Polycarbonate Ultracentrifuge Tube Re-Use in Proteomic Analyses of Extracellular Vesicles
Summary
A detailed protocol is provided for cleaning and re-using polycarbonate ultracentrifuge tubes to perform extracellular vesicle isolation suitable for proteomics experiments.
Abstract
Single-use laboratory plastics exacerbate the pollution crisis and contribute to consumable costs. In extracellular vesicle (EV) isolation, polycarbonate ultracentrifuge (UC) tubes are used to endure the associated high centrifugal forces. EV proteomics is an advancing field and validated re-use protocols for these tubes are lacking. Re-using consumables for low-yield protein isolation protocols and downstream proteomics requires reagent compatibility with mass spectroscopy acquisitions, such as the absence of centrifuge tube-derived synthetic polymer contamination, and sufficient removal of residual proteins.
This protocol describes and validates a method for cleaning polycarbonate UC tubes for re-use in EV proteomics experiments. The cleaning process involves immediate submersion of UC tubes in H2O to prevent protein drying, washing in 0.1% sodium dodecyl sulfate (SDS) detergent, rinsing in hot tap water, demineralized water, and 70% ethanol. To validate the UC tube re-use protocol for downstream EV proteomics, used tubes were obtained following an experiment isolating EVs from cardiovascular tissue using differential UC and density gradient separation. Tubes were cleaned and the experimental process was repeated without EV samples comparing blank never-used UC tubes to cleaned UC tubes. The pseudo-EV pellets obtained from the isolation procedures were lysed and prepared for liquid chromatography-tandem mass spectrometry using a commercial protein sample preparation kit with modifications for low-abundance protein samples.
Following cleaning, the number of identified proteins was reduced by 98% in the pseudo-pellet versus the previous EV isolation sample from the same tube. Comparing a cleaned tube against a blank tube, both samples contained a very small number of proteins (≤20) with 86% similarity. The absence of polymer peaks in the chromatograms of the cleaned tubes was confirmed. Ultimately, the validation of a UC tube cleaning protocol suitable for the enrichment of EVs will reduce the waste produced by EV laboratories and lower the experimental costs.
Introduction
Extracellular vesicles (EVs) are lipid-bilayer-delimited particles released from cells that carry biologically active cargo, such as protein, and participate in various biological processes, including cell-cell communication and the formation of biologic mineralization1. These particles are found in all body fluids and tissues, and their biological activities and uses are a rapidly evolving field of scientific research. Isolation and validation of these nanoparticles present various challenges due to their small size and bio-similarity to other particles, such as liposomes and protein aggregates. The most recent International Society of Extracellular Vesicles guidelines, Minimal information for studies of extracellular vesicles 2018 (MISEV2018), are the recognized standard for EV scientific research2.
Various methods, often in tandem, must be used for EV isolation depending on the upstream source and the downstream applications. As of 2015, the most common primary isolation method for EVs was differential ultracentrifugation (UC)2. In principle, initial lower speed centrifugation separates larger or denser unwanted components, such as cells and cell debris, leaving EVs in the supernatant. Subsequently, UC uses very high centrifugal force to pellet out and thus separate and purify EVs from other particles that are smaller or less dense but which may also contain dense non-EV particles. Most protocols will often use, at one step or another, UC to isolate EVs from a fluid3. Further, UC is used in other methods of EV isolation, such as density gradient ultracentrifugation (DGUC), which uses mediums such as OptiprepTM iodixanol and centrifugal force to separate EVs according to their buoyant density4. Other methods of EV isolation exist3.
Considering the rapidly evolving understanding of the biological processes dictated by EVs and their potential as biomarkers and information regarding pathophysiology, discovery-based analyses such as proteomics have gained traction5,6,7,8. EVs are small, and, depending on the source, have a low yield of protein (<1 μg) when compared to whole tissue or cell lysate. Isolation of EVs for proteomics analysis requires special considerations, such as the removal of non-EV protein contaminants from upstream liquid or tissue, consideration of EV protein degradation during the isolation process, and the utilization of methods that create protein solutions that are chemically compatible with peptide preparation and mass spectrometry analyses.
Research laboratory consumables are often plastic and disposable in nature. These single-use materials contribute to the global plastic pollution crisis and consumable costs. Specialized polycarbonate and polystyrene UC tubes are designed to withstand the high centrifugal forces required to pellet EVs in laboratory applications. While it is possible to sterilize and disinfect UC tubes for re-use, proteomic analysis, especially those of low protein yields such as EVs, requires special attention. Not only is sufficient removal of residual protein from the previous use paramount, chemical compatibility with mass-spectroscopy and plastic-derived polymer contamination must also be considered.
Here we present a cleaning protocol of polycarbonate tubes using detergents suitable for mass-spectrometry and perform experiments to validate its successful removal of residual protein below limits of detection and lack of detectable polymer contaminants. To validate the cleaning protocol for EV proteomic applications using both UC and DGUC purposes, we obtained tubes from isolations of human vasculature tissue EVs with a combined UC and DGUC protocol. Tubes were cleaned using the protocol described, and the experimental process was repeated without samples comparing a blank never-used UC tube and a cleaned UC tube. Ultimately, the validation of a UC tube cleaning protocol suitable for the enrichment of EVs will reduce the waste produced by EV laboratories and lower the cost associated with such experiments.
Protocol
1. Tube cleaning
NOTE: The EV isolation procedure uses both capped and uncapped polycarbonate UC tubes (detailed below). The same procedure was followed for both capped and uncapped tubes. In the case of capped tubes, the lid parts were cleaned individually and reassembled post drying and pre-storage.
- Following the initial use of the polycarbonate tubes and removal of the sample, immediately submerge UC tubes in tap water to prevent drying of the sample to the side of the tube.
- Transfer and submerge tubes in 0.1% sodium dodecyl sulfate (SDS) in tap water for 10 min.
- One tube at a time, decant most, but not all, of the SDS solution from the tube and use a cotton swab to thoroughly wipe the area of the tube where the EV pellet was located.
NOTE: Do not use anything with bristles. This protocol used normal cotton swabs; however, specialized, low-lint cotton swabs can be used for cleaning the tubes. - Rinse at least 3x with hot tap water (~50 °C). Make sure to fill and decant the tube completely.
- Rinse 1x with demineralized water using a spray bottle or a narrow-mouthed wash bottle.
- Rinse 1x with 70% ethanol using a spray bottle.
- Leave the tube upside down to dry.
NOTE: Tube racks for fluorescence activated cell sorting work well for drying the UC tubes. - Place the completely dried tubes in a clean jar; they are ready for reuse.
NOTE: Currently, the protocol is validated for re-use up to 2x.
2. EV enrichment
NOTE: The following EV isolation and proteomics protocol was used for both the original tissue EV isolation, as well as the "mock samples" used to obtain the blank (never-used) and cleaned tube samples. The original samples were resuspended tissue-entrapped EVs from carotid plaques of patients who underwent carotid endarterectomies. Surgical specimens were collected from the University Health Network (Toronto, Canada) and was approved by the institutional ethics committee. All patients provided informed consent for sample collection and data analysis, in accordance with the Declaration of Helsinki. The mock samples were pseudo-pellets obtained from treating blank and cleaned tubes with the same solutions and processing steps as the EV isolation samples.
- Defrost EV samples or mock samples in PBS on ice.
- Make NTE buffer (10 mM Tris-HCl, 1 mM EDTA, 0.137 M NaCl pH 7.4) and sterile filter.
- Spin the samples at 10,000 × g for 10 min at 4 °C.
- Move the supernatant into a 1 mL open-top polycarbonate tube.
- Centrifuge the supernatant at 100,000× g for 1 h at 4 °C.
- Resuspend the pellet in 150 µL of sterile-filtered NTE buffer with protease inhibitor.
- Transfer to a 1.5 mL microcentrifuge tube and add NTE buffer to reach a total sample volume of 1.5 mL.
- Keep the samples on ice.
3. Preparation of density gradients and density gradient separation
NOTE: This EV isolation protocol has been previously described by Blaser et al.9 In brief:
- Prepare five solutions of Iodixanol Density Gradient Medium (10%, 15%, 20%, 25%, and 30%) with NTE buffer as a diluent.
- Layer the five density solutions sequentially in capped ultracentrifuge tube(s) with 30% at the bottom and 10% on the top.
- Secure the caps on the tube(s) and move gently to a stable horizontal position for 1 h at room temperature.
- After 1 h, move the tube(s) gently to a vertical position on ice for 30 min to cool the gradient.
- Add 1.5 mL of the sample or mock sample (NTE buffer) to the top of the prepared density gradient tube(s).
- Ultracentrifuge tube(s) at 250,000 × g for 40 min at 4 °C.
- Remove the top fractions of the density gradient (2.4 mL) to uncapped ultracentrifuge tube(s). Fill to 9 mL with sterile-filtered NTE buffer.
- Pellet the EVs by ultracentrifuging at 100,000 × g for 1 h at 4 °C.
NOTE: Circle the location of the visible EV pellet or expected location with a marker. This will allow you to clean the area of the pellet with the swab directly. - Aspirate the supernatant while moving the tube into an upside-down position for 3 min. Remove the remaining droplets with a cotton swab before turning the tube right side up.
- Resuspend the EV pellet in 12 µL of lyse buffer in the referenced kit.
4. Peptide preparation for proteomics
NOTE: Proteomics sample preparation was performed according to the kit protocol with in-house modifications for low-abundance protein samples. The modified protocol is as follows:
- Lyse, denature, reduce, and alkylate the samples by heating in lyse buffer at 95 °C for 10 min.
- Digest the samples using the Lys-C/trypsin digest solution for 2 h at 37 °C using 10 µL of lyse solution from the sample and 10 µL of the digest solution.
- Purify peptides through a cartridge by washing hydrophilic and hydrophobic contaminants out and eluting off purified peptides using kit solutions.
- Dry the purified peptides in a speed vacuum at 45 °C for 45 min or until dry.
NOTE: Avoid overdrying of the peptides. - Store samples at -80 °C until the time of mass spectrometry sequencing.
- Add 17 µL of LC-load solution to the dried peptides.
- Leave the samples on ice for 1 h.
- Vortex the samples briefly and then sonicate in an ultrasonic waterbath for 5 min.
- Centrifuge the LC-load samples at 16,000 × g for 1 min, aspirate 15 µL from the top, and separate into a new tube.
5. Liquid chromatography and tandem mass spectrometry sequencing
- Inject 2 µL of LC-load peptide solution onto a mass spectrometer.
- Fractionate the peptides using a dual-column setup.
- Heat the column at a constant temperature of 45 ˚C.
- Run the analytical gradient at 300 nL/min from 5% to 21% Solvent B (acetonitrile/0.1% formic acid) for 60 min, followed by 10 min of 21% to 30% Solvent B, and another 10 min of a 95%-5% jigsaw wash.
- Set the MS1 to 120 K resolution (maximum injection time, 25 ms), subject the top precursor ions (within a 3.0 s cycle time) to HCD isolation width 1.2 m/z, dynamic exclusion enabled (60 s), and set the MS2 resolution to 60 K (maximum injection time, auto; normalized collision energy, 24%, 26%, and 28%).
- Set the MS1 and MS2 acquisition range to 400 m/z-1,200 m/z and 120 m/z-1,200 m/z, respectively.
6. Peptide/protein identification
- Search the acquired peptide spectra with a proteomic search software using a commercial search algorithm against a human uniProt database.
- Set the digestion enzyme to trypsin and allow up to two missed cleavages.
- Set the precursor tolerance to 10 ppm and the fragment tolerance window to 0.02 Da.
- Set methionine oxidation and N-terminal acetylation as variable modifications and cysteine carbamidomethylation as fixed modification.
- Filter the peptides based on a false discovery rate (FDR) threshold of 1.0% as calculated by the Percolator algorithm.
- Only consider peptides assigned to one given protein group and not detected in any other protein group as unique and use them for further analyses.
- Only include a minimum of two unique peptides for each protein in the analyses.
- Maximize IDs with precursor ion detection.
- Perform chromatographic alignment with a maximum retention time shift of 10 min, mass tolerance of 10 ppm, and signal-to-noise minimum of 5.
- Normalize peptide abundance by total peptide amount.
Representative Results
To validate the cleaning protocol (Figure 1), two experiments were performed. First, the proteome of the "mock sample" from the cleaned tube was compared against the proteome of the tissue EV sample from the tube's initial use to determine the carryover of identified proteins. Representative chromatograms show a reduction in peak heterogeneity following cleaning of the tubes (Figure 2). In the original EV isolation, 806 proteins were identified with two or more unique peptides. Following cleaning, the number of identified proteins was reduced by 98% to 19 proteins. Twelve proteins were shared in both, and seven proteins were unique to the cleaned tube (Figure 3A). Using a literature review, a list of characteristic EV proteins was compiled as previously published9. Of the listed 24 characteristic EV proteins identified in the original sample, only one was identified after cleaning (Table 1). Of the shared proteins, MFGE8, a characteristic EV protein was the most differentially enriched protein in the EV sample. In comparison two contaminant proteins, KRT14 and KRT5, were the most differentially enriched in the post-cleaned sample (Figure 3B).
To confirm the removal of protein from the tubes after washing, an SDS-PAGE gel was performed with a BSA standard curve from 1 µg (typical low end of EV isolation yield) to 16.5 ng, and then blank and cleaned tube "mock" EV pellets in triplicate. All standard curve bands were visible however the samples were not, indicating cleaning removal of at least 98% of protein and no observable difference between the blank and cleaned tube (Figure 4A). To further assess the specific proteins identified in the cleaned tubes compared to a blank never-used tube, a direct comparison was conducted (N = 4, group). The chromatograms of the blank and cleaned tubes were very similar (Figure 4B). In the blank never-used tubes, 19 proteins were identified. Similarly, in the cleaned tube, 20 proteins were identified, with 18 shared within the two groups (Figure 4C). Ranking all the proteins by their abundance, contaminant keratin proteins KRT1, KRT9, and KRT10 are the most abundant in both groups (Figure 4D). In the cleaned tube, one characteristic EV protein was identified: lactadherin (MFGE8). There was high similarity between the clean versus blank tubes in the number of triggered MS2 sequencing events at each 5 min retention time bin (Figure 4E), the number of peptides sequenced, and the confidence in which they were sequenced (Figure 4F), as well as large overlap between the peptides (Figure 4G). Taken together, these results indicate no discernable impact of tube re-use with this cleaning protocol on the proteome.
The monomer of polycarbonate is 254.3 MH1+, which is below the MS1 acquisition range. Chromatograms were assessed for the presence of singly charged dimers and trimers, which were not detected (data not shown, N = 4).
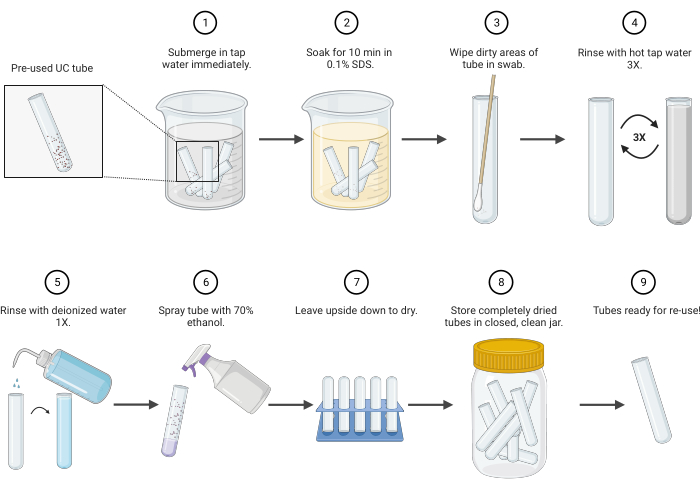
Figure 1: Polycarbonate ultracentrifuge tube cleaning protocol schematic. Abbreviations: UC = ultracentrifuge; SDS = sodium dodecyl sulfate. Please click here to view a larger version of this figure.
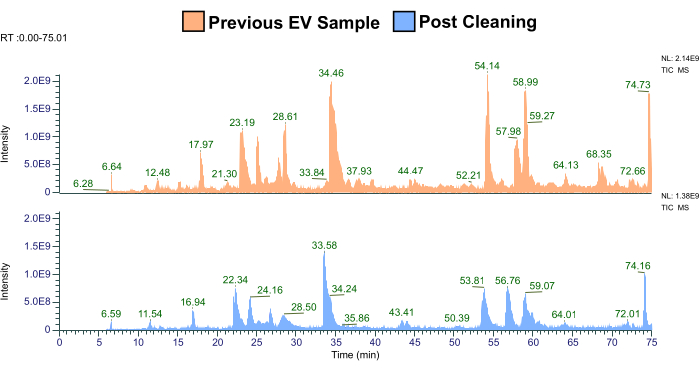
Figure 2: Representative comparison of the previous EV sample proteome to the "mock sample" from the cleaned tube (N = 2). Abbreviations: EV = extracellular vesicle; TIC-MS = total ion chromatography-mass spectrometry. Please click here to view a larger version of this figure.
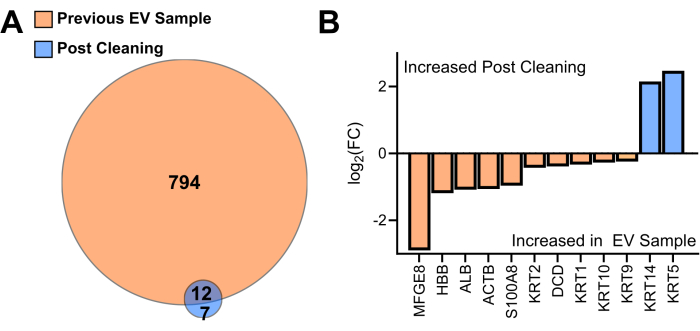
Figure 3: Comparison of proteins identified in the EV sample to the "mock" sample obtained from the cleaned tubes. (A) Venn diagram protein identifications in the previous EV sample proteome to the "mock sample" from the cleaned tube (N = 2). (B) Differential enrichment of shared proteins ranked by log 2-fold change. Abbreviation: EV = extracellular vesicle, FC = fold change Please click here to view a larger version of this figure.
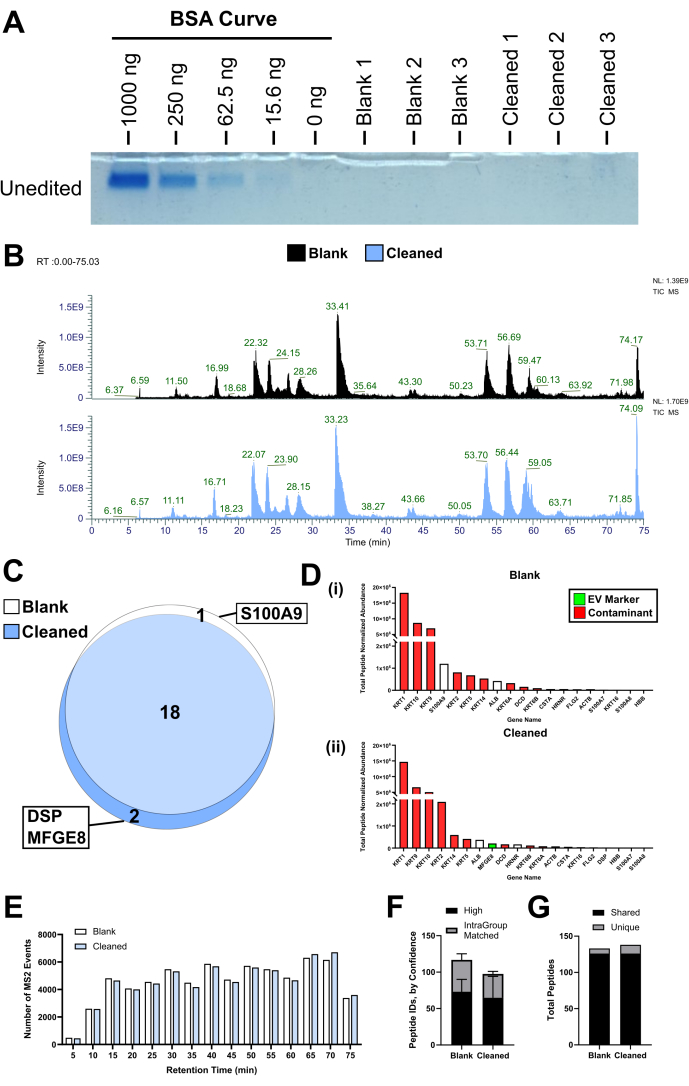
Figure 4: Comparison of blank never-used tubes to cleaned tubes. (A) An SDS-PAGE gel was performed with a BSA standard curve from 1 µg to 16.5 ng, and then blank and cleaned tube "mock" EV pellets in triplicate. (B) Representative chromatograms for blank and cleaned tubes. (C) Venn diagram of the shared or unique proteins from blank and cleaned tubes (N = 4). (D) All proteins identified in each group ranked according to abundance for (i) blank and (ii) cleaned tubes. Median values are plotted. (E) Histogram of all triggered MS2 events by retention time summed by group (N = 4). (F) Number of peptide IDs in each group by confidence, where high confidence peptides are distinct from intragroup matched peptides, where the identification confidence is lower but bolstered by matching to other spectra, mean/SD. (G) Number of unique and shared peptides between groups. Abbreviations: EV = extracellular vesicle; TIC-MS = total ion chromatography-mass spectrometry. Please click here to view a larger version of this figure.
Table 1: Characteristic EV proteins found in pre versus post UC tube cleaning analysis. Abbreviations: UC = ultracentrifuge; EV = extracellular vesicle. Please click here to download this Table.
Discussion
Here we describe and validate a protocol for cleaning polycarbonate UC tubes for EV enrichment and proteomic applications. We demonstrated the successful removal of residual protein from the previous UC tube sample compared with a cleaned pseudo-pellet analysis below the limit of detection of this mass spectrometry acquisition protocol and showed the proteomic similarity of blank never-used UC tube compared to cleaned UC tube pseudo pellets.
First, to prevent the inadvertent adsorption of protein to the inner polycarbonate wall, UC tubes were immediately submerged in H2O10. Hand washing the polycarbonate UC tubes is recommended by the manufacturers11. Other detergents, such as Lipsol, should be used with caution. It is a proprietary laboratory detergent composed of a mixture of detergent, surfactant, and chelating agents with a non-neutral pH. Therefore, for this step, a mass spectrometry-compatible detergent, 0.1% SDS, was used and washing was not performed for more than 10 min11. In our experience, the use of bristle cleaning tools can lead to tube scratching if caution is not taken. To mitigate potential scratches, which could provide a reservoir for residual protein, a cotton swab was used to wipe the inside of the polycarbonate tube, with a focus on the area of EV pellet formation. Repeated autoclaving can reduce the polycarbonate tube life11; instead, three brief hot water rinses were performed. It should also be noted that although the manufacturers suggest non-thermal sterilization methods, a specific temperature to be avoided is not described. Tubes should always be inspected for signs of deformation or cracking before re-use, as per manufacturers' recommendations. Flammable substances are not recommended to be used near operating centrifuges; thus, we dry the tubes overnight after the final 70% ethanol rinse step.
No significant polymer contamination was observed by manual assessment of chromatograms. SDS is not expected to degrade polycarbonate, and brief exposures to hot water (<60 °C) have no effect, while longer exposures do12. Further, previous studies have indicated that polycarbonate species elute at a retention time prior to collection based on our acetonitrile gradient13. While we cannot be certain that polymers are not present, with our acquisition methods, we are confident that the proteome is not substantively impacted.
When re-using labware for label-free proteomics experiments, residual protein contaminants from the previous use must be specifically considered. Further, EV proteomics are often conducted with very low-yielding samples, sometimes yielding protein amounts in the <100 ng range. These amounts of starting protein are suitable for mass-spectrometry but substantially increase the impact even small amounts of residual protein will have on the overall proteome. Given the sensitivity of advancing mass spectrometry sequencing, contaminants from human handling and manufacturing will be identified. In this protocol, we show a substantial reduction in the number of identified proteins by 98% following cleaning. The remaining proteins were almost exclusively shared between the blank never-used tubes and the clean tubes. These results suggest that the identified proteins were contaminants from the proteomics preparation itself or residual elements from tube manufacturing, not carryover from the previous use. By far, the most abundant three identified proteins by orders of magnitude in the blank and cleaned tubes were human keratin type I (KRT1, KRT10, KRT9), found in hair and skin, generally recognized in these types of preparations as contaminants from human handling (https://www.thegpm.org/crap/). Nine of the shared 18 proteins were other keratins. Also included as a recognized human contaminant is dermcidin (DCD), a protein found in human sweat14. Other identified proteins are also found in the epidermis (CSTA15, HRNR16, FLG217) or previously identified in dust mites (S100A7, S100A8, S100A9)18,19. It should be noted that a substantial component of "dust" is human skin20. The two unique proteins identified in the cleaned tubes have both been implicated in EVs, however, are very minor compared to other identified EV proteins1,21. The proteins identified in the samples are consistent with the preparation and indicate a very clear overlap between blank and cleaned tubes, supporting re-use.
EVs have an important role in biological processes, including but not limited to cell-cell communication and the formation of biologic mineralization. EVs are ubiquitous in the human body and are a rapidly growing area of basic science due to their potential as biomarkers, therapeutics, and sources of information regarding (patho)physiology. Enriching these particles for experimentation using differential UC involves specialized polycarbonate tubes. Single use and disposal of these thermoplastics contribute to higher experimental consumable costs and the global plastic pollution burden. Herein, we presented and validated a protocol for cleaning and re-using polycarbonate UC tubes to perform EV isolation suitable for proteomics experiments. Ultimately, this will reduce the waste produced by EV laboratories and lower the cost associated with such experiments.
Divulgaciones
The authors have nothing to disclose.
Acknowledgements
This study was supported by a research grant from National Institutes of Health grants (NIH) R01HL147095, R01HL141917, and R01HL136431, Kowa Company, Ltd., and the European Union's Horizon 2020 Research and Innovation Program under the Marie Skłodowska-Curie Grant Agreement No. 101023041 (R. Cahalane). Figure 1 was created with Biorender.com. The current cleaning protocol was developed by modifying a recommended tube cleaning protocol presented at the International Society of Extracellular Vesicles 2023 Education Day (https://www.youtube.com/watch?v=DOebcOes6iI). Many thanks to Dr. Kathryn Howe and Dr. Sneha Raju from the University Health Network (University of Toronto, Canada) for the original carotid tissue EV samples.
Materials
| 10 mL Open-Top Thickwall Polycarbonate Tube | Beckman Coulter Life Sciences | 355630 | uncapped ultracentrifuge tube(s) |
| 10.4 mL Polycarbonate Bottle with Cap Assembly | Beckman Coulter Life Sciences | 355603 | capped ultracentrifuge tube(s) |
| an Acclaim PepMap 100 C18 HPLC Columns, 75 µm x 70 mm; and an EASY-Spray HPLC Column, 75 µm x 250 mm | ThermoFisher Scientific | 164946 and ES902 | Dual column setup |
| Critical Swab Swab, Cotton Head | VWR | 89031-270 | cotton swab |
| Exploris 480 fronted with EASY-Spray Source, coupled to an Easy-nLC1200 HPLC pump. | ThermoFisher Scientific | BRE725533 | Mass spectrometer |
| Human UniProt database (101043 entries, updated January 2022) | NA | NA | Human database |
| MilliQ water | water | ||
| PreOmics iST kit | PreOmics | P.O.00027 | commercial protein sample preparation kit |
| Proteome Discoverer package (PD, Version 2.5) | ThermoFisher Scientific | NA | Proteomic search software |
| SEQUEST-HT search algorithm | NA | NA | Search algorithm |
| Sodium Dodecyl Sulfate (20%) | Boston BioProducts | BM-230 | detergent |
Referencias
- van Niel, G., D’Angelo, G., Raposo, G. Shedding light on the cell biology of extracellular vesicles. Nature Reviews Molecular Cell Biology. 19 (4), 213-228 (2018).
- Théry, C., et al. Minimal information for studies of extracellular vesicles 2018 (MISEV2018): a position statement of the International Society for Extracellular Vesicles and update of the MISEV2014 guidelines. Journal of Extracellular Vesicles. 7 (1), 1535750 (2018).
- Witwer, K. W., et al. Standardization of sample collection, isolation and analysis methods in extracellular vesicle research. Journal of Extracellular Vesicles. 2 (1), 20360 (2013).
- Konoshenko, M. Y., Lekchnov, E. A., Vlassov, A. V., Laktionov, P. P. Isolation of extracellular vesicles: general methodologies and latest trends. BioMed Research International. 2018, e854347 (2018).
- Kowal, J., et al. Proteomic comparison defines novel markers to characterize heterogeneous populations of extracellular vesicle subtypes. Proceedings of the National Academy of Sciences. 113 (8), E968-E977 (2016).
- Karimi, N., et al. Detailed analysis of the plasma extracellular vesicle proteome after separation from lipoproteins. Cellular and Molecular Life Sciences. 75 (15), 2873-2886 (2018).
- Lischnig, A., Bergqvist, M., Ochiya, T., Lässer, C. Quantitative proteomics identifies proteins enriched in large and small extracellular vesicles. Molecular & Cellular Proteomics. 21 (9), 100273 (2022).
- van Herwijnen, M. J. C., et al. Comprehensive proteomic analysis of human milk-derived extracellular vesicles unveils a novel functional proteome distinct from other milk components. Molecular & Cellular Proteomics. 15 (11), 3412-3423 (2016).
- Blaser, M. C., et al. Multiomics of tissue extracellular vesicles identifies unique modulators of atherosclerosis and calcific aortic valve stenosis. Circulation. 148 (8), 661-678 (2023).
- Kristensen, K., Henriksen, J. R., Andresen, T. L. Adsorption of cationic peptides to solid surfaces of glass and plastic. PLOS ONE. 10 (5), e0122419 (2015).
- Use and care of centrifuge tubes and bottles. Beckman Coulter Available from: https://www.beckman.com/supplies/tubes-and-bottles (2014)
- Wang, L., Zhang, J., Hou, S., Sun, H. A simple method for quantifying polycarbonate and polyethylene terephthalate microplastics in environmental samples by liquid chromatography-tandem mass spectrometry. Environmental Science & Technology Letters. 4 (12), 530-534 (2017).
- Schittek, B., et al. Dermcidin: a novel human antibiotic peptide secreted by sweat glands. Nature Immunology. 2 (12), 1133-1137 (2001).
- Blaydon, D. C., et al. Mutations in CSTA, encoding Cystatin A, underlie exfoliative ichthyosis and reveal a role for this protease inhibitor in cell-cell adhesion. American Journal of Human Genetics. 89 (4), 564-571 (2011).
- Henry, J., et al. Hornerin is a component of the epidermal cornified cell envelopes. FASEB Journal. 25 (5), 1567-1576 (2011).
- Bolling, M. C., et al. Generalized ichthyotic peeling skin syndrome due to FLG2 mutations. The Journal of Investigative Dermatology. 138 (8), 1881-1884 (2018).
- Jin, S., et al. DAMP molecules S100A9 and S100A8 activated by IL-17A and house-dust mites are increased in atopic dermatitis. Experimental Dermatology. 23 (12), 938-941 (2014).
- Gläser, R., et al. The antimicrobial protein psoriasin (S100A7) is upregulated in atopic dermatitis and after experimental skin barrier disruption. Journal of Investigative Dermatology. 129 (3), 641-649 (2009).
- Molhave, L., et al. House dust in seven Danish offices. Atmospheric Environment. 34, 4767-4779 (1999).
- Buschow, S. I., et al. MHC class II-associated proteins in B-cell exosomes and potential functional implications for exosome biogenesis. Immunology and Cell Biology. 88 (8), 851-856 (2010).

