Phthalic Acid Ester-Binding DNA Aptamer Selection, Characterization, and Application to an Electrochemical Aptasensor
Summary
A protocol for the in vitro selection and characterization of group-specific phthalic acid ester- binding DNA aptamers is presented. The application of the selected aptamer in an electrochemical aptasensor is also included.
Abstract
Phthalic acid esters (PAEs) areone of the major groups of persistent organic pollutants. The group-specific detection of PAEs is highly desired due to the rapid growing of congeners. DNA aptamers have been increasingly applied as recognition elements on biosensor platforms, but selecting aptamers toward highly hydrophobic small molecule targets, such as PAEs, is rarely reported. This work describes a bead-based method designed to select group-specific DNA aptamers to PAEs. The amino group functionalized dibutyl phthalate (DBP-NH2) as the anchor target was synthesized and immobilized on the epoxy-activated agarose beads, allowing the display of the phthalic ester group at the surface of the immobilization matrix, and therefore the selection of the group-specific binders. We determined the dissociation constants of the aptamer candidates by quantitative polymerization chain reaction coupled with magnetic separation. The relative affinities and selectivity of the aptamers to other PAEs were determined by the competitive assays, where the aptamer candidates were pre-bounded to the DBP-NH2 attached magnetic beads and released to the supernatant upon incubation with the tested PAEs or other potential interfering substances. The competitive assay was applied because it provided a facile affinity comparison among PAEs that had no functional groups for surface immobilization. Finally, we demonstrated the fabrication of an electrochemical aptasensor and used it for ultrasensitive and selective detection of bis(2-ethylhexyl) phthalate. This protocol provides insights for the aptamer discovery of other hydrophobic small molecules.
Introduction
Along with rapid economic development, acceleration of industrialization, and urban construction, environmental pollution is more severe than ever. Typical environmental pollutants include heavy metal ions, toxins, antibiotics, pesticides, endocrine disruptors, and persistent organic pollutants (POPs). Besides metal ions and toxins, other pollutants are small molecules that quite often consist of a variety of congeners. For example, the most toxic POPs include polychlorinated biphenyls (PCBs), polycyclic aromatic hydrocarbons (PAHs), polybrominated biphenyl ethers (PBDEs), polychlorinated dibenzo-p-dioxin (PCDDs), polychlorinated dibenzofuran (PCDFs), and phthalic acid esters (PAEs)1,2, which all consist of many congeners. Small molecule detection has been mainly performed by chromatography/mass spectrometry-based techniques due to their diversity of applications3,4,5,6. For on-site detections, antibody-based methods have recently been developed7,8,9. However, since these methods are highly specific for a certain congener, multiple tests must be performed. What is more serious is that the novel congeners grow so fast that their antibodies can't be generated in time. Therefore, the development of group-specific biosensors to monitor the total levels of all congeners in one test may provide an invaluable metric for assessing environmental pollution status.
Recently, nucleic acid aptamers have been widely applied as recognition elements in various biosensing platforms due to their capability of recognizing a wide variety of targets, from ions and small molecules to proteins and cells10,11,12. Aptamers are identified through an in vitro method called systematic evolution of ligands by exponential enrichment (SELEX)13,14. SELEX begins with the random synthetic single strand oligonucleotide library, which contains approximately 1014-1015 sequences. The size of the random library ensures the diversity of the RNA or DNA candidate structures. The typical SELEX process consists of multiple rounds of enrichment until the library is enriched in sequences with high affinity and specificity to the target. The final enriched pool is then sequenced, and the dissociation constants (Kd) and selectivity against potential interfering substances are determined by different techniques such as filter binding, affinity chromatography, surface plasmon resonance (SPR), etc.15
Due to the extremely poor water solubility and lack of functional groups for surface immobilization, the aptamer selection of POPs is theoretically difficult. Significant advances for SELEX have speeded up the discovery of aptamers. However, the selection of group-specific aptamers for POPs has not yet been reported. So far, only PCB-binding DNA aptamers with high specificity for a certain congener have been identified16. PAEs are mainly used in polyvinyl chloride materials, changing polyvinyl chloride from a hard plastic to an elastic plastic, thus acting as a plasticizer. Some PAEs have been identified as endocrine disruptors, can cause serious damage to liver and kidney function, reduce the motility of male sperm, and may result in abnormal sperm morphology and testicular cancer17. Neither the compound- nor group-specific PAE-binding aptamers have been reported.
The goal of this work is to provide a representative protocol for selecting group-specific DNA aptamers to highly hydrophobic small molecules such as PAEs, a representative group of POPs. We also demonstrate the application of the selected aptamer for environmental pollution detection. This protocol provides guidance and insights for the aptamer discovery of other hydrophobic small molecules.
Protocol
1. Library and Primer Design and Synthesis
- Design the initial library and primers.
Library (Pool0): 5'-TCCCACGCATTCTCCACATC-N40-CCTTTCTGTCCTTCCGTCAC-3'
Forward primer (FP): 5'-TCCCACGCATTCTCCACATC-3'
Phosphorylated reverse primer (PO4-RP): 5'-PO4-GTGACGGAAGGACAGAAAGG-3' - Synthesize Pool0, FP, and PO4-RP using standard phosphoramidite chemistry18,19,20, and purify all the DNAs by standard high-performance liquid chromatography (HPLC)21.
- Reconstitute Pool0, FP, and PO4-RP in nuclease-free grade water at 100 µM, aliquot, and store them at -20 or -80 °C for up to one year.
NOTE: It is strongly suggested to store Pool0, FP, and PO4-RP in 10-20 µL aliquots to avoid possible cross contamination.
2. Synthesize the Anchor Target and Its Immobilization onto Epoxy-Activated Agarose Bead
- Synthesize the amino group functionalized dibutyl phthalate (DBP-NH2) as the anchor target.
NOTE: The experimental details on the synthesis and characterization of DBP-NH2 have been described in the literature22. - Prepare the anchor target stock solutions in a medium appropriate for the target solubility. For this study, prepare 100 mM DBP-NH2 stock solution in dimethyl sulfoxide (DMSO) and store the solution at 4 or -20 °C for up to one year.
- Immobilize DBP-NH2 onto epoxy-activated agarose beads according to a modified protocol from the manufacturer.
- Weigh out 0.1 g freeze dried powder of epoxy-activated agarose beads and suspend it in distilled water. Wash the medium for 1 h by using 20 mL distilled water added in several aliquots. Wash the medium with 0.2 M Na2CO3 (pH 12).
NOTE: The medium swells immediately after adding the water into it. - Dissolve DBP-NH2 (46.7 mg, 0.15 mmol) in the 500 µL coupling buffer (0.2 M Na2CO3 buffer).
- Mix the coupling solution containing the ligand with the medium. Use a shaker at 5 rpm for 48 h at room temperature.
- Repeat washing the product three times, sequentially using 0.1 M acetate buffer (0.5 M NaCl, pH 4.5) and 0.2 M carbonate buffer (0.5 M NaCl, pH 12) in each wash cycle. Wash and suspend the product in water to make the final volume of 500 µL.
- Store the product at 4 °C prior to use.
- Confirm the success of the target immobilization by elemental analysis.
- Weigh out 0.1 g freeze dried powder of epoxy-activated agarose beads and suspend it in distilled water. Wash the medium for 1 h by using 20 mL distilled water added in several aliquots. Wash the medium with 0.2 M Na2CO3 (pH 12).
3. SELEX
- Prepare 500 mL of PAE binding buffer in ultra-pure water or nuclease-free grade water with 20 mM Tris·HCl, 100 mM NaCl, 2 mM MgCl2, 5 mM KCl, 1 mM CaCl2, 1% polyoxyethy-lene (20) sorbaitan monolaurate, and 0.03% non-ionic surface active agent (pH 7.9). Filter the buffer through a sterile 0.22 µm nitrocellulose filter and store it at a temperature higher than 20 °C for months.
NOTE: It is critical to maintain a good dispersion status of PAEs in PAE binding buffer. Multiple types of surfactants are needed to enhance the solubility of PAEs in aqueous solution. However, the number of surfactants should be kept to a minimum to avoid the formation of surfactant micelles that would encapsulate PAE molecules. Non-ionic, surface active agent is easy to precipitate at temperatures lower than 20 °C, and therefore the buffer should be stored at a temperature higher than 20 °C. - Dilute 10 µL of 100 µM Pool0(~1.0 nmol for 1014-1015 sequences) in 490 µL of PAE binding buffer. Heat the Pool0 aliquots (5 × 100 µL) in the thin-wall centrifuge tubes in the water bath set at 95 °C for 10 min. Place the tubes on ice for 5 min and subsequently incubate the tubes for 5 min at room temperature (~25 °C in this work).
- The first round of selection: incubation of Pool0 and DBP−NH2−coated medium.
- Wash 200 µL of DBP−NH2−coated medium three times with 500 µL of PAE binding buffer. Mix DBP−NH2−coated medium with Pool0 and incubate the mixture at room temperature for 1 h under mild shaking.
- Separate the DBP−NH2−coated medium bound with aptamers from the unbound DNAs by ultrafiltration usingan ultrafiltration tube with a molecular cut-off of 100 kDa. Wash the medium three times with 500 µL of PAE binding buffer and centrifuge at 9,168× g for 10 min at 4 °C.
- The first round of selection: aptamer elution from DBP−NH2−coated medium.
- Add 50 µL of PAE binding buffer to the washed medium and heat the mixture at 90 °C for 9 min under shaking.
- Collect the supernatant containing the eluted DNAs by ultrafiltration. Repeat this elution process three times to recover more bound DNAs.
- The first round of selection: PCR
- Small scale PCR
- Perform a small scale PCR23 (4 × 20 µL aliquots) using ~15-30 cycles. Set up a PCR reaction24 as follows: 6.5 µL of RNase-free water, 1 µL of 10 µM each primer (FP, PO4-RP, 0.01 nmol each), 1.5 µL eluted pool from section 3.4 (or RNase-free water as negative control), and 10 µL of premixed reaction mixture containing dNTPs, hot start polymerase, and 2 × PCR reaction buffer (20 mM tris·HCl, 100 mM KCl, 3 mM MgCl2, pH 8.3).
- Run PCR using a thermal cycler with the following settings: 1 cycle of 95 °C, 1 min; 30 cycles of [95 °C, 30 s; 56 °C, 30 s; 72 °C, 30 s]; 1 cycle of 72 °C, 2 min and hold at 4 °C.When the instrument is running the 20th s of the extension steps at the 15, 20, 25, and 30 cycles, press the pause key, open the instrument cover and take one tube out, then press the pause key again to resume PCR.
- Prepare 12% denatured polyacrylamide gel electrophoresis (PAGE)25,26. Mix 4.8 g of urea, 6 mL of ultra-pure water, 1 mL of 10 × tris/boric acid/ethylene diamine tetraacetic acid (TBE), 3 mL of 40% acrylamide, 10 µL of tetramethylethylenediamine (TEMED), and 100 µL 10% ammonium persulfate (APS), in that order. Stir well and wait for 1.5-2 h to ensure the gel completely solidifies.
- Analyze the results of the cycle optimization: combine 1 µL of each PCR product with 5 µL of RNA loading buffer (62.5% formamide, 0.4 M formaldehyde, 1.25×3-(N-morpholino) propansulfonic acid buffer, 0.02% xylene cyanol FF, 0.02% bromophenol blue), and 4 µL of RNase-free water; heat the mixture at 95 °C for 10 min, and quickly cool it to 0 °C on ice; incubate for 5 min at room temperature; load into 5 separate wells of the 12% denatured PAGE; run electrophoresis by a vertical electrophoresis system in 1 × TBE buffer at 170 V for 45 min.
- After electrophoresis, stain the gel with 3 µL of 1× nucleic acid gel stain in 30 mL of 1 × TBE buffer for 10 min, and take an image of the gel.
- Large scale PCR: select the appropriate number of cycles to produce the highest intensity band at the correct size marker without unwanted by-products. Perform a large scale PCR (20 × 100 µL aliquots) utilizing the conditions and appropriate number of cycles determined in section 3.5.1.
NOTE: The final amount of single-stranded library that can be obtained is proportional to the total volume of the PCR, not the concentration of template. Typically, to obtain 1 nmol of library, the single-stranded pool needs 2 mL PCR. PCR is very sensitive and easily contaminated by outside DNAs. Steps should be taken to reduce the chance of contamination, such as wearing gloves, using sterilized tips, not spitting in the tubes, and not opening the cap of the PCR tube prior to centrifugation.
- Small scale PCR
- The first round of selection: purification of PCR product by ethanol precipitation.
- Combine two tubes of PCR mixture into one tube (200 µL each). Repeat this for all tubes.
- Add 15 µL of sodium acetate (3 M, pH 5.2) solution, 4 µL of DNA/RNA precipitation carrier solution, and 2.5 times the volume of pre-cooled ethanol at -20 °C into each tube. Mix them evenly.
NOTE: DNA/RNA precipitation carrier is widely used in ethanol precipitation experiments to significantly improve the recovery percentage of DNA/RNA at low concentrations. It doesn't interfere with downstream applications such as PCR and sequencing. - Centrifuge the mixture at 20,627 × g for 20 min at 4 °C and carefully discard the supernatant and leave white precipitate.
- Add 400 µL of 70% pre-cooling ethanol solution at -20 °C into each tube to wash the precipitation. Centrifuge the mixture at 20,627 × g for 5 min at 4 °C, and carefully discard the supernatant and leave white precipitate. Repeat the wash step two more times.
- Open the tube cover; prick the sealing membrane; let ethanol volatilize at 40 °C on the heating block until the white precipitate becomes transparent. Store the precipitate at -20 °C.
NOTE: The purified PCR product can be stored at -20 °C.
- The first round of selection: single-stranded DNA generation (generation of enriched pool1) by performing the λ exonuclease reaction to digest the phosphorylated reverse strand.
- Small scale λ exonuclease reaction
- Add 100 µL of RNase-free water to one tube containing the purified PCR product from section 3.6. Vortex the tube to completely dissolve the precipitate in the tube.
- Prepare five centrifuge tubes and add 5 µL of the above solution, 11 µL of RNase – free water, and 2 µL of 10 × reaction buffer (670 mM glycine-KOH, 25 mM MgCl2 ,0.1% (v/v) Triton X-100 pH 9.4) into each tube.
- Add 2 µL of water or diluted λ exonuclease solution containing 2 U, 5 U, 8 U, and 10 U λ exonuclease into these tubes, respectively. Mix the solution well by gently pipetting. The 10 × reaction buffer is provided along with λ exonucleasefrom the provider.
- Incubate the tubes for 35 min at 37 °C, 15 min at 80 °C, and hold at 4 °C.
- Prepare the 12% native PAGE. Mix 6 mL of ultra-pure water, 1 mL of 10 × TBE, 3 mL of 40% acrylamide, 10 µL of TEMED, and 100 µL of 10% APS, in that order. Stir well and wait for 60-90 min to ensure the gel completely solidifies.
- Analyze the results of the reactions by combining 5 µL of each reaction with 1 µL of loading dye and 4 µL of RNase-free water, and loading these mixtures into 5 separate wells of 12% native PAGE (150 V for 45 min).
NOTE: A 20 bp double-stranded DNA ladder can also be loaded on the gel, but it may not be necessary since the distinct bands of PCR product and the generated single stranded library show on the gel.
NOTE: The λ exonuclease reaction has an excellent structural selectivity. The double-stranded PCR product (the anti-sense strand is labeled with a phosphate group at the 5' end) can be completely digested into a single-stranded library in the presence of λ exonuclease in a quite wide concentration range (Figure 4). The minimum amount of λ exonuclease that can completely generate single-stranded DNA in the small scale λ exonuclease reaction is also used for the large scale λ exonuclease reaction. A higher concentration of λ exonuclease may also be used, since λ exonuclease works well in a wide concentration range without losing its structure selectivity and product yield. The λ exonuclease reaction is inhibited by a high concentration of salt. The incomplete digestion of the PCR product usually implies that too much salt exists in the PCR product. The purification of PCR products by ethanol precipitation (section 3.6) is usually compatible with this step, while the PCR product purified by isopropanol precipitation quite often contains too much salt.
- Large scale λ exonuclease reaction: Choose the minimum amount of λ exonuclease that can completely generate single-stranded DNAs for the large scale λ exonuclease reaction. The reaction is proportionally scaled up according to this condition. For example: if 2 U is the minimum amount of λ exonuclease required, mix 38 µL of Rnase-free water, 10 µL of 10 x buffer, 50 µL of DNA pool1 and 2 µL of 10 U/µL λ exonuclease.
- Small scale λ exonuclease reaction
- The first round of selection: purification of the single-stranded Pool1 by ethanol precipitation.
- Follow the instructions of step 3.6.
- Determine the concentration of purified Pool1 by UV absorption at 260 nm.
- Reconstitute the 90% purified Pool1 in an appropriate volume of PAE binding buffer. If there is not enough DNA obtained for the next round of selection, repeat sections 3.5-3.8.2.
NOTE: The volume of PAE binding buffer usually varies from 200 to 500 µL according to the amount of library and the desired final concentration of library in the next round of SELEX.
- The second, third, fourth, and fifth rounds of SELEX.
- Conduct the second, third, fourth, and fifth rounds of SELEX subsequently following the same procedure described above (section 3.2 – 3.8), except that the ~300 pmol library was input to increase the selection stringency.
- To minimize the nonspecific absorption of DNAs on the medium, incubate the pool with unmodified medium on a rotary shaker for 30 min in the third and fourth round of selection prior to incubation with the DBP−NH2−coated medium.
NOTE: The SELEX process ended at the fifth round due to the serious crosslink between DNAs revealed by the large amount of high molecular weight product that didn't migrate into the gel.
4. High-Throughput Sequencing
- Prepare Pool4 from section 3.9.2 for sequencing by PCR amplification with compatible primers.
- Design and synthesize an indexed forward primer with 6 nt at the 5′ end to facilitate the sequence analysis.
Indexed forward primer (FP-sequencing): 5'-agtacgaTCCCACGCATTCTCCACATC-3' - Set up a 1,000 µLPCR reaction as follows: 380 µL of RNase-free water, 50 µL of each 10 µM primer (FP-sequencing, PO4-RP, 0.5 nmol each), 20 µL of eluted pool from section 3.9.2, and 500 µL of premixed reaction mixture containing dNTPs, hot start polymerase, and 2 × PCR reaction buffer. Aliquot the mixture into 10 PCR tubes. Use the same PCR conditions as described in section 3.5.1.2.
- Design and synthesize an indexed forward primer with 6 nt at the 5′ end to facilitate the sequence analysis.
- Purify the PCR product to meet the sequencing facility's requirements. For example, use the PAGE-gel DNA recovery kit according to the manufacturer's instructions.
- Send the purified PCR product (~2 µg) to a sequencing facility under the shipping conditions required by the sequencing facility. For example, ship the PCR product in a centrifuge tube with caps thoroughly sealed with ice packs to the sequencing facility.
NOTE: Received the results that ranked the sequences according to the copy number of each sequence to assess the enrichment of the sequenced pool.The top sequence was named DBP-1 (5'-CTTTCTGTCCTTCCGTCACATCCCACGCATTCTCCACAT-3'). Its affinity and selectivity were measured by the following protocol (section 5 and 6). Its application in an electrochemical bio-sensing was subsequently demonstrated in section 7.
5. Kd Determination of Selected Aptamer Candidates using Magnetic Bead-Based Quantitative PCR (qPCR)
- Synthesize DBP-1-qPCR and primers using standard phosphoramidite chemistry and purify it by standard HPLC.
Aptamer candidate (DBP-1-qPCR): 5'-ATACCAGCTTATTCAATTCTTTCTGTCCTTCCGTCACATC CCACGCATTCTCCACATAGATAGTAAGTGCAATCT-3'
Forward primer for qPCR (FP-qPCR): 5'-ATACCAGCTTATTCAATT-3'
Reverse primer for qPCR (RP-qPCR): 5'-AGATTGCACTTACTATCT-3'
NOTE: The primer regions of the tested sequence are different from those of pool0 used in section 3. The purpose of changing the primer regions is to test the affinity of the core sequence alone, excluding the primer regions. - Immobilize DBP-NH2 onto carboxylic acid-coated magnetic beads following the manufacturer's instructions.
- Wash the carboxylic acid-coated magnetic beads twice with 25 mM 2-(N-morpholino)ethanesulfonic acid (MES) (pH 5.0), using an equal volume of magnetic beads (100 µL) pipetted out of the vial, for 10 min with good mixing (end-over-end or similar).
- Immediately before use, prepare 50 mg/mL of 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide hydrochloride (EDC) solution with cold 25 mM MES (pH 5.0).
- Immediately before use, prepare 50 mg/mL of N-hydroxysuccinimide (NHS) solution with cold 25 mM MES (pH 5.0).
- Mix 100 µL of EDC solution and 100 µL of NHS solution with the washed beads and incubate for 30 min at room temperature with slow tilt rotation.
- Put the tube on a magnet for 4 min after incubation, discard the supernatant, and wash beads twice with 100 µL of cold 25 mM MES (pH 5.0).
- Incubate 6 µL of 100 mM DBP-NH2 stock solution and 25 mM MES (pH 5.0, 290 µL) with the activated beads under rotation for at least 30 min at room temperature or 2 h at 4 °C, with slow tilt rotation at 5 rpm.
- Put the tube on the magnet for 4 min after incubation and discard the supernatant. Wash the coated beads 4 times and resuspend beads with 200 µL of PAE binding buffer and store at 4 °C until use.
- Collect titration curve to determine Kd.
- Prepare a series of DBP-1 solutions (500 µL) at varied concentrations in PAE binding buffer from 1 pM to 300 nM.
- Add 10 µL of DBP-NH2-coated magnetic beads as described in section 5.2.7 to each DBP-1 solution. Incubate for 1 h at room temperature under rotation.
- Place the tubes on a magnet for 4 min and remove the supernatant. Wash the beads 4 times using 200 µL of PAE binding buffer.
- Add 60 µL of PAE binding buffer to each tube. Incubate the tubes at 95 °C for 15 min. Collect the supernatant containing the eluted DBP-1 by magnetic separation. Repeat this elution process to recover more bound DBP-1.
- Dilute the elute solution 100-fold and take 3 µL for real-time qPCR. Set up a qPCR reaction as follows: 3 µL of RNase-free water, 2 µL of 10 µM each primer (FP, PO4-RP, 0.01 nmol each), 3 µL of eluted and diluted DBP-1, and 10 µL of premixed reaction mixture containing dNTPs, polymerase, and 2x PCR reaction buffer.
- Determine the numbers of DBP-1 in each sample by running qPCR using a thermal cycler with the following settings: 1 cycle of 95 °C, 30 s; 30 cycles of [95 °C, 30 s; 45 °C, 30 s; 72 °C, 30 s]; 1 cycle of 72 °C, 30 s and hold at 4 °C.
- Plot the titration curve and determine Kd by nonlinear fitting of the curve assuming a 1:1 binding ratio27.
6. Relative Affinity and Specificity Test by Competitive Assays
- Add 10 µL of DBP-NH2-coated magnetic beads to DBP-1 solution (500 µL, 1 µM). Incubate for 1 h at room temperature under rotation. Place the tubes on a magnet for 4 min and remove the supernatant. Wash the beads 4 times using 200 µL of PAE binding buffer and resuspend it in 10 µL of PAE binding buffer.
- Add 10 µL of DBP-NH2-coated medium bound with DBP-1 to 110 µL of each 10 µM tested sample (DBP-NH2, DBP, DEHP, butyl benzyl phthalate(BBP), heavy metal ions, etc.) Incubate at room temperature for 1 h. Collect the supernatant by magnetic separation.
- Dilute the supernatant 100-fold and take 3 µL for qPCR quantification. Set up a qPCR reaction as follows: 3 µL of RNase-free water, 2 µL of 10 µM each primer (FP, PO4-RP, 0.01 nmol each), 3 µL of eluted and diluted DBP-1, and 10 µL of premixed reaction mixture containing dNTPs, polymerase, and 2 x PCR reaction buffer.
- Determine the numbers of DBP-1 in each sample by running qPCR using a thermal cycler with the following settings: 1 cycle of 95 °C, 30 s; 30 cycles of [95 °C, 30 s; 45 °C, 30 s; 72 °C, 30 s]; 1 cycle of 72 °C, 30 s and hold at 4 °C.
- Calculate the relative affinity by dividing the number of DBP-1 released from the beads in the presence of the sample by the number of DBP-1 released from the beads in the PAE binding buffer: Relative affinity = Nsample/Nbuffer.
7. Fabrication and Electrochemical Measurements of DEHP Electrochemical Biosensors
- Synthesize the thiolated core sequence (HS-DBP-1) and the signaling probe (DBP-1-C-Fc) using standard phosphoramidite chemistry and purify it by standard HPLC.
HS-DBP-1: 5'- HS-(CH2)6-CTTTCTGTCCTTCCGTCACATCCCACGCATTCTCCACAT -3'
DBP-1-C-Fc: 5'-GATGTGACGGAAGGTTTTTTTT-(CH2)6Fc -3'
NOTE: The rational design of the sensor probes was described in our previous pbulications18,22. - Reconstitute HS-DBP-1 and DBP-1-C-Fc in nuclease-free grade water at 100 µM, aliquot, and store them at -20 or -80 °C for up to one year.
- Polish the gold electrode carefully to a mirror-like surface with 1, 0.3, and 0.05 µm Al2O3 powder on a microcloth for 5 min and sonicate it in ultrapure water for 5 min to remove residual Al2O3. Then clean the electrode by electrochemical polishing with 35 successive cyclic voltammetry (CV) scans from 0.4 to +1.2 V (vs. Hg-Hg2SO4) in 0.5 M H2SO4 at 100 mV/s.
- Prepare a mixture of 0.5 µM HS-DBP-1 and 0.5 µM DBP-1-C-Fc in 100 µL 10 mM phosphate buffer containing 1 M NaCl, pH 7.4 (PBS). Heat the mixture in the thin-wall centrifuge tube in the water bath set at 95 °C for 10 min, and slowly cool the mixture to room temperature.
- Add 1 µL of tris-(2-carboxyethyl)-phosphine hydrochloride (TCEP, 10 mM, stock solution) into the mixture. Keep at room temperature for 1 h.
- Immerse the clean gold electrode in the above solution for 12 h, or overnight at room temperature.
- Rinse the electrode with PBS and immerse the electrode in 1 mM [S(CH2)2(OCH2CH2)6OCH3]2 (HS-EG6-OMe) in PBS for 1 h. Rinse the electrode thoroughly using PAE binding buffer and immerse the electrode in PAE binding buffer.
- Take a background square wave voltammetry (SWV) scan in target-free PAE binding buffer.
- Clean all the experimental equipment: electrolytic cells, gold working electrode, platinum counter electrode, and saturated calomel reference electrode (SCE).
- Activate the installed software; set the experimental parameters for SWV measurements.
NOTE: The following experimental parameters were used for the SWV experiments: potential range: from -0.2 to 0.7 V; modulation amplitude: 25 mV; pulse width: 5 ms; scan rate: 50 mV/s; step potential: 1 mV. - Connect the gold working electrode, platinum counter electrode, and SCE to a potentiostat, and put these three electrodes into an electrolytic cell containing PAE binding buffer.
- Run SWV measurement.
NOTE: Once the SWV measurement starts, a SWV curve will be displayed on the screen of the computer. The scan is repeated until the scans are constant and no further changes in peak height or shape are observed.
- Immerse the working electrode in the PAE binding buffer containing a certain concentration of DEHP (e.g., 10 pM) for 30 min at room temperature. Rinse the electrode thoroughly using PAE binding buffer and immerse the electrode in PAE binding buffer with the counter electrode and SCE. Collect the SWV curve under the same condition as described in section 7.8.
- Repeat step 7.9, except that the concentration of DEHP (e.g. 100 pM, 1 nM, 10 nM, 100 nM, 1 µM) was sequentially increased. Plot the titration curve.
- Specificity tests: Repeat steps 7.3-7.9, except that the PAE bind buffer containing DEHP is replaced by the PAE binding buffer containing the potential interfering substance at the desired concentration.
Representative Results
We designed and synthesized the amino group functionalized dibutyl phthalate (DBP-NH2) as the anchor target (Figure 1F). We then performed the DNA aptamer selection of PAEs using DBP-NH2 as the anchor target and following the classical target immobilization-based method (Figure 2). In each round, a pilot PCR was performed using the denatured PAGE to optimize the cycle number of PCR (Figure 3). The denatured PAGE, instead of native PAGE, is strongly suggested because we and other colleagues have found that the suspected high molecular weight by-products shown on the native gels at the higher cycle numbers are actually crosslinking complexes formed by sequences of the right size. Thus, the band intensity of the denatured PAGE can truly reflect the amount of the right product. The single stranded DNA generation is another critical step, and many methods for this step have been reported28. The λ exonuclease digestion method is the cheapest method. The success of the single-stranded DNA generation can be conveniently checked by native PAGE, where the PCR product is shown as single (first several rounds of SELEX) or multiple bands, while only one band is shown after the digestion (Figure 4).
The SELEX was stopped after the fifth round of the selection because of the significant quantity of DNA accumulated at the top of the PAGE, even after the denaturation treatment (wherein DNAs are heated at 95 °C for 10 min in 2 × TBE–containing 8.3 M urea), which suggested serious cross-linking between DNAs and also indicated that the sequences were enriched. Therefore, the pool4 was sent out for the high-throughput sequencing. The high-throughput result showed that the top 100 most frequently occurring sequences were highly conserved and were from one family. The top sequence DBP-1 displayed nanomolar affinity (Figure 5) and good group-specificity to PAE congeners (DBP, BBP, DEHP) (Figure 6) via the convenient qPCR assays, as described in the protocol section.
DBP-1 has been used to construct a strand displacement-based electrochemical aptasenor accordingly to our previously reported work29. The DEHP sensor can sensitively and selectively respond to DEHP (Figure 7). The common environmental pollutants such as heavy metal ions, antibiotics, and small molecules with similar functional groups showed very weak response to DBP-1.
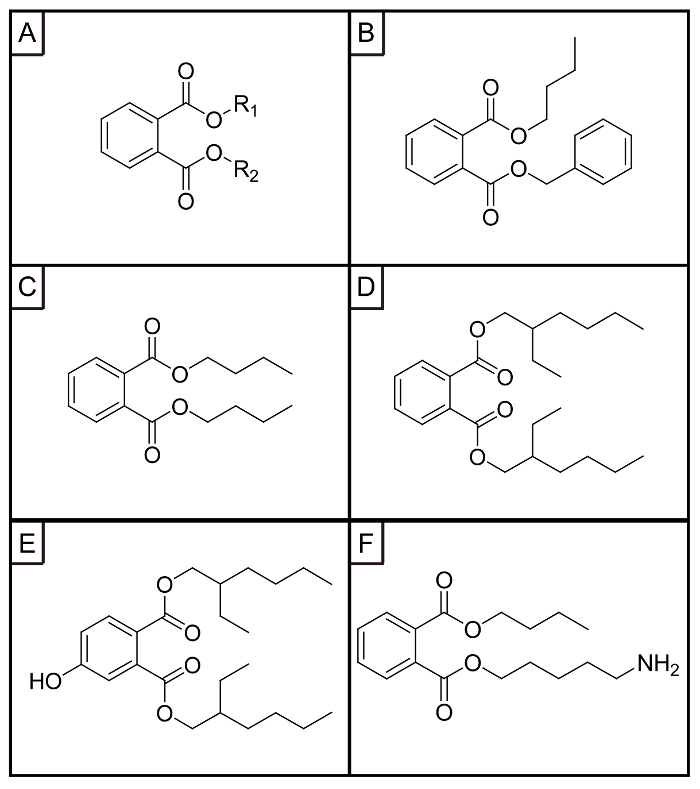
Figure 1: Chemical structures of (A) PAEs, (B) BBP, (C) DBP, (D) DEHP, (E) 4-OH-DEHP, (F) DBP−NH2. This figure has been reprinted with permission from Han Y. et al.22 Please click here to view a larger version of this figure.
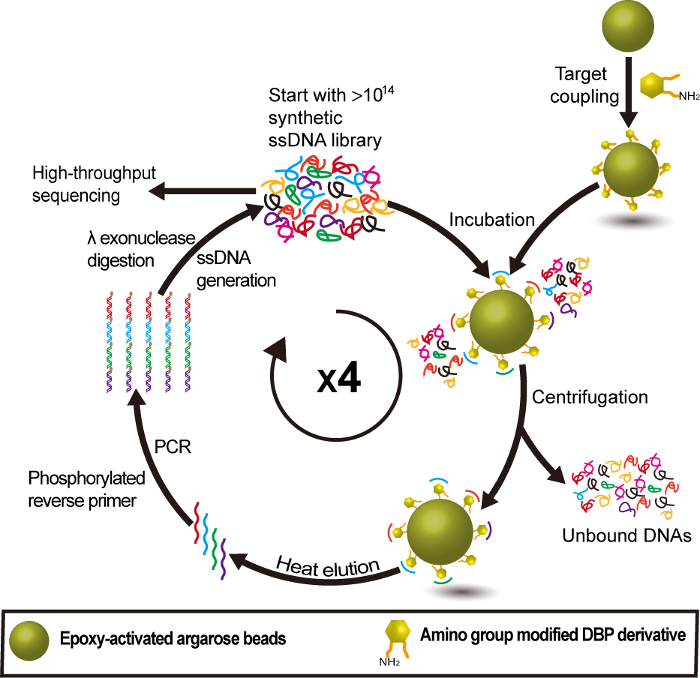
Figure 2: Group-specific DNA aptamer selection procedure for PAEs using DBP-NH2 as an anchor target. This figure has been reprinted with permission from Han Y. et al.22 Please click here to view a larger version of this figure.
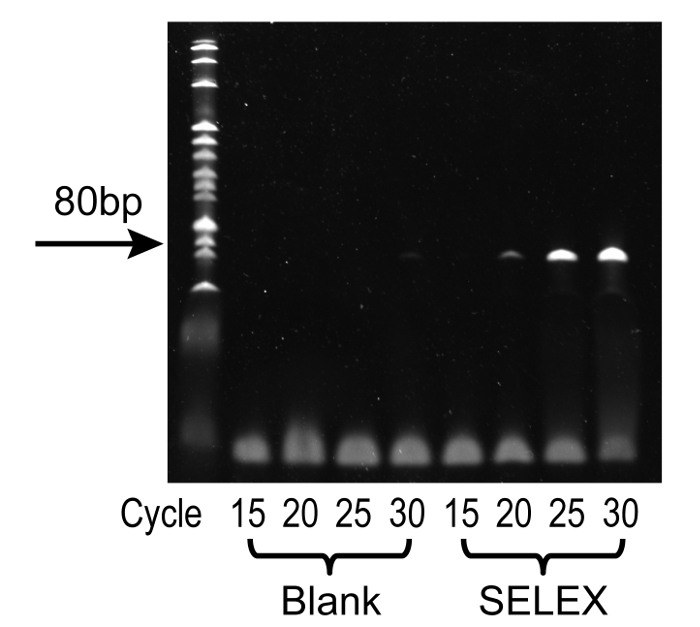
Figure 3: Representative results from PCR cycle optimization using denatured PAGE. From left to right, the lanes represent: 20 bp DNA ladder standard, 15 cycles (no DNA template), 20 cycles (no DNA template), 25 cycles (no DNA template), 30 cycles (no DNA template), 15 cycles, 20 cycles, 25 cycles, and 30 cycles. Optimal conditions were observed at 25 cycles, where the single product band was at high intensity without unwanted by-products, furthermore, no band was observed in the corresponding blank control. Please click here to view a larger version of this figure.
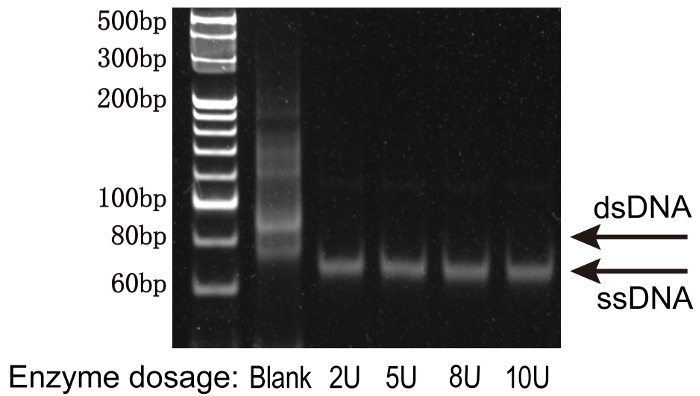
Figure 4: The optimization of λ exonuclease reaction to digest the phosphorylated strand using native PAGE. From left to right, the lanes represent: 20 bp DNA ladder standard, negative (no λ exonuclease), 2U, 5U, 8U, and 10U. The minimum amount of enzyme was observed at 2U, where the double PCR product becomes single-stranded DNAs. Please click here to view a larger version of this figure.
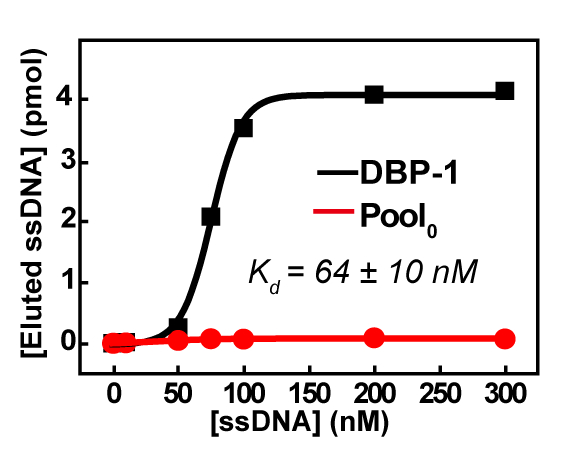
Figure 5: Affinity measurements of DBP-1 by qPCR-based assays. The error (standard deviation, SD) of the Kd 's was calculated from three measurements of the same sample. This figure has been reprinted with permission from Han Y. et al.22
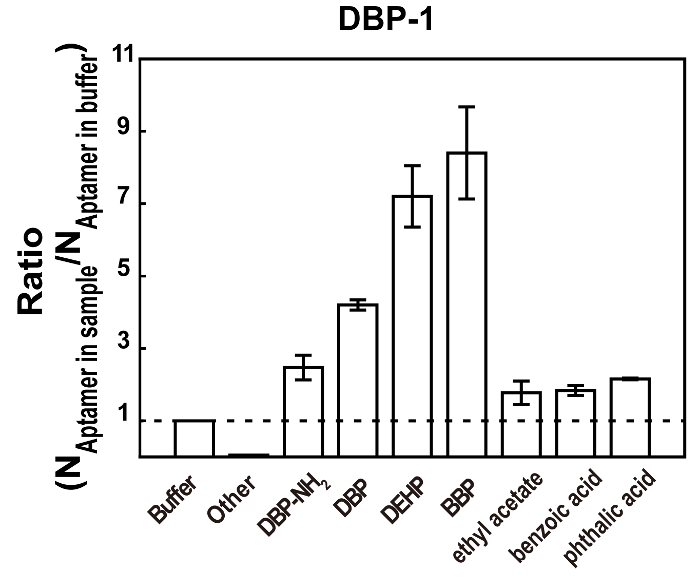
Figure 6: Relative affinity measurements of DBP-1 for free DBP-NH2 (10 µM), DBP (10 µM), DEHP (10 µM), BBP (10 µM), ethyl acetate (10 µM), benzoic acid (10 µM), phthalic acid (10 µM), and other via competition assays. Other: a mixture of potential interferences (glucose, kanamycin, ampicillin, and ethanol) all at 10 µM. The ratio (the relative affinity) was calculated by dividing the number of aptamers released from the beads in the presence of the sample by the number of aptamers released from the beads in the PAE binding buffer. The bars represent mean ± SD. The SDs were calculated from three individual measurements. Please click here to view a larger version of this figure.
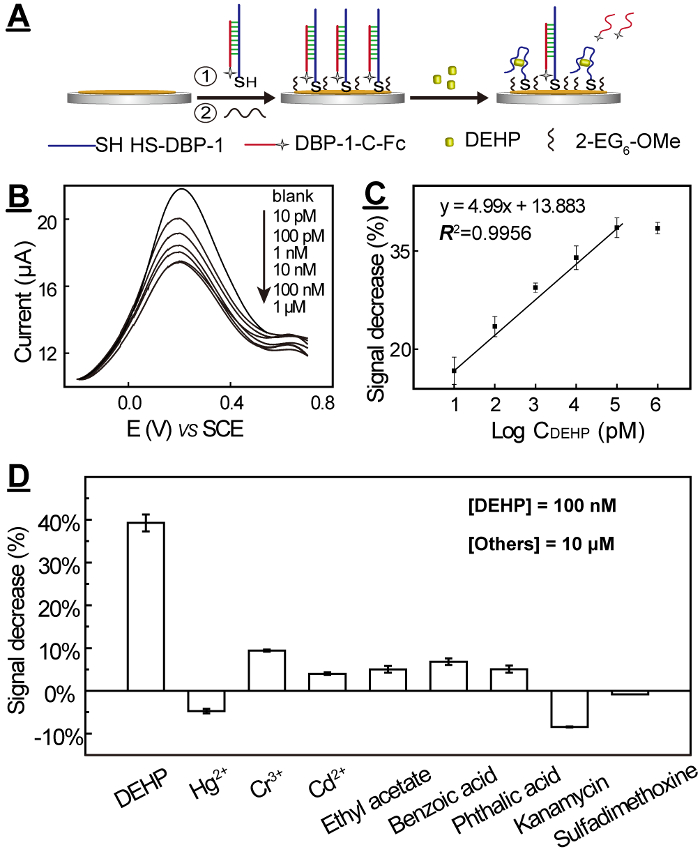
Figure 7: DEHP electrochemical biosensor for ultrasensitive and ultraselective detection of DEHP: mechanism (A), SWV curves (B), calibration curve (C), and selectivity tests (D). The errors (SDs) were calculated from three individual measurements. This figure has been reprinted with permission from Han Y. et al.22 Please click here to view a larger version of this figure.
Discussion
One outstanding benefit of aptamers is that they are identified through the in vitro method SELEX, while antibodies are generated via in vivo immunoreactions. Therefore, aptamers can be selected with desired target specificity under well-designed experimental conditions, whereas antibodies are limited to physiological conditions.
To facilitate the separation of bound sequences from free sequences, several modified SELEX have recently been reported, in which capillary electrophoresis30, microfluidics31, magnetic / acrylic beads / agarose beads14, etc., have replaced nitrocellulose filters or affinity columns to achieve more efficient separation. Among these techniques, the bead-based methods have been most widely used for the aptamer selection of small molecule targets due to their simple set-up, easy operation, and being amenable to small molecule analyses.
There are two groups of methods that are commonly used for aptamer selection of small molecule targets: the target13 and library14 immobilization-based methods. In the former group of methods, the small molecule targets are immobilized on the solid phase such as functionalized magnetic beads or agarose beads via covalent coupling reactions. Libraries are incubated with small-molecule coated beads, and those sequences that do not bind or bind weakly to the target on the solid phase are removed simply by performing washing and centrifugation or magnetic separation steps. The bound sequences are subsequently eluted and amplified by PCR. The small molecule targets must have at least one functional group available for the coupling reaction. For those without suitable functional groups, the functional group has to be incorporated into the original target through organic synthesis. The synthesis of the carefully designed target could involve multiple steps, and sometimes is quite challenging as well. The structural modification of the targets could also strongly affect the binding sites and affinity with their aptamers. To avoid these problems, library immobilization-based methods have been developed, where the library is hybridized to complementary DNA capture probes on magnetic beads, and the binding sequences fall off the beads upon binding with the targets. In this group of methods, the targets are added in the binding buffer and no functional groups are required. PAEs are the ester derivatives of phthalate acid, and a typical PAE consists of a phthalate acid ester group and one or two alkyl chains (Figure 1A-D). PAEs have no functional groups available for solid phase immobilization. Thus, the library immobilization-based methods seem more attractive for the aptamer selection of PAEs.
A well-controlled dispersion status is critical to ensure the desired enrichment of the library via SELEX. However, PAEs are extremely hydrophobic and easily form aggregates in aqueous solutions at a concentration higher than several µM, even in the presence of multiple surfactants to facilitate dispersion. Thus, the dispersion status of PAEs is hard to control, and it would be difficult to select aptamers that specifically bind to individual PAE molecules. To solve the solubility problem, target-immobilization methods were chosen, in which PAEs were immobilized on the hydrophilic beads via covalent bonding. By using this immobilization strategy, the targets should be dominantly present on the bead surface in a single molecular state, instead of aggregates.
The structural design of the anchor target is also quite critical for the successful selection of group-specific aptamers. Three factors need to be considered for the structural design. The first factor is the purpose of the aptamer selection. Considering the purpose of selecting group-specific aptamers, it is essential to expose the common group of PAEs for aptamer binding and prevent the rest of the parts from participating in the aptamer binding. Thus, the functional group should be introduced on the terminal of the side chain. At the beginning, we designed OH-functionalized DEHP (4-OH−DEHP) (Figure 1E) as the anchor target and immobilized it on the carboxylic acid magnetic beads16. Thus, the alkyl chains were exposed on the surface of the matrix. We tried to choose magnetic beads with carboxyl groups as the solid matrix. No obvious affinity improvement was observed after five rounds of selection, and the nonspecific absorption onthe bare matrix was kept strong.
Therefore, DBP-NH2 was later designed based on the following thoughts: (1)The phthalate group is more likely to interact specifically with the aptamer through the π-π stack and the hydrogen bond than the alkyl chain; (2) NHS-mediated carbodiimide reaction is mild and has a very efficient coupling efficiency, so -NH2 is introduced at the end of an alkyl chain of DBP; (3) It is easy to operate with the magnetic beads as a solid phase partition, but the nonspecific adsorption of the library is strong. Although the loss of the agarose beads during separation is higher than the magnetic beads, the nonspecific adsorption of the library is much lower. Thus, DBP-NH2 was immobilized on the epoxy-activated agarose beads, and the phthalic acidester group was exposed on the surface of the matrix.
The choice of selection buffer is important, especially for small molecule targets with diverse solubility. The binding buffer needs to be carefully prepared to avoid aggregation and ensure a good dispersion state of POPs during the whole aptamer selection and characterization process. In our study, we found that DEHP can't dissolve in regular buffers without surfactants, showing two distinct layers. The layer disappears upon addition of multiple surfactants in the optimized quantity. The clear solutions need to be stored at temperatures higher than 20 °C to maintain its status. Details are provided in protocol step 3.1.
Single-stranded DNA generation from double-stranded PCR products is one critical step in the SELEX process. Several different methods have currently been described in the literature32,33, including asymmetric PCR, λ exonuclease digestion, magnetic separation with streptavidin-coated beads, and size separation by denaturing urea-polyacrylamide gel.
Different methods have their own strengths and weaknesses. Currently, the most commonly used method for the generation of ssDNA is magnetic separation with streptavidin-coated beads. The outstanding advantage of this method is its time-saving and simple operation. The drawback is its high cost compared with other methods. In contrast, the size separation-based method, using reverse primers with GC-rich stem–loop structure, is one of the cheapest methods, while the yield of single-stranded DNA is the lowest among these methods32. In this protocol, we described the use of λ exonuclease digestion to generate single-stranded DNA, which is one of the cheapest methods. We found that the yield of single-stranded DNA is comparable with the two methods of magnetic separation and streptavidin-coated beads. Furthermore, we found that the exonuclease reaction was inhibited by the high concentration of salt. The incomplete digestion of the PCR product reported in the literature33 was likely due to too much salt existing in the PCR product. In addition, the λ exonuclease is highly active and inexpensive (Figure 4).
The characterization and validation of aptamers is laborious, time-consuming, and embodies a major bottleneck in the aptamer discovery pipeline33. Most techniques are mass-sensitive methods which work well for the larger aptamer binding partners (>10,000 amu), but are not sufficiently sensitive to measure the interactions with low molecular weight targets (<1,000 amu)15. The characterization and validation of aptamers of hydrophobic small molecules like PAEs is even more difficult. Their poor water solubility results in the unsaturation of the titration curve, which prevents the Kd's determination in solution or immobilizes aptamers on surface. Therefore, we determined the Kds of the identified aptamers by high throughput sequencing by immobilizing DBP-NH2 on the hydrophilic magnetic beads to avoid the solubility problem. The relative affinities and selectivity of the aptamers to other PAEs were then determined by the competitive assays, where the aptamer candidates were pre-bounded to the DBP-NH2 attached magnetic beads and released to the supernatant upon incubation with the tested PAEs or other potentially interfering substances. The competitive assay was applied because it provided a facile affinity comparison among PAEs that had no functional groups for surface immobilization. In addition, magnetic bead-based fluorescent assays are suitable for the affinity study of small molecule-aptamer interactions34. However, we found that the magnetic beads sometimes cause fluorescence quenching for unknown reasons. Thus, the qPCR assays were used for the affinity measurements.
One critical technique tip for the electrochemical biosensor described in this study is the surface passivation of the electrode35. Due to the high hydrophobicity of DEHP, it has a strong tendency to be nonspecifically absorbed onto the gold electrode, leading to the failure of the detection. The most commonly used surface passivation agent, 6-mercapto-1-hexanol (MCH)36,37, is not sufficient to prevent the nonspecific absorption of DEHP, while we found that HS-(CH2)2-[OCH2CH2]6-OCH3 was effective enough to enable the sensitive detection of DEHP38.
This procedure describes a protocol for selecting group-specific DNA aptamers of highly hydrophobic small molecules and an application of the selected aptamer in an electrochemical biosensor. The protocol helps with the selection of other hydrophobic small molecules and provides insights on sensor development of highly hydrophobic small molecules as well. The aptamer selection process belongs to the category of target immobilization-based methods. The limitations of this type of method also exist for this protocol, for example the need for complicated synthesis of anchor targets and the impacts of the solid phase on aptamer binding. The attractive advantages of the electrochemical biosensors described in this protocol include their simple design and high sensitivity. The major drawback is their limited precision due to their extremely broad dynamic range. Therefore, the biosensors described here are more suitable for screening tests, instead of quantitative measurements of the targets.
Divulgaciones
The authors have nothing to disclose.
Acknowledgements
We are grateful for financial support from the National Natural Science Foundation (21675112), Key project of science and technology plan of Beijing Education Commission (KZ201710028027) and Yanjing Young Scholar Program of Capital Normal University.
Materials
| UV-2550 | Shimadzu,Japan | protocol, section 3.8.2 | |
| DNA Engnine Thermal cycler,PTC0200 | BIO-RAD | section 3.5.1.2 and 3.5.2 | |
| C1000 Touch | BIO-RAD | section 5.3.6 and 6.3 | |
| VMP3 multichannel potentiostat | Bio-Logic Science, Claix, France | section 7.4,7.8 and 7.11 | |
| Epoxy-activated Sepharose 6B | GE Healthcare (Piscataway, NJ, USA) | 10220020 | argarose beads, section 2.3 and 3.3 |
| Dynabeads M-270 carboxylic acid magnetic beads | Invitrogen, USA | 420420 | magnetic beads,section 5.2. and 5.3 |
| Premix Taq Hot Start Version | Takara,Dalian,China | R028A | polymerase, section 3.5.1.1 |
| PARAFILM Sealing Membrane | Bemis, USA | PM-996 | section 3.6.5 |
| Lambda Exonuclease | Invitrogen, USA | EN0561 | section3.7.1.2.The 10 × reaction buffer is provided along with λ exonuclease by the provider. |
| Dr. GenTLE Precipitation Carrier |
Takara,Dalian,China | 9094 | section 3.6.2 and 3.8.1 |
| UNIQ-10 PAGE DNA recovery kit | Sangon Biotech (Shanghai) | B511135 | section 4.2 |
| SYBR Gold nucleic acid gel stain | Invitrogen, USA | 1811838 | nucelic acid stain dye, section 3.5.1.5 |
| SYBR Premix Ex Taq II | Takara,Dalian,China | RR820A | polymerase mix contaning polymerase and dNTPs, section 5.3.5 |
| 2-(N-Morpholino)ethanesulfonic acid (MES) | Sigma-Aldrich | CAS: 1132-61-2 | section 5.2.1 |
| 1-ethyl-3-(3-dimethylaminopropyl) carbodiimide hydrochloride (EDC) | Invitrogen, USA | CAS: 25952-53-8 | section 5.2.2 |
| N-hydroxysuccinimide (NHS) | Sigma-Aldrich | 6066-82-6 | section 5.2.3 |
| mercaptohexanol (MCH) | Sigma-Aldrich | CAS: 1633-78-9 | section 7.7 |
| Gold electrode | Shanghai Chenhua | CHI101 | section 7.4. – 7.11 |
| tris(2-carboxyethyl) phosphine hydrochloride (TCEP) | Sigma-Aldrich | CAS: 51805-45-9 | section 7.5 |
| O-(2-Mercaptoethyl)-O'-methyl-hexa-(ethylene glycol) | Sigma-Aldrich | CAS: 651042-82-9 | section 7.7 |
| diethylhexyl phthalate (DEHP) | National Institute of Metrology, China | CAS: 117-81-7 | section 7.11 |
| Tween 20 | Sigma-Aldrich | CAS: 9005-64-5 | polyoxyethy-lene(20) sorbaitan monolaurate |
| Triton X-100 | Sigma-Aldrich | CAS: 9002-93-1 | non-ionic surface active agent |
| PBS | Sigma-Aldrich | P5368 | 10 mM phosphate buffer containing 1 M NaCl, pH 7.4 |
Referencias
- Vorkamp, K., Riget, F. F. A review of new and current-use contaminants in the Arctic environment: Evidence of long-range transport and indications of bioaccumulation. Chemosphere. 111, 379-395 (2014).
- Net, S., Sempere, R., Delmont, A., Paluselli, A., Ouddane, B. Occurrence, fate, behavior and ecotoxicological state of phthalates in different environmental matrices. Environ Sci Technol. 49 (7), 4019-4035 (2015).
- Xie, Q. L., Liu, S. H., Fan, Y. Y., Sun, J. Z., Zhang, X. K. Determination of phthalate esters in edible oils by use of QuEChERS coupled with ionic-liquid-based dispersive liquid-liquid microextraction before high-performance liquid chromatography. Anal BioanalChem. 406 (18), 4563-4569 (2014).
- Ierapetritis, I., Lioupis, A., Lampi, E. Determination of phthalates into vegetable oils by isotopic dilution gas chromatography mass spectrometry. Food Anal Methods. 7 (7), 1451-1457 (2014).
- Sun, J. Z., He, H., Liu, S. H. Determination of phthalic acid esters in Chinese white spirit using dispersive liquid-liquid microextraction coupled with sweeping beta-cyclodextrin-modified micellar electrokinetic chromatography. J Sep Sci. 37 (13), 1679-1686 (2014).
- Yilmaz, P. K., Ertas, A., Kolak, U. Simultaneous determination of seven phthalic acid esters in beverages using ultrasound and vortex-assisted dispersive liquid-liquid microextraction followed by high-performance liquid chromatography. J Sep Sci. 37 (16), 2111-2117 (2014).
- Sun, R., Zhuang, H. An ultrasensitive gold nanoparticles improved real-time immuno-PCR assay for detecting diethyl phthalate in foodstuff samples. Anal Biochem. 480, 49-57 (2015).
- Sun, R. Y., Zhuang, H. S. A sensitive heterogeneous biotin-streptavidin enzyme-linked immunosorbent assay for the determination of di-(2-ethylhexyl)phthalate (DEHP) in beverages using a specific polyclonal antibody. Anal Methods. 6 (24), 9807-9815 (2014).
- Zhou, L., Lei, Y., Zhang, D., Ahmed, S., Chen, S. An ultra-sensitive monoclonal antibody-based enzyme-linked immunosobent assay for dibutyl phthalate in human urinary. Sci Total Environ. 541, 570-578 (2016).
- Shen, J., Li, Y., Gu, H., Xia, F., Zuo, X. Recent development of sandwich assay based on the nanobiotechnologies for proteins, nucleic Acids, small Molecules, and ions. Chem Rev. 114 (15), 7631-7677 (2014).
- Yin, X. -. B. Functional nucleic acids for electrochemical and electrochemiluminescent sensing applications. TrAC, Trends Anal Chem. 33, 81-94 (2012).
- Nguyen, V. -. T., Kwon, Y. S., Gu, M. B. Aptamer-based environmental biosensors for small molecule contaminants. Curr Opin Biotechnol. 45, 15-23 (2017).
- Groher, F., Suess, B. In vitro selection of antibiotic-binding aptamers. Methods. 106, 42-50 (2016).
- Yang, K. -. A., Pei, R., Stojanovic, M. N. In vitro selection and amplification protocols for isolation of aptameric sensors for small molecules. Methods. 106, 58-65 (2016).
- Jing, M., Bowser, M. T. Methods for measuring aptamer-protein equilibria: a review. Anal. Chim. Acta. 686 (1-2), 9-18 (2011).
- Mehta, J., et al. Selection and characterization of PCB-binding DNA aptamers. Anal Chem. 84 (3), 1669-1676 (2012).
- Matsumoto, M., Hirata-Koizumi, M., Ema, M. Potential adverse effects of phthalic acid esters on human health: A review of recent studies on reproduction. Regul Toxicol Pharm. 50 (1), 37-49 (2008).
- Goodchild, J. Conjugates of oligonucleotides and modified oligonucleotides: a review of their synthesis and properties. Bioconjug Chem. 1 (3), 165-187 (1990).
- Brown, D. M. A brief history of oligonucleotide synthesis. Protocols for Oligonucleotides and Analogs: Synthesis and Properties. , 1-17 (1993).
- Reese, C. B. Oligo-and poly-nucleotides: 50 years of chemical synthesis. Org Biomol Chem. 3 (21), 3851-3868 (2005).
- Sproat, B., Colonna, F., Mullah, B., et al. An efficient method for the isolation and purification of oligoribonucleotides. Nucleos Nucleot Nucl. 14 (1-2), 255-273 (1995).
- Han, Y., et al. Selection of group-specific phthalic acid esters binding DNA aptamers via rationally designed target immobilization and applications for ultrasensitive and highly selective detection of phthalic acid esters. Anal Chem. 89 (10), 5270-5277 (2017).
- Bartlett, J. M. S., Stirling, D. A short history of the polymerase chain reaction. PCR protocols. , 3-6 (2003).
- Saiki, R. K., Scharf, S., Faloona, F., et al. Enzymatic amplification of beta-globin genomic sequences and restriction site analysis for diagnosis of sickle cell anemia. Science. 230, 1350-1354 (1985).
- Albright, L. M., Slatko, B. E. Denaturing polyacrylamide gel electrophoresis. Current Protocols in Nucleic Acid Chemistry. , A. 3B 1-A. 3B 5 (2001).
- Summer, H., Grämer, R., Dröge, P. Denaturing urea polyacrylamide gel electrophoresis (Urea PAGE). JoVE. (32), e1485 (2009).
- Jing, M., Bowser, M. T. Methods for measuring aptamer-protein equilibria: a review. Anal Chim Acta. 686 (1), 9-18 (2011).
- Sharma, T. K., Bruno, J. G., Dhiman, A. ABCs of DNA aptamer and related assay development. Biotechnol Adv. 35 (2), 275-301 (2017).
- Liu, R., et al. Signaling-probe displacement electrochemical aptamer-based sensor (SD-EAB) for detection of nanomolar kanamycin A. Electrochim Acta. 182, 516-523 (2015).
- Mendonsa, S. D., Bowser, M. T. In vitro evolution of functional DNA using capillary electrophoresis. J Am Chem Soc. 126, 20-21 (2004).
- Lou, X. H., et al. Micromagnetic selection of aptamers in microfluidic channels. Proc Natl Acad Sci USA. 106 (9), 2989-2994 (2009).
- Cho, M., et al. Quantitative selection of DNA aptamers through microfluidic selection and high-throughput sequencing. Proc. Natl. Acad. Sci. U.S.A. 107, 15373-15378 (2010).
- Cho, M., et al. Quantitative selection and parallel characterization of aptamers. Proc Nat Acad Sci USA. 110 (46), 18460-18465 (2013).
- Song, K. -. M., et al. Gold nanoparticle-based colorimetric detection of kanamycin using a DNA aptamer. Anal Biochem. 415 (2), 175-181 (2011).
- Yang, Z., Ding, X., Guo, Q., et al. Second generation of signaling-probe displacement electrochemical aptasensor for detection of picomolar ampicillin and sulfadimethoxine. Sens Actuators B. 253 (2017), 1129-1136 (2017).
- Lou, X., Zhao, T., Liu, R., Ma, J., Xiao, Y. Self-assembled DNA monolayer buffered dynamic ranges of mercuric electrochemical sensor. Anal Chem. 85 (15), 7574-7580 (2013).
- Zhao, T., et al. Nanoprobe-enhanced, split aptamer-based electrochemical sandwich assay for ultrasensitive detection of small molecules. Anal Chem. 87 (15), 7712-7719 (2015).
- Lou, X., He, L. A. Surface passivation using oligo(ethylene glycol) in ATRP-assisted DNA detection. Sens Actuators,B. 129 (1), 225-230 (2008).

