Generation of Shear Adhesion Map Using SynVivo Synthetic Microvascular Networks
Summary
Flow chambers used in adhesion experiments typically consist of linear flow paths and require multiple experiments at different flow rates to generate a shear adhesion map. SynVivo-SMN enables the generation of shear adhesion map using a single experiment utilizing microliter volumes resulting in significant savings in time and consumables.
Abstract
Cell/particle adhesion assays are critical to understanding the biochemical interactions involved in disease pathophysiology and have important applications in the quest for the development of novel therapeutics. Assays using static conditions fail to capture the dependence of adhesion on shear, limiting their correlation with in vivo environment. Parallel plate flow chambers that quantify adhesion under physiological fluid flow need multiple experiments for the generation of a shear adhesion map. In addition, they do not represent the in vivo scale and morphology and require large volumes (~ml) of reagents for experiments. In this study, we demonstrate the generation of shear adhesion map from a single experiment using a microvascular network based microfluidic device, SynVivo-SMN. This device recreates the complex in vivo vasculature including geometric scale, morphological elements, flow features and cellular interactions in an in vitro format, thereby providing a biologically realistic environment for basic and applied research in cellular behavior, drug delivery, and drug discovery. The assay was demonstrated by studying the interaction of the 2 µm biotin-coated particles with avidin-coated surfaces of the microchip. The entire range of shear observed in the microvasculature is obtained in a single assay enabling adhesion vs. shear map for the particles under physiological conditions.
Introduction
Current assays to study to cell-cell and particle-cell interactions typically involve static well plate format in which particles or cells are incubated on protein matrices or adherent cells. At the end of the specified incubation time, the numbers of adherent particles or cells are quantified using microscopy1. Even though these assays provide significant insight into the biochemical processes behind these interactions, a key limitation is the lack of physiological fluid flow (typical of microcirculation) and its impact on particle adhesion.
To overcome this limitation, in vitro flow chambers have been developed in recent years. A common element of these flow chambers is a transparent apparatus perfused at low Reynolds numbers to match wall shear rates observed in blood vessels in vivo2. The vessel wall is modeled by either coating of biomolecules or growth of cells on one surface of the flow chamber3. Particles4-7 or cells8-16 are then flowed in at desired range of flow rates to quantify the number of adhering particles under various shear rates.
However, the use of parallel plate flow chambers to study and validate the biochemical phenomena is rather expensive and time consuming. This is mainly due to the fact that multiple experiments need to be conducted for generating a map of the fluidic shear vs. the number of particles/cells adhered. In addition, plate flow chambers require large volumes of reagents due to their large size (height > 250 µm and width > 1 mm). Finally, these devices do not accurately model geometrical features (e.g., bifurcations) and flow conditions (e.g., converging vs. diverging flows) that are present in vivo.
Recent advances in lithography based microfabrication17-19 have accelerated the field of lab-on-a-chip devices20-21. These devices have been instrumental in developing a miniaturized version of the parallel plate flow chamber with dimensions in the micrometer regime. The reduction in dimension also yields significant benefits in terms of volumes of reagents, cells or particle required for experiments. However, a key limitation of the currently available devices is the use of linear channels to model microvessels, which does not mimic the complex microvasculature observed in vivo.
We have recently developed a novel methodology for recreating microvascular networks onto disposable plastic substrates resulting in synthetic representation of the in vivo conditions. These devices termed SynVivo-Synthetic Microvascular Networks (SMN) are developed using PDMS based soft-lithography process. SynVivo-SMN devices can be used to obtain shear adhesion map of cell/particle adhesion22, study targeted drug delivery23 and have been validated against in vivo data24-25. In this paper, we present a protocol that enables generation of the shear adhesion map from a single experiment in volumes as small as 1-5 µl thereby resulting in significant savings of resources and time.
Protocol
1. Priming the SynVivo-SMN Microfluidic Device
- Each port (inlet/outlet) of the device is comprised of two parallel ports – one for flowing in surface coating moieties (adhesion molecules, growth matrices, etc.) and/or cells for seeding and the other for running the assay (Figure 1A).
- Completely submerge the SynVivo-SMN microfluidic device (Figure 1B) in a Petri dish containing sterile deionized (DI) water and place the dish into a vacuum desiccator. Allow the desiccator to run until all of the air is removed from the channels of the device. This should take approximately 15 min.
- Before removing the device from the water, place Tygon tubing (O.D. of 0.06″ and I.D. of 0.02″) primed with water into each port of the device with fine-point forceps. The tubing should be approximately 1 inch in length. The device can now be removed from the water. Figure 1C shows image of the device with the tubing.
2. Coating the Microfluidic Device with Desired Protein (e.g., Avidin)
- Using a pipette, place a drop of water (approximately 100 µl) around the base of the one inlet port tubing. Carefully remove the tubing used to prime the device. The drop of water will prevent air from entering the device.
- Prepare a 1 ml syringe loaded with avidin at a concentration of 20 µg/ml. Connect the syringe to a 24 G stainless steel needle and tubing. Insert the tubing to one of the inlet ports of the device. Clamp the inlet port not being utilized with a jaw clamp.
- Inject avidin at a flow rate of 1 µl/min for 10 min to allow complete perfusion of the device. At the end of the flow time, clamp the tubing with the jaw clamp and place the device at 4 °C overnight.
3. Flowing the Biotinylated Particles for Adhesion Experiments
- Allow the device to come to room temperature. Place the device on an inverted fluorescence microscope equipped with a motorized stage and a high performance camera.
- Prepare a solution of 2 µm biotinylated particles at a concentration of 5 x 106 particles/ml in Phosphate Buffer Saline (PBS). Load the particles into a 1 ml syringe. Prepare a second 1 ml syringe of PBS only. Load each syringe on a syringe pump and connect to needle and tubing.
- Using a pipette, place a drop of water (approximately 100 µl) around the base of the inlet port tubing. Carefully remove the tubing used to coat the device. The drop of water will prevent air from entering the device.
- Carefully insert the tubing’s for biotinylated particles and PBS from step 3.2 into each of the inlet ports. Figure 2A shows image of the set-up.
- Start injecting biotinylated particles at a flow rate of 2.5 µl/min. Monitor the inlet port on the microscope. At the first sign of particles, begin the timer and continue flow for 3 min.
- At the end of the 3 min, stop the flow of biotinylated particles while simultaneously staring the flow of PBS at a flow rate of 2.5 µl/min. Allow PBS to flow in the device for 3 min to wash off unbound particles.
4. Acquiring Images and Making Area of Interest (AOI) Measurements Using Imaging Software (NIKON Elements)
- Use the “scan large image” function in the imaging software to acquire the image of the entire device.
- Sequentially number the bifurcations in the device and create a circular AOI with twice the diameter of the channels. In this case, set the AOI diameter to 200 µm since the channel diameter is 100 µm.
- Use the automated count function in the imaging software to export the number of particles in each AOI to an MS Excel sheet.
- Likewise, use the automated count feature to export the number of particles in the entire device.
5. Particle Flux Analysis Using Computational Fluid Dynamics (CFD) Models
- CFD simulations are run using commercially available software (CFD-ACE+, ESI Inc.) for the SynVivo-SMN device topology. The results are stored in a database for analyzing experimental observations. The simulation results store information on wall shear rates, velocity, particle flux, and adhesion in the device.
- The simulation results are used to determine the number of particles entering each AOI based on a given inlet particle concentration.
6. Generating Shear Adhesion Map
- Calculate % of adhesion by dividing the adhered particles in the bifurcation by the particles flowing in the bifurcation as shown in equation 6.1.
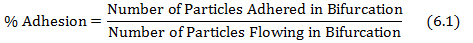
where the number of particles adhered and the particles flowing are obtained from protocol steps 4.3 and 5.2, respectively. - Plot the shear adhesion map using the shear rate at each bifurcation of the networks obtained from the database in step (5.1) and the % adhesion values obtained from equation 6.1.
Representative Results
Figure 1A shows a schematic and a bright field image of SynVivo-SMN device. Figure 1B shows the SynVivo-SMN device mounted on a glass slide. Figure 1C shows the device with tubing following priming with water in a vacuum desiccator.
Figure 2A shows an image of the experimental-set up. Figure 2B shows a typical avidin-coated SynVivo-SMN device following binding of 2 µm biotinylated particles. Note that particles preferentially adhere near the bifurcations in the network.
Figure 3A shows the numbered bifurcations in the SynVivo-SMN network. Figure 3B shows sample wall shear map generated by the CFD model of the device. The shear rate varies in the device ranging from 250 sec-1 to 15 sec-1 as observed in the microvasculature in vivo. Note that these varying shear rates cannot be obtained simultaneously in linear microfluidic channels as they provide a constant shear rate for each flow rate. Figure 3C shows a plot of the values of the shear rate at each of the bifurcations of the network. The data shows that the shear rate patterns are complex and cannot be obtained with a simple analytical relationship unlike linear flow channels. Furthermore, multiple bifurcations (AOIs) are present in the network that fall in the same shear bin, adding to the statistical confidence of the data.
Figure 4A shows sample results of the CFD analysis for particle fluxes in the network which are used to compute the particle fluxes in the branches and bifurcations of the experimental network. Figure 4B shows the shear adhesion map computed from the single SynVivo-SMN experiment. The shear adhesion maps follows the inverse relationship as observed from shear adhesion map obtained from linear channel experiments. However, in contrast using a SynVivo assay, a single experiment allows generation of this shear map unlike multiple experimental runs required from the linear channels resulting in significant savings of time and resources. Note that experiments are run in triplicates for maximal statistical analysis.
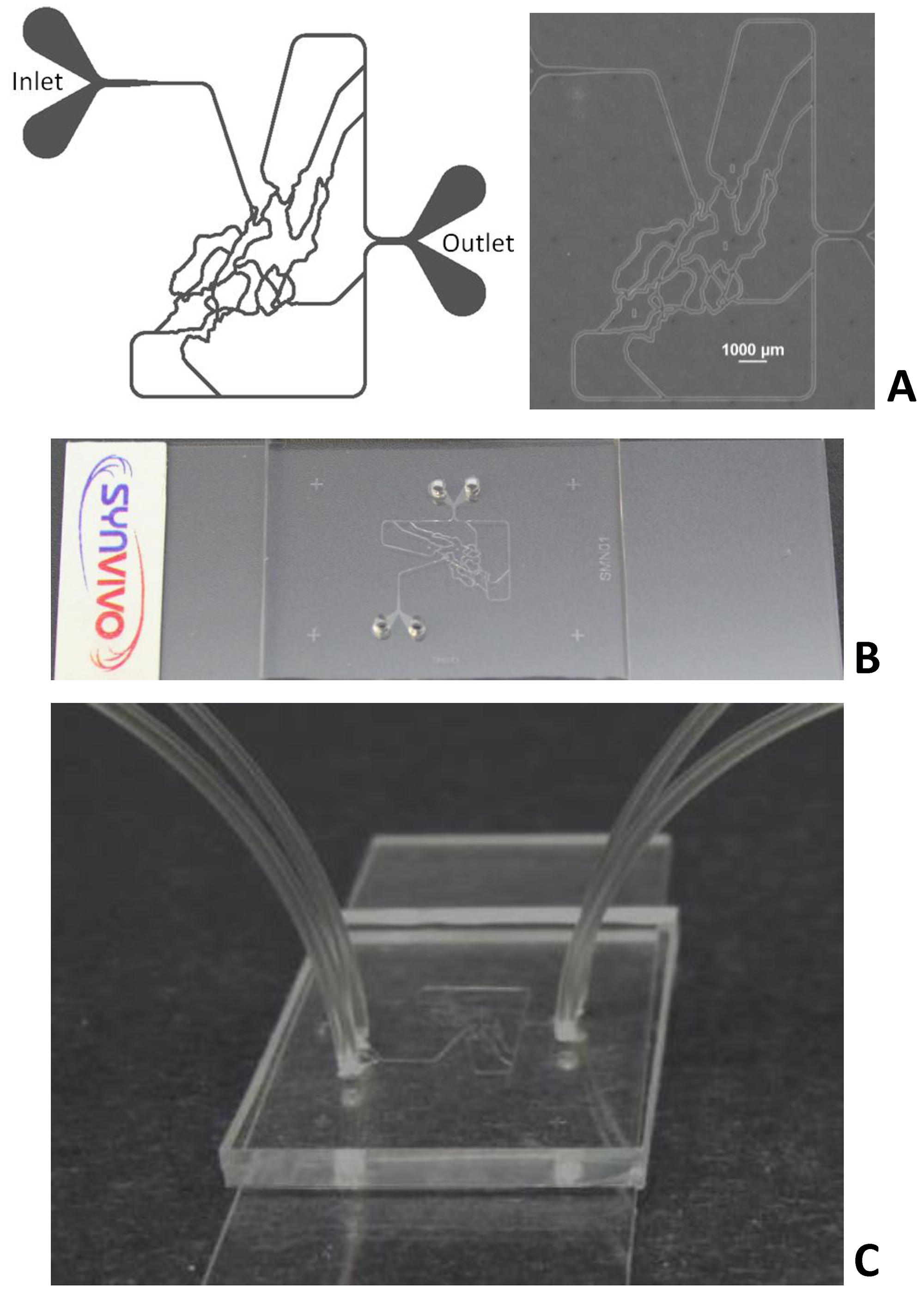
Figure 1. SynVivo-SMN devices. Left panel (Figure 1A) shows a schematic of the device. Right panel (Figure 1A) shows bright field image of the device. Figure 1B shows the SynVivo-SMN device mounted on microscope glass slide. Figure 1C shows the device with Tygon™ tubing attached to the inlet/outlet ports following priming with water.
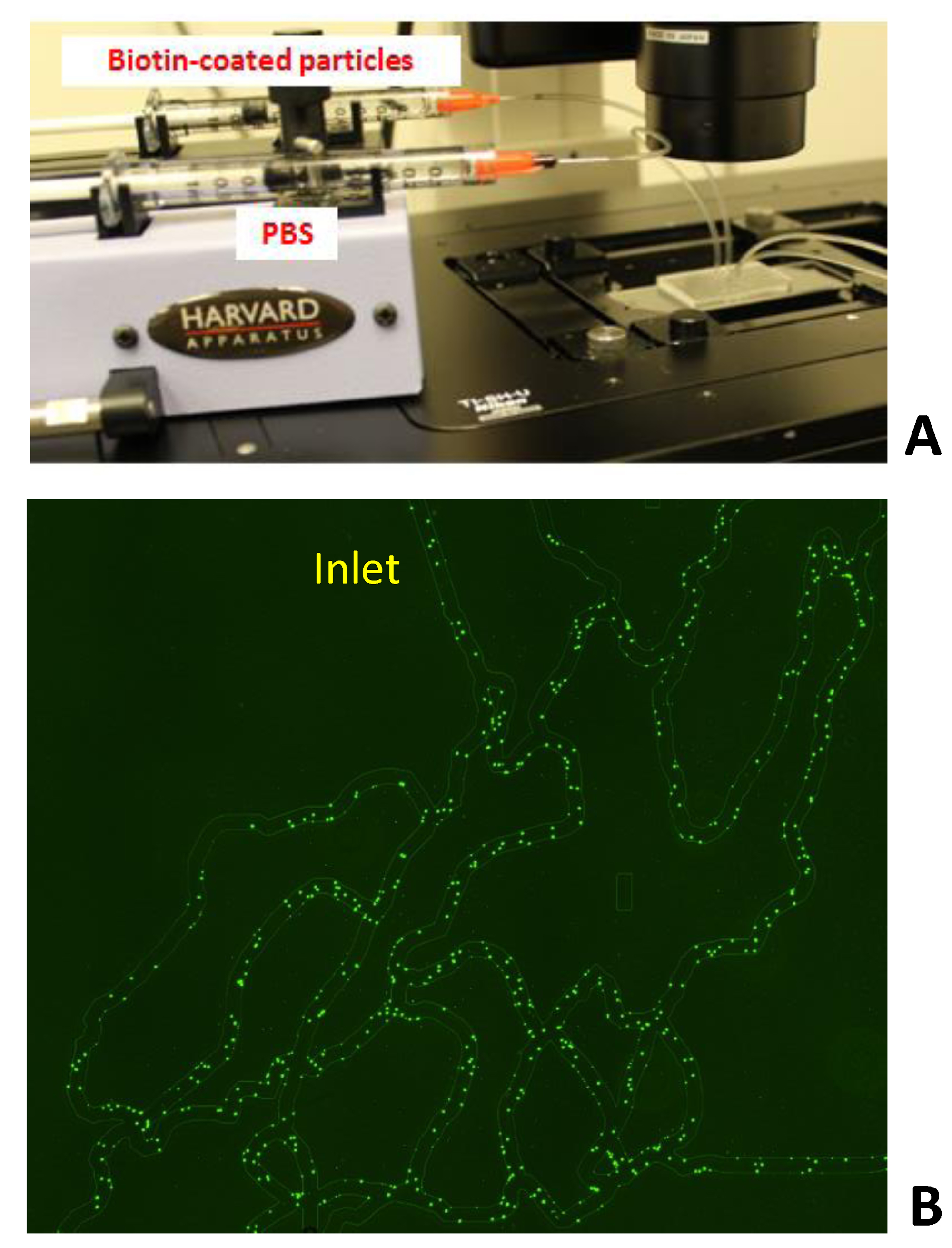
Figure 2. Typical adhesion assay in SynVivo-SMN. Figure 2A shows an image of the experimental-set up. Figure 2B shows biotinylated particles following binding in the device. Note that particle adhesion is found to be localized near bifurcation compared to the branches of the network.
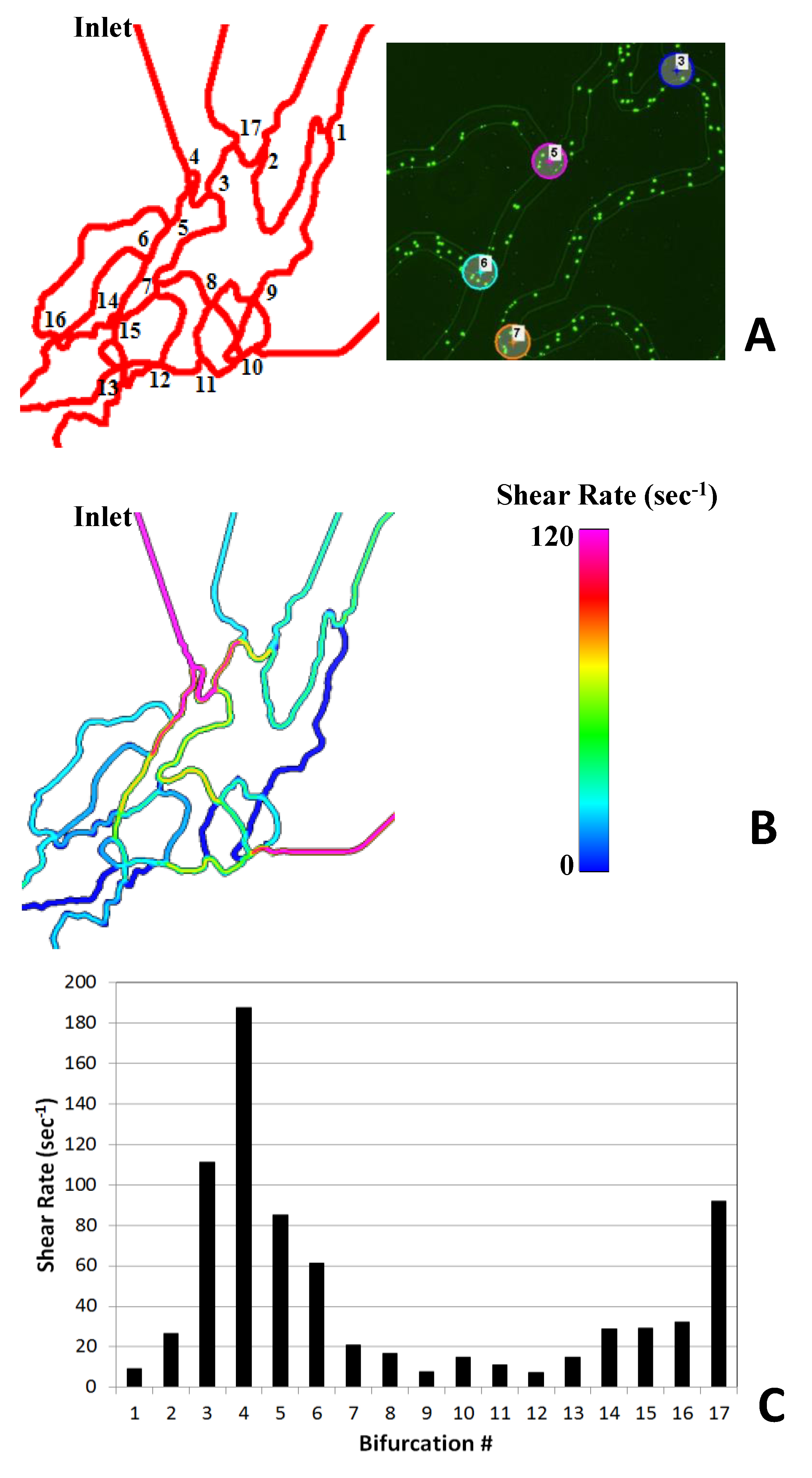
Figure 3. Shear analysis in SynVivo-SMN. Figure 3A shows all the bifurcations numbered in the network and a magnified view of bifurcations from the experiments. Figure 3B shows the qualitative wall shear map in the bifurcations in the network. Figure 3C shows the quantitative information on shear in the branches in the network. Note that the shear patterns in the network are complex and span the physiological ranges of shear rates (0-240 sec-1) found in vivo.
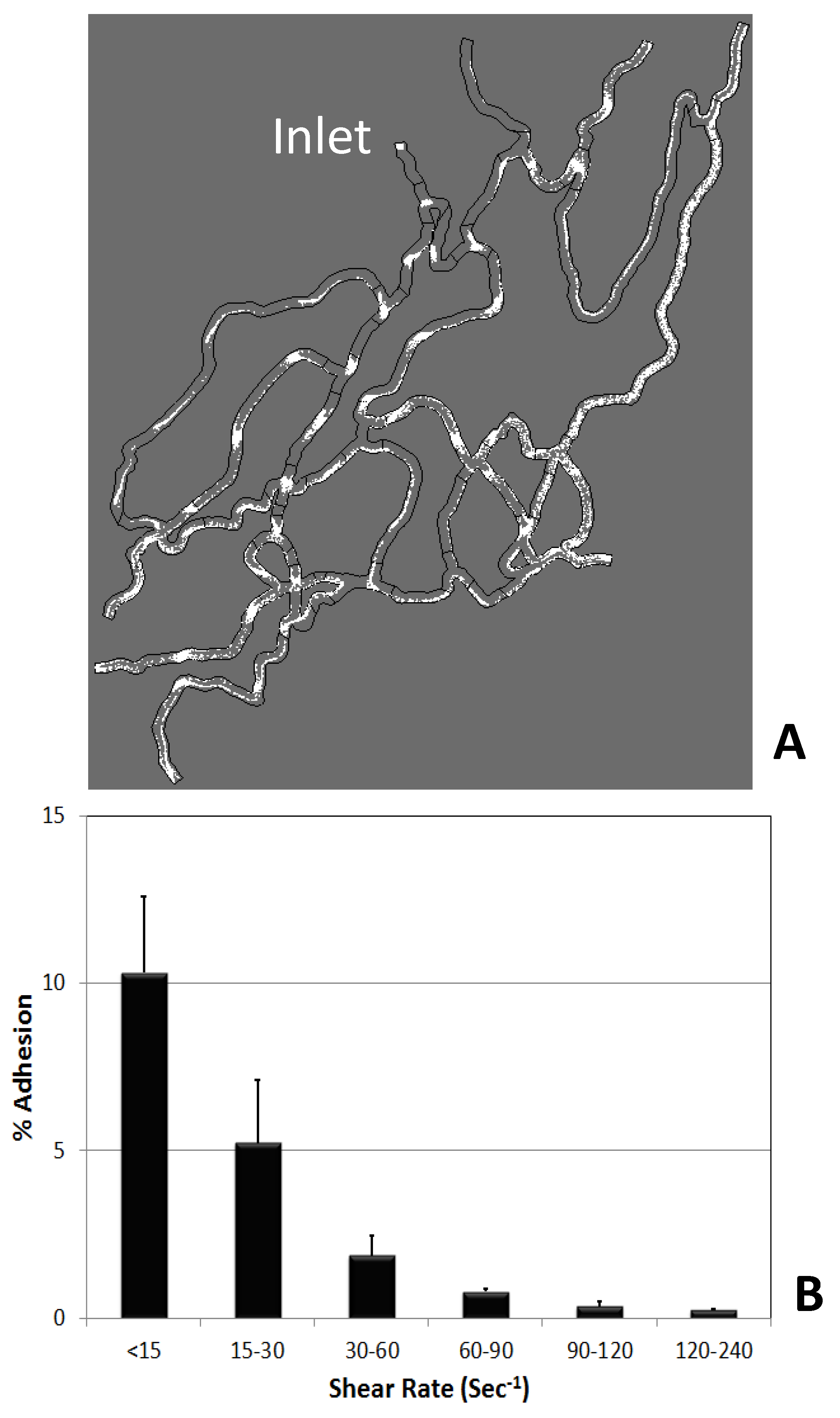
Figure 4. Generation of shear adhesion map. Figure 4A highlights the particle trajectories (shown in white) in the networks obtained from the CFD simulation. These trajectories are post-processed to obtain particle fluxes in the different branches and the bifurcations. Figure 4B shows the shear adhesion map obtained from a single SynVivo-SMN experiment.
Discussion
Parallel plate flow chambers, while providing significant insights into cell-cell and cell-particle interactions, suffer from several limitations such as high consumption of reagents and the need for multiple experimental runs to generate a shear adhesion map. The use of SynVivo-Synthetic Microvascular Networks (SynVivo-SMNs) enables the generation of a shear adhesion map from a single experiment in conditions mimicking in vivo conditions. In addition, significant savings (>95%) in reagents is also obtained.
The most important step in running particle adhesion experiments with SynVivo-SMN is to ensure bubble free conditions in the chip prior to experimentation. Introduction of bubbles would lead to flow instability and lack of access to “air locked” regions of the chip. Hence, great care needs to be taken during insertions and extractions of tubings from the ports of the device. In case of a bubble found in the device during the priming step, one can repeat the priming step again to remove the bubbles. Alternatively, one can flow in media/reagents at a low flow rates (0.1 µl/min or less) to ensure bubble free priming of the device.
The main limitation of the protocol is the chip being rendered unusable following the presence of a bubble that cannot be removed. However, with careful practice one can perform experiments with near 100% success. Also, the generation of shear adhesion map requires information from CFD models. However, a pre-computed database of CFD results for different flow conditions can easily overcome the need to perform the CFD simulations.
Although, the methodology presented here used the avidin-biotin system for ease of demonstration, any ligand-receptor combination on particles or cells can be used to be study particle-surface or cell-surface interactions in real-time in the device. Furthermore, cells can be cultured in the device to study particle-cell and cell-cell interactions. Culture of cells will require coating of the channels of the device with matrix suitable for desired cell types. For example, endothelial cells can be cultured on fibronectin coated channels. Following confluence, the endothelial cells can be activated using TNF-α or other relevant cytokines. White Blood Cells (WBCs) can be injected and their interactions on activated endothelial cells can be studied real-time. Similar to the protocol demonstrated in this study, a shear adhesion map of white blood cells can be easily generated from a single experiment.
The protocol mentioned in this paper can be used to study effect of network morphology on particle/cell adhesion as a model of in vivo conditions. In addition, the effect of flow on particle/cell adhesion and cellular behavior for modeling physiological conditions can also be readily evaluated. Finally, the protocol can be used to study drug delivery and efficacy for targeted delivery to desired cellular populations.
Divulgations
The authors have nothing to disclose.
Acknowledgements
SynVivo technology was developed under grant #2R44HL076034 from the NHLBI.
Materials
| SynVivo-SMN | CFD Research | SMN-001 | Exclusive at CFDRC |
| CFD-ACE+ | ESI Inc. | N/A | |
| Avidin | Invitrogen | 43-4401 | Any avidin source will work for this assay |
| Biotinylated Particles | Polysciences | 24173-1 | Any source of biotinylated particles will work for the assay |
| Tygon Tubing | VWR | 63018-044 | Size is typical for use with SynVivo-SMN |
| NIKON Elements | NIKON Instruments | N/A | Any other imaging software can be used |
References
- Weitz-Schmidt, G., Chreng, S. Cell adhesion assays. Methods Mol Biol. 757, 15-30 (2012).
- Parsons, S. A., Jurzinsky, C., Cuvelier, S. L., Patel, K. D. Studying leukocyte recruitment under flow conditions. Methods Mol Biol. 946, 285-300 (2013).
- Luscinskas, F. W., Gimbrone, M. A. Jr Endothelial-dependent mechanisms in chronic inflammatory leukocyte recruitment. Annu Rev Med. 47, 413-421 (1996).
- Adriani, G., et al. The preferential targeting of the diseased microvasculature by disk-like particles. Biomaterials. 33, 5504-5513 (2012).
- Decuzzi, P., et al. Flow chamber analysis of size effects in the adhesion of spherical particles. Int J Nanomedicine. 2, 689-696 (2007).
- Zou, X., et al. PSGL-1 derived from human neutrophils is a high-efficiency ligand for endothelium-expressed E-selectin under flow. Am J Physiol Cell Physiol. 289, 415-424 (2005).
- Sakhalkar, H. S., et al. Leukocyte-inspired biodegradable particles that selectively and avidly adhere to inflamed endothelium in vitro and in vivo. Proc Natl Acad Sci U S A. 100, 15895-15900 (2003).
- Van Kruchten, R., Cosemans, J. M., Heemskerk, J. W. Measurement of whole blood thrombus formation using parallel-plate flow chambers – a practical guide. Platelets. 23, 229-242 (2012).
- Ganguly, A., Zhang, H., Sharma, R., Parsons, S., Patel, K. D. Isolation of human umbilical vein endothelial cells and their use in the study of neutrophil transmigration under flow conditions. J Vis Exp. 66 (66), (2012).
- Shirure, V. S., Reynolds, N. M., Burdick, M. M. Mac-2 binding protein is a novel E-selectin ligand expressed by breast cancer cells. PLoS One. 7, (2012).
- Ploppa, A., Schmidt, V., Hientz, A., Reutershan, J., Haeberle, H. A., Nohé, B. Mechanisms of leukocyte distribution during sepsis: an experimental study on the interdependence of cell activation, shear stress and endothelial injury. Crit Care. 14, 201 (2010).
- Oh, H., Diamond, S. L. Ethanol enhances neutrophil membrane tether growth and slows rolling on P-selectin but reduces capture from flow and firm arrest on IL-1-treated endothelium. J Immunol. 181, 2472-2482 (2008).
- Resto, V. A., Burdick, M. M., Dagia, N. M., McCammon, S. D., Fennewald, S. M., Sackstein, R. L-selectin-mediated lymphocyte-cancer cell interactions under low fluid shear conditions. J Biol Chem. 283, 15816-15824 (2008).
- Enders, S., Bernhard, G., Zakrzewicz, A., Tauber, R. Inhibition of L-selectin binding by polyacrylamide-based conjugates under defined flow conditions. Biochim Biophys Acta. 1770, 1441-1449 (2007).
- Prabhakarpandian, B., Goetz, D. J., Swerlick, R. A., Chen, X., Kiani, M. F. Expression and functional significance of adhesion molecules on cultured endothelial cells in response to ionizing radiation. Microcirculation. 8, 355-364 (2001).
- Brown, D. C., Larson, R. S. Improvements to parallel plate flow chambers to reduce reagent and cellular requirements. BMC Immunology. 2, 9 (2001).
- Zheng, W., Zhang, W., Jiang, X. Precise control of cell adhesion by combination of surface chemistry and soft lithography. Adv Healthc Mater. 2, 95-108 (2013).
- Qian, T., Wang, Y. Micro/nano-fabrication technologies for cell biology. Med Biol Eng Comput. 48, 1023-1032 (2010).
- Biswas, A., Bayer, I. S., Biris, A. S., Wang, T., Dervishi, E., Faupel, F. Advances in top-down and bottom-up surface nanofabrication: techniques, applications & future prospects. Adv Colloid Interface Sci. 170, 2-27 (2012).
- Whitesides, G. M., Ostuni, E., Takayama, S., Jiang, X., Ingber, D. E. Soft lithography in biology and biochemistry. Annu Rev Biomed Eng. 3, 335-373 (2001).
- McDonald, J. C., et al. Fabrication of microfluidic systems in poly(dimethylsiloxane). Electrophoresis. 21, 27-40 (2000).
- Prabhakarpandian, B., et al. Synthetic microvascular networks for quantitative analysis of particle adhesion. Biomed Microdevices. 10, 585-595 (2008).
- Rosano, J. M., et al. A physiologically realistic in vitro model of microvascular networks. Biomed Microdevices. 11, 1051-1057 (2009).
- Tousi, N., Wang, B., Pant, K., Kiani, M. F., Prabhakarpandian, B. Preferential adhesion of leukocytes near bifurcations is endothelium independent. Microvasc Res. 80, 384-388 (2010).
- Prabhakarpandian, B., et al. Bifurcations: focal points of particle adhesion in microvascular networks. Microcirculation. 18, 380-389 (2011).

