Examination of Proteins Bound to Nascent DNA in Mammalian Cells Using BrdU-ChIP-Slot-Western Technique
Summary
In this protocol, we describe a novel BrdU-ChIP-Slot-Western technique to examine proteins and histone modifications associated with newly synthesized or nascent DNA.
Abstract
Histone deacetylases 1 and 2 (HDAC1,2) localize to the sites of DNA replication. In the previous study, using a selective inhibitor and a genetic knockdown system, we showed novel functions for HDAC1,2 in replication fork progression and nascent chromatin maintenance in mammalian cells. Additionally, we used a BrdU-ChIP-Slot-Western technique that combines chromatin immunoprecipitation (ChIP) of bromo-deoxyuridine (BrdU)-labeled DNA with slot blot and Western analyses to quantitatively measure proteins or histone modification associated with nascent DNA.
Actively dividing cells were treated with HDAC1,2 selective inhibitor or transfected with siRNAs against Hdac1 and Hdac2 and then newly synthesized DNA was labeled with the thymidine analog bromodeoxyuridine (BrdU). The BrdU labeling was done at a time point when there was no significant cell cycle arrest or apoptosis due to the loss of HDAC1,2 functions. Following labeling of cells with BrdU, chromatin immunoprecipitation (ChIP) of histone acetylation marks or the chromatin-remodeler was performed with specific antibodies. BrdU-labeled input DNA and the immunoprecipitated (or ChIPed) DNA was then spotted onto a membrane using the slot blot technique and immobilized using UV. The amount of nascent DNA in each slot was then quantitatively assessed using Western analysis with an anti-BrdU antibody. The effect of loss of HDAC1,2 functions on the levels of newly synthesized DNA-associated histone acetylation marks and chromatin remodeler was then determined by normalizing the BrdU-ChIP signal obtained from the treated samples to the control samples.
Introduction
Defective DNA repair and/or DNA replication are a major cause of genome instability, which can trigger cell death. A single unrepaired double strand break is sufficient to cause cell death1. Chromatin organization is transiently altered during both replication and repair2,3, and failure to maintain epigenetic information during these processes will result in a threat to genome integrity. Loss of HDAC3 or HDAC1,2 function impedes S-phase progression, DNA replication and repair leading to genotoxic stress (DNA damage) and cell death4-9. It is therefore a practical strategy to use selective HDAC inhibitors to disrupt replication, repair and chromatin in cancer cells and cause DNA damage, which in turn can stop growth and induce cell death selectively in rapidly growing cancer cells.
During DNA replication, chromatin is rapidly disassembled and then reassembled following DNA duplication. Newly synthesized histone H4 is acetylated at K5 and K12 residues (H4K5K12ac) and following deposition onto chromatin, they are rapidly deacetylated by histone deacetylases (HDACs)10,11. Failure of cells to maintain nascent chromatin integrity by histone deacetylation can lead to fork collapse, which in turn can result in DNA damage and cell death. We recently showed that selective inhibition of HDAC1,2 increases histone acetylation (H4K5ac, H4K12ac and H4K16ac) and inhibits SMARCA5 chromatin remodeler activity on nascent chromatin, which correlates with reduced replication fork progression, increased fork collapse and increased replication stress-induced DNA damage6. Thus, HDAC inhibitor treatment can alter nascent chromatin structure to trigger DNA damage and death very quickly in cancer cells, as these cells cycle rapidly and pass many times through the S-phase. It is therefore important to understand how HDACs function to regulate histone acetylation and protein binding to maintain DNA replication in mammalian cells.
To quantitatively measure the amount of histone acetylation and SMARCA5 chromatin remodeler associated with nascent DNA during DNA replication, we devised a modified ChIP assay called the BrdU-ChIP-Slot-Western technique. Following chromatin immunoprecipitation (ChIP) of a desired protein or histone modification, the amount of nascent DNA (labeled using thymidine analog, BrdU) in the ChIP sample can be detected using western analysis of BrdU-labeled ChIP DNA transferred onto a membrane using a Slot Blot apparatus. Using this technique, we showed that H4K16ac (a mark involved in chromatin packaging) and the ISWI family member SMARCA5 (SWI/SNF-related matrix-associated actin-dependent regulator of chromatin) chromatin remodeler are associated with nascent DNA in S-phase cells6. We also found that the nascent H4K16ac mark is deacetylated by HDAC1,2 during DNA replication6. H4K16ac inhibits chromatin remodeler activity of the ISWI family member SMARCA512. Hence, using the BrdU-CHIP-Slot-Western technique, we could connect the functions for HDAC1,2 in regulating chromatin remodeling during DNA replication. Hence, the BrdU-ChIP-Slot-Western Blot technique is a powerful approach to quantitatively measure the association and dissociation dynamics of proteins or their post-translational modifications that are bound to nascent DNA.
Protocol
1. Chromatin immunoprecipitation (ChIP) following BrdU Labeling
- Culture NIH3T3 cells in the presence of either DMSO or HDAC1,2-selective inhibitors as described previously6.
- Following treatment with DMSO or HDAC1,2 selective inhibitors, add BrdU to the cells in a sterile tissue culture hood. Incubate 2 x 106 NIH3T3 cells plated in 10 ml NIH3T3 media with a 20 µM final concentration of bromodeoxyuridine (BrdU) for 60 min in a sterile 37 °C tissue culture incubator.
- Crosslink intracellular proteins to DNA by adding formaldehyde directly to the culture media at a final concentration of 1%. Incubate 2 x 106 NIH3T3 cells with formaldehyde at RT for 10 min on a shaker.
- Quench the cross linking by the addition of glycine to a final concentration of 125 mM. Incubate at RT for 5 min on a shaker.
- Remove the media and collect cells in PBS in a 15 ml centrifuge tube. Spin cells at 956 x g for 5 min at 4 °C. Wash the cell pellet twice with ice-cold PBS. Remove PBS from the tube. Store the cross-linked cell pellet without PBS (the supernatant) at -80 °C until use.
- Prepare the lysate from the crosslinked cell pellet by adding 500 µl FA140 buffer (50 mM of HEPES KOH pH 7.5, 140 mM of NaCl, 1 mM of EDTA (pH 8.0), 1% Triton-X-100 and 0.1% sodium deoxycholate) supplemented with protease inhibitor cocktail.
- Shear the chromatin by sonicating the samples five times at 35% power and with a 15 sec pulse, 0.9 sec on and 0.1 sec off setting. Keep the tubes on ice after every round of sonication to avoid heating of the samples.
- Check the shearing on a DNA gel as the shearing efficiency will vary between different sonicators. To check sonication, reverse crosslink by adding NaCl to a concentration of 0.16 M and perform steps 1.17.1. – 1.22. Run eluted 10 µl DNA on a 1.5% agarose gel to check whether the shearing is between 300 – 700 bp with an average intensity at 500 bp.
- Following sonication, centrifuge samples for 10 min at 17,949 x g at 4 °C and transfer the supernatant to a new tube.
- Perform protein estimation using the commercial protein assay kit according to manufacturer's protocol. Use equal amount of lysate for ChIP of control and experimental samples. Generally, use 1 mg of total lysate per ChIP reaction.
- Prepare 50% slurry of protein A-agarose using FA140 buffer containing protease inhibitor cocktail. Add 1/10 volume of the protease inhibitor cocktail. Calculate the volume of beads needed based on the number of samples in total.
- Wash Protein A-agarose beads with FA140 buffer twice to remove the storage buffer and add an equal volume to FA140 buffer to the beads to make 50% slurry.
- To remove proteins that non-specifically bind to the beads, add 50 µl of 50% slurry of Protein A-agarose and incubate the samples at 4 °C with constant end-over-end rotation in a rotisserie mixer for 30 min.
- Spin the samples at 956 x g at 4 °C for a minute. Collect the supernatant and perform immunoprecipitation by adding either 8 μl H4K16ac antibody or 5 μl (5 μg) of 1 mg/ml SMARCA5 antibody to the supernatant. Incubate at 4 °C O/N with constant end-over-end rotation in a rotisserie mixer.
- Save 1/10th of the extract for 'input' control and set it as aside at -20 °C. The input needs to be reverse cross-linked along with the IP samples.
- Use equal amount of rabbit IgG (5 μl of 1 mg/ml) as the negative control immunoprecipitation sample.
- The following day, add 50 μl of 50% protein A-agarose slurry to the extract and incubate for 1 hr at 4 °C with rotation.
- Pellet agarose beads by centrifugation at 956 x g for 2 min at 4 °C, carefully remove the supernatant and wash the beads with the following buffers sequentially. In buffer preparations, add protease inhibitors right before use.
- Wash with Low Salt Immune Complex Wash Buffer (FA140; final concentration of 50 mM of HEPES KOH pH 7.5, 140 mM of NaCl, 1 mM of EDTA (pH 8.0), 1% Triton-X-100 and 0.1% Sodium deoxycholate) for 5 min at 4 °C with end-over-end rotation.
- Wash with High Salt Immune Complex Wash Buffer (FA500; final concentration of 50 mM of HEPES KOH pH 7.5, 500 mM of NaCl, 1 mM of EDTA (pH 8.0), 1% Triton-X-100 and 0.1% Sodium deoxycholate) for 5 min at 4 °C with end-over-end rotation.
- Wash with LiCl Immune Complex Wash Buffer (final concentration of 10 mM of Tris-Cl pH 8.0, 250 mM of LiCl, 0.5% NP40 and 0.5% Sodium deoxycholate) for 5 min at 4 °C with end-over-end rotation.
- Following washes as described above, add 200 µl elution solution (final concentration of 1% SDS and 100 mM Sodium bicarbonate) and incubate at RT for 15 min with end-over-end rotation at RT.
- Spin the samples at 956 x g at RT and transfer the eluate to a new 1.5 ml tube. Repeat step 1.15 with additional 200 µl elution solution.
- To 400 µl of the eluate, add NaCl to a concentration of 0.16 M and mix well.
- In addition, add 400 µl elution solution to 10 µl input that was set aside (step 1.11) and add NaCl to a concentration of 0.16 M to reverse crosslink input samples as well. Reverse crosslink the samples by incubating in 65 °C water bath for 5 hr.
- Add 1 ml of 100% EtOH to the reverse-crosslinked eluate. Mix well and incubate at -80 °C for 2 – 3 hr or at -20 °C O/N. Spin the samples at 17,949 x g for 15 min and discard ethanol.
- Add 800 µl 70% ethanol to wash the pellet. Spin at 17,949 x g for 15 min at 4 °C and discard ethanol. Spin briefly to remove any residual ethanol. Air-dry the samples at 37 °C for 10 min to remove residual ethanol.
- Dissolve DNA in 90 µl RNase and DNase free water and add 2 µl 10 mg/ml RNase at a 0.2 mg/ml final concentration. Incubate at 37 °C in water bath for 30 min.
- Add 10 µl 10x Proteinase K Buffer (100 mM Tris-Cl pH 8.0, 50 mM EDTA pH 8.0, 5% SDS). Mix well and add 1 µl Proteinase K. Incubate at 42 °C for 1 hr.
- Purify input and ChIP DNA using commercial PCR purification kit according to manufacturer's protocol and elute DNA in 50 µl water. Store DNA at -20 °C until further use.
2. Slot Blot Analysis of ChIP DNA
- Measure the yield of input and ChIP DNA eluted in 50 µl water as described in step 1.22 using a spectrophotometer at 280 nm according to manufacturer's protocol.
- Take equal volumes (50 µl) of input or immunoprecipitated DNA that are within the linear range of detection.
- For instance, if input DNA yield is 1.5 ng/µl, then take 30 µl (or 45 ng total DNA) and make volume to 50 µl with water. For factor ChIP, take the entire 50 µl eluate from Step 2.1 and proceed to step 2.3.
- Denature 50 µl input or immunoprecipitated DNA by adding 2.5 volumes of 0.4 N NaOH, briefly vortex and centrifuge for 956 x g for 10 sec, and incubate at RT for 30 min.
- Neutralize the denatured DNA by adding 175 µl of 1 M Tris-HCl, pH 6.8, vortex, centrifuge for 956 x g for 10 sec and place the samples on ice.
- Prepare a serial dilution of input and immunoprecipitated DNA for the Slot Blot assay as described below.
- Follow steps 2.3 – 2.4 to get a total sample volume of 350 µl (i.e., 50 µl Input/ChIP DNA, 125 µl 0.4 N NaOH, 175 µl 1 M Tris-HCl, pH 6.8).
Note: For the instance stated above, 45 ng of input DNA will end up being 0.129 ng/ µl. - For input DNA, label three tubes as 100, 50 and 25 and add 100 µl, 50 µl and 25 µl of the denatured and neutralized DNA sample. Make the final volume to 100 µl with nuclease-free water. For ChIP DNA, label three tubes as 200, 100 and 50 and add 200 µl, 100 µl and 50 µl of the denatured and neutralized DNA sample. Make final volume to 200 µl with nuclease free water.
- Follow steps 2.3 – 2.4 to get a total sample volume of 350 µl (i.e., 50 µl Input/ChIP DNA, 125 µl 0.4 N NaOH, 175 µl 1 M Tris-HCl, pH 6.8).
- Cut the membrane to the size of the Slot Blot apparatus. Pre-wet the membrane and blotting papers in 20x SSC before and prior to adding DNA. Assemble them into the Slot Blot apparatus according to the manufacturer's protocol.
- Pipet DNA from step 2.5.2 into appropriate slots. Apply vacuum slowly to pull the DNA onto the zeta-probe membrane. Stop vacuum after all samples have passed through the apparatus.
- Disassemble the apparatus and immediately immobilize the DNA onto the membrane using a UV crosslinker. Have the DNA side up in the UV crosslinker and use the autocrosslink setting in the crosslinker. Follow manufacturer's protocol.
- Detect BrdU on the slot blot by performing western blotting.
- Block the membrane with 5% non-fat dry milk in PBS containing 0.1% Tween-20 (PBST) for 1 hr at RT to prevent the non-specific binding of antibodies to the membrane.
- Add primary anti-BrdU antibody (1:500 dilution) diluted in 1% non-fat dry milk in PBST and incubate the blot with the primary antibody for 3 hr at RT on a shaker. Wash the membrane with PBST three times after the primary antibody incubation.
- Add HRP-conjugated anti-mouse secondary antibody to the blot (1:5,000 dilution) and incubate at RT for 1 hr. Wash the membrane with PBST three times after the secondary antibody incubation.
NOTE: Use enough volume of the antibodies to completely cover the membrane during primary and secondary antibody incubations.
- Prepare the ECL mix and add it to the blot according to the manufacturer's protocol. Incubate the membrane with ECL mix for 5 min at RT.
- Place the membrane in an X-ray cassette, expose the membrane to the X-ray film and develop the blot using the manufacturer's protocol.
- Quantify the signal obtained for input and ChIPed DNA after scanning the blots using the Image J software according to manufacturer's protocol. For quantitation, use the signal that is within the linear range of detection in order to ensure accuracy of measurement.
Representative Results
To determine the specificity of HDAC1,2-selective inhibitors, Hdac1,2FL/FL and Hdac3FL/FL fibrosarcoma cells were used. Adenovirus-containing Cre recombinase (Ad-Cre) was used to delete Hdac1,2 and Hdac3 in these cells. Following Ad-Cre infection of Hdac1,2FL/FL cells, a robust increase in H4K5ac was observed. Treatment of Hdac1,2 knockout cells with 233 or 898 did not result in any further increase in H4K5ac confirming that 233 and 898 inhibit Hdac1,2 and not Hdac3 in these cells. In contrast, addition of 233 or 898 to Hdac3 knockout cells resulted in a significant increase in H4K5ac compared to the increase seen in Hdac3 knockout cells due to an additive effect on H4K5ac levels as a result of inhibition of Hdac1,2 and 3. Hence, 233 and 898 are Hdac1, 2-selective inhibitors. These results are shown in Figure 1.
To validate the BrdU-ChIP-Slot technique, BrdU-pulse chase analysis was performed to look at the kinetics of PCNA loading on to the nascent DNA in HeLa cells. Our results showed that PCNA association with nascent DNA occurs rapidly within 15 min and disappears after a 30 min chase in agreement with previously published results13. These results are shown in Figure 2.
BrdU-H4K16ac ChIP-Slot-Western technique was used to determine the amount of H4K16ac associated with nascent DNA in the absence of HDAC1,2 function. A robust enrichment in H4K16ac associated with nascent DNA was observed when compared to the rabbit IgG control (Figures 3A and 3B). The increase in nascent DNA-associated H4K16ac following inhibition of HDAC1,2 activities or knockdown of Hdac1,2 is shown in Figure 3C, 3D, and 3E.
The level of SMARCA5 chromatin remodeler on nascent DNA was determined using BrdU-SMARCA5 ChIP-Slot-Western technique. Our results showed that SMARCA5 associates with nascent DNA in mammalian cells. Furthermore, HDAC1,2 inhibition or knockdown of Hdac1,2 did not change the amount of nascent DNA-associated SMARCA5 chromatin remodeler as shown in Figure 4.
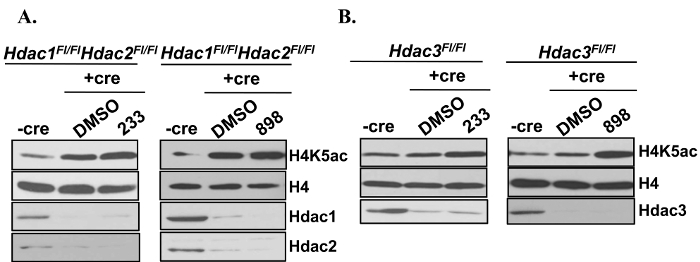
Figure 1. Confirmation of the Specificity of HDAC1,2-selective Inhibitors. Western analysis of whole cell lysates prepared from Hdac1FL/FL Hdac2FL/FL or Hdac3FL/FL fibrosarcoma cells following Ad-Cre infection and treatment with 898 or 233. Cells were treated with 3 µM 898 or 233 for 24 hr following a 48 hr Ad-Cre infection. This figure is derived from our previous published work6. Please click here to view a larger version of this figure.
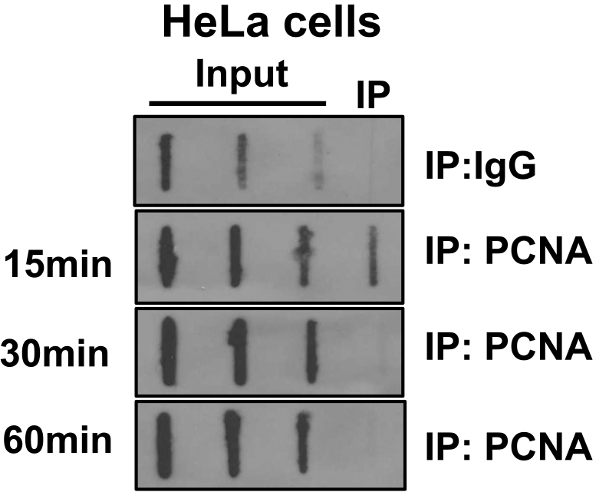
Figure 2. Dynamics of PCNA Association with the Nascent DNA in HeLa Cells. HeLa cells were labeled with bromodeoxyuridine (BrdU) for 30 min. Cells were then washed to remove unincorporated BrdU and cultured in media without BrdU for indicated periods of time (chase). Chromatin immunoprecipitation (ChIP) was performed with anti-proliferating cell nuclear antigen (PCNA) antibody. BrdU labeled DNA present in input DNA and those associated with PCNA were assessed in slot blot analysis using an anti-BrdU antibody as shown in Figure 2. This figure is derived from our previous published work6. Please click here to view a larger version of this figure.
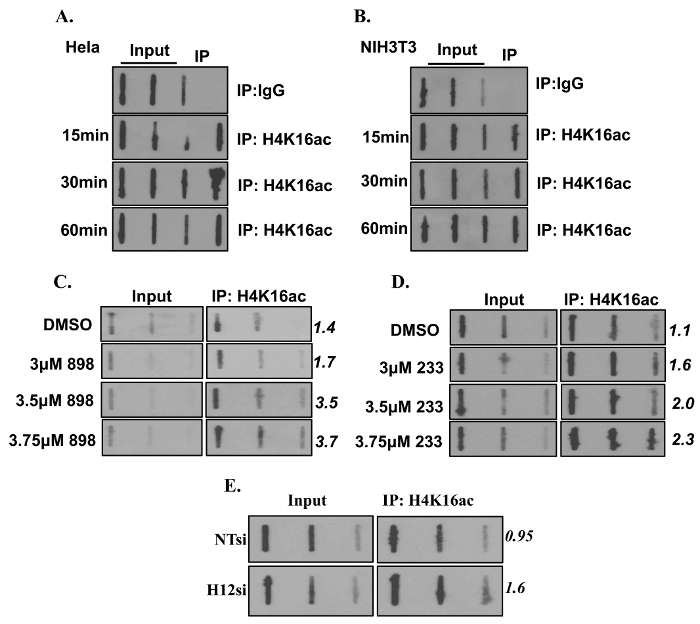
Figure 3. Loss of Histone Deacetylase 1 and 2 Increases H4K16ac on Nascent DNA. (A-B) Bromodeoxyuridine (BrdU) pulse chase was performed in HeLa and NIH3T3 cells to determine association of H4K16ac with nascent DNA at the indicated time points. (C-D) NIH3T3 cells were treated with either DMSO or a HDAC1,2-selective inhibitor (898 or 233) for 24 hr. (E) NIH3T3 cells were either transfected with non-targeting (NT) or Hdac1,2 (H12) siRNA. Cells from (C – E) were labeled with BrdU and used for ChIP with anti-H4K16ac followed by Slot blotting. The membrane was probed with anti-BrdU antibody. Numbers indicate to Image J quantitation of the average BrdU signal of high and medium volume of ChIP DNA spotted. This figure is derived from our previous published work6. Please click here to view a larger version of this figure.
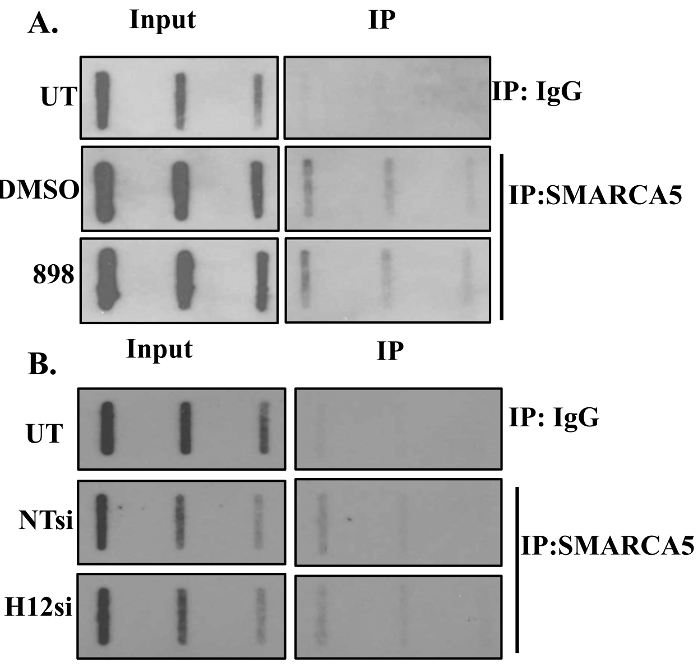
Figure 4. SMARCA5 Associates with Nascent DNA. The amount of SMARCA5 on nascent chromatin following loss of HDAC1,2 function is shown. NIH3T3 cells were either treated with an HDAC1,2-selective inhibitor (898) or transfected with non-targeting (NT) or Hdac1,2 (H12) siRNA. Cells were labeled with BrdU following the above-mentioned treatments and ChIP with anti-SMARCA5 or rabbit IgG (negative control) was performed. Increasing volumes of ChIP DNA was spotted on the slot blot and the membrane was probed with anti-BrdU antibody. This figure is derived from our previous published work6. Please click here to view a larger version of this figure.
Discussion
The protocol described in this manuscript is a relatively quick method to demonstrate the presence of proteins or their post-translationally modified forms on newly replicated or nascent DNA. Additionally, this technique permits one to measure the association-dissociation kinetics of a protein or its modified form with nascent DNA. This technique is complementary to the elegant iPOND technology13. In the iPOND technology, newly synthesized DNA is labeled with ethyl deoxyuridine (EdU). A biotin conjugate is then added onto EdU using the click chemistry. Biotin-tagged nascent DNA is then immunoprecipitated using streptavidin beads and co-purifying proteins are detected by western blotting. On the other hand, in our BrdU-ChIP-Slot-Western technique, a protein or its modified form associated with nascent DNA is immunoprecipitated using protein-specific or modified form-specific antibody and the amount of BrdU-labeled nascent DNA bound to the protein is then determined quantitatively by Slot-Western blotting using an anti-BrdU antibody.
There are critical steps in this protocol that needs special attention. It is critical to ensure that the antibody of interest works well in standard ChIP assays with a high efficiency. It is important to test the efficiency of an antibody in ChIP assays by quantitative-PCR before performing the BrdU-CHIP-Slot-Western assay. In order for the quantitation of the BrdU-CHIP-Slot-Western technique to be accurate, a serial dilution of the BrdU-labeled DNA should be applied to slot blot and western analysis using the anti-BrdU antibody should be initially performed in order to determine the linear range of detection. Determining the DNA concentration of ChIP and Input DNA enables one to ensure that the amount of BrdU-labeled DNA is within the linear range of detection in Western blotting. For example, we determined 50 ng to 12.5 ng of input DNA to be in the linear range in our pilot experiment. Also, if the concentration of the ChIP samples is very high, it is imperative to dilute the ChIP DNA before performing slot. The limitation of this technique is its resolution, which is dependent on the efficiency of chromatin shearing. The shearing of chromatin by sonication determines the proximity of a given protein or its modified form to the newly synthesized DNA.
In the past, the study of changes in the chromatin and chromatin-modifying enzymes during DNA replication in mammalian cells remained a difficult question to address due to the unavailability of appropriate techniques. Moreover, since Hdac1,2-null cells arrest in G1 phase, it was difficult to examine functions for Hdac1,2 within S-phase. In our study, we showed functions for these two Hdacs during DNA replication using a combination of first-in-class selective inhibitors to inhibit their activities within S-phase and the novel BrdU-ChIP-Slot-Western technique. In future, this technique could also be used to study the dynamics of DNA damage response and DNA repair proteins at the stalled fork in addition to studying the histone modifications and chromatin-modifying enzyme at the fork. Moreover, this technique is not limited to mouse NIH3T3 cells. This technique can be successfully used in other cell lines in order to investigate how other inhibitors of enzymes or proteins affect DNA replication and replication stress-induced chromatin changes at the replication fork.
Divulgations
The authors have nothing to disclose.
Acknowledgements
The work in this manuscript was supported by funds from Dept. of Radiation Oncology and Huntsman Cancer Institute and the National Institute of Health grant (R01-CA188520) to SB. I thank Danielle Johnson and Steven Bennett in my lab for demonstrating the protocol and explaining its benefits. I am grateful to Dr. Mahesh Chandrasekharan (Huntsman Cancer Institute) for critical comments on the manuscript.
Materials
| Anti-BrdU (westerns) | BD Biosciences | B555627 | |
| Anti-SMARCA5 | Abcam | ab3749 | |
| Anti-H4K16ac | Active Motif | 39167 | |
| Zeta-Probe GT Membrane | Bio-Rad | 162-0197 | |
| Formaldehyde | Fisher | BP531-500 | |
| Protein A agarose | Millipore | 16-156 | |
| Rnase A | Qiagen | 19101 | |
| Proteinase K | Sigma | P4850 | |
| PCR Purification Kit | Qiagen | 28106 | |
| Glycine | Sigma | G7403 | |
| Protease inhibitor cocktail | Roche | 11836170001 | |
| BrdU | Sigma | B9285 | |
| Rabbit IgG | Millipore | 12-370 | |
| HEPES | Sigma | H3375 | |
| NaCl | Sigma | S-3014 | |
| Triton-X-100 | Sigma | 93443 | |
| NP40 | USB Corporation | 19628 | |
| Sodium deoxycholate | Sigma | D6750 | |
| Sodium bicarbonate | Sigma | S-4019 | |
| Ethanol | Decon Laboratories | 04-355-222 | |
| DEPC-treated water | Sigma | 95284 | |
| Tris | Fisher | BP-152-5 | |
| EDTA | Invitrogen | 15575-020 | |
| SDS | Ambion | AM9820 | |
| Sodium hydroxide | Mallinckrodt GenAR | MAL7772-06 | |
| 20X SSC | Life Technologies | 15557-036 | |
| Non-fat dry milk | Lab Scientific Inc | M0841 | |
| ECL | Thermo Fisher | 80196 | |
| X-ray film | Genesee Sci | 30-101 | |
| Developer | Konica Minolta | SRX-101A | |
| UV Crosslinker | Stratagene | XLE-1000 | |
| Sonicator | Branson Sonifier Digital Ultrasonic Cell Disruptor | Model: 450 | |
| Centrifuge | Eppendorf | Model: 5810R | |
| Water bath | Fisher Scientific Isotemp | Model: 2322 | |
| NanoDrop 1000 spectrophotometer | ThermoScientific | Model: 1000 | |
| Slot Blot apparatus | Schleicher and Schuell Minifold II | 44-27570 | |
| Tissue Culture Incubator | Thermo Scientific | Series II 3110 Water-Jacketed CO2 Incubators |
References
- Podhorecka, M., Skladanowski, A., & Bozko, P. H2AX Phosphorylation: Its Role in DNA Damage Response and Cancer Therapy. J nucleic acids. 2010 (2010).
- Groth, A., Rocha, W., Verreault, A., & Almouzni, G. Chromatin challenges during DNA replication and repair. Cell. 128, 721-733 (2007).
- Demeret, C., Vassetzky, Y., & Mechali, M. Chromatin remodelling and DNA replication: from nucleosomes to loop domains. Oncogene. 20, 3086-3093 (2001).
- Bhaskara, S. et al. Deletion of histone deacetylase 3 reveals critical roles in S phase progression and DNA damage control. Mol Cell. 30, 61-72 (2008).
- Bhaskara, S., & Hiebert, S. W. Role for histone deacetylase 3 in maintenance of genome stability. Cell Cycle. 10, 727-728 (2011).
- Bhaskara, S. et al. Histone deacetylases 1 and 2 maintain S-phase chromatin and DNA replication fork progression. Epigenetics Chromatin. 6, 27 (2013).
- Bhaskara, S. et al. Hdac3 is essential for the maintenance of chromatin structure and genome stability. Cancer Cell. 18, 436-447 (2010).
- Johnson, D. P. et al. HDAC1,2 inhibition impairs EZH2- and BBAP-mediated DNA repair to overcome chemoresistance in EZH2 gain-of-function mutant diffuse large B-cell lymphoma. Oncotarget. 6, 4863-4887 (2015).
- Bhaskara, S. Histone deacetylases 1 and 2 regulate DNA replication and DNA repair: potential targets for genome stability-mechanism-based therapeutics for a subset of cancers. Cell Cycle. 14, 1779-1785 (2015).
- Taddei, A., Roche, D., Sibarita, J. B., Turner, B. M., & Almouzni, G. Duplication and maintenance of heterochromatin domains. J Cell Biol. 147, 1153-1166 (1999).
- Alabert, C., & Groth, A. Chromatin replication and epigenome maintenance. Nat Rev Mol Cell Biol. 13, 153-167 (2012).
- Clapier, C. R., & Cairns, B. R. Regulation of ISWI involves inhibitory modules antagonized by nucleosomal epitopes. Nature. 492, 280-284 (2012).
- Sirbu, B. M. et al. Analysis of protein dynamics at active, stalled, and collapsed replication forks. Genes Dev. 25, 1320-1327 (2011).

