Simultaneous Flow Cytometric Characterization of Multiple Cell Types Retrieved from Mouse Brain/Spinal Cord Through Different Homogenization Methods
Summary
We present a flow cytometry method to identify simultaneously different cell types retrieved from mouse brain or spinal cord. This method could be exploited to isolate or characterize pure cell populations in neurodegenerative diseases or to quantify the extent of cell targeting upon in vivo administration of viral vectors or nanoparticles.
Abstract
Recent advances in viral vector and nanomaterial sciences have opened the way for new cutting-edge approaches to investigate or manipulate the central nervous system (CNS). However, further optimization of these technologies would benefit from methods allowing rapid and streamline determination of the extent of CNS and cell-specific targeting upon administration of viral vectors or nanoparticles in the body. Here, we present a protocol that takes advantage of the high throughput and multiplexing capabilities of flow cytometry to allow a straightforward quantification of different cell subtypes isolated from mouse brain or spinal cord, namely microglia/macrophages, lymphocytes, astrocytes, oligodendrocytes, neurons and endothelial cells. We apply this approach to highlight critical differences between two tissue homogenization methods in terms of cell yield, viability and composition. This could instruct the user to choose the best method depending on the cell type(s) of interest and the specific application. This method is not suited for analysis of anatomical distribution, since the tissue is homogenized to generate a single-cell suspension. However, it allows to work with viable cells and it can be combined with cell-sorting, opening the way for several applications that could expand the repertoire of tools in the hands of the neuroscientist, ranging from establishment of primary cultures derived from pure cell populations, to gene-expression analyses and biochemical or functional assays on well-defined cell subtypes in the context of neurodegenerative diseases, upon pharmacological treatment or gene therapy.
Introduction
Gene and drug delivery technologies (such as viral vectors and nanoparticles) have become a powerful tool that can be applied to gain better insights on specific molecular pathways altered in neurodegenerative diseases and for development of innovative therapeutic approaches1,2,3. Optimization of these tools relies on quantification of: (1) the extent of penetration in the CNS upon different routes of administration and (2) targeting of specific cell populations. Histological analyses are usually applied to visualize fluorescent reporter genes or fluorescently-tagged nanoparticles in different CNS areas and across different cell types, identified by immunostaining for specific cell markers4,5. Even though this approach provides valuable information on the biodistribution of the administered gene or drug-delivery tools, the technique can be time-consuming and labor-intense since it requires: (1) tissue fixation, cryopreservation or paraffin-embedding and slicing; (2) staining for specific cellular markers sometimes requiring antigen retrieval; (3) acquisition by fluorescence microscopy, which usually allows the analysis of a limited number of different markers within the same experiment; (4) image processing to allow proper quantification of the signal of interest.
Flow cytometry has become a widely used technique which takes advantage of very specific fluorescent markers to allow not only a rapid quantitative evaluation of different cell phenotypes in cell suspensions, based on expression of surface or intracellular antigens, but also functional measurements (e.g., rate of apoptosis, proliferation, cell cycle analysis, etc.). Physical isolation of cells through fluorescent activated cell sorting is also possible, allowing further downstream applications (e.g., cell culture, RNAseq, biochemical analyses etc.)6,7,8.
Tissue homogenization is a critical step necessary to obtain a single cell suspension to allow reliable and reproducible downstream flow cytometric evaluations. Different methods have been described for adult brain-tissue homogenization, mainly with the aim to isolate microglia cells9,10,11; they can be overall classified in two main categories: (1) mechanical dissociation, which uses grinding or shearing force through a Dounce homogenizer (DH) to rip apart cells from their niches and form a relatively homogenized single cell suspension, and (2) enzymatic digestion, which relies on incubation of minced tissue chunks at 37 °C in the presence of proteolytic enzymes, such as trypsin or papain, favoring the degradation of the extracellular matrix to create a fairly homogenized cell suspension12.
Regardless of which method is utilized, a purification step is recommended after tissue homogenization to remove myelin through centrifugation on a density gradient or by magnetic selection9,12, before moving to the downstream applications.
Here, we describe a tissue processing method based on papain digestion (PD) followed by purification on a density gradient, optimized to obtain viable heterogeneous cell suspensions from mouse brain or spinal cord in a time-sensitive manner and suitable for flow cytometry. Moreover, we describe a 9-color flow cytometry panel and the gating strategy we adopted in the laboratory to allow the simultaneous discrimination of different CNS populations, live/dead cells or positivity for fluorescent reporters such as green fluorescent protein or rhodamine dye. By applying this flow cytometric analysis, we can compare different methods of tissue processing, i.e., PD versus DH, in terms of preservation of cellular viability and yields of different cell types.
The details we provide herein can instruct decision on the homogenization protocol and the antibody combination to use in the flow cytometry panel, based on the specific cell type(s) of interest and the downstream analyses (e.g., temperature-sensitive applications, tracking of specific fluorescent markers, in vitro culture, functional analyses).
Protocol
All methods described here have been approved by the Institutional Animal Care and Use Committee (IACUC) of Dana Farber Cancer Institute (protocol number 16-024).
1. Preparation of solutions needed for the experiment
- Prepare 1x Hank’s balanced salt solution (HBSS) by diluting 10x HBSS with sterile water. Pre-chill the solution on ice. At least 25 mL of solution are needed for each sample.
- Prepare isotonic Percoll solution (IPS) by mixing 10x sterile HBSS 1:10 with density gradient medium (i.e., Percoll). Pre-chill on ice.
NOTE: IPS can be stored for up to 30 days at 4 °C. - Prepare flow cytometry (FACS) blocking (BL) solution (1% bovine serum albumin [BSA], 5% fetal bovine serum [FBS] in phosphate-buffered saline [PBS]). Pre-chill on ice.
2. Animal euthanasia by intracardiac perfusion and tissue dissection
NOTE: Eight-week-old C57BL/6J mice, either sex, were used in the experiments. Perfusion with PBS solution is performed to eliminate blood contamination from organs, before proceeding with tissue digestion.
- Anesthetize the mouse by using a mixture of ketamine/xylazine (90−200 mg/kg ketamine, 10 mg/kg xylazine). Place the mouse on its back and tape each limb down to the support. Verify adequate depth of anesthesia by checking the withdrawal reflex.
- Make a midline skin incision at the level of the thoracic inlet to expose the sternum. Use forceps to grasp the tip of the sternum, then make one 1 cm incision on each side of the rib cage. Finally cut through the diaphragm and open the sternum widely enough to visualize the heart.
- Use forceps to gently grasp the heart by the right ventricle and lift it to the midline and slightly out of the chest.
- Insert a 23 G butterfly needle into the tip of the left ventricle, towards the aorta and hold firmly.
- Start the perfusion with 1x PBS. Pierce through the right auricle using scissors to allow the perfusate to exit the circulation. Set the flow rate of PBS at 3 mL/min. Perfuse with at least 15 mL of 1x PBS to ensure tissues are clear.
NOTE: Blanching of the liver and mesenteric blood vessels are signs of good perfusion. If necessary, the volume of prefusion can be increased up until the fluid exiting the heart is clear of blood, at which point the flush line can be stopped. - After perfusion, sever the brain from the spinal cord and remove the brain from the skull with scissors and forceps. Remove the fur to increase visibility and control during the dissection and to avoid carrying over hair contaminants. Flush the spinal cord out of its column by using a 3 mL syringe filled with PBS.
- Transfer each tissue in a well of a 6-well multi-well plate prefilled with 2 mL of ice-cold 1x HBSS and keep on ice until digestion.
- Divide the brain and the spinal cord into two halves, along the longitudinal line.
NOTE: One half of each tissue is homogenized (see sections below) to allow flow cytometric analyses; the other half can be assigned to different processing for alternative analyses (e.g., dipped in paraformaldehyde fixative solution for histology).
3. Enzymatic digestion of brain and spinal cord
NOTE: Volumes described in this section are enough for digestion of one-half brain or spinal cord.
- Use a pair of scissors to mince the tissues into 1−2 mm thick pieces.
- Cut the tip of a 1000 µL pipette with a pair of scissors to make it sufficiently large to allow the collection of the tissue pieces. Pre-rinse the pipette tip with 1x HBSS. Then use the pipette to transfer the 2 mL of HBSS solution containing the minced tissue to a 15 mL conical tube.
NOTE: Pre-rinsing of the pipette tip is important to prevent stickiness of the tissue pieces inside the tip. - Wash the well with additional 2 mL of ice-cold 1x HBSS and transfer the solution to the corresponding 15 mL conical tube containing the tissue pieces.
- Centrifuge each sample for 5 min at 250 x g at 4 °C.
- Prepare enzyme mix 1 of the neural tissue dissociation kit (NTDK; Table of Materials) by mixing 50 µL of enzyme P with 1900 µL of buffer X per sample. Warm enzyme mix 1 at 37 °C in a water bath. Incubate enzyme mix 1 at 37 °C for at least 10 min before use in order to allow for the full activation of the enzyme.
- Aspirate the supernatant from the 15 mL conical tube and add 1.95 mL of enzyme mix 1 to each sample. Gently vortex to make sure the pellet is resuspended.
- Incubate the samples on a wheel or shaker for 15 min at 37 °C.
- In the meanwhile, prepare enzyme mix 2 of the NTDK by mixing 10 µL of enzyme A with 20 µL of buffer Y per sample; pre-warm the solution at 37 °C in a water bath.
- At the end of the incubation with enzyme mix 1, add 30 µL of enzyme mix 2 to each sample.
- Gently mix the samples by pipetting up and down with a 1000 µL pipette tip pre-rinsed with 1x HBSS.
- Incubate the sample on a wheel or shaker for 15 min at 37 °C.
- After incubation, add 10 mL of ice-cold 1x HBSS to each tube to inactivate enzyme mix 1 and enzyme mix 2.
- Centrifuge each sample for 10 min at 320 x g at 4 °C.
- Discard the supernatant; add ice-cold 1x HBSS to each tube up to a final volume of 7 mL and gently resuspend the pellet by vortexing.
- Continue to section 5 for debris removal.
4. Mechanical homogenization of brain and spinal cord
NOTE: Volumes described in this section are enough for homogenization of one-half brain or spinal cord. The protocol described in this section can be used as a method alternative to the one described in section 3, depending on user need as discussed below.
- Pre-chill the glass mortar of the Dounce tissue grinder (Table of Materials) set on ice.
- Add 3 mL of pre-chilled 1x HBSS to the mortar.
- Transfer the tissue (brain or spinal cord) from the well of the 6-well plate into the glass mortar making sure it is dipped in 1x HBSS and sits at the bottom of the mortar.
- Gently smash the tissue with 10 strokes of pestle A followed by 10 strokes of pestle B. Transfer the homogenized mix into a new 15 mL conical tube.
- Fill the tube to a final volume of 10 mL by using pre-chilled 1x HBSS and centrifuge for 10 min at 320 x g at 4 °C.
- Aspirate the supernatant and add ice-cold 1x HBSS to each tube up to a final volume of 7 mL and gently resuspend the pellet by vortexing.
- Continue to section 5 for debris removal.
5. Debris removal
NOTE: Removal of debris, composed mainly of undigested tissue and myelin sheaths, is a critical step to allow efficient staining of the tissue homogenate for subsequent flow cytometric analyses.
- Filter each sample through a 70 µm cell strainer to remove any undigested tissue chunk. This step is particularly important especially when working with spinal cord tissues since these samples are more likely to contain undigested nerve fragments or meninges that could affect the subsequent steps.
- Make sure that the final volume is 7 mL in each sample tube. If not, fill with ice-cold 1x HBSS up to 7 mL.
- Add 3 mL of pre-chilled IPS to each sample to make a final volume of 10 mL of a solution containing density gradient medium at 30% final concentration. Gently vortex the samples to make sure they are homogenously mixed.
- Centrifuge samples for 15 min at 700 x g at 18 °C making sure to set the acceleration of the centrifuge to 7 and the brake to 0.
NOTE: Centrifugation should take approximately 30 min. - Delicately remove the samples from the centrifuge.
NOTE: A whitish disk composed of debris and myelin should be visible floating on the surface of the solution. A pellet (containing the cells of interest) should be visible at the bottom of the tube. - Carefully aspirate all the whitish disk of debris and then the rest of the supernatant making sure not to dislodge the pellet. Leave about 100 µL of solution on top of the cell pellet to avoid the risk of inadvertently dislodging it.
- Add 1 mL of FACS BL, resuspend the pellet by pipetting up and down with a 1000 µL pipette tip and transfer samples to 1.5 mL tubes.
- Centrifuge for 5 min at 450 x g at room temperature (RT).
- Gently aspirate the supernatant and resuspend the pellet in appropriate buffer compatible with downstream analyses (see section 6 for the protocol used for flow cytometric evaluation of multiple cell types).
6. Staining for flow cytometric evaluation of multiple cell types
- Resuspend the pellet obtained in step 5.9 with 350 µL of FACS BL. Add Fc-block to each sample at a final concentration of 5 µg/mL.
NOTE: At least 100 µL of the sample should be used for one staining, to make sure to process enough cells to allow reliable analyses. - Incubate the sample for 10 min at 4 °C before proceeding with the staining.
- Prepare an antibody mix according to Table 1.
- Add antibody mix to each tube, vortex for 5 s and incubate the samples for 15 min at 4 °C in the dark.
- Add 1 mL of PBS to each tube, vortex and centrifuge for 5 min at 450 x g at RT.
- Meanwhile prepare streptavidin mix according to Table 1.
- Discard the supernatant and resuspend the pellet in the streptavidin mix prepared in step 6.6. For each sample, use the same volume as the one used for the staining in step 6.4.
- Vortex for 5 s and incubate the samples for 10 min at 4 °C in the dark.
- Add 1 mL of PBS to each tube, vortex and centrifuge for 5 min at 450 x g at RT.
- Discard the supernatant and resuspend the pellet in FACS BL. Use 300 µL of FACS BL for each 100 µL of sample stained.
- Add 7-amino-actinomycin D (7-AAD) solution to each sample. Use 5 µL of 7-AAD for each 300 µL of sample prepared in step 6.10.
- Store samples at 4 °C in the dark until cytofluorimetric analysis. Perform the analysis within 16 h from sample preparation, to guarantee >60% cell viability.
Representative Results
We compared two different homogenization methods (DH versus PD) applied to mouse brain and spinal cord, to test the efficiency in retrieving different viable cell types suitable for downstream applications. To do so, we exploited a 9-color flow cytometry panel designed to characterize, in the same sample, different CNS cell types including microglia, lymphocytes, neurons, astrocytes, oligodendrocytes and endothelium.
Brain and spinal cord tissues were retrieved from different mice (n ≥ 6), split in two halves longitudinally, weighed and processed in parallel by applying either mechanical disruption using the Dounce homogenizer (DH method) or gently minced and digested enzymatically using the commercial NTDK based on papain (PD method) (Figure 1A). After debris removal, cells from the brain or the spinal cord were diluted 1/10 or 1/2−1/5, respectively, in Trypan blue to determine cell yield and viability with a Neubauer chamber (Figure 1B,C). The DH method overall produced a higher cell yield from both brain and spinal cord. However, majority of the cells retrieved were dead, resulting in only 13.8% ± 3.3% of viable cells in the brain and 10.5% ± 1.5% in the spinal cord (Figure 1B). Many of the dead cells formed aggregates (Figure 1C); this phenomenon could be due to the presence of highly interconnected cell networks (like the endothelial and glial cells lining the CNS vasculature) that could not be disaggregated by the shearing force applied with the DH. These aggregates of death cells were likely not removed by the density gradient and ended up in the final cell pellet used for cytofluorimetric analysis. On the contrary, the PD method determined an overall better preservation of cellular viability (90.6% ± 0.6% in the brain and 85.2% ± 2.8% in the spinal cord). Papain is able to digest the extracellular matrix and cell-to-cell junctions efficiently, leading to a more uniform single cell suspension. Some of the cells that die during the mincing process could be further digested by papain leading to formation of cell debris that are more efficiently separated through the density gradient. Overall, this likely determined a better preservation of cell viability with PD method, despite a slightly lower cell yield as compared with the DH method.
An aliquot of 100 µL from the brain and spinal cord cell suspensions was stained with the antibody mix (Table 1) and analyzed by flow cytometry with a 9-color panel. Figure 2A shows the gating strategy used to identify the different cell types from the brain and spinal cord cell suspensions. Briefly, the first gate identifies the general population according to forward scatter (FSC) and side scatter (SSC), excluding small cell debris. Then live (7-AAD-) cells are identified. Within total live cell population, CD45+ and CD45- cells are highlighted. Within the CD45+ gate, CD45+CD11b+ microglia/macrophages and CD45+CD11b- lymphocytes are identified. Within CD45- gate, cells are discriminated according to positivity for ACSA2 (astrocytes) or O4 (oligodendrocytes). CD45-ACS2-O4- cells are further subdivided according to positivity for Thy1 (neurons) or CD31 (endothelium). Remaining Thy1-CD31- cells are classified as “other cell types”, not accounted by our antibody mix.
As shown in Figure 2B, with DH method about 38% of the viable cells retrieved from the brain and about 32% of the viable cells retrieved from the spinal cord were of hematopoietic origin (CD45+). On the other hand, PD method allowed to retrieve a significantly high yield of viable cells in both tissues, with a very large fraction represented by non-hematopoietic CD45- cells (about 82% in the brain and 92% in the spinal cord). Remarkably, CD45+CD11b+ microglia/macrophages represented the most abundant viable cell fraction with the DH method (Figure 2C). However, PD method produced a more heterogeneous representation of cell types, including ACSA+ astrocytes, O4+ oligodendrocytes, CD31+ endothelial cells and Thy1+ neurons (Figure 2C). Interestingly, viable neurons and endothelial cells were hardly detectable with the DH method.
The DH method relies on mechanical grinding of the tissue between the glass pestle and mortar of the Dounce homogenizer to obtain tissue homogenization. This could cause some shear stress that will likely damage and affect viability of large or very sensitive cells such as neurons or cells of the neurovasculature. We evaluated the cellular viability (percentage of 7-AAD- cells) within each cell subpopulation identified through the antibody panel (Figure 3). Hematopoietic cells (CD45+) isolated from brain and spinal cord, including microglia/macrophages (CD45+CD11b+) and other non-myeloid cells (CD45+Cd11b-), displayed very high viability independently from the homogenization method that was used (Figure 3A). On the contrary, the DH method determined a significant reduction of viability of CD45- populations (Figure 3B) whereas the PD method determined an extensive preservation of different CNS cell types. In detail, neurons and endothelial cells were the subpopulations most significantly affected by DH and preserved by the PD method.
A schematic presentation of the critical steps required for proper sample preparation and efficient debris removal are summarized in Figure 4.
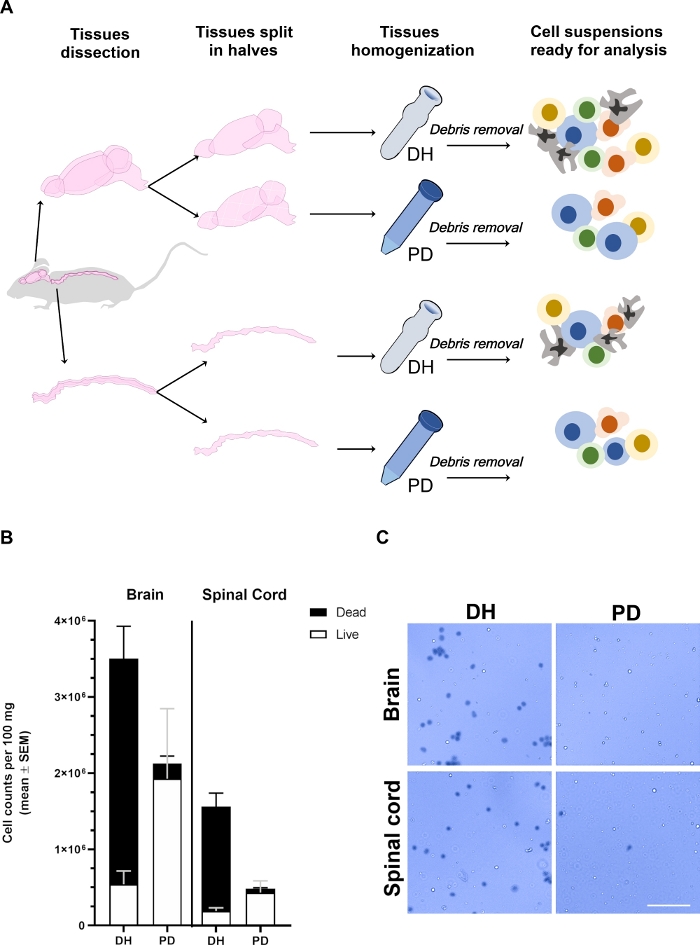
Figure 1: Yield of cells retrieved from brain and spinal cord is affected by the homogenization method.
(A) Experimental outline. Mice were anesthetized and intracardiacally perfused with PBS to remove intra-vascular circulating blood cells. The brain and spinal cord were carefully dissected and split in two halves longitudinally. Tissues were homogenized using either Dounce homogenizer (DH) or papain digestion (PD) as detailed in the main text. Myelin and tissue debris were then removed by centrifugation in a 30% density gradient medium solution resulting in a heterogeneous cell suspension containing different cell types that could be analyzed by flow cytometry. (B) Histograms showing the yield of cells retrieved from the brain or the spinal cord upon tissue homogenization with the DH or PD method. The mean ± SEM of at least 6 animals per condition is represented. (C) Representative brightfield microscope photomicrographs of Trypan blue positive (dead) and negative (live) cells retrieved from brain or spinal cord by the two methods. Scale bar = 100 µm. Please click here to view a larger version of this figure.
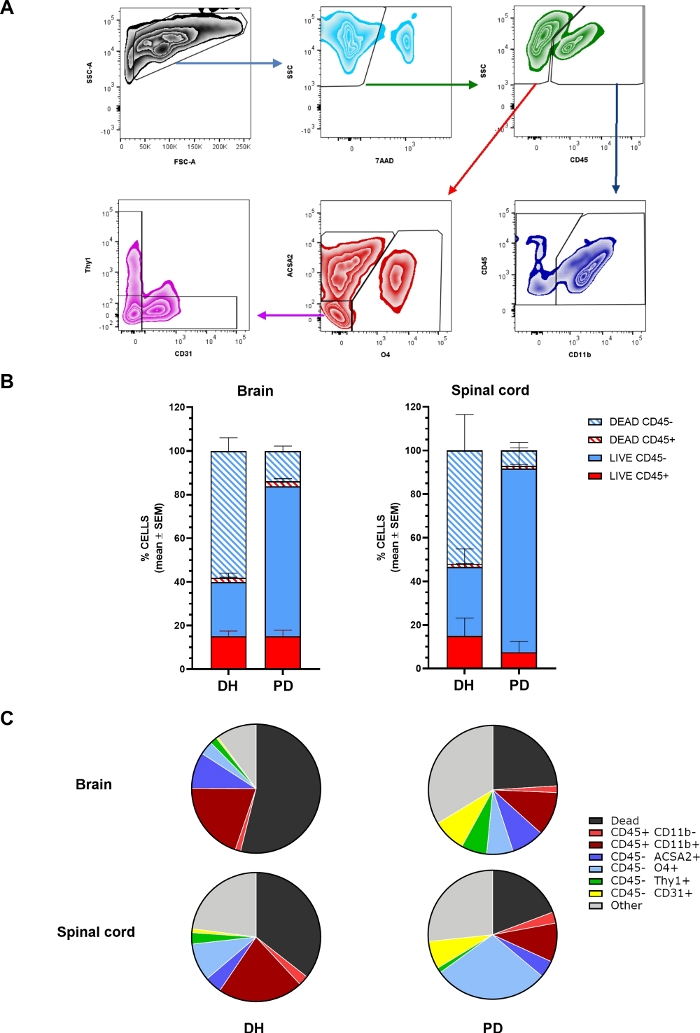
Figure 2: Relative proportions of different cell types retrieved from the CNS are affected by the tissue homogenization method.
(A) Representative flow cytometry plots showing the gating strategy to identify different cell subpopulation within cell preparations obtained from brain or spinal cord: cell population is gated on FSC and SSC physical parameters, followed by selection for 7-AAD- live cells; then cells are discriminated according to positivity for CD45 marker; microglia/macrophages are identified as CD11b+ cells within the CD45+ fraction whereas lymphocytes are CD11b-. Astrocytes, oligodendrocytes, endothelial and neuronal cells are identified as ACSA2+, O4+, CD31+ or Thy1+ cells within CD45-, respectively. (B) Histograms showing the percentage of CD45+ and CD45- cells within total live or dead cell populations, in brain or spinal cord upon homogenization with the DH or PD method. The statistical analysis of the results shown in the graphs is reported in Table 2. (C) Pie charts showing the percentage of different viable cell subtypes within total cell population, in brain or spinal cord upon homogenization with the DH or PD method. The percentage of total dead cells is also reported. N ≥ 6. CD45+CD11b+ = microglia/macrophages; CD45+CD11b- = lymphocytes/non-myeloid cells; CD45-ACSA2+ = astrocytes; CD45-O4+ = oligodendrocytes; CD45-Thy1+ = neurons; CD45-CD31+ = endothelial cells; Other = cells negative for all above-mentioned markers. The statistical analysis of the results shown in the graphs is reported in Table 2. Please click here to view a larger version of this figure.
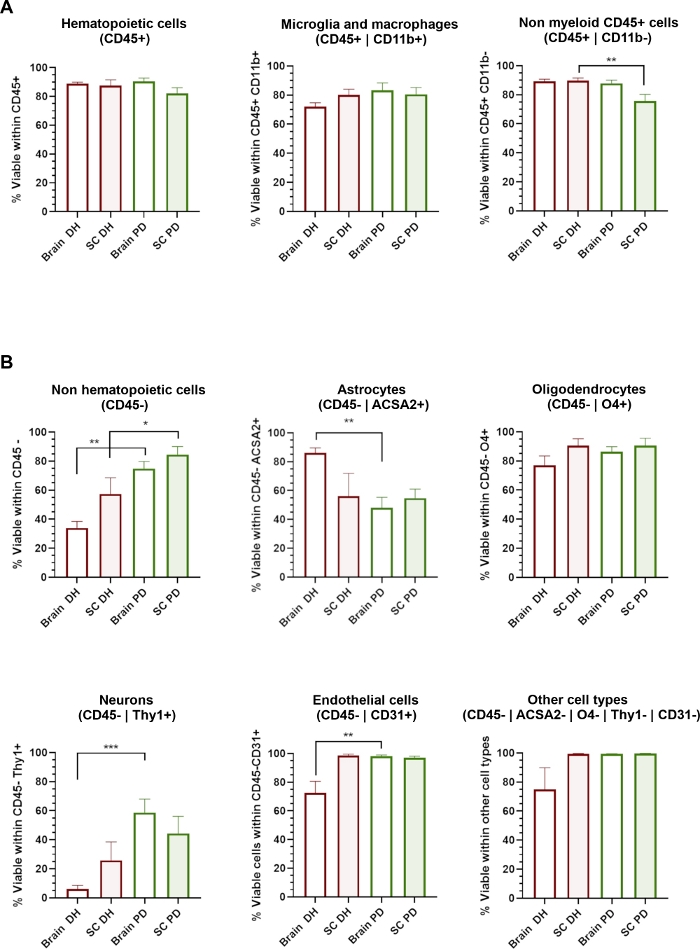
Figure 3: Cellular viability of different CNS cell types is affected by the homogenization method applied.
(A) Histograms showing the percentage of 7-AAD- live cells within CD45+ hematopoietic populations including CD11b+ microglia/macrophages and CD11b- non-myeloid cells. (B) Histograms showing the percentage of 7-AAD- live cells within CD45- non-hematopoietic populations including astrocytes, oligodendrocytes, neurons, endothelial and other cell types. * = p < 0.05, ** = p < 0.01, Mann-Whitney between DH and PD. Please click here to view a larger version of this figure.
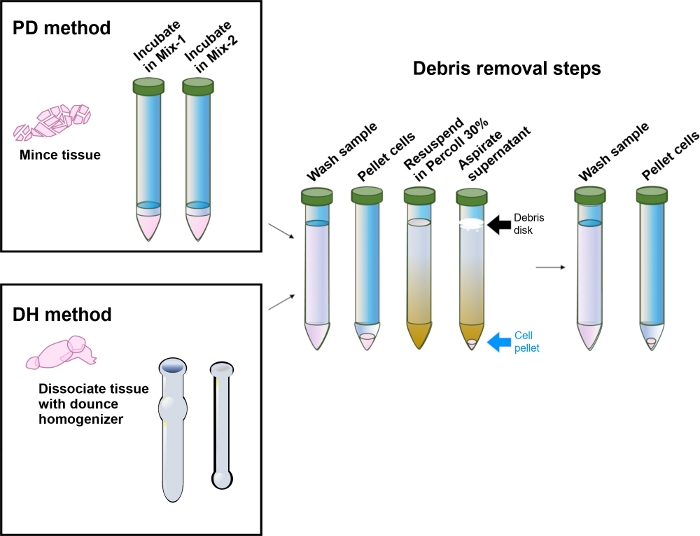
Figure 4: Schematic representation of the critical steps required for proper tissue processing.
A list of the most critical steps required for proper tissue processing and efficient removal of debris is shown. It is important to identify properly the debris disk (black arrow) and the cell pellet (blue arrow) formed after centrifugation of the samples on the 30% density gradient. The debris disk, together with the rest of the supernatant, must be carefully removed by aspiration without dislodging the cell pellet to avoid sample loss. Please click here to view a larger version of this figure.
| Antibody mix | Initial concentration (µg/mL) | Final concentration (µg/mL) | Dilution factor |
| anti CD45/BV510 | 200 | 2 | 100 |
| anti CD11b/APC.780 | 200 | 2 | 100 |
| anti CD31/BV421 | 200 | 2 | 100 |
| anti ACSA2/APC | 150 | 0.75 | 200 |
| anti O4/biotin | na | na | 40 |
| anti CD90.2/PE.Cy7 | 200 | 2 | 100 |
| Streptavidin mix | Initial concentration (µg/mL) | Final concentration (µg/mL) | Dilution factor |
| Streptavidin/Alexa 680 | 1000 | 1 | 1000 |
Table 1: Recipe for preparation of mixes for flow cytometry staining. The table describes the optimal concentrations of antibodies and streptavidin used to allow flow cytometric analyses of multiple cell types. Please refer to Table of Materials for details on catalogue numbers of each reagent mentioned in the table.
| Statistics for Figure 2B | ||||||||||
| BRAIN (% cells) | ||||||||||
| CD45+ | CD45- | |||||||||
| LIVE | DEAD | LIVE | DEAD | |||||||
| Method | mean | ± SEM | mean | ± SEM | mean | ± SEM | mean | ± SEM | ||
| DH | 15.20 | ± 2.32 | 1.90 | ± 0.30 | 24.78 | ± 4.045 | 51.58 | ± 6.033 | ||
| PD | 15.20 | ± 2.65 | 2.33 | ± 1.10 | 68.53 | ± 3.618 | 13.93 | ± 2.180 | ||
| Mann-Whitney | ns | ns | *** | ** | ||||||
| p-value | 0.9989 | 0.738 | 0.0006 | 0.0015 | ||||||
| SPINAL CORD (% cells) | ||||||||||
| CD45+ | CD45- | |||||||||
| LIVE | DEAD | LIVE | DEAD | |||||||
| Method | mean | ± SEM | mean | ± SEM | mean | ± SEM | mean | ± SEM | ||
| DH | 15.00 | ± 8.21 | 1.41 | ± 0.11 | 31.64 | ± 8.21 | 51.95 | ± 16.52 | ||
| PD | 7.49 | ± 4.99 | 1.15 | ± 0.68 | 84.27 | ± 9.39 | 7.09 | ± 3.75 | ||
| Mann-Whitney | ns | ns | * | ns | ||||||
| p-value | 0.5548 | 0.7236 | 0.0438 | 0.1144 | ||||||
| Statistics for Figure 2C | |||||||||||||||||
| BRAIN (% cells) | |||||||||||||||||
| CD45+ | CD45- | ||||||||||||||||
| CD11b+ | CD11b- | ACSA2 | O4 | Thy1 | CD31 | Other | Dead | ||||||||||
| Method | mean | ± SEM | mean | ± SEM | mean | ± SEM | mean | ± SEM | mean | ± SEM | mean | ± SEM | mean | ± SEM | mean | ± SEM | |
| DH | 19.32 | ± 3.88 | 1.17 | ± 0.27 | 9.52 | ± 2.68 | 3.41 | ± 1.01 | 1.39 | ± 0.77 | 0.48 | ± 0.29 | 10.52 | ± 4.49 | 53.83 | ± 5.79 | |
| PD | 10.88 | ± 2.03 | 1.65 | ± 0.48 | 8.17 | ± 2.66 | 6.54 | ± 0.76 | 6.37 | ± 1.76 | 8.27 | ± 1.25 | 33.28 | ± 6.34 | 23.72 | ± 5.31 | |
| Mann-Whitney | ns | ns | ns | * | ** | *** | ** | ** | |||||||||
| p-value | 0.1206 | 0.4819 | 0.5894 | 0.0264 | 0.0093 | 0.0003 | 0.0084 | 0.0022 | |||||||||
| SPINAL CORD (% cells) | |||||||||||||||||
| CD45+ | CD45- | ||||||||||||||||
| CD11b+ | CD11b- | ACSA2 | O4 | Thy1 | CD31 | Other | Dead | ||||||||||
| Method | mean | ± SEM | mean | ± SEM | mean | ± SEM | mean | ± SEM | mean | ± SEM | mean | ± SEM | mean | ± SEM | mean | ± SEM | |
| DH | 21.23 | ± 6.25 | 2.51 | ± 0.57 | 4.26 | ± 2.34 | 9.40 | ± 1.89 | 2.82 | ± 1.51 | 0.97 | ± 0.50 | 22.74 | ± 9.04 | 35.28 | ± 1.89 | |
| PD | 9.63 | ± 1.67 | 2.77 | ± 0.48 | 4.23 | ± 1.59 | 28.62 | ± 3.57 | 1.26 | ± 0.49 | 6.94 | ± 2.14 | 26.39 | ± 8.17 | 19.09 | ± 4.76 | |
| Mann-Whitney | ns | ns | ns | * | ns | * | ns | ns | |||||||||
| p-value | 0.1905 | >0.9999 | 0.7302 | 0.0159 | 0.7302 | 0.0317 | 0.7302 | 0.1111 | |||||||||
Table 2: Statistical analysis of different populations retrieved by applying the DH or PD method. The table describes the statistics for the graphs shown in Figure 2B and Figure 2C. The average and SEM of at least six independent samples is represented. The p value and details on statistical test applied for each comparison are also reported.
Discussion
Herein we describe a protocol for the co-purification and concurrent flow cytometric analysis of some of the most relevant CNS cells from mouse brain and spinal cord. Traditionally, histological analyses have been applied to describe the distribution of nanoparticles or the transduction efficiency of viral vectors in the CNS5,13, or to provide insights on morphological and molecular changes occurring in specific cell types during a pathology or upon pharmacological treatment14. However, histology lacks processivity and it does not allow comprehensive examination of multiple features in the same histological samples, due to the limited number of markers that can be concurrently analyzed. Our approach can be complementary to traditional histologic analyses and it can be coupled with several downstream applications (sorting, primary culture, biochemical or next-generation-sequencing analyses) to expand the compilation of information that can be obtained from individual samples. However, some key factors listed below must be considered as they can critically impact the success of this approach:
- Amount of starting tissue. We have optimized the cell separation to start with as little as half spinal cord or half brain. In our experience, processing half brain or half spinal cord from an adult 8-week-old mouse with the PD method yields 1−6 x 106 viable cells from the brain and about 0.1−0.5 x 106 viable cells from the spinal cord after the density gradient step. We have not measured cells yield after applying the PD method on tissues from newborn or mice younger than 8 weeks. However, in our hands, the outcome is proportional to the weight of the starting tissue. Thus, for younger animals (e.g., 10-day-old pups) the entire brain or spinal cord should be processed to guarantee a good cell yield for reliable downstream analyses. If necessary, pooling tissues from multiple animals would help increasing cell yields. In our experience, this protocol can also be applied without modifications to isolate cells from the brain or spinal cord of rats, as far as the proportion of reagents used per weight of starting tissue is respected. For tissues more than 250 mg of weight, scaling reagents up or splitting the tissue in multiple samples (each weighting <250 mg) is suggested.
- Removing meninges/residual tissue chunks before the density gradient. In our experience, the PD method hereby described is not able to digest efficiently meninges or very highly myelinated tissues such as the nerves and nerve roots emerging from the spinal cord. When these tissues are present, some undigested sticky pieces may remain in the cell suspension retrieved after the enzymatic digestion steps, before debris removal. Removing these chunks by filtering the solution through a 70 µm cell strainer (as suggested in step 5.1) is critical for a successful cell preparation. In fact, if not removed, meninges or tissue pieces will hinder efficient density gradient separation resulting in poor cell yields.
- Timing, temperature and sterility. It is very important to perform all steps in a timely fashion using the right temperature settings and incubations as suggested. This is critical to ensure high cell viability and integrity of the sample. Depending on the downstream application, performing all the steps under a sterile hood and with sterile reagents might be necessary (e.g., establishing primary cell cultures). Extended incubation in the enzymatic digestive solution beyond the suggested time (section 3) could result in a drop of cell viability. The epitopes of some surface antigens could be sensitive to papain resulting in loss of signal at flow cytometry. For specific applications requiring additional markers other than the ones described in this article, testing the performance of different antibody clones is recommended before starting the experiment. It has been reported that the density gradient medium could contain some traces of endotoxins that may trigger activation of immune cells (microglia/macrophages). Appropriate internal controls should always be added in the experiments whenever these populations are analyzed, to exclude possible effects induced by the procedure. Sticking with the suggested temperatures and washing density gradient medium leftovers immediately after the debris-removal step is usually enough to avoid overt immune cells activation. However, in case the downstream applications (e.g., RNAseq or functional analyses) are affected by this step, the user should switch to a low-endotoxin density gradient medium preparation (suggested in Table of Materials).
- FACS antibodies and machine. The flow cytometry staining protocol hereby presented makes use of antibody concentrations and color conjugations that worked best with cell yields retrieved in our laboratory experience and with FACS machines available in our institute. The user should titer the antibodies in his/her hands before starting a new experiment, as the concentration might need to be slightly adjusted. Besides, we encourage to use always single-color staining controls in each experiment to verify that all antibodies and the compensation set-up of the FACS machine is working adequately. It has to be noticed that the CD90 (Thy) antigen used for detecting neurons exists in two different isotypes, namely CD90.2 or CD90.1 depending on the mouse strain: the most commonly used mouse strains, such as C57BL6/J express CD90.2; mouse strains such as AKR/J, PL, and FVB/N express CD90.1. Thus, the user should carefully verify the mouse strain and choose the appropriate anti-CD90 antibody (as suggested in Table of Materials) before starting the experiments.
In summary, the protocol here presented takes advantage of a gentle enzymatic digestion followed by a 9-color staining allowing efficient simultaneous flow cytometric evaluation of different cell types from mouse brain and spinal cord. The protocol could be exploited to monitor in a streamline and comprehensive manner the efficiency of cell targeting by nanoparticles or viral vectors administered in the CNS15. Moreover, the protocol could be easily adopted for very delicate downstream applications, such as cell sorting, ex vivo subculture, single cell RNAseq, resulting of utmost importance not only for preclinical assessment of cell-targeting by therapeutics but also for in-depth characterization of pathological processes in neurodegenerative diseases.
A fraction of the whole CNS cell population is not discriminated by this protocol (see “other” cell types in Figure 2); this can be explained by the presence of other cell subtypes that are present in the CNS but are not captured by the antibodies we used. In our preliminary analyses, about 14% of the “other cells” fraction is positive for CD73, a mesenchymal cell marker enriched in the neurovasculature and involved in several neuroinflammatory processes16,17. Moreover, we hypothesize that the “other cells” fraction could also comprise less differentiated cells, like progenitors at different stages of maturation, such as nestin+ neural stem cells, nestin+ vimentin+ radial glia progenitors, doublecortin+ neural progenitors, NG2+ oligodendrocyte precursor cells, among others. These cell sub-types could be easily investigated by applying our flow cytometry protocol, since we chose a configuration of fluorescent dyes that allows to accommodate up to two additional cell markers conjugated with either the fluorescein isothiocyanate (FITC) or phycoerythrin (PE) fluorophores.
Overall, our approach could provide a new tool for more comprehensive investigations in the context of the CNS (in health and disease) taking advantage of a well-consolidated technology allowing both qualitative and high-throughput quantitative assessments such as flow cytometry.
Divulgations
The authors have nothing to disclose.
Acknowledgements
This study was funded by Boston Children’s Hospital start-up funds to A.B., ALSA grant nr. 17-IIP-343 to M.P., and the Office of the Assistant Secretary of Defense for Health Affairs through the Amyotrophic Lateral Sclerosis Research Program under Award No. W81XWH-17-1-0036 to M.P. We acknowledge DFCI Flow Cytometry Core for technical support.
Materials
| 10X HBSS (Calcium, Magnesium chloride, and Magnesium Sulfate-free) | Gibco | 14185-052 | |
| 70 mm Cell Strainer | Corning | 431751 | |
| ACSA/ACSA2 anti-mouse antibody | Miltenyi Biotec | 130-117-535 | APC conjugated |
| Bovine Serum Albumin | Sigma Aldrich | A9647-1KG | |
| CD11b rat anti-mouse antibody | Invitrogen | 47-0112-82 | APC-eFluor 780 conjugated |
| CD31 rat anti-mouse antibody | BD Bioscience | 562939 | BV421 conjugated |
| CD45 rat anti-mouse antibody | Biolegend | 103138 | Brilliant Violet 510 conjugated |
| CD90.1/Thy1.1 rat anti-mouse antibody | Biolegend | 202518 | PE/Cy7 conjugated |
| CD90.2/Thy1.2 rat anti-mouse antibody | Biolegend | 1005325 | PE/Cy7 conjugated |
| Conical Tubes (15 mL) | CellTreat | 229411 | |
| Conical Tubes (50 mL) | CellTreat | 229422 | |
| Dounce Tissue Grinder set (Includes Mortar as well as Pestles A and B) | Sigma-Aldrich | D9063-1SET | |
| Fc (CD16/CD32) Block rat anti-mouse antibody | BD Pharmingen | 553142 | |
| Fetal Bovine Serum | Benchmark | 100-106 | |
| Neural Tissue Dissociation Kit (P) | Miltenyi Biotec | 130-092-628 | |
| O4 anti mouse/rat/human antibody | Miltenyi Biotec | 130-095-895 | Biotin conjugated |
| Percoll | GE Healthcare | 10266569 | sold as not sterile reagent |
| Percoll | Sigma | 65455529 | sterile reagent (to be used for applications requiring sterility) |
| Percoll PLUS | Sigma | GE17-5445-01 | reagent containing very low traces of endotoxin |
| Streptavidin | Invitrogen | S3258 | Alexa Fluor 680 conjugated |
References
- Deverman, B. E., Ravina, B. M., Bankiewicz, K. S., Paul, S. M., Sah, D. W. Y. Gene therapy for neurological disorders: progress and prospects. Nature Reviews Drug Discovery. 17 (9), 641-659 (2018).
- Teleanu, D., Negut, I., Grumezescu, V., Grumezescu, A., Teleanu, R. Nanomaterials for Drug Delivery to the Central Nervous System. Nanomaterials. 9 (3), 371 (2019).
- Chen, S., et al. Recombinant Viral Vectors as Neuroscience Tools. Current Protocols in Neuroscience. 87 (1), 67 (2019).
- Alves, S., et al. Ultramicroscopy as a novel tool to unravel the tropism of AAV gene therapy vectors in the brain. Scientific Reports. 6 (1), 28272 (2016).
- Peviani, M., et al. Lentiviral vectors carrying enhancer elements of Hb9 promoter drive selective transgene expression in mouse spinal cord motor neurons. Journal of Neuroscience Methods. 205 (1), 139-147 (2012).
- Baumgarth, N., Roederer, M. A practical approach to multicolor flow cytometry for immunophenotyping. Journal of Immunological Methods. 243 (1-2), 77-97 (2000).
- Sykora, M. M., Reschke, M. Immunophenotyping of Tissue Samples Using Multicolor Flow Cytometry. Methods in Molecular Biology. 1953, 253-268 (2019).
- Legroux, L., et al. An optimized method to process mouse CNS to simultaneously analyze neural cells and leukocytes by flow cytometry. Journal of Neuroscience Methods. 247, 23-31 (2015).
- Lee, J. K., Tansey, M. G. Microglia Isolation from Adult Mouse Brain. Methods in Molecular Biology. 1041, 17-23 (2013).
- Grabert, K., McColl, B. W. Isolation and Phenotyping of Adult Mouse Microglial Cells. Methods in Molecular Biology. 1784, 77-86 (2018).
- Nikodemova, M., Watters, J. J. Efficient isolation of live microglia with preserved phenotypes from adult mouse brain. Journal of Neuroinflammation. 9 (1), 635 (2012).
- Garcia, J. A., Cardona, S. M., Cardona, A. E. Isolation and analysis of mouse microglial cells. Current Protocols in Immunology. 104 (1), 1-15 (2014).
- Papa, S., et al. Selective Nanovector Mediated Treatment of Activated Proinflammatory Microglia/Macrophages in Spinal Cord Injury. ACS Nano. 7 (11), 9881-9895 (2013).
- Peviani, M., et al. Neuroprotective effects of the Sigma-1 receptor (S1R) agonist PRE-084, in a mouse model of motor neuron disease not linked to SOD1 mutation. Neurobiology of Disease. 62, 218-232 (2014).
- Peviani, M., et al. Biodegradable polymeric nanoparticles administered in the cerebrospinal fluid: Brain biodistribution, preferential internalization in microglia and implications for cell-selective drug release. Biomaterials. 209, 25-40 (2019).
- Meng, F., et al. CD73-derived adenosine controls inflammation and neurodegeneration by modulating dopamine signalling. Brain. 142 (3), 700-718 (2019).
- Nedeljkovic, N. Complex regulation of ecto-5′-nucleotidase/CD73 and A2AR-mediated adenosine signaling at neurovascular unit: A link between acute and chronic neuroinflammation. Pharmacological Research. 144, 99-115 (2019).

