In Vitro Establishment of a Genetically Engineered Murine Head and Neck Cancer Cell Line using an Adeno-Associated Virus-Cas9 System
Summary
Development of murine models with specific genes mutated in head and neck cancer patients is required for understanding of neoplasia. Here, we present a protocol for in vitro transformation of primary murine tongue cells using an adeno-associated virus-Cas9 system to generate murine HNC cell lines with specific genomic alterations.
Abstract
The use of primary normal epithelial cells makes it possible to reproducibly induce genomic alterations required for cellular transformation by introducing specific mutations in oncogenes and tumor suppressor genes, using clustered regulatory interspaced short palindromic repeat (CRISPR)-based genome editing technology in mice. This technology allows us to accurately mimic the genetic changes that occur in human cancers using mice. By genetically transforming murine primary cells, we can better study cancer development, progression, treatment, and diagnosis. In this study, we used Cre-inducible Cas9 mouse tongue epithelial cells to enable genome editing using adeno-associated virus (AAV) in vitro. Specifically, by altering KRAS, p53, and APC in normal tongue epithelial cells, we generated a murine head and neck cancer (HNC) cell line in vitro,which is tumorigenic in syngeneic mice. The method presented here describes in detail how to generate HNC cell lines with specific genomic alterations and explains their suitability for predicting tumor progression in syngeneic mice. We envision that this promising method will be informative and useful to study tumor biology and therapy of HNC.
Introduction
HNC is a common malignancy worldwide1. Modeling the genesis of HNC neoplasia is currently at a scientific turning point2. While many genetic mutations have been identified in HNC2,3,4 (e.g., TP53, PIK3CA, NOTCH1, FAT1, and RAS) the specific combinations of genomic alterations required in concert to induce HNC remain unclear.
The current use of human HNC cell lines has significantly helped to elucidate the mechanisms associated with pathogenesis and treatment3. However, the study of human cell lines in immunocompromised murine systems has its limitations, because these systems do not address the in vivo neoplastic process, the role of specific gene mutations, and treatment responses in an immune microenvironment. Hence, the development and establishment of murine cell lines with specific genetic alterations are of primary importance to study how different genes contribute to the transformation process and to test novel molecule-based therapies in immunocompetent mice.
Gene function studies in biomedical research have been significantly affected by advances in DNA editing technologies, by introducing double-strand breaks (DSBs), for example5. These technologies, including the use of zinc finger nucleases, transcription activator-like effector nucleases, and clustered regulatory interspaced short palindromic repeats (CRISPR/Cas9), allow for the manipulation of any gene of interest at its locus. The latest CRISPR/Cas9 systems is comprised of a guide RNA (gRNA) that directs the Cas9 nuclease to generate a DSB at a specific site in the genome. This system has gained extensive recognition in modifying endogenous genes in any cell or target tissue, even in the most traditionally difficult-to-treat organisms5. It has multiple advantages over other systems due to its simplicity, speed, and efficiency.
In oncology, CRISPR/CAS9 technology has fulfilled the need to effectively mimic cancer cells. To establish this system in HNC, we manipulated the potent KRAS oncogene and two important tumor suppressor genes, APC and p536. According to the GENIE database7, this combination is rare in HNC. RAS mutations (HRAS, NRAS, and KRAS) are present in only ~7% of all HNC populations. These tumors tend to be resistant to therapy8,9.
Delivery of Cas9 and its gRNA is achieved through viral transduction using either AAVs or lentiviruses. Recombinant AAV is often the preferred method for delivering genes to target cells owing to its high titer, mild immune response, ability to transduce a broad range of cells, and overall safety. Using an AAV system, various tissue-specific mouse cell lines have been generated, and new cell lines are still being developed10,11,12. However, an efficient genomic editing system that can generate murine HNC cell line models cells remains to be developed. In this study, we sought to develop an in vitro AAV-Cas9-based system for transforming primary murine tongue cells into a tumorigenic state. This unique CRISPR/Cas9 transformation protocol and the established tumor cell line can be used to better understand the biology of HNC induced by a diversity of genomic alterations.
Protocol
This study was approved by the Ben Gurion University of the Negev Animal Care and Use Committee. Animal experiments were approved by the IACUC (IL.80-12-2015 and IL.29-05-2018(E)). All aspects of animal testing, housing, and environmental conditions used in this study were in compliance with The Guide for the Care and Use of Laboratory Animals13.
1. Adeno-associated virus production
- Day 1: Cell culture
- Seed 4 x 106 HEK293T cells per 14.5 cm plate in 15 mL of DMEM. Prepare 10 plates for transfection with polyethylenimine (PEI) (1 µg/µL).
- Day 2: Transfection of HEK293T cells using PEI
- Remove media and refeed with warm DMEM 1 h before the transfection.
- Prewarm transfection reagents and DMEM.
- Use 10 µg of the plasmid of interest containing AAV ITR (AAV pCM109 EFS Cre sg APC sg KRAS sg P53- KRAS HDR), 10 µg of the AAV 2/9n capsid plasmid (with the rep gene of AAV2 and the cap gene of AAV9, and 10 µg of the helper plasmid (pAdDelta F5 helper) per plate (See Table of Materials).
NOTE: A new generation of helper plasmid – pAdDeltaF614,15,16 that can also be used for AAV production is available. - Mix the plasmids in 1 mL of plain DMEM per plate. Then add 90 µL of polyethylenimine (total plasmid: PEI concentration = 1:3) to the plasmid mix and vortex briefly.
- Incubate the PEI-plasmid DNA mix for 20 min at room temperature (RT).
- Add the mix dropwise on top of the HEK293T cells and transfer the plates to the cell culture incubator at 37 °C with 5% CO2 for 24 h.
- Day 3: Medium change after transfection
- Remove the medium completely and add 15 mL of fresh DMEM.
- Day 5: Harvesting the virus
- Prepare a dry ice/ethanol bath.
- Collect the cells with a cell scraper and transfer them into 50 mL tubes (2 plates per 50 mL tube).
- Spin tubes at 800 x g for 15 min at room temperature.
- Discard the supernatant from all the tubes. Add 0.5 mL of the lysis buffer (150 mM NaCl, 50 mM Tris-HCl, pH = 8.5) per plate (i.e., 5 mL for 10 plates) to the first tube only. Resuspend the cells and transfer the total volume to the next tube and continue until the last tube.
- Transfer the cell suspension to a fresh 50 mL tube. Wash the tubes with the same volume of lysis buffer (0.5 mL per plate) using the same transfer method as described in step 1.4.4.
- Subject the cell suspension to three rounds of ~10 min freeze/thaw cycles between a dry ice/ethanol bath and a 37 °C water bath. Vortex briefly after each thawing.
- If the purification is carried out on the same day, set the equilibration and elution buffer (see Table 1 for the recipe) at RT.
- Add 50 units of benzonase per plate and incubate at 37 °C for 1.5 h.
NOTE: Benzonase is used to digest residual nucleic acids from the host producer cells and the plasmid DNA present in the cell suspension. - Spin the tubes at 3,000 x g for 15 min at 4 °C.
- Collect the supernatant in a syringe using a 18 G steel needle and push the solution through a 0.45 µm filter into a 15 mL tube to obtain the crude lysate.
- Store the crude lysate at 4 °C for a few weeks until purification or continue purification.
2. AAV purification
- Day 5 or later
- Place the equilibration and elution buffer at RT.
- Wash chromatography columns with 10 mL of the equilibration buffer.
- Add 0.5 mL per plate of heparin-agarose to the column17 and then add the equilibration buffer (4x the volume of the heparin-agarose). Mix the solutions by inverting the column and let the agarose sediment.
- Elute the equilibration buffer from the column by gravity.
NOTE: Leave some equilibration buffer to prevent the agarose from drying out. - Load the crude lysate onto the column and incubate for 2 h at 4 °C with constant agitation. Bring the column into an upright position and allow the agarose to sediment.
- Elute the crude lysate by gravity.
- Wash the column with equilibration buffer (4x the volume of the heparin-agarose).
- Place a 100 kDa centrifugal protein filter below the column and elute the virus using an elution buffer (3x the volume of the heparin-agarose) into the filter.
NOTE: The elution buffer should not exceed 15 mL. - Centrifuge the filter at 3,000 x g for ~30 min until less than 1 mL is left in the filter.
- Fill up the filter with PBS and centrifuge at 3,000 x g for ~30 min until less than 1 mL is left in the filter. Repeat this step 2x to remove all salts.
- Concentrate the virus solution in the filter by centrifugation at 3,000 x g to arrive at a volume as small as possible (less than 200 µL).
- Aspirate the virus solution with a needle and syringe and push the solution through a 0.22 µm filter into a tube.
- Make aliquots of concentrated viral particles for storage.
NOTE: The aliquots should be ~20 µL for short-term storage at 4 °C and ~5 µL for long-term storage at -80 °C. - Determine the viral titer and viral transduction efficiency as described previously14,18,19,20.
3. Isolation and culture of primary cells
- Day 1
- Euthanize a 6-week-old male or female B6;129-Gt(ROSA)26Sortm1(CAG-cas9*,-EGFP)Fezh/J by CO2 asphyxiation or any other IACUC approved protocol.
NOTE: These CRISPR/Cas9 knockin mice have Cre recombinase-dependent expression of cas9 endonuclease and EGFP directed by a CAG promoter. The upstream Lox-Stop-Lox (LSL) sequence present in the genome of these mice prevent the expression of Cas9 and EGFP in the absence of Cre recombinase. When used in combination with single guide RNAs (sgRNAs) and a Cre source, they allow editing of single or multiple mouse genes in vivo or ex vivo14. - Dissect the tongue from the euthanized mice using surgical scissors.
- Manually dissociate the tissue by mincing the tongue tissue into very small fragments using a scalpel. Collect the tissue fragments in a 15 mL tube containing 4.5ml of RPMI plain medium (with out serum).
- Add the triple enzyme mix (200 µl) (see Table 1 for the recipe) to the tissue fragments.
- Incubate the tissue-enzyme mix at 37 °C for 30 min and tap the tube every 10 min to enhance enzymatic dissociation of the tissue.
- Add 5% fetal bovine serum (FBS) containing HBSS/PBS to the tissue-enzyme mix to stop the enzyme action.
- Filter the above cell suspension through a sterile 70 µm nylon mesh to separate the dispersed cells and larger tissue fragments.
- Wash the filtered cell suspension by centrifugation for 300 x g for 5 min in HBSS/PBS at RT.
- Resuspend the pellet in culture medium (10% FBS in RPMI/DMEM) and grow in 60 mm culture dishes until distinct cell colonies are formed.
- Culture cell aggregates retained on top of the filter in a 60 mm culture dish containing 3 mL of complete media (10% FBS in RPMI/DMEM) until cell colonies are formed.
- Microscopically examine the primary cells for the presence of fibroblast contamination after 1 week of culture. Treat the primary cells developed from aggregates and cell suspensions with 0.25 trypsin 0.02% EDTA solution at 37 °C for 1 min to remove fibroblasts.
NOTE: Usually the primary culture from cell aggregates produces more colonies compared to single-cell suspensions. The cells from these colonies provide better transduction efficiency with AAV transduction.
- Euthanize a 6-week-old male or female B6;129-Gt(ROSA)26Sortm1(CAG-cas9*,-EGFP)Fezh/J by CO2 asphyxiation or any other IACUC approved protocol.
4. AAV transduction of primary cells
- Day 10
- Seed 2 x 105 primary cells per well in 6 well plates in 2 mL of complete media (10% FBS in RPMI/DMEM).
- The next day, transduce the cells with 1012 viral genome/mL (1010 transducing units/mL) and incubate the cells in viral particle-containing media for 48 h at 37 °C.
- Remove the viral particle-containing media and feed the AAV-transduced cells with complete media.
NOTE: Only cells that underwent transduction will express GFP and start to proliferate. Cells that did not undergo transduction will eventually die within 2 weeks. - After 2 weeks of culture expansion, seed the cells for validation and the in vivo tumorigenic experiments.
NOTE: Genomic DNA fromAAV-transduced cells and normal primary cells from Cas9 mice were extracted for validating specific genome editing using standard protocols. Sequencing of the extracted DNA and analysis of the sequenced data (Table 2) were performed using a hybridization capture-based next-generation sequencing assay (e.g., MSK-IMPACT platform) as described previously21.
5. Validating the transformation of normal cells to tumorigenic cells using immunofluorescence and western blotting
- Immunofluorescence
- Seed 2 x 105 cells on 12 mm glass coverslips and place them in an incubator overnight.
- Fix cells in 4% paraformaldehyde in PBS (pH = 7.4) and process for immunofluorescence labeling.
- Incubate the fixed cells with the first primary antibody in 1% BSA or 1% serum in PBST in a humidified chamber for 1 h at RT or overnight at 4 °C. The primary antibodies used are the rabbit monoclonal anti-KRT 14, rabbit monoclonal anti-E-cadherin antibody, and Cas9 mouse mAb.
- Remove the primary antibody solution from the coverslips by washing the cells 3x for 5 min with 1x PBS.
- Incubate the cells with Cy3 donkey anti-rabbit IgG and/or Cy3 goat anti-mouse IgG secondary antibodies in 5% BSA in PBST for 1 h at RT in the dark.
- Remove the secondary antibody solution from the coverslips and wash the cells 3x as described in step 5.1.4.
- Mount the coverslips with a drop of mounting medium containing DAPI (DAPI Fluoromount).
- Store in the dark at 4 °C until the slides are imaged using fluorescence microscopy.
- Western blotting
- Seed 1 x 106 transformed cells in a 100 mm culture dish and place them in an incubator overnight.
- Wash and scrape the transformed cells into 200 µl ice cold PBS.
- Centrifuge the tube containing the cell suspension for 10 min at 2,000 x g at 4 °C to pellet the cells.
- Aspirate the supernatant and lyse the cells using lysis buffer (see Table 1) containing phosphatase inhibitor cocktails and a protease inhibitor for 10 min at 4 °C. Centrifuge the lysates for 10 min at 10,000 x g and 4 °C and collect the cleared lysates.
- Use a commercially available Bradford assay kit to determine the protein concentration following the manufacturer's protocol. Use 4x sample buffer (500 mM Tris pH = 6.8, 40% glycerol, 8% SDS, 20% H2O, 0.02% bromophenol blue) to adjust the protein samples to 0.5 or 1 µg/µL. Boil at 96 °C for 5 min.
NOTE: The samples can be stored at -80 °C or until a Western blot analysis is performed. - Separate equal amounts of total lysate (30 µg) by 10% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE). Ensure that the dye reaches the bottom of the gel. Transfer the protein to the nylon membrane by semi-dry blotting at 25 V for 30 min.
- Pour 5% BSA in Tris-buffered saline (TBS)-0.1% Tween (TBST) over the membrane to cover it completely. Block the membrane for 1 h at RT. Incubate the mebrane with primary antibodies (anti-Cre, anti-Cas9, anti-GFP, anti-β catenin, anti p53, anti-phospho-ERK, and anti-β actin diluted in 5% BSA TBST) at 4 °C overnight.
- After incubation, wash the membranes for 10 min with 1x TBST making sure that the solution covers the membrane completely. Perform this wash 3x, and then add horseradish peroxidase (HRP)-conjugated secondary antibody (diluted 1:10,000 in 5% BSA TBST) to the membrane. Incubate for 1 h at RT.
- After incubation, wash the membranes for 10 min with 1x TBST making sure that the solution covers the membrane completely. Perform the chemiluminescence imaging (see Table of Materials) to expose the bands, and capture images accordingly.
- Validating the tumorigenic potential of transformed cells in immunocompetent mice
NOTE: Mice were maintained and treated according to the institutional guidelines of the Ben-Gurion University of the Negev. NOD.CB17-Prkdc-scid/NCr Hsd (NOD.SCID) and C57BL/6 mice were used for the in vivo studies. Mice were housed in air-filtered laminar flow cabinets with a 12 hour light/dark cycle and were fed food and water ad libitum.- Use 6-8 week-old female NOD.CB17-Prkdc-scid/NCr Hsd (NOD.SCID) and C57BL/6 mice for the study.
- Trypsinize the AAV-Cas9 transformed cells. Stop the trypsinization using DMEM prewarmed at 37 °C and collect in a 50 mL tube.
- Centrifuge the tube at 800 x g for 10 min at room temperature. Discard the supernatant and resuspend the cell pellet in DMEM medium without FBS. Perform the centrifugation again and resuspend the cell pellet in 1x PBS.
NOTE: Do not keep the cells in 1x PBS for too long. Always keep the cell suspension on ice to prevent cell clumping. - Use an automatic cell counter to count the cells and dilute the cells to the desired concentration (2.5 x 107 cells/mL) using 1x PBS. Generate tumors through a subcutaneous injection of the AAV-Cas9 transformed cell suspension in the right flank of each NOD.CB17-Prkdc-scid/NCr Hsd (NOD.SCID) mouse (2 x 106 cells/mouse). To generate an orthotropic model in syngeneic mice, inject 0.5 × 106 primary cells or the AAV-Cas9 transformed cells into the tongue of C57BL/6 immunocompetent mice.
- Euthanize the animals 2 weeks postinjection by CO2 asphyxiation, and dissect the tumors from the euthanized mice for immunohistochemistry analysis.
Representative Results
Using the AAV system to transform normal Cas9 cells
Figure 1 provides a detailed vector map of the AAV transgene plasmid used in this study. Figure 2 outlines the design and working of the AAV-Cas9 based-system. To produce viral particles, the HEK293T cells were transfected with the AAV transgene vector and other viral packaging vectors using the PEI transfection method. After transfection, the virus-containing cells were collected and lysed, and using the heparin-agarose purification process, the viral particles were purified and concentrated (Figure 2A). These purified viral particles were used for the in vitro transformation of primary cells isolated from Cas9 mice to check the efficacy. In our system, the AAV transgene expression cassette is introduced through homologous recombination into the Cre-dependent Cas9 Rosa26 targeting template within mouse cells that carry an ubiquitous CAG promoter (pCAG), a LoxP-Stop-LoxP cassette (LSL), a Cas9 nuclease gene, a self-cleaving P2A peptide, and the reporter gene (EGFP) flanked by two homologous arms14. The Cre recombinase activity of the transduced cells excises the LSL and induces Cas9 and GFP expression (Figure 2B). Cas9 nuclease activity introduces a double-stranded break at the target sites of interest, directed through gRNAs, and helps in gene editing. The AAV system used in this study helps in delivering multiplexed gRNAs (sgRNAs for KRAS, p53, and APC) and also in expressing an oncogenic KRASG12D allele through homology-directed repair (HDR) in the transduced cells.
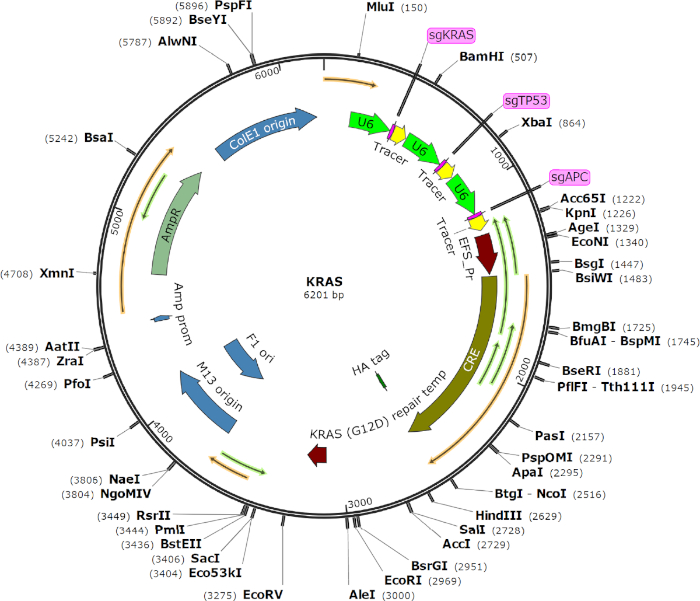
Figure 1: Vector map of the AAV transgene plasmid used in this study. This AAV system has an AAV backbone that expresses Cre recombinase and three U6-driven sgRNAs targeting KRAS, p53, and APC. It also contains a KRAS G12D homology-directed repair (HDR) donor. Please click here to view a larger version of this figure.
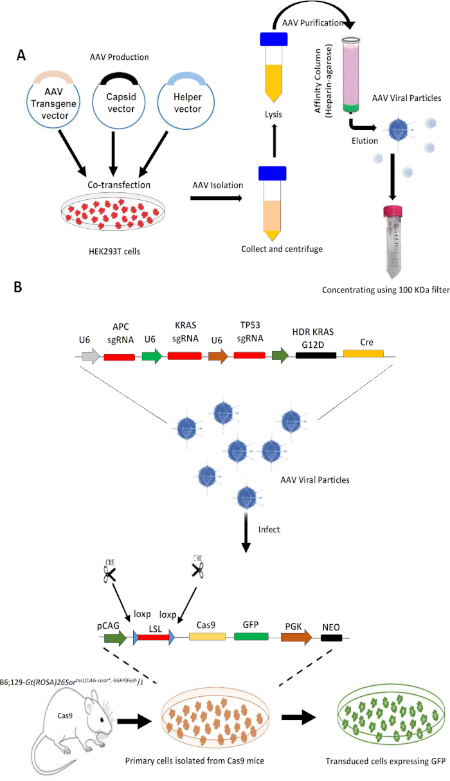
Figure 2: Design and working of a AAV-Cas9 system in transforming normal cells. (A) Outline of the production and purification of AAV particles from HEK293T after transfecting them with the AAV transgene and packaging plasmids. (B) Viral transduction of primary cells with AAV particles and the mechanism by which Cre-dependent Cas9 expression enables CRISPR editing and the transformation of primary cells to tumorigenic cells. A single AAV vector integrating gRNAs for three different genes with a knockin of the KRAS G12D allele through HDR with a Cre recombinase expression cassette was delivered into Cre-dependent Cas9 mouse primary cells. Cre recombinase activity in the transduced cells leads to excision of the LoxP-Stop-LoxP cassette and induces Cas9 and GFP expression. Please click here to view a larger version of this figure.
Validation of the AAV-Cas9 system in vitro in isolated mouse embryonic fibroblast cells
Primary mouse embryonic fibroblast (MEF) cells have been used exclusively to study the consequences of gene ablation in cellular growth and proliferation22. To validate the efficiency of the AAV-Cas9 system in transforming normal cells, we transduced MEF cells isolated from Cas9 mouse embryos (E14) with AAV. The MEF cells were isolated as described previously23,24. MEFs from 14-day-old Cas9 mouse embryos were used because they are easy to culture and transduce in vitro. In brief, to transform MEFs via AAV transduction, primary MEFs at approximately 30% confluence were considered ready for transduction. One hour before transduction, the MEF culture media was changed. The purified AAV was thawed on ice and the MEF culture media was aspirated. Viral supernatant with a titer varying from 106-1014 viral genome/mL was added to each well and incubated overnight at 37 °C. The viral supernatant was removed after 48 h, replaced with 2 mL of MEF culture medium, and then incubated at 37 °C until GFP expression, about 5 days after transduction. GFP-expressing MEFs were subcultured and validated. Note that for this vector we found that 1012 viral genomes/mL (equivalent to 1010 transducing units/mL) could efficiently transform primary MEF cells to tumorigenic MEFs (Figure 3A). The transduced cells expressed Cre, Cas9, and GFP (Figure 3B,C). To access the tumorigenic potential of the AAV transformed MEF cells, 0.5 x 106 cells (primary/AAV-transduced MEF) were injected into the tongue, lip, and subcutaneously in NOD-SCID mice. Transformed cells formed tumors in these mice, compared with the primary MEF, which did not form tumors (Figure 3D-F).
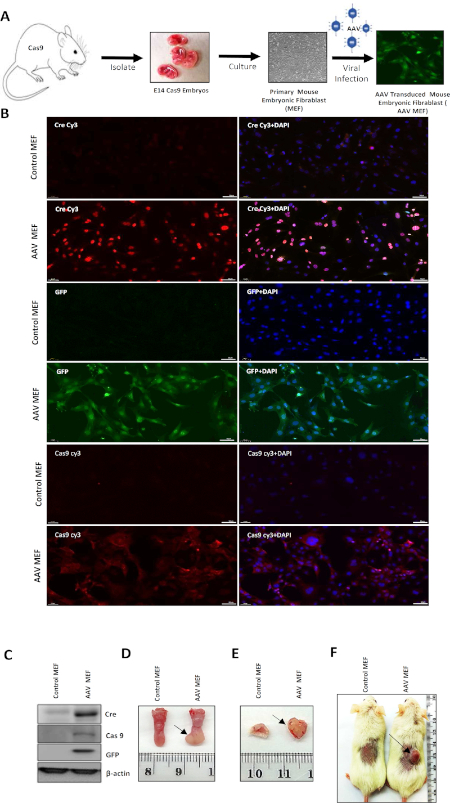
Figure 3: Validation and in vitro transformation of primary MEF cells using the AAV-Cas9 system. (A) Outline of the in vitro transformation of fibroblasts from embryos collected from Cas9 mice using AAV particles. (B) Representative immunofluorescence image of primary MEF and AAV-transduced MEF cells showing the expression of GFP, Cre, and Cas9. DAPI was used to stain the nucleus. (C) Western blotting showing Cre, Cas9, GFP, and β-actin in embryonic normal fibroblasts and genetically edited fibroblasts. (D) Representative image of a tongue tumor that developed in an NOD-SCID mouse after injecting it with transformed fibroblast cells. (E) Representative image of a lip tumor that formed in an NOD-SCID mouse after injecting it with transformed fibroblast cells. (F) Representative image of a flank tumor that developed in an NOD-SCID mouse after injecting it with transformed fibroblast cells. The arrows indicate a tumor. Please click here to view a larger version of this figure.
In vitro generation of the tumorigenic murine tongue epithelial cell line from primary tongue epithelial cells using the AAV-Cas9 system
After validating the transforming property of the AAV system using the MEF cells, we isolated primary tongue epithelial cells from Cas9 mice and cultured them in vitro. The primary tongue cells were transduced with AAV viral particles (1010 cfu/mL), and after one week cells started to express GFP. The tumorigenic transformation of primary tongue epithelial cells (Figure 4A) was confirmed using immunofluorescence and Western blotting (Figure 4B,C). The transformed cells exhibit enhanced KRT 14, E-cadherin, and GFP expression (Figure 4B). The tumorigenic transformation of primary tongue epithelial cells was confirmed by the expression of GFP (i.e., the Cre-dependent expression of Cas9 and GFP) (Figure 4C). The expression of KRT14 and E-cadherin confirms their epithelial cell characteristics. Protein validation confirmed enhanced β-catenin, phospho-ERK, and reduced p53 expression (Figure 4C), indicating the effective downregulation of APC and p53 and the incorporation of mutant KRAS into the tongue cells, thus making them tumorigenic. Genomic analysis by deep sequencing of the DNA isolated from the transformed cells confirmed the effective gene editing and frame-shift of the TP53 and APC genes by the AAV-Cas9 system (Table 2). The tumorigenic property of the AAV-transformed tongue cell line was further assessed by injecting it into the tongue of immunocompetent wild type C57BL/6 mice. The transformed tongue cells, designated as KRAS mutant epithelial cells of the tongue (KRAS Mut EpT), form tumors efficiently in C57BL/6 mice (Figure 4D). KRAS Mut EpT tumors were evaluated by an oral and maxillofacial pathologist by routine hematoxylin and eosin (H&E) staining and by immunohistochemistry. The tumors exhibited a spindle cell morphology with prominent nuclear atypia: pleomorphism, hyperchromatism, giant nuclei, several nucleoli, and a high mitotic rate with atypical mitotic figures. Perivascular invasion and angioinvasion were also noted. These morphological features were accompanied by very limited to nonexistent tumor-associated inflammation. Immunohistochemically, the neoplastic cells were cytoplasmically positive for E-cadherin and KRT 14, both markers of epithelial tissue. KRT 14 is typical of squamous epithelium, supporting the squamous origin of the tumor cells and thus the diagnosis of squamous cell carcinoma (Figure 4E). Hence, this approach provides a versatile methodology for transforming primary cells in vitro with a desired gene alteration. In addition, this methodology facilitates the use of these developed cell lines in vivo in syngeneic mice for better understanding of neoplasia in a real tumor microenvironment milieu.
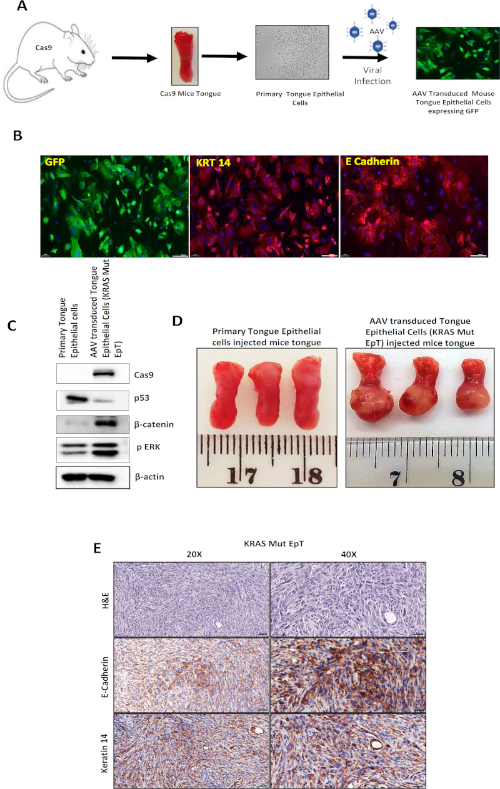
Figure 4: In vitro transformation of primary tongue epithelial cells using the AAV-Cas9 system. (A) Schematic representation of the in vitro transformation of normal epithelial cells obtained from the tongue of Cas9 mice. (B) Representative immunofluorescence image of AAV-transduced tongue epithelial cells showing the expression of KRT14, E-cadherin, and GFP. (C) Western blot showing Cas9, β-catenin, and phospho-ERK upregulation with p53 protein level downregulation, showing the genetically edited epithelial cells. (D) Representative image of a tumor that developed in the tongue of the immunocompetent mice after injecting them with transformed tongue epithelial cells (KRAS Mut EpT). (E) Representative images of tumor tissue sections for H&E, E-cadherin, and KRT 14 staining at 20x and 40x magnification. Please click here to view a larger version of this figure.
| Cell lysis buffer for viral harvesting | Volume |
| 150 mM NaCl | 876.6mg |
| 50 mM Tris-HCl | 605.7mg |
| Use HCl to adjust pH 8.5 | |
| Molecular grade water | Make up to 100 mL |
| Equilibration buffer | |
| 1 mM MgCl2 | 9.52 mg |
| 2.5 mM KCl | 18.64mg |
| Mix all in PBS 1X and adjust pH to 7.2 | Make up to 100 mL |
| Elution buffer | |
| 0.5 M NaCl | 2.92g |
| Mix all in PBS 1X and adjust pH to 7.2 | Make up to 100 mL |
| 10X Triple Enzyme Stock Solution: | |
| Collagenase | 1 g final conc. [10 mg/ml] |
| Hyaluronidase | 100 mg [1 mg/ml] |
| DNase | 20,000 Units final conc. [200 mg/ml] |
| PBS 1X | Make up to 100 mL |
| Plasmid mix (for one 14.5cm Plate) | |
| AAV pCM109 EFS Cre sg APC sg KRAS sg P53- KRAS HDR | 10µg |
| AAV 2/9 capsid vector | 10µg |
| pAD Delta F5 helper | 10µg |
| PEI (1µg/µl Stock) | 90µl |
| DMEM | 1 mL |
Table 1: Recipes of buffers used in this study.
Table 2: Genome analysis data of Kras Mut EpT cells. Please click here to view this table (Right click to download).
Discussion
Several methods have previously been used to transform primary cells in culture with variable degrees of success25,26,27,28. Most of these methods use mouse fibroblast cells for transformation14,17,18,19 or use carcinogens such as 4-nitroquinoline-1-oxide (4-NQO)21,22 for developing murine cell line models. Use of knockout and transgenic mouse models has also greatly enhanced our understanding of the various molecular pathways that govern the development and differentiation of HNC. Use of murine oral epithelial cell lines with a unique genetic pattern would certainly complement these studies, and their subsequent biochemical assays would contribute towards the treatment and diagnosis of HNC. However, only a few reports have described the establishment of mouse oral epithelial cells28,29,30,31,32,33 and these cell lines are not in widespread use.
The major limitation of these murine oral epithelial cell lines is that they are not characterized extensively at the molecular level and were found to express uncommon gene patterns. Thus, the limited molecular understanding of these murine cell line models and the lack of an efficient methodology prompted us to harness the unique potential of the AAV-CRISPR genome editing technology in Cas9 knock-in mice for establishing murine cell lines with specific gene alterations. In this study, we introduced and developed an in vitro AAV-Cas9 system for transforming primary murine cells into a tumorigenic state to confirm the genetic alterations that give rise to carcinoma cells. We successfully established murine tongue epithelial cells with specific gene mutations with high efficiency using AAV-mediated CRISPR-induced somatic gene editing by several sgRNA combinations.
This methodology offers a new experimental approach for better understanding the cell-autonomous etiology of HNC. Starting with murine primary tongue epithelial cells, we were able to transform such cells to tumorigenicity by deleting a limited set of genes and introducing a single KRAS mutation. Using this methodology, it is now possible to investigate whether other genes known to be implicated in other carcinomas can produce malignancies that more closely resemble HNC. The morphological characteristics of the tumors from KRAS Mut EpT confirmed squamous cell carcinoma. The histopathological analysis of KRAS Mut EpT tumors in syngeneic mice is associated with and supportive of the aggressive biological behavior of head and neck carcinoma. Thus, it may be possible to link the genotypes of cancer cells to specific clinical and histopathological features of the tumors.
The main advantages of this methodology are the ability to develop cell lines from primary cells from any organ using specific gene mutations and the use of AAV-Cas9, which makes the system more robust and stable. The endogenous Cas9 system makes it possible to utilize other gene knockouts. The major disadvantage of the current methodology is the time required to isolate and establish the primary culture as well as the high titer of viral particles needed to transduce the primary cells. However, this could be overcome by standardizing the isolation process for each cell type. The problem of requiring a high AAV viral titer for efficient transduction could be alleviated by use of the latest generation of capsid and helper plasmids for viral packaging and a better purification process.
In the future, a better and more efficient AAV system could be designed and extrapolated in vivo to make organ-specific genetically modified tumor models. In conclusion, establishing the murine cell lines by this approach will serve as a valuable in vitro cell culture-based tool to test several hypotheses in the context of oncology.
Divulgations
The authors have nothing to disclose.
Acknowledgements
We wish to thank Dr. Daniel Gitler for providing us with the pAd Delta 5 Helper plasmid. This work was funded by the Israel Science Foundation (ISF, 700/16) (to ME), the United States-Israel Binational Science Foundation (BSF, 2017323) (to ME and MS), the Israel Cancer Association (ICA, 20170024) (to ME), the Israel Cancer Research Foundation (ICRF, 17-1693-RCDA) (to ME), and the Concern Foundation (#7895) (to ME). Fellowship: the Alon fellowship to ME and BGU Kreitman fellowships to SJ and MP.
Materials
| Antibodies | |||
| Anti mouse HRP | Jackson ImmunoResearch | 115-035-146 | |
| Anti rabbit HRP | Jackson ImmunoResearch | 711-035-152 | |
| Cas9 Mouse mAb | Cell Signaling Technology | 14697 | |
| Cre | BioLegend | 900901 | |
| Cy3-AffiniPure Goat Anti-Mouse IgG | Jackson ImmunoResearch | 115-165-062 | |
| Cy-AffiniPure Goat Anti-Rabbit IgG | Jackson ImmunoResearch | 111-165-144 | |
| GFP | Santa Cruz Biotechnology | sc-9996 | |
| Phospho-p44/42 MAPK (Erk1/2) | Cell Signaling Technology | 4370 | |
| Rabbit monoclonal anti E cadherin | Cell Signaling Technology | 3195S | |
| Rabbit monoclonal anti-KRT 14 | Abcam | AB-ab181595 | |
| β actin | MP Biomedicals | 691001 | |
| β catenin | Cell Signaling Technology | 9582S | |
| Cell lines | |||
| HEK93T | ATCC | CRL-3216 | |
| Culture Media, Chemicals and Reagents | |||
| Bradford Reagent | Bio-Rad | 30015484 | |
| BSA | Amresco | 0332-TAM-50G | |
| DAPI fluoromount | Southern Biotech | 0100-20 | |
| DMEM | Biological Industries Israel Beit-Haemek Ltd. | 01-055-1A | |
| ECL (Westar Supernova and Westar Nova 2.0) | Cyanagen | XLS3.0100 and XLS071.0250 | |
| FBS | Biological Industries Israel Beit-Haemek Ltd. | 04-127-1A | |
| HBSS | Sigma | H6648 | |
| Heparin – Agarose | Sigma | H6508 | |
| Isolate II Genomic DNA Kit | Bioline | BIO-52066 | |
| MgCl2 | Panreac AppliChem | 300283 | |
| NaCl | Bio Lab Ltd | 1903059100 | |
| PBS | Biological Industries Israel Beit-Haemek Ltd. | 02-023-1A | |
| PEI | Polysciences | 23966-1 | |
| Pen Strep Solution | Biological Industries Israel Beit-Haemek Ltd. | 03-031-1B | |
| PFA | Santa Cruz Biotechnology | 30525-89-4 | |
| Phosphatase inhibitor cocktail | Biotool | B15001A/B | |
| Protease inhibitor cocktail | MilliporeSigma | P2714-1BTL | |
| Tris buffer | MERCK Millipore | 648311-1KG | |
| Enzymes | |||
| Benzonase | Sigma | E1014 | |
| Collagenase IV | Thermo Fisher Scientific | 17104019 | |
| DNAse | Thermo Fisher Scientific | 18047019 | |
| Hyaluronidase | MilliporeSigma | H3506 | |
| Trypsin | Biological Industries Israel Beit-Haemek Ltd. | 03-050-1B | |
| Glass wares | |||
| Cover slips | Bar Naor | BNCB00130RA1 | |
| Slides | Bar Naor | BN9308C | |
| Mouse strains | |||
| C57BL/6 | Envigo | ||
| B6;129-Gt(ROSA)26Sortm1(CAG-cas9*,-EGFP)Fezh/J | Jackson labs | 24857 | |
| NOD.CB17-Prkdc-scid/NCr Hsd (Nod.Scid) | Envigo | ||
| Plasmids | |||
| AAV pCM109 EFS Cre sg APC sg Kras sg P53- Kras HDR | Broad Institute of MIT | Kind gift from Dr Randall J Platt and Dr. Joseph Rosenbluh, Broad Institute of MIT and Harvard, Cambridge, MA 02142, USA | |
| AAV 2/9 capsid vector | Addgene | 112865 | |
| pAD Delta F5 helper | Ben Gurion University of the Negev | Provided by Dr Daniel Gitler, Department of Physiology and Cell Biology, Faculty of Health Sciences, and Zlotowski Center for Neuroscience, Ben-Gurion University of the Negev, Beer-Sheva 84105, Israel. | |
| Plastic wares | |||
| Amicon-ULTRA filter 100 KDa | Millipore | UFC910024 | |
| 0.22 µm sterile filters, 4 mm | Millex | SLGV004SL | |
| 0.45 µm sterile filters, 13 mm | Millex | SLHV013SL | |
| Culture plates | Greiner Bio-One | ||
| Falcon tubes | Greiner Bio-One |
References
- Reyes-Gibby, C. C., et al. Genome-wide association study identifies genes associated with neuropathy in patients with head and neck cancer. Scientific Reports. 8 (1), 8789 (2018).
- Riaz, N., Morris, L. G., Lee, W., Chan, T. A. Unraveling the molecular genetics of head and neck cancer through genome-wide approaches. Genes & Diseases. 1 (1), 75 (2014).
- Leemans, C. R., Snijders, P. J. F., Brakenhoff, R. H. The molecular landscape of head and neck cancer. Nature Reviews Cancer. 18 (5), 269-282 (2018).
- Jiang, X., Ye, J., Dong, Z., Hu, S., Xiao, M. Novel genetic alterations and their impact on target therapy response in head and neck squamous cell carcinoma. Cancer Management and Research. 11, 1321-1336 (2019).
- Im, W., Moon, J., Kim, M. Applications of CRISPR/Cas9 for Gene Editing in Hereditary Movement Disorders. Journal of Movement Disorders. 9 (3), 136-143 (2016).
- Li, H., et al. Genomic analysis of head and neck squamous cell carcinoma cell lines and human tumors: a rational approach to preclinical model selection. Molecular Cancer Research: MCR. 12 (4), 571-582 (2014).
- AACR Project GENIE Consortium, T.A.P.G.. AACR Project GENIE: Powering Precision Medicine through an International Consortium. Cancer Discovery. 7 (8), 818-831 (2017).
- Suh, Y., Amelio, I., Guerrero Urbano, T., Tavassoli, M. Clinical update on cancer: molecular oncology of head and neck cancer. Cell Death & Disease. 5 (1), e1018 (2014).
- Anderson, J. A., Irish, J. C., Ngan, B. Y. Prevalence of RAS oncogene mutation in head and neck carcinomas. The Journal of Otolaryngology. 21 (5), 321-326 (1992).
- Ryals, R. C., Boye, S. L., Dinculescu, A., Hauswirth, W. W., Boye, S. E. Quantifying transduction efficiencies of unmodified and tyrosine capsid mutant AAV vectors in vitro using two ocular cell lines. Molecular Vision. 17, 1090-1102 (2011).
- Smith-Arica, J. R., et al. Infection Efficiency of Human and Mouse Embryonic Stem Cells Using Adenoviral and Adeno-Associated Viral Vectors. Cloning and Stem Cells. 5 (1), 51-62 (2003).
- Li, J., Xiao, X., Kenniston, T., Kudlow, J., Giannoukakis, N. AAV Vectors. Molecular Therapy. 3 (5), S174-S192 (2001).
- Institute of Laboratory Animal Resources (U.S.). . Guide for the care and use of laboratory animals. , (1996).
- Platt, R. J., et al. CRISPR-Cas9 knockin mice for genome editing and cancer modeling. Cell. 159 (2), 440-455 (2014).
- Zhang, Y., Chirmule, N., Gao, G. P., Wilson, J. CD40 ligand-dependent activation of cytotoxic T lymphocytes by adeno-associated virus vectors in vivo: role of immature dendritic cells. Journal of Virology. 74 (17), 8003-8010 (2000).
- Xiao, X., Li, J., Samulski, R. J. Production of high-titer recombinant adeno-associated virus vectors in the absence of helper adenovirus. Journal of Virology. 72 (3), 2224-2232 (1998).
- Auricchio, A., Hildinger, M., O’Connor, E., Gao, G. P., Wilson, J. M. Isolation of Highly Infectious and Pure Adeno-Associated Virus Type 2 Vectors with a Single-Step Gravity-Flow Column. Human Gene Therapy. 12 (1), 71-76 (2001).
- Lock, M., et al. Characterization of a Recombinant Adeno-Associated Virus Type 2 Reference Standard Material. Human Gene Therapy. 21 (10), 1273 (2010).
- Challis, R. C., et al. Systemic AAV vectors for widespread and targeted gene delivery in rodents. Nature Protocols. 14 (2), 379-414 (2019).
- Rabinowitz, J. E., et al. Cross-packaging of a single adeno-associated virus (AAV) type 2 vector genome into multiple AAV serotypes enables transduction with broad specificity. Journal of Virology. 76 (2), 791-801 (2002).
- Cheng, D. T., et al. Memorial Sloan Kettering-Integrated Mutation Profiling of Actionable Cancer Targets (MSK-IMPACT). The Journal of Molecular Diagnostics. 17 (3), 251-264 (2015).
- Sun, H., Taneja, R. Analysis of Transformation and Tumorigenicity Using Mouse Embryonic Fibroblast Cells. Cancer Genomics and Proteomics. 383, 303-310 (2007).
- Durkin, M., Qian, X., Popescu, N., Lowy, D. Isolation of Mouse Embryo Fibroblasts. BIO-PROTOCOL. 3 (18), (2013).
- Jozefczuk, J., Drews, K., Adjaye, J. Preparation of Mouse Embryonic Fibroblast Cells Suitable for Culturing Human Embryonic and Induced Pluripotent Stem Cells. Journal of Visualized Experiments. (64), e3854 (2012).
- Boehm, J. S., Hession, M. T., Bulmer, S. E., Hahn, W. C. Transformation of Human and Murine Fibroblasts without Viral Oncoproteins. Molecular and Cellular Biology. 25 (15), 6464 (2005).
- Gupta, T., Sáenz Robles, T. M., Pipas, J. M. Cellular transformation of mouse embryo fibroblasts in the absence of activator E2Fs. Journal of Virology. 89 (9), 5124-5133 (2015).
- Leong, H., Blewitt, M. Retrovirus Mediated Malignant Transformation of Mouse Embryonic Fibroblasts. BIO-PROTOCOL. 3 (15), (2013).
- Parikh, N., Nagarajan, P., Sei-ichi, M., Sinha, S., Garrett-Sinha, L. A. Isolation and characterization of an immortalized oral keratinocyte cell line of mouse origin. Archives of Oral Biology. 53 (11), 1091-1100 (2008).
- Badarni, M., et al. Repression of AXL expression by AP-1/JNK blockage overcomes resistance to PI3Ka therapy. JCI Insight. 5, 125341 (2019).
- Chen, Y. F., et al. Establishing of mouse oral carcinoma cell lines derived from transgenic mice and their use as syngeneic tumorigenesis models. BMC Cancer. 19 (1), 281 (2019).
- Hoover, A. C., et al. The Role of Human Papillomavirus 16 E6 in Anchorage-Independent and Invasive Growth of Mouse Tonsil Epithelium. Archives of Otolaryngology-Head & Neck Surgery. 133 (5), 495 (2007).
- Smahel, M., et al. Metastatic MHC class I-negative mouse cells derived by transformation with human papillomavirus type 16. British Journal of Cancer. 84 (3), 374-380 (2001).
- He, L., et al. Increased Growth of a Newly Established Mouse Epithelial Cell Line Transformed with HPV-16 E7 in Diabetic Mice. PloS One. 11 (10), e0164490 (2016).

