Purification and microRNA Profiling of Exosomes Derived from Blood and Culture Media
Summary
The presence of stable microRNAs (miRNAs) in exosomes has generated immense interest as a novel mode of intercellular communication, for their potential utility as biomarkers and as a route for therapeutic intervention. Here we demonstrate exosome purification from blood and culture media followed by quantitative PCR to identify miRNAs being transported.
Abstract
Stable miRNAs are present in all body fluids and some circulating miRNAs are protected from degradation by sequestration in small vesicles called exosomes. Exosomes can fuse with the plasma membrane resulting in the transfer of RNA and proteins to the target cell. Their biological functions include immune response, antigen presentation, and intracellular communication. Delivery of miRNAs that can regulate gene expression in the recipient cells via blood has opened novel avenues for target intervention. In addition to offering a strategy for delivery of drugs or RNA therapeutic agents, exosomal contents can serve as biomarkers that can aid in diagnosis, determining treatment options and prognosis.
Here we will describe the procedure for quantitatively analyzing miRNAs and messenger RNAs (mRNA) from exosomes secreted in blood and cell culture media. Purified exosomes will be characterized using western blot analysis for exosomal markers and PCR for mRNAs of interest. Transmission electron microscopy (TEM) and immunogold labeling will be used to validate exosomal morphology and integrity. Total RNA will be purified from these exosomes to ensure that we can study both mRNA and miRNA from the same sample. After validating RNA integrity by Bioanalyzer, we will perform a medium throughput quantitative real time PCR (qPCR) to identify the exosomal miRNA using Taqman Low Density Array (TLDA) cards and gene expression studies for transcripts of interest.
These protocols can be used to quantify changes in exosomal miRNAs in patients, rodent models and cell culture media before and after pharmacological intervention. Exosomal contents vary due to the source of origin and the physiological conditions of cells that secrete exosomes. These variations can provide insight on how cells and systems cope with stress or physiological perturbations. Our representative data show variations in miRNAs present in exosomes purified from mouse blood, human blood and human cell culture media.
Here we will describe the procedure for quantitatively analyzing miRNAs and messenger RNAs (mRNA) from exosomes secreted in blood and cell culture media. Purified exosomes will be characterized using western blot analysis for exosomal markers and PCR for mRNAs of interest. Transmission electron microscopy (TEM) and immunogold labeling will be used to validate exosomal morphology and integrity. Total RNA will be purified from these exosomes to ensure that we can study both mRNA and miRNA from the same sample. After validating RNA integrity by Bioanalyzer, we will perform a medium throughput quantitative real time PCR (qPCR) to identify the exosomal miRNA using Taqman Low Density Array (TLDA) cards and gene expression studies for transcripts of interest.
These protocols can be used to quantify changes in exosomal miRNAs in patients, rodent models and cell culture media before and after pharmacological intervention. Exosomal contents vary due to the source of origin and the physiological conditions of cells that secrete exosomes. These variations can provide insight on how cells and systems cope with stress or physiological perturbations. Our representative data show variations in miRNAs present in exosomes purified from mouse blood, human blood and human cell culture media
Introduction
Short noncoding miRNAs modulate gene expression by binding to the target mRNA. Seed sequence complementarity of ~7 base pairs enables miRNA to bind to the target mRNA resulting in the inhibition of translation or in reduction in the stability of the mRNA, both of which can result in decreased expression of the target protein 1. Research over the last decade has unequivocally proven a fundamental role for miRNAs in mediating cellular functions. There has also been considerable effort directed towards dissecting miRNA mediated molecular changes underlying various diseases 2,3. Furthermore, recent identification of stable miRNAs in bodily fluids 4-6 paved the way for their use as novel biomarkers amenable to clinical diagnosis.
One mode of miRNA transport in bodily fluids is via exosomes, small vesicles that carry mRNAs, proteins, lipid mediators, and miRNAs to recipient cells via systemic blood circulation 7-14. This results in modulation of gene expression in recipient cells and represents a novel mechanism of cellular communication. For instance, cells can modulate immune-regulatory processes by secreting and/or absorbing exosomes containing biomolecules involved in inflammation such as interleukin-1β (IL1β), tumor necrosis factor-α (TNFα), transforming growth factor-β5 (TGFβ5), and the miRNAs that regulate these genes 13. As aberrant miRNA expression is a common feature in a variety of human diseases, these molecules offer exciting new opportunities for the discovery and validation of novel therapeutic targets 2.
Circulating miRNAs are present in all body fluids and it is known that the composition of exosomes is different based on the source cells from which they were released. Thus they offer an avenue to study the physiological status of the cells and how the cells alter signaling events in response to stress including diseases. Studying alterations in exosome composition can provide insight into signal transduction and investigate their potential utility as biomarkers or therapeutic intervention routes.
Here we will demonstrate the purification of exosomes from multiple sources based on published protocols. These exosomes will be used for RNA isolation followed by qPCR to identify and measure the levels of miRNAs present in the exosomes.
Protocol
All experiments using blood samples from human and rodents were executed in compliance with all relevant guidelines, regulations and regulatory agencies. Human subjects were enrolled after giving informed consent as approved by the Drexel University College of Medicine Institutional Review Board and all procedures for studies performed using animals were approved by Drexel’s Institutional Animal Care and Use Committee.
1. Exosome Purification from Blood (~5.5 hr)
TIPS
- We resuspend exosomes from one tube of blood (~10 ml human blood and ~2 ml mouse blood) in 300 μl RNA lysis buffer from the mirVana miRNA isolation kit.
- To purify exosomes from cell culture media, the media is spun at 500 x g for 10 min to remove cells and also filtered through a 0.22 μm filter after centrifugation at 12,000 x g. The exosome purification from cell culture media does not require a dilution in PBS, the PBS wash or the second ultracentrifugation step.
- If the purified exosomes will be used for proteomics studies, adding a sucrose gradient centrifugation step after the ultracentrifugation step will decrease the amount of protein aggregates and contaminants 15.
- The observed morphology of exosomes can differ depending on both the source and the processing steps for TEM. We did not observe the cup shaped morphology for exosomes which has been previously attributed to differences in sample processing 9,15. Our TEM images including the immunogold labeling are similar to previous reports from different sources including blood, biliary and cortical culture derived exosomes 9,16,17.
- Invert EDTA coated tube containing the blood 5x and place upright at room temperature for 10-60 min.
- Centrifuge at 2,000 x g for 15 min at 4 °C.
- Collect the top layer, the plasma, in 15 ml tubes and keep on ice.
- Dilute fluid with an equal volume of PBS. Centrifuge 30 min at 2,000 x g at 4 °C.
- Transfer to centrifuge tubes, add 1X PBS for a total volume of 24 ml, and centrifuge 45 min at 12,000 x g at 4 °C.
- Transfer to ultracentrifuge tubes and centrifuge 2 hr at 110,000 x g at 4 °C.
- Resuspend pellet in PBS and centrifuge 1 hr at 100,000 x g at 4 °C.
- Resuspend pellet in appropriate buffer (RNA lysis buffer).
- Resuspend in PBS or RIPA buffer for electron microscopy and western blot analysis respectively.
2. RNA Purification Using a Modified mirVana miRNA Isolation Kit (~1 hr)
TIPS
- Prepare the area for isolating RNA by wearing personal protective equipment (PPE), wiping surfaces and pipettes with RNase Zap, and using sterile filter tips and tubes.
- Though the protocol from the vendor gives the option of purifying only miRNA, we always isolate total RNA including mRNA and miRNA so that we can study both from the same sample.
- Adding an on-column DNase treatment step removes traces of DNA that may be present. Depending on the source of exosomes, DNA has been reported to be absent in exosomes purified from some sources including mouse and human mast cell lines 8 but chromosomal DNA sequences have been detected in exosomes purified from media of cultured cardiomyocytes 18.
- We reduced the volume of wash solution 1 to 350 μl (Step 2.4 below) because the vendor provides only enough Wash solution 1 for one 700 μl addition. If the volume is not reduced there is not sufficient solution left for using the entire kit contents. Remaining steps were performed according to manufacturer’s recommendations.
- Add 1/10 volume of miRNA Homogenate Additive (30 μl) and incubate on ice for 10 min.
- Extract with a volume of acid-phenol:chloroform equal to the initial lysate volume. Centrifuge and recover the upper phase in a fresh tube.
- Add 1.25 volumes 100% ethanol (375 μl). Vortex. Apply to filter cartridge and centrifuge 15 sec at 10,000 x g.
- Add half of recommended volume miRNA Wash Solution 1 (350 μl) and centrifuge 15 sec at 10,000 x g.
- Add 10 μl DNase 1 to 70 μl buffer RDD (Qiagen) for each sample. Add 80 μl to column and incubate 15 min at room temperature.
- Add Wash Solution 1 (350 μl) and centrifuge 15 sec at 10,000 x g.
- Add Wash Solution 2/3 (500 μl) and centrifuge 15 sec at 10,000 x g. REPEAT.
- Elute with 40 μl 95 °C nuclease-free H2O.
- Determine the concentration of RNA and dilute the RNA to 100 ng/ μl.
3. miRNA Profiling of Exosomes from Blood Using Taqman Low Density Array (TLDA) Cards (~6.5 hr)
TIPS
- Prepare the area for isolating RNA by wearing PPE, wiping surfaces and pipettes with RNase Zap, and using sterile filter tips and tubes.
- Pre-amplification (pre-amp) step is recommended for 1-350 ng RNA (350-1,000 ng samples do not require pre-amp step after cDNA synthesis). It is important to determine whether pre-amp is necessary for your sample because you must treat all samples under equal conditions. Pre-amp step adds cost and the Ct values have a lower acceptable cut off (32 instead of 40).
- It is useful to prepare the cDNA reaction in strip tubes instead of a plate depending on the number of cards that can be run in a single day. This will minimize freeze-thaw of cDNA samples that will not be processed on the same day.
- Exosomes collected from different sources have different amounts of RNA. This protocol allows a maximum of 3 μl RNA template for the cDNA synthesis reaction. Depending on the source, the amount of exosomal RNA that can be isolated maybe limited. While a typical tissue would yield enough RNA that cDNA would be made from 100 ng RNA in 3 μl volume, exosomes from human blood or human cell culture media often contain less RNA and cDNA must be made from less starting material. Mouse blood exosomes contain higher levels of RNA and at least 100 ng in 3 μl can be obtained.
- To prepare cDNA from purified RNA using Megaplex Pools reverse transcriptase primers (Pool A and Pool B contain primers for the miRNAs on microfluidic cards Array A and B), thaw the reaction components on ice.
- Label two RNase-free 1.5 ml tubes for RT reaction with Pool A primers or Pool B primers and add components in chart order. Add at least one extra volume of components for 2 or more reactions or make a master mix as recommended by the vendor.
| RT reaction Component | Component Concentration | Volume for 1 sample(total V/sample = 4.5 μl) |
| Nuclease-free water | 0.20 μl | |
| Megaplex RT buffer (10X) | 1X | 0.80 μl |
| dNTPs (100 mM) | 4.4 mM | 0.20 μl |
| MgCl2 (25 mM) | 5 mM | 0.90 μl |
| RNase Inhibitor (20U/μl) | 2 U | 0.10 μl |
| Megaplex RT Primers A or B (10X) | 1X | 0.80 μl |
| MultiScribe Reverse Transcriptase | 75 U | 1.50 μl |
- Invert tubes 6 times and centrifuge briefly. Then, pipette 4.5 μl RT reaction mix into MicroAmp 8-tube Strips or a 96-well MicroAmp optical reaction plate.
- Add 100 ng of RNA and adjust the volume to 3 μl with DEPC treated water or add 3 μl total RNA, stir with the pipette tip, and seal the tubes or plate. Spin briefly and incubate on ice for 5 min.
- Load the reaction into the PCR machine and run it under the following conditions (as recommended by the Applied Biosystems Megaplex Pools protocol):
| Stage | Temp | Time |
| Cycle (40x) | 16 °C | 2 min |
| 42 °C | 1 min | |
| 50 °C | 1 sec | |
| Hold | 85 °C | 5 min |
| Hold | 4 °C | ∞ |
- Store cDNA at -20 °C (good for at least one week) or continue to pre-Amp step which can be stored for one week.
- Thaw PreAmp primers for Pool A and Pool B on ice and invert to mix. Swirl Taqman Pre-Amp master mix and keep on ice also.
- Follow the chart adding at least one extra volume of components for 2 or more reactions or make a master mix as recommended by the vendor.
| Pre-Amp Reaction Component | Volume for 1 sample |
| Nuclease-free water | 7.5 μl |
| Taqman PreAmp Master Mix (2X) | 12.5 μl |
| Megaplex PreAmp Primers A or B (10X) | 2.5 μl |
- Pipette 2.5 μl RT product into MicroAmp 8-tube Strips or a 96-well MicroAmp optical reaction plate and dispense 22.5 μl PreAmp reaction mix into each well.
- Seal the plate, incubate on ice 5 min and then load the sample into the PCR machine. Run under the following conditions.
| Stage | Temp | Time |
| Hold | 95 °C | 10 min |
| Hold | 55 °C | 2 min |
| Hold | 72 °C | 2 min |
| Cycle (12x) | 95 °C | 15 sec |
| 60 °C | 4 min | |
| Hold | 99.9 °C | 10 min |
| Hold | 4 °C | ∞ |
- After the preamp reaction is finished, briefly centrifuge the tubes or plate and then add 75 μl 0.1X TE pH 8.0 to each well or tube.
- Seal and mix and spin briefly. Store the diluted product for one week at -20 °C or continue on to load the plate.
- Combine the following in an RNase-free 1.5 ml tube on ice.
| Component | Volume for 1 card |
| Taqman Universal PCR Master Mix, No AmpErase UNG (2X) | 450 μl |
| Diluted preAmp product | 9 μl |
| Nuclease-free water | 441 μl |
- Invert the tube to mix and centrifuge briefly. Dispense 100 μl PCR reaction mix into each port of the Taqman MicroRNA Array card.
- Centrifuge the card 2 x 1,000 x g for 1 min. Seal the card and run it using the template provided with the primers in the Applied Biosystems manual and the 384 well Taqman Low Density Array settings. We use Applied Biosystems 7900HT Fast Real-Time PCR System.
4. mRNA Analysis by Taqman (~1.5 hr)
- To prepare qPCR reaction, mix the reaction components in the order in the chart. All samples will be analyzed with primer probes for the gene of interest as well as 18S rRNA as the normalizer gene.
| Taqman Reaction Component | Volume for 1 sample |
| Nuclease-free water | 7 μl (adjust based on individual sample) |
| Taqman Fast Master Mix (2X) | 10 μl |
| Taqman Primer probes (20x) | 1 μl |
| cDNA from 100 ng RNA | 2 μl (adjust based on individual sample) |
- Run the 96 well plate using the Fast 96-well block as recommended by the manufacturer (Applied Biosystems 7900HT Fast Real-Time PCR System).
Representative Results
After isolating exosomes from blood or cell culture media, the purity of the exosomes can be tested by electron microscopy (EM) and western blot (Figures 1A and 1B). We confirmed our exosome preparations from various sources with EM and western blot using multiple antibodies. Figure 1A shows EM images confirming that exosomes are intact with a diameter of ~30 -100 nm and contain CD81 by immunogold labeling. Commonly used exosomal markers are Hsp70 and tetraspannin family glycoproteins CD63, CD81 and CD9 14. After confirming the integrity of the purified exosomes, we performed western blot for Hsp70 in Figure 1B and showed that exosomes contain Hsp70 while media alone (negative control) does not. The Bioanalyzer trace shows that exosomes isolated from RAW cell culture media contain a variety of RNAs, but do not contain large amounts of ribosomal RNAs that are typical in RNA from whole cell (Figure 1C). Figure 1D shows qPCR analysis of mRNA from whole blood and exosomes derived from blood. We are able to isolate 500 ng of exosomal RNA from ~10 ml human blood using mirVana miRNA isolation kit. The qPCR results indicate the presence of mRNAs encoding TNFα and vascular endothelial growth factor A (VEGFA) in purified exosomes as well as in whole blood.
In addition to qPCR for gene expression studies, total RNA was also used for miRNA profiling. The miRNA TLDA card version 3.0 contains primer probes for ~758 miRNAs including the endogenous control RNA U6. With the addition of the pre-amplification step, the vendor recommends 30 ng RNA to run a TLDA card. This is useful for analyzing RNA from exosomes purified from serum or human blood which typically have lower yields than those purified from mouse blood. The miRNA analysis shown in Figure 2 was performed with 250 ng RNA from mouse blood exosomes and 15-30 ng exosomal RNA from human blood or human aortic endothelial cells (HAOEC). Our results indicate the presence of 89 miRNAs in exosomes collected from mouse blood, 209 miRNAs in human blood exosomes, and 199 miRNAs in exosomes from human cell culture media. Representative data for six miRNAs is shown as delta Ct value normalized to the endogenous U6 RNA for rodent blood (Figure 2A), human blood (Figure 2B) and HAOEC cell culture media (Figure 2C). While miR-126 and miR-200c were absent in exosomes from rodent blood or HAOEC media respectively, the remaining four miRNAs are present in all three samples in varying amounts. Relative to exosomes purified from cell culture media, miR-223 is expressed at higher levels in exosomes from human and rodent blood samples.
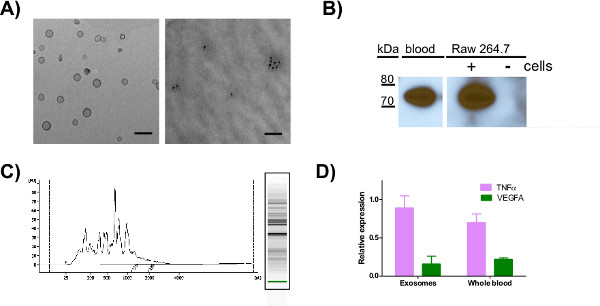
Figure 1. Transmission electron microscopy (TEM), western blot analysis and qRT-PCR for mRNAs using exosomes purified from various sources. A) TEM image of mouse blood exosomes resuspended in 1% glutaraldehyde (left) or RAW cell derived exosomes resuspended in 4% paraformaldehyde, labeled with 10 nm gold and rabbit-anti CD81 (right) and spotted on formvar carbon-coated grids for EM analysis (scale bar = 100 nm). B) Western analysis of HSP70 in exosomes purified from mouse blood or exosome-free media +/- 24 hr incubation with RAW 264.7 murine cells and resuspended in RIPA buffer. C) Bioanalyzer analysis of total RNA purified from exosomes derived from RAW 264.7 cell culture media D) Total RNA purified from whole blood and exosomes obtained from a representative human control was used to compare the expression levels of TNFα and VEGFA. Click here to view larger figure.
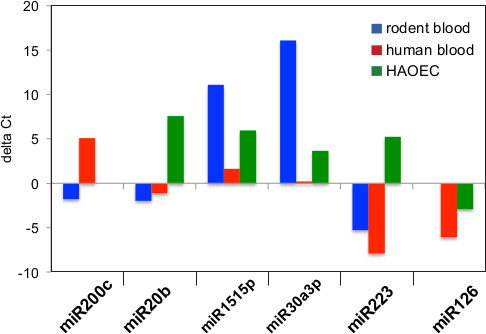
Figure 2. Relative Expression of miRNA fraction found in exosomes. Threshold cycle (CT value) is a relative measure of the concentration of an individual miRNA in the PCR reaction and lower CT value indicates higher expression. qPCR values for six detectable exosomal miRNAs were normalized to U6 RNA and the delta Ct values were graphed for exosomes from three sources. A negative value indicates higher expression in this figure. Exosomes from rodent blood (A) and human blood (B) expressed hsa-miR-223 at higher levels than control, while exosomes from HAOEC cell culture media (C) expressed hsa-miR-223 at lower levels than control. Hsa-miR-226 was absent from rodent blood exosomes and hsa-miR-200c was absent from HAOEC exosomes.
Discussion
In this protocol, we show the quantification of miRNAs and mRNAs from exosomes purified by differential centrifugation from blood and culture media. Exosomes have diverse components dependent upon their origin and are involved in a number of biological functions, including immune response, antigen presentation, intracellular communication, and the transfer of RNA and proteins 9,11,12,19,20. While size and shape is a determinant of exosome purity, a number of papers showing EM data of exosomes indicate that these vesicles can range in size and morphology based on the source from which they are secreted as well as the method used for fixation and imaging 9,15. The accepted size for exosomes of 30-100 nm is an average range that includes a small population of exosomes with a larger diameter depending on the source 21,22, one reason that multiple methods are used to validate exosomal purity. Secreted miRNAs have many requisite features of good biomarkers 23 in addition to being potential therapeutic targets for various diseases 2,3,24. This purification method provides a noninvasive way to characterize temporal biological changes in exosomal content from a variety of bodily fluids while miRNA profiling by qPCR provides a comprehensive overview of miRNA content 2. Over 4,500 proteins, 1,639 mRNAs, 764 miRNAs and 194 lipids are known to associate with exosomes 25 (www.exocarta.org) and in some cases, changes in the levels of these biomolecules has already been linked to disease states. In our studies, the exosomes purified from human blood had many miRNAs in common with exosomes purified from media from cultured human cell lines but significant differences from exosomal miRNAs found in mouse blood. While there are examples of miRNAs that target only one mRNA, many miRNAs function to partially reduce the levels of multiple targets leading to a strong physiological response. Characterizing both mRNA and miRNA that reside in exosomes provides unique insight into exosome-mediated information transfer and the role of circulating miRNAs in mediating changes in gene expression. Purification and characterization of exosomes from a variety of sources will be beneficial in identifying molecular signatures associated with these secretory vesicles.
Divulgations
The authors have nothing to disclose.
Acknowledgements
This study was supported by funds from Rita Allen Foundation grant to Seena Ajit. The authors would like to acknowledge Erika Balogh and Dr. Soumitra Ghoshroy from the University of South Carolina Electron Microscopy Center for instrument use, scientific and technical assistance.
Materials
| Name of Reagent/Material | Company | Catalog Number | Comments |
| EDTA coated vacutainer (10 ml tubes) | BD Diagnostics | 366643 | For exosome purification from human blood |
| EDTA coated vacutainer (2 ml tubes) | BD Diagnostics | 367841 | For exosome purification from mouse blood |
| PAXgene Blood RNA Tube | BD Diagnostics | 762165 | For miRNA isolation from total blood |
| miRVana microRNA isolation kit | Ambion | AM1561 | |
| Acid-Phenol: CHCl3 | Ambion | 9721G | |
| DNase 1 | Qiagen | 79254 | |
| TaqMan Universal PCR Master Mix, No AmpErase UNG | Applied Biosystems | 4326614 | For TLDA cards |
| Megaplex RT rodent Pool Set v3.0 | Applied Biosystems | 4444746 | |
| Megaplex preamp rodent Pool set v3.0 | Applied Biosystems | 4444747 | |
| Taqman Array Rodent MicroRNA A+B set V3.0 | Applied Biosystems | 4444909 | |
| Megaplex RT Human Pool set V3.0 | Applied Biosystems | 4444745 | |
| Megaplex Preamp human pool set v3.0 | Applied Biosystems | 4444748 | |
| Taqman Array Human MicroRNA A+B Cards Set v3.0 | Applied Biosystems | 4444913 | |
| Taqman MicroRNA RT kit | Applied Biosystems | 4366596 | |
| Taqman fast universal PCR master mix | Applied Biosystems | 4366072 | For mRNA qRT-PCR |
| Tumor necrosis factor (primer probe) | Applied Biosystems | Hs01113624_g1 | |
| Vascular endothelial growth factor A (primer probe) | Applied Biosystems | Hs00900055_m1 | |
| Maxima First Strand cDNA Synthesis Kit for RT-qPCR | Thermo Scientific | K1642 | For mRNA |
| HSP70 antibody | Abcam | ab94368 | For western blot |
| Anti-rabbit IgG-Gold | Sigma | G7402 | For electron microscopy |
| Rabbit-anti CD81 | Sigma | SAB3500454 | |
| Nickel 300 mesh carbon formvar grids | Electron Microscopy Sciences | FCF300-Ni | |
| Copper 300 mesh carbon formvar grids | Electron Microscopy Sciences | FCF300-Cu | |
| Table 1. Table of specific reagents. |
References
- Bartel, D. P. MicroRNAs: target recognition and regulatory functions. Cell. 136, 215-233 (2009).
- Mendell, J. T., Olson, E. N. MicroRNAs in stress signaling and human disease. Cell. 148, 1172-1187 (2012).
- Esteller, M. Non-coding RNAs in human disease. Nature reviews. Genetics. 12, 861-874 (2011).
- Mitchell, P. S., et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proceedings of the National Academy of Sciences of the United States of America. 105, 10513-10518 (2008).
- Chen, X., et al. Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell research. 18, 997-1006 (2008).
- Gilad, S., et al. Serum microRNAs are promising novel biomarkers. PloS one. 3, e3148 (2008).
- Mittelbrunn, M., et al. Unidirectional transfer of microRNA-loaded exosomes from T cells to antigen-presenting cells. Nature. 2, 282 (2011).
- Valadi, H., et al. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat Cell Biol. 9, 654-659 (2007).
- Gyorgy, B., et al. Membrane vesicles, current state-of-the-art: emerging role of extracellular vesicles. Cell Mol Life Sci. 68, 2667-2688 (2011).
- Mathivanan, S., Ji, H., Simpson, R. J. Exosomes: extracellular organelles important in intercellular communication. J. Proteomics. 73, 1907-1920 (2010).
- Ramachandran, S., Palanisamy, V. Horizontal transfer of RNAs: exosomes as mediators of intercellular communication. Wiley Interdiscip Rev. RNA. , (2011).
- Record, M., Subra, C., Silvente-Poirot, S., Poirot, M. Exosomes as intercellular signalosomes and pharmacological effectors. Biochem Pharmacol. 81, 1171-1182 (2011).
- Thery, C., Ostrowski, M., Segura, E. Membrane vesicles as conveyors of immune responses. Nat Rev Immunol. 9, 581-593 (2009).
- Ludwig, A. K., Giebel, B. Exosomes: small vesicles participating in intercellular communication. The international journal of biochemistry & cell biology. 44, 11-15 (2012).
- Thery, C., Amigorena, S., Raposo, G., Clayton, A. Isolation and characterization of exosomes from cell culture supernatants and biological fluids. Curr. Protoc. Cell Biol. Chapter 3, Unit 3 22 (2006).
- Masyuk, A. I., et al. Biliary exosomes influence cholangiocyte regulatory mechanisms and proliferation through interaction with primary cilia. American journal of physiology. Gastrointestinal and liver physiology. 299, G990-G999 (2010).
- Fauré, J., et al. Exosomes are released by cultured cortical neurones. Molecular and Cellular Neuroscience. 31, 642-648 (2006).
- Waldenstrom, A., Genneback, N., Hellman, U., Ronquist, G. Cardiomyocyte microvesicles contain DNA/RNA and convey biological messages to target cells. PloS one. 7, e34653 (2012).
- Simpson, R. J., Lim, J. W., Moritz, R. L., Mathivanan, S. Exosomes: proteomic insights and diagnostic potential. Expert Rev Proteomics. 6, 267-283 (2009).
- Turchinovich, A., Weiz, L., Langheinz, A., Burwinkel, B. Characterization of extracellular circulating microRNA. Nucleic acids research. 39, 7223-7233 (2011).
- El-Andaloussi, S., et al. Exosome-mediated delivery of siRNA in vitro and in vivo. Nature. 7, 2112-2126 (2012).
- Singh, P. P., Smith, V. L., Karakousis, P. C., Schorey, J. S. Exosomes isolated from mycobacteria-infected mice or cultured macrophages can recruit and activate immune cells in vitro and in vivo. J. Immunol. 189, 777-785 (2012).
- Etheridge, A., Lee, I., Hood, L., Galas, D., Wang, K. Extracellular microRNA: A new source of biomarkers. Mutat. Res. , (2011).
- Stenvang, J., Silahtaroglu, A. N., Lindow, M., Elmen, J., Kauppinen, S. The utility of LNA in microRNA-based cancer diagnostics and therapeutics. Seminars in cancer biology. 18, 89-102 (2008).
- Mathivanan, S., Fahner, C. J., Reid, G. E., Simpson, R. J. ExoCarta 2012: database of exosomal proteins, RNA and lipids. Nucleic acids research. 40, D1241-D1244 (2012).

