium phosphate buffer (pH = 7.0), 90 mM NaCl, and 0.01% non-ionic detergent in distilled water. Also prepare 3x binding buffer, which contains the same components, except at threefold concentrations. If using 3x binding buffer as a stock solution for the 1x binding buffer, prepare volumes > 500 mL; otherwise, prepare 250 mL.
NOTE: This composition was optimized for transcription factor stability and to prevent glutathione S-transferase (GST) dimerization. - Measure with the microscopy setup described in step 1 the FA of 200 µL of binding buffer containing 0.8 nM labeled reference DNA in the presence of different amounts of TF in a glass bottom microscopy 96-well plate (5-6 wells with different TF concentrations) to determine the TF concentration to use. Perform a titration series with increasing amounts of TF and choose for the assay the concentration for which the curve reaches a plateau, indicating complete binding of the DNA reference oligomer.
NOTE: The optimal TF concentration depends on the values of the TF-DNA dissociation constants. Generally, lower KDs require lower concentrations.
3. Oligomer Annealing
- To anneal the DNA oligomers of the labeled reference DNA (sequence determined in the previous step), mix 7 µL of a 10 mM dye-labeled forward single-stranded DNA solution and 7 µL of a 10 mM concentration of its unlabeled reverse complement in 186 µL of water.
- For the competitor DNA sequences, mix 20 µL of 100 mM solutions (in water, provided by the manufacturer) of forward single-stranded DNA with 20 µL of 100 mM of the corresponding reverse single-stranded DNA for each individual competitor sequence to be measured.
- Perform the annealing separately in a standard PCR cycler by heating up the solutions to 70 °C for 3 min and decreasing the temperature to RT at a rate of 0.1 K/s. If the PCR machine used does not support temperature gradients at that rate, simply do stepwise incubations with decreasing temperatures (tested were 99 cycles of 3 s with -0.4 K per cycle).
4. Gel Preparation
NOTE: The following section explains the preparation of two different kinds of gels: 1) the titration wells contain gels with protein and are used to determine the KDs for the respective competitor DNA sequences, and 2) the calibration wells make use of NB to determine the DNA concentration at every given time point and acquisition height. The focus is on the preparation of the experiment in a 96-well plate, but the corresponding volumes for a 384-well plate format are also indicated.
- Dissolve 0.5% w/v low melting point agarose in the binding buffer by boiling it in a laboratory microwave oven. After complete dissolution, adjust the volume again with ddH2O to compensate for possible evaporation.
NOTE: For convenience, prepare a stock of 10-20 10 mL aliquots of the gels and melt them at 75°C when they are needed. Gel stocks can be stored at RT.
CAUTION: Be careful to avoid superheating the gel solution in the microwave oven. Short heating time intervals with shaking in between are preferable. - To prepare titration and calibration wells, first melt two 10 mL gel stock aliquots at 75 °C under shaking.
- Use 240 µL (including 20% overhead) for each competitor (n = number of competitor sequences).
- Use the same volume of gel for the NB calibration well to ensure an equal temperature and viscosity of both gels.
- Then set the temperature to 35 °C and wait for the temperature to equilibrate.
- For the titration wells, add 1.4 nM (final concentration) hybridized reference DNA (obtained in step 3), TF protein (final concentration CTF = 20-60 nM, as determined in step 2.6), DTT (0.2 mM), and binding buffer in a total volume of n x 200 µL or n x 13 µL in a 96- or 384-well plate format, respectively (plus overhead). Mix thoroughly by inverting/shaking (do not vortex).
- Slowly add 200 µL per well in 96-well plate format (13 µL/well for 386 wells) of the gel solution prepared in the previous step into the titration wells of the well plate.
- For the calibration wells, first, add 5 nM NB to the melted gel outside the wells (total volume depending on the well plate format used and on the number of calibration wells; usually 5-6 per well plate is enough).
- Pipette 200 µL (13 µL for 384-well plate format) of NB-containing gel slowly within the titration wells of the well plate and make sure to avoid air bubbles.
NOTE: The usage of electronic pipets or robotics significantly increases reproducibility. - Let the gel solidify for 10 min at RT, and another 10 min at 4 °C (remove condensation from the glass afterward if necessary). Make sure to conduct all these steps on a perfectly horizontal surface to avoid inhomogeneous gel surfaces.
NOTE: The protein containing gels are usually stable for at least several hours at 4 °C.
5. Adding the Competitor DNA Solution
NOTE: The following solutions should be prepared before starting the titration and are added on top of the calibration and titration wells simultaneously.
- Add the annealed labeled reference DNA and protein in 3x binding buffer at 3 times higher concentrations than the gel stock aliquots.
- Mix 20 µL of the obtained solution with 40 µL of each annealed competitor DNA solution obtained in step 3.
- For each calibration well, mix 20 µL of 3x binding buffer containing 15 mM NB solution with 40 µL of annealed competitor DNA (any sequence of the same length is suitable).
NOTE: For the 384-well plates, use 21 µL instead of 60 µL in total.
- Optionally, check the homogeneity of the gel height levels in the different wells of the plate spectroscopically by measuring the absorbance at 380 nm, using a multi-well plate reader (the absorbance values are proportional to the gel heights).
- Add 50 µL (7 µL for 384-well plate format) of the mixed competitor DNA solutions (annealed in step 3) on top of the gels. Try to add all the competitor solutions as simultaneously as possible by using electronic multichannel pipettes or a 96-channel pipetting head, if available. After the addition of the competitor solutions, place the plate on the microscope stage and start the measurements immediately.
6. Image Acquisition
- Sequentially acquire times series of z-stacks (e.g., use 12 planes and 100-300 ms of illumination time). Avoid taking images too close to the well surface (< ~1.4 µm with the plates used herein) to exclude any polarization bias.
- Perform 10-25 cycles of measurements until complete unbinding of the labeled reference DNA from the TF. The endpoint is typically reached after 1-2 h, depending on the binding kinetics and diffusivity of the competitor DNA.
Representative Results
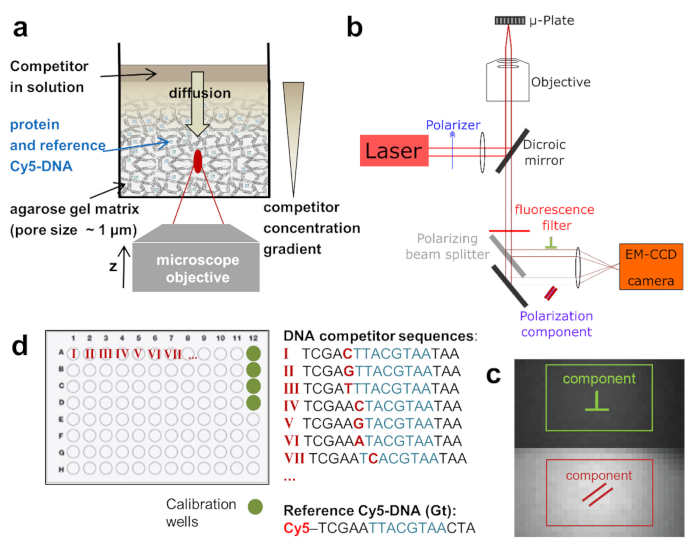
Figure 1: Schematic depictions of the HIP-FA assay and experimental setup. (a) Gel delivery system for titrating competitor DNA in single wells. (b) HIP-FA microscopy setup.Customized automated widefield microscope with polarized fluorescence light detection on an EM-CCD camera. (c) Raw fluorescence image with the two regions of interest used to determine the parallel (red) and the perpendicularly (green) polarized components. (d) Typical layout of a 96-well plate.
Divulgaciones
The authors have nothing to disclose.
Materials
| Cy5-labled 16- / 18-bp DNA-oligomers | Eurofins | Custom synthesis | |
| 16- / 18-bp DNA-oligomers | Eurofins | Custom synthesis | |
| Nile Blue A | Sigma | N5632-25G | |
| Sensoplate plus microplate 96- or 384-well, PS | Greiner | 655891 | |
| 384-Well Sensoplate, black | Greiner | 788896 | |
| Agarose, low gelling temperature | Sigma | A9414-50G | |
| Sodium Chloride | Merck | 1.06404.1000 | |
| Tween-20 | Sigma | P1379-1L | |
| Di-Potassium hydrogen phosphate trihydrate | Merck | 1.05099.1000 | |
| Potassium dihydrogen phosphate | Merck | 1.04873.1000 | |
| Q-POD Element | Merck Millipore | ZMQSP0DE1 | |
| Millipak 40 Gamma Gold Filter | Merck Millipore | MPGL04GK2 | |
| Milli-Q Integral 3 Water Purification System | Merck Millipore | ZRXQ003WW | |
| Quantum TIX | Merck Millipore | QTUMOTIX1 | |
| DL-Dithiothreitol | Sigma | 43815-1G | |
| Mastercycler gradient | Eppendorf | Z316083 | |
| SafeSeal tube 1.5 mL | Sarstedt | 72.706.200 | |
| Tube 15 mL | Sarstedt | 62.554.502 | |
| Multiply-Pro cup 0.2 mL PP | Sarstedt | 72.737.002 | |
| MICROSCOPY SETUP: | |||
| Automated widefield microscope | LEICA | DMI6000 | |
| Long distance objective | LEICA | HCX PL FLUOAR L 60x/0.60 N.A. Dry | |
| 638 nm line continuous diode laser | Omicron | PHOxX 638-40, 40mW | |
| Back-illuminated EM-CCD Camera | Andor | iXon DV897 | |
| Dichroic mirror | AHF | 640nm cut-off | |
| Bandpass filter | AHF | ET bandpass 700/75 | |
| Linear polarizer | Thorlabs | LPVISC050-MP2 | |
| Polarizing beam splitter | Thorlabs | BS010 | |
| Achromatic lens | Thorlabs | 200 mm focal length | |
| Multimode optical fiber | Optronis | FVP600660710 | |
| ROBOTIC SYSTEM: | |||
| Our robotic system includes a Biomek NXP workstation with a 96-channel head and Span-8 pipettors, connected with a servo-shuttle, which are used for all liquid transfer steps. In addition, the system is equipped with orbital shakers and a microplate reader (Paradigm, Molecular device) served by the Span-8 gripper | Beckman Coulter | Biomek NXP | |
| SOFTWARE: | |||
| Programming language | National Instruments | Labview 9.0 | |
| Script for the HiP-FA software available at | https://github.com/ GeneCenterMunich/HiP-FA |

