Electromechanical Assessment of Optogenetically Modulated Cardiomyocyte Activity
Summary
We present a protocol for evaluating the electromechanical effects of GtACR1 activation in rabbit cardiomyocytes. We provide detailed information on cell isolation, culturing and adenoviral transduction, and on functional experiments with the patch-clamp and carbon-fiber techniques.
Abstract
Over the past two decades, optogenetic tools have been established as potent means to modulate cell-type specific activity in excitable tissues, including the heart. While Channelrhodopsin-2 (ChR2) is a common tool to depolarize the membrane potential in cardiomyocytes (CM), potentially eliciting action potentials (AP), an effective tool for reliable silencing of CM activity has been missing. It has been suggested to use anion channelrhodopsins (ACR) for optogenetic inhibition. Here, we describe a protocol to assess the effects of activating the natural ACR GtACR1 from Guillardia theta in cultured rabbit CM. Primary readouts are electrophysiological patch-clamp recordings and optical tracking of CM contractions, both performed while applying different patterns of light stimulation. The protocol includes CM isolation from rabbit heart, seeding and culturing of the cells for up to 4 days, transduction via adenovirus coding for the light-gated chloride channel, preparation of patch-clamp and carbon fiber setups, data collection and analysis. Using the patch-clamp technique in whole-cell configuration allows one to record light-activated currents (in voltage-clamp mode, V-clamp) and AP (current-clamp mode, I-clamp) in real time. In addition to patch-clamp experiments, we conduct contractility measurements for functional assessment of CM activity without disturbing the intracellular milieu. To do so, cells are mechanically preloaded using carbon fibers and contractions are recorded by tracking changes in sarcomere length and carbon fiber distance. Data analysis includes assessment of AP duration from I-clamp recordings, peak currents from V-clamp recordings and force calculation from carbon fiber measurements. The described protocol can be applied to the testing of biophysical effects of different optogenetic actuators on CM activity, a prerequisite for the development of a mechanistic understanding of optogenetic experiments in cardiac tissue and whole hearts.
Introduction
ChR-mediated photocurrents were first recorded in the eyespot of unicellular green algae1,2. Soon after genetic cloning and heterologous expression of Chlamydomonas reinhardtii ChR1 and ChR2, ChR were used as tools to alter the membrane potential in Xenopus oocytes and mammalian cells by light3,4. Cation non-selective ChR depolarize the membrane of cells with a resting membrane potential that is negative to the reversal potential of ChR. They can thus be used to elicit AP in excitable cells, including neurons and CM, allowing optical pacing5,6.
Complementary to cation ChR, light-driven proton, chloride and sodium pumps7,8,9 have been used to inhibit neuronal activity10,11,12. However, the latter have limitations, requiring high light intensities and sustained illumination, as one ion is transported per absorbed photon. In 2014, two independent studies by Wietek et al. and Berndt et al. described the conversion of cation-conducting ChR into ACR via mutations in the channel pore13,14. One year later, natural ACR were discovered in the cryptophyte Guillardia theta (GtACR)15. As engineered ACR showed residual cation conductance, they were replaced by natural ACR, characterized by a large single-channel conductance and high light sensitivity15. GtACR were used to silence neuronal activity by polarizing the membrane potential towards the reversal potential of chloride16,17. Govorunova et al. applied GtACR1 to cultured rat ventricular CM and showed efficient photoinhibition at low light intensity levels that were not sufficient to activate previously available inhibition tools, such as the proton pump Arch18. Our group recently reported that GtACR1-mediated photoinhibition of CM is based on depolarization and that GtACR1 can also be used, therefore, for optical pacing of CM19.
Here, we present a protocol for studying the electrophysiological and mechanical effects of GtACR1 photoactivation on cultured rabbit ventricular CM. We first describe cell isolation, culturing and transduction. Electrophysiological effects are measured using whole-cell patch-clamp recordings. Light-mediated currents at a given membrane voltage are assessed in V-clamp mode. Membrane potential dynamics are measured while electrically or optically pacing CM (I-clamp mode). Optical inhibition of electrically triggered AP is tested using sustained light application. Mechanical effects are measured using carbon fibers in combination with imaging-based tracking of sarcomere length. To do so, optically paceable cells are mechanically preloaded by attaching two carbon fibers to the plasma membrane near opposite cell ends. Sarcomere length changes are recorded during optical or electrical pacing. Finally, photoinhibition is measured during electrical field stimulation of the cells, and generated forces are analyzed.
The protocol includes the following steps shown in the flowchart in Figure 1: rabbit deep anesthesia, thiopental overdose injection, heart excision, Langendorff-perfusion and tissue digestion, mechanical dissociation of the tissue to release cells, microscopic analysis of CM yield, culturing of CM, transduction with adenovirus type 5, followed by incubation and functional experiments.
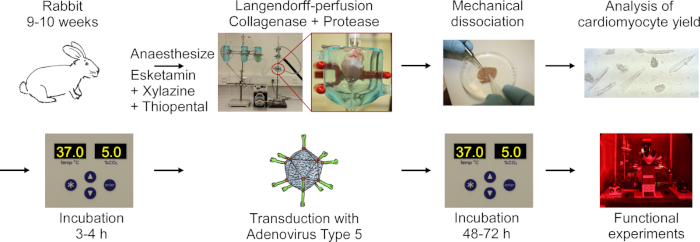
Figure 1: Flowchart of the protocol used to obtain electrically and optically paceable CM. Hearts are excised from rabbits 9-10 weeks old, and cardiac tissue is digested while being perfused using a Langendorff setup. Cells are released by mechanical agitation. The CM yield is counted under a microscope. CM are cultured, transduced with adenovirus type 5 and functional experiments are performed 48-72 hours post-transduction. Please click here to view a larger version of this figure.
Protocol
All rabbit experiments were carried out according to the guidelines stated in Directive 2010/63/EU of the European Parliament on the protection of animals used for scientific purposes and approved by the local authorities in Baden-Württemberg (Regierungspräsidium Freiburg, X-16/10R, Germany).
1. Solutions for cell isolation
- Prepare the solutions for the cell isolation with water of the following requirements (Table 1) and according to the ionic compositions listed in Table 2.
NOTE: CaCl2 and MgCl2 are added from 1 M stock solutions.
| Water requirements | |
| Conductivity [µS/cm] at 25 °C | 0.055 |
| Pyrogen [EU/mL] | < 0.001 |
| Particle (size > 0.22 µm) [1/mL] | ≤ 1 |
| Total organic carbon [ppb] | < 5 |
| Microorganisms [CFU/mL] | ≤ 1 |
| RNase [ng/mL] | < 0.01 |
| DNase [ng/mL] | < 4 |
Table 1: Water requirements.
| Physiological saline solution (1) | Low calcium, high potassium solution (2) | Enzyme solution (3) | Blocking solution | |
| NaCl [mM] | 137 | 137 | 137 | 137 |
| KCl [mM] | 4 | 14 | 14 | 14 |
| HEPES [mM] | 10 | 10 | 10 | 10 |
| Creatine [mM] | 10 | 10 | 10 | 10 |
| Taurine [mM] | 20 | 20 | 20 | 20 |
| Glucose [mM] | 10 | 10 | 10 | 10 |
| MgCl2 [mM] | 1 | 1 | 1 | 1 |
| Adenosine [mM] | 5 | 5 | 5 | 5 |
| L-Carnitine [mM] | 2 | 2 | 2 | 2 |
| CaCl2 [mM] | 1 | – | 0.1 | 0.1 |
| Na-Heparin [IU/L] | 5000 | – | – | – |
| EGTA [mM] | – | 0.096 | ||
| Collagenase type 2, 315 U/mg [g/L] | – | – | 0.6 | – |
| Protease XIV [g/L] | – | – | 0.03 | – |
| Bovine serum albumin [%] | – | – | – | 0.5 |
| Osmolarity [mOsmol/L] | 325 ± 5 | 345 ± 5 | 345 ± 5 | 345 ± 5 |
Table 2: Solutions for CM isolation.
- Adjust all solutions to pH 7.4 at 37 °C and check osmolarity.
NOTE: Dissolve the enzymes (Collagenase type 2 and Protease XIV) directly before heart excision. Oxygenate all solutions prior to use.
2. Preparation of the Langendorff-perfusion setup
NOTE: The used setup is custom-made. As depicted in Figure 2, the setup consists of three water jacketed reservoirs (1-3), one spiral counter-flow heat exchanger (4) and a water jacketed perfusion vessel (5).
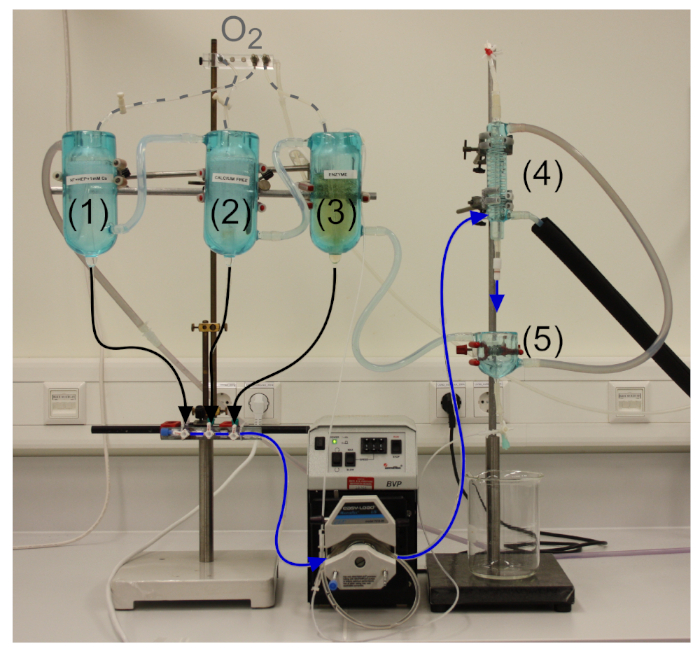

Figure 2: Langendorff-perfusion setup optimized for rabbit cell isolation. (1-3) Water jacketed reservoirs with (1) physiological saline solution, (2) low calcium, high potassium solution and (3) enzyme-containing cardioplegic solution. (4) Spiral counter-flow heat exchanger and (5) water jacketed collecting tank. The inflow of the water jacketed system is the spiral heat exchanger (temperature of solutions leaving the perfusion cannula at the end of the heat exchanger should be constant at 37 °C), followed by the perfusion vessel and the three reservoirs. All solutions are oxygenated (dashed line). Please click here to view a larger version of this figure.
- Switch on the pump of the water bath to circulate water at 38 °C in the heat exchange system and preheat all solutions to 37 °C.
NOTE: The temperature at the outflow of (4) must be controlled and constant at 37 °C. - Fill the three reservoirs with the respective solution and wash each line (black) with the corresponding solution. Fill the main (blue) line at the end without air bubbles using solution (1).
NOTE: Oxygenate the solutions prior (10 min) and during use. Fill the line from reservoir (3) to the tap with low calcium, high potassium solution. - Prepare a suture to tie the heart around the aorta at the cannula.
3. Cell isolation
- Prepare the following syringes.
- For sedation/anesthesia: Mix 0.5 mL/kg body weight esketamine hydrochloride (25 mg/mL) and 0.2 mL/kg body weight xylazine hydrochloride (2%).
- Fill two syringes with 12 mL of NaCl solution (0.9%).
- Prepare 6 mL of 12.5 mg/mL Na-thiopental, dissolved in 0.9% NaCl solution.
- Fill 0.2 mL of esketamine hydrochloride (25 mg/mL) in a syringe.
- Dilute 0.2 mL of Na-heparin (5,000 IU/mL) in 1 mL of 0.9% NaCl solution (end-concentration 1,000 IU/mL).
- Sedate/anaesthetize rabbits (9-10 weeks, New Zealand white rabbit, female or male, ~2 kg) via intramuscular injection of esketamine hydrochloride and xylazine hydrochloride (step 3.1.1).
NOTE: Rabbits need at least 10 min to be fully anaesthetized; exact duration depends on their body weight. Confirm anesthesia with the loss of the righting reflex. - Shave the chest and the ears where the veins are located.
- Insert a flexible cannula into the ear vein, fix it with tape and flush it with 0.9% NaCl solution.
- Inject 1 mL of Na-heparin solution intravenously and flush with 0.9% NaCl solution.
- Inject 0.2 mL of esketamine hydrochloride, flush again with 0.9% NaCl solution and inject Na-thiopental until apnea.
NOTE: Rabbit should not respond to the pedal withdrawal reflex. - Open the chest at the left side and remove the pericardium.
- Start the timer when the heart is excised and wash the heart twice in physiological saline solution.
NOTE: Use scissors with round tips to prevent accidental damage to cardiac tissue. - Cannulate the aorta in a bath with physiological saline solution and keep all tissue in solution. Switch on the Langendorff-perfusion system (physiological saline solution (1), speed 24 mL/min).
- Transfer the heart to the Langendorff-perfusion setup, connect the aorta to the perfusate nozzle, and tightly tie the heart with the suture around the aorta to the cannula (< 1 min).
NOTE: Pre-fill the cannula with physiological saline solution, ensure that no air bubbles enter the cannula during transportation from the cannulation site to the Langendorff setup, connect bubble-free. - Perfuse the heart until all blood is washed out (2-3 min).
- Switch to low calcium, high potassium solution (2). Perfuse for 2 more min after the heart has stopped beating and switch to enzyme solution.
- Start recirculating the enzyme solution, after 2 min from start of digestion, back into the reservoir. Decrease speed to 16 mL/min after 5 min of digestion.
- When the tissue appears soft (40-50 min of digestion), cut the heart off the cannula and separate the left ventricle.
- Release cells by mechanical dissociation (gently pulling apart the tissue with a pipette and a fine forceps to hold the tissue) in blocking solution.
- Filter the cell suspension through a mesh (pore size of 1 mm2) and centrifuge for 2 min at 22 x g (gravitational acceleration).
- Remove the supernatant containing non-myocytes and re-suspend CM in blocking solution.
4. Culturing of CM
NOTE: Perform the following steps under sterile conditions.
- Dilute Laminin (from Engelbreth-Holm-Swarm murine sarcoma basement membrane, 1 mg/mL) 1:10 in sterile phosphate buffered saline (without Ca2+/Mg2+) to a final concentration of 100 µg/mL.
- Prepare culture medium in M199-Medium with the supplements as indicated in Table 3.
| Cell culture medium in M199-Medium | |
| Creatine [mM] | 5 |
| L-Carnitine hydrochloride [mM] | 2 |
| Taurine [mM] | 5 |
| Na-Pyruvat [mM] | 1 |
| Insulin (bovine pancreas) [U/L] | 0.25 |
| Cytosine-β-D-arabinofuranoside [mM] | 0.01 |
| Gentamycin [mg/mL] | 0.05 |
Table 3: Cell culture medium.
- Sterile-filter solution (0.22 µm) and add 5% Fetal Bovine Serum.
- For patch-clamp experiments autoclave coverslips ø 16 mm, thickness No. 0, coat them with 100 µg/mL laminin directly before culturing.
- For carbon fiber experiments, coat the Petri dish surface with poly(2-hydroxyethyl methacrylate) (poly-HEMA, 0.12 g/mL in 95:5 EtOH:H20) and let it solidify.
NOTE: Cells do not 'stick' to poly-HEMA coated Petri dishes; this is crucial for their friction-less contraction in cell mechanics studies. - After re-suspended CM have settled (~10-15 min), remove the supernatant, and then re-suspend CM in culture medium.
- Count CM with a Neubauer chamber and seed at a target density of 17,500 cells/mL either on laminin coated coverslips or in poly-HEMA coated Petri dishes.
- Incubate cells at 37 °C, 5% CO2 for 3-4 hours. Exchange the medium (37 °C) of coverslip seeded cells.
- Add adenovirus (type 5) coding for GtACR1-eGFP at a multiplicity of infection (MOI) of 75 and start functional experiments after 48 hours.
NOTE: After transduction keep the cells in the dark. Use red illumination when working with blue or green light-activated proteins. A commercially available adenoviral delivery system (see Table of Materials) is used to clone the genes encoding GtACR1-eGFP into the adenoviral vector. The insert of interest, here GtACR1-eGFP, is PCR amplified and then combined with an adenoviral vector including a CMV promoter in an IN-Fusion Cloning reaction. The CMV (human cytomegalovirus) promoter is commonly used to drive overexpression of transgenes in mammalian cells. eGFP is an enhanced green fluorescent protein derived from Aequorea victoria with an excitation maximum of 488 nm and an emission maximum at 507 nm. Adenovirus (type 5) was externally produced at Charité-Universitätsmedizin Berlin, Institut für Pharmakologie, Berlin, Prof. Dr. Michael Schupp.
CAUTION: Adenoviral transduction is categorized as BSL-2 safety level work, and appropriate safety measures are legally required.
5. Functional experiments
NOTE: Recordings are performed using an inverted fluorescence microscope. Filter the transmission light by a red band-pass filter (630/20 nm) in the condenser to avoid co-activation of GtACR1.
- Patch-clamp setup
- Use an amplifier in combination with an analogue-to-digital converter. Use a data acquisition software to record current and voltage data (see Table of Materials).
NOTE: The recorded data are digitized at 10 kHz and filtered at 5 kHz.
- Use an amplifier in combination with an analogue-to-digital converter. Use a data acquisition software to record current and voltage data (see Table of Materials).
- Carbon fiber setup
- Use a camera to detect carbon fiber position and sarcomere length by tracking changes in optical contrast (carbon fibers appear as darker structures, overlaid on the striated cell pattern). A schematic representation of the setup is shown in Figure 3.
NOTE: Sarcomere length is calculated in real time using a fast Fourier transform (FFT) of the power spectrum of the striation pattern.
- Use a camera to detect carbon fiber position and sarcomere length by tracking changes in optical contrast (carbon fibers appear as darker structures, overlaid on the striated cell pattern). A schematic representation of the setup is shown in Figure 3.
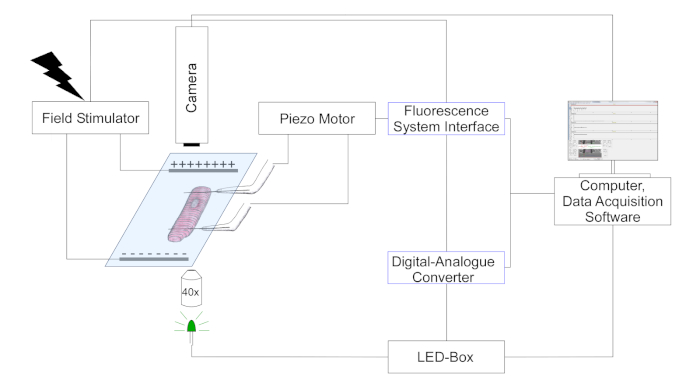
Figure 3: Scheme depicting experimental setup for carbon fiber measurements. (Drawing is not at scale). Two carbon fibers are attached on a cell and their position is controlled by a piezo positioner. The pacer is used for electrical field stimulation. Multi-color LEDs are coupled into the epifluorescence port of the inverted microscope for illumination of cells in the object plane. LED power is controlled via a dedicated control box, which receives digital pulses via the digital output of the digital-analogue-converter (DAC). The DAC communicates via analogue output with the fluorescence system interface. A black-and-white camera (774 pixels by 245 lines) for cellular imaging is connected to the computer to track sarcomere length and carbon fiber bending. Please click here to view a larger version of this figure.
- Timed illumination
- Provide light for fluorescence microscopy and activation of light-gated ion channels via an external custom-built LED control box, comprising three LEDs of different color (460 nm, 525 nm, 640 nm, see Table of Materials).
- Modify the microcontroller and graphical user interface (GUI) code for the control box to allow control of the LED via external Time to Live (TTL) pulses, generated in data acquisition software protocols (see Table of Materials). Transmit TTL pulses to the LED control box via the digital-analogue-converter.
- Drive the LED and choose the number of pulses via the GUI. Upon receiving the command from the GUI the microcontroller starts a process on a new core. In this process the TTL input as well as a control switch set from the GUI will be continuously checked.
- When the TTL input is positive, the microcontroller turns the LED on and then resumes checking the TTL input. Once the TTL signal returns to zero, the microcontroller turns the LED off and reduces the number of pulses left by one. If at any point the control switch is false or the number of pulses is zero, the microcontroller stops this process until a new command is received from the GUI.
- Directly couple the LEDs into the backport of the microscope.
- Determination of light intensity in the object plane
- Measure the illuminated area with a stage micrometer (objective magnification 40x, A = 0.8 mm2).
- Use an optical power meter (see Table of Materials).
- Define the settings for the experiments: excitation wavelength (525 nm), objective magnification (40x), excitation filter (530/20 nm) or mirror, and read out the light power [W] at various LED-input voltages.
- Calculate the light intensity [W/mm2] by dividing the light power [W] by the illuminated area [mm2] (here: 0.8 mm2).
NOTE: Measure the actual light power with the respective protocols in step 5.6 to check if short light pulse durations of 10 ms reach and long durations hold the set value (Supplemental Figure 1).
- Preparation for patch-clamp experiments
- Prepare the following external and internal solutions (Table 4; for water requirements see Table 1).
- Adjust the osmolarity with glucose to 300 ± 5 mOsmol/L. Aliquot the internal solution and store at -20 °C.
NOTE: Keep the internal solution on ice for the day of the recording. Keep the external solution at room temperature. The here described patch-clamp solutions were based on previously used solutions and the Cl– concentration was changed to lower, more physiological levels7. For characterization of ion selectivity of the respective optogenetic actuator, we suggest to vary the concentrations of major ions (e.g., Cl–, Na+, K+, H+) in the extra- and intracellular solutions19. - Take off the recording electrode from the pipette holder and remove the silver chloride layer from the silver wire with very fine sandpaper.
NOTE: Do this step at the beginning of each measuring day. - Connect the wire to the positive pole of a 1.5 V battery and immerse in 3 M KCl solution for silver chloride coating for 10 min.
NOTE: The negative pole is connected to a reference silver wire immersed in the 3 M KCl solution. - Prepare the measuring chamber: put silicon grease on the frame of the measuring chamber and place a coverslip (diameter: 50 mm, thickness No. 0) on the top of the frame that the chamber is sealed.
- Put a reference silver/silver-chloride pellet electrode in the bath and connect it with the head stage.
- Pull 1.7 – 2.5 MΩ patch pipettes from soda lime glass capillaries (outer diameter: 1.55 mm, inner diameter: 1.15 mm) with a micropipette puller (see Table of Materials).
- Start data acquisition software and adjust the membrane test (pulse 10 mV for 15 ms, baseline 0 mV).
| External bath solution | Internal pipette solution | |
| NaCl [mM] | 140 | – |
| KCl [mM] | 5.4 | 11 |
| CaCl2 [mM] | 1 | – |
| MgCl2 [mM] | 2 | 2 |
| Glucose [mM] | 10 | – |
| HEPES [mM] | 10 | 10 |
| K-Aspartate [mM] | – | 119 |
| Mg-ATP [mM] | – | 3 |
| EGTA [mM] | – | 10 |
| pH | 7.4 (NaOH) | 7.2 (KOH) |
| Osmolarity (adjust with Glucose) [mOsmol/L] | 300 ± 5 | 300 ± 5 |
Table 4: Patch-clamp solutions.
- Protocols for patch-clamp measurements
- Record photoactivation protocol in the V-clamp mode at a holding potential of -74 mV. Use light pulses of 300 ms.
NOTE: We suggest performing V-clamp recordings close to the resting membrane potential of cultured CM (established in I-clamp; in our hands between -79 mV and -77 mV both for transduced and non-transduced CM19). Freshly isolated cells show a mean resting membrane potential of -79 mV (Supplemental Figure 2, all values after correction for liquid junction potential). - Record AP in I-clamp mode at 0 pA.
- For electrical pacing, inject current pulses of 10 ms (ramp from 0 pA to the set value within 10 ms), 0.25 Hz and find the threshold to elicit AP. Record AP by current injections of 50% more than the threshold.
- For optical pacing use light pulses of 10 ms, 0.25 Hz at the minimal light intensity to elicit reliable AP.
- Record photoinhibition in I-clamp mode at 0 pA. Elicit AP as described in step 5.6.2.1 and apply sustained light for 64 s at 4 mW/mm2 after 15 electrically triggered AP.
NOTE: Figure 6F shows a photoinhibition protocol where during sustained light higher current injections are applied. Starting from 1.5 times the threshold (here: 0.7 nA) the injected current was increased in steps of 0.1 nA (final level: 2.2 nA). At all tested current amplitudes, sustained light application inhibited AP generation.- As a control experiment, pause electrical stimulation for 64 s without light application.
- Record photoactivation protocol in the V-clamp mode at a holding potential of -74 mV. Use light pulses of 300 ms.
- Patch-clamp experiments
NOTE: Perform the following experiments in the dark (red light can be used for blue/green light-activated tools).- Place coverslip with cells in measuring chamber with external solution and select fluorescent CM.
NOTE: eGFP-positive cells can be detected using a blue LED (460 nm) in combination with a band-pass excitation filter (450 nm – 490 nm), a 510 nm dichroic mirror and a 515 nm long-pass emission filter. If other fluorescent tags are used, use corresponding LED and fluorescence filter sets. If a high transduction efficiency is achieved (in our hands >99% with the GtACR1 adenovirus), there is no need to check eGFP fluorescence before the functional experiments; this avoids potential GtACR1 pre-activation. - Fill the patch pipette with internal solution. Ensure that there are no air bubbles in the tip.
- Attach pipette to the pipette holder, inserting the recording silver-chloride coated silver wire in the internal solution.
- After reaching the cell-attached configuration, switch to whole-cell mode in the data acquisition software with a holding potential of -74 mV. Rupture the membrane by gently applying negative pressure to access the whole-cell configuration. This is indicated by an immediate increase in the measured capacitance.
- Run the protocols described in section 5.6.
- Place coverslip with cells in measuring chamber with external solution and select fluorescent CM.
- Carbon fiber technique
- Produce carbon fibers.
- Use glass capillaries with the following parameters: outer diameter: 2.0 mm, inner diameter: 1.16 mm, length: 100 mm (see Table of Materials). Using a micropipette puller, pull the glass capillary into two pipettes of the same length (total taper length ~11 mm, Figure 5) to a final inner diameter of ~30 µm.
NOTE: Settings used for the first and second pull are 85.2% (proportion to the maximum output of the puller) and 49.0%, respectively (will depend on the puller, type and age of the filament). - Bend the pipettes up to 45° with a self-made micro forge using settings of 12 V, 24 A (see Figure 4 for details of the pipette bending setup).
- Align the capillary (2) on the red line in the orientation circle (5), keep the positioning of the capillary constant so the length of the bend part is always the same after the center of the orientation circle (radius of 4.5 mm).
- Bend the capillary up to 45° (green line) by pushing down the tip of the capillary with the bender (3) and forge by heating-up the filament (4) until the capillary captures the 45° angle even after the bender is removed.
- Fit the carbon fibers (provided from Prof. Jean-Yves Le Guennec) into the fine tip of the glass capillary under a stereo microscope. Use fine forceps with soft tubing at the end to increase grip and decrease the risk of damaging the fibers.
NOTE: These fibers are characterized by microstructures, which increase the contact surface between fibers and cells, thereby improving adhesion20. - Cut the carbon fibers at a length of 2 mm and use super glue (cyanoacrylate) to fix the fiber to the front section of the capillary.
NOTE: The longer the fibers are, the more they bend upon application of the same force.
- Use glass capillaries with the following parameters: outer diameter: 2.0 mm, inner diameter: 1.16 mm, length: 100 mm (see Table of Materials). Using a micropipette puller, pull the glass capillary into two pipettes of the same length (total taper length ~11 mm, Figure 5) to a final inner diameter of ~30 µm.
- Calibrate carbon fibers.
- Calibrate the carbon fibers using a force transducer with a sensitivity of 0.05 mN/V and a force range of 0 – 0.5 mN (see Table of Materials).
NOTE: This setup is custom-made in order to measure compression instead of "pull". - Attach the capillary with the carbon fiber to a holder that is controlled by a micromanipulator and a piezo motor.
- Place the tip of the fiber in contact with the force sensor, but without producing any force and move the piezo motor in steps of 10 µm (total movement of 60 µm) towards the sensor and read out the measured voltage (E) in Volt.
NOTE: Make sure the force transducer is contacted by the very tip of the free end of the carbon fiber. - Repeat these measurements three times.
- Use Formula 1 to calculate the force for each piezo position (ΔΕ
 difference of measured voltage between the piezo motor steps):
difference of measured voltage between the piezo motor steps):

NOTE: The sensitivity of the force transducer is dependent on the model of the transducer (here: 0.05 mN/V = 50 µN/V). The gain can be set at the controller. - Plot the force [µN] against the piezo position. The slope corresponds to the fiber stiffness [µN/µm].
- Calibrate the carbon fibers using a force transducer with a sensitivity of 0.05 mN/V and a force range of 0 – 0.5 mN (see Table of Materials).
- Record force of contracting CM.
NOTE: Perform the following experiments in the dark (red light can be used for blue/ green light-activated tools).- Coat the surface of the measuring chamber with poly-HEMA. Fill the measuring chamber with external bath solution and put a few drops of the cultured cell suspension in the chamber (step 4.9).
- Attach capillaries with carbon fibers to the stage micromanipulator. Select GtACR1-expressing CM by checking the ability to induce contractions by application of short green-light pulses. Align the carbon fibers near-horizontally to the surface of the measuring chamber.
- Lower the first fiber onto the cell surface. Attach the second fiber parallel to the first fiber at the other end of the CM. The ideal alignment is near-perpendicular to the cell axis.
NOTE: Attach the fiber by gently pushing the cell to the bottom surface. Release the pressure before attaching the second fiber. Do not stretch the cell by attaching the second fiber. - After both fibers are attached on the cell, lift the fibers, so the cell has no contact to the chamber surface anymore and is able to contract without any friction.
- Focus the sarcomeres in the data acquisition software (see Table of Materials) and set the sarcomere length tracking window (Figure 7A I (3)) between the fibers.
NOTE: Resulting FFT power spectrum (Figure 7A I (2)) shows ideally one sharp peak, representing the average sarcomere length. - Track fiber bending using the edge detection module. Set the detection areas with the red and green window and define a threshold (red and green horizontal line) at the first derivative of the light intensity trace (Figure 7A III).
- Start to optically pace the cell at 0.25 Hz (if possible, try faster pacing rates) and track sarcomere length and fiber bending.
NOTE: Fiber holder position, LED trigger and electric stimulation pulses are controlled via the data acquisition software (see Table of Materials). - After recording at least 15 optically elicited contractions, field-stimulate the cell electrically (see Table of Materials). Find the threshold to elicit contractions and record by applying 1.5 times of the threshold voltage.
- For the inhibition protocol, apply electrical stimuli to elicit contractions and then expose to sustained light of 64 s (at various light intensities).
- Produce carbon fibers.
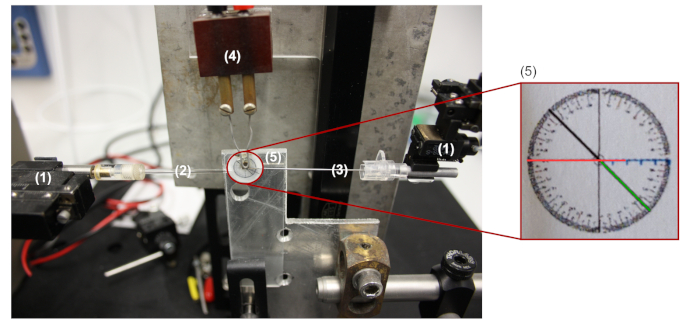

Figure 4: Pipette bending setup. (1) The micromanipulator on the left side is used to control the position of the capillary, and a second micromanipulator on the right is used to bend it. (2) Capillary. (3) Bender. (4) Microforge. (5) Orientation circle. Please click here to view a larger version of this figure.
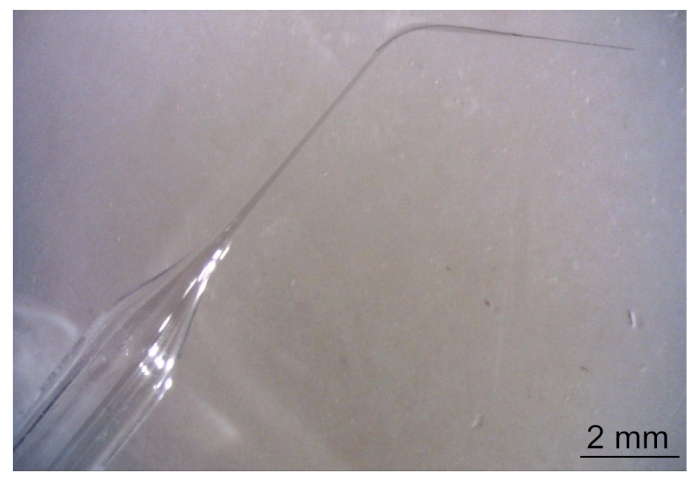

Figure 5: Pipette with carbon fiber. Please click here to view a larger version of this figure.
6. Data analysis
- Patch-clamp recordings
NOTE: Correct all recorded and command voltages for the liquid junction potential after the experiment. Determine liquid junction potential in the data acquisition software by using the tool junction potential calculator (for the stated patch-clamp solutions in Table 4: 14.4 mV at 21 °C). Subtract the liquid junction potential from the recorded/command voltage.- For I-clamp AP recordings, check electrical pacing versus optical pacing. Calculate the AP duration (APD) at 20 and 90% repolarization with a custom-written script (Supplemental Material). Determine resting membrane potential and AP amplitude.
NOTE: Determine the APD as described in Wang K. et al.21 The script to load .abf files is generally accessible via the following link: https://de.mathworks.com/matlabcentral/fileexchange/22114-fcollman-abfload. Average APD values for at least 6 AP. - For V-clamp photoactivation, check if baseline is at 0 pA. If not adjust baseline to zero. Analyze the recorded current triggered by 300 ms light pulses at -74 mV. Transfer the data to data analysis software and determine the peak and average stationary current.
- For I-clamp AP recordings, check electrical pacing versus optical pacing. Calculate the AP duration (APD) at 20 and 90% repolarization with a custom-written script (Supplemental Material). Determine resting membrane potential and AP amplitude.
- Carbon fiber experiments
- Contraction recordings during optical pacing: Load the recorded data in the data acquisition software and read out the baseline and maxima of the carbon fiber bending and the sarcomere length changes.
NOTE: Average values for 10 contractions of a stable recording. - Measure the cell width and calculate the cross-sectional area of the cell assuming an elliptical cross-section (Figure 7B II).
NOTE: The formula for the area of an ellipse is A = π·a·b (Formula 2) where a is the distance from the center to the vertex and b is the distance from the center to the co-vertex. In our case this means a = (width of the cell)/2 and b = (thickness of the cell)/2. According to Nishimura et al.22 the thickness of CM can be estimated to be one third of the cell width so that A = π·(1/2)·width·(1/2)·thickness = π·(1/4)·width·(1/3)·width = π·(1/12)·width2. - Calculate the end-systolic force (F):
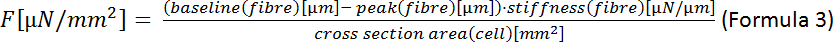
- Calculate the end-systolic cell deformation (ESD):

NOTE: Further contractile parameters can be analyzed: resting sarcomere length, time to peak, time to 90% relaxation, fractional sarcomere shortening, maximum velocity of contraction and relaxation (see software acquisition manual).
- Contraction recordings during optical pacing: Load the recorded data in the data acquisition software and read out the baseline and maxima of the carbon fiber bending and the sarcomere length changes.
Representative Results
GtACR1-eGFP was expressed in cultured rabbit CM (Figure 6 insert) and photocurrents were measured with the patch-clamp technique. Photoactivation of GtACR1 shows large inward directed currents at -74 mV. In Figure 6A peak current (IP) at 4 mW/mm2 is 245 pA. AP were triggered either electrically (Figure 6B) or optically (Figure 6C) with current injections 1.5 times the threshold, or short depolarizing light pulses of 10 ms, respectively. Analyzing APD values, electrically paced CM show an APD 20 of 0.24 ± 0.08 s and an APD 90 of 0.75 ± 0.17 s, whereas optically paced CM show an APD 20 of 0.31 ± 0.08 s and an APD 90 of 0.81 ± 0.19 s (SE, n = 5, N = 2, in the here presented example APD 20electrical = 0.17 s; APD 20optical = 0.27 s and APD 90electrical = 0.61 s; APD 90optical = 0.68 s; Figure 6D). Optically paced CM show a slower AP onset (Figure 6D). CM activation was inhibited upon sustained illumination (for 64 s, 4 mW/mm2) by polarizing the membrane potential towards the reversal potential of chloride, here -58 mV (Figure 6E). Higher current injections than 1.5 times the threshold do not elicit AP generation (Figure 6F). Generated peak forces were determined from carbon fiber bending (Figure 7B,C,E). The CM generated 232 µN/mm2 upon electrical pacing (Figure 7B) and 261 µN/mm2 following optical pacing (Figure 7C). Prolonged green-light pulses inhibit contractions (Figure 7E). Following optical inhibition for 64 s, reoccurring contractions generate a lower contractile force, and force values recover towards baseline after ~10 contractions (pacing at 0.25 Hz, Figure 7D) in keeping with diastolic calcium loss from rabbit CM.
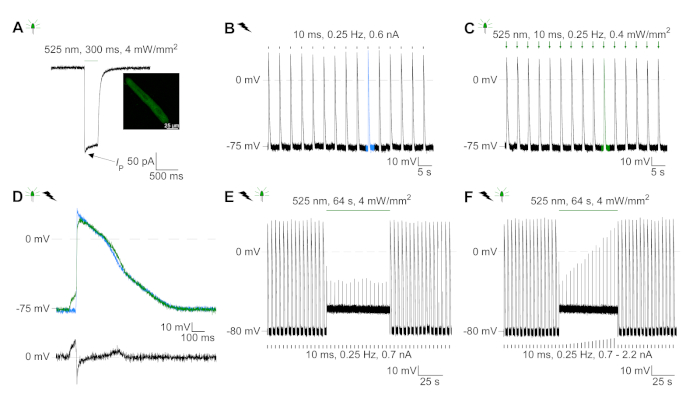
Figure 6: Representative patch-clamp recordings of electrically and optically paced/inhibited CM. (A) Representative photocurrent at -74 mV using a light pulse of 300 ms, 4 mW/mm2. IP indicates the peak current. The insert shows a GtACR1-eGFP positive cell. (B) Representative AP recording at 0 pA using a current ramp of 10 ms, 0.6 nA to electrically pace the CM. (C) Representative AP recording at 0 pA using light pulses of 10 ms, 0.4 mW/mm2. (D) Top graph shows the overlay of the 10th AP of electrically (blue) and optically (green) activated CM. AP were aligned by the maximum change in membrane potential (dV/dt max). Bottom graph shows the difference of membrane potential between optically and electrically triggered AP (Eoptical–Eelectrical). (E) Electrically triggered AP were inhibited under sustained light of 64 s, 4 mW/mm2. (F) AP are inhibited by higher current injections than 1.5 times the threshold (from 0.7 nA in steps of 0.1 nA to 2.2 nA) under sustained light. Please click here to view a larger version of this figure.
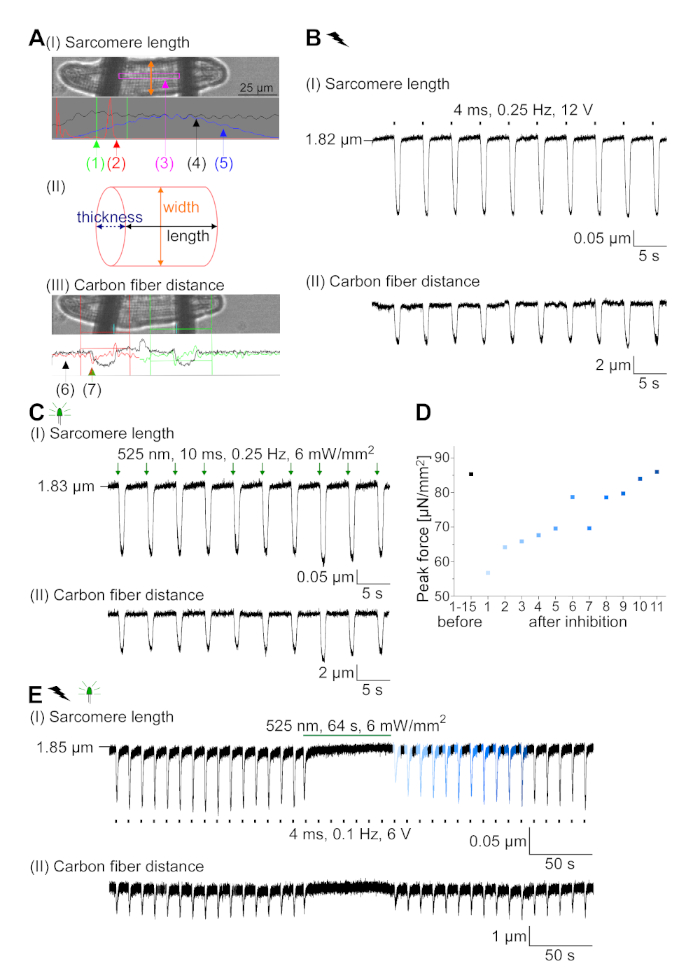
Figure 7: Representative data from carbon fiber recordings of optically and electrically paced/inhibited CM. (A) Display in the data acquisition software. Image (I) shows the measured CM with the window for calculating sarcomere length. Cell width is labelled in orange. (1) Range of relevant frequencies. (2) FFT power spectrum shows the frequency of the sarcomere spacing on the cell. The average sarcomere length is calculated from the peak frequency. (3) Sarcomere length tracking window. (4) Intensity trace. (5) The intensity trace multiplied by a Hamming window is the windowed intensity trace. Scheme (II) shows the elliptical cross-section of the cell. Width in orange and thickness in dashed blue. Image (III) shows the position of the carbon fibers with the respective detection boxes, left in red and right in green. (6) Intensity trace. (7) First derivative of intensity trace (see data acquisition software manual). (B) Representative trace of electrically elicited contractions. Panel (I) shows the sarcomere length shortening, panel (II) the distance between the two carbon fibers. (C) Representative trace of optically elicited contractions (525 nm, 0.25 Hz, 10 ms, 6 mW/mm2). Panel (I) shows the sarcomere length shortening, panel (II) the distance between the two carbon fibers. (D) Generated peak force from contraction 1 to 11 after a pause caused by inhibition of AP generation. (E) Representative trace of optical inhibition of contractions under sustained illumination (525 nm, 64 s, 6 mW/mm2). Panel (I) shows the sarcomere length shortening, panel (II) the length between the two carbon fibers. Please click here to view a larger version of this figure.
| A | area |
| ACR | anion channelrhodopsin |
| AP | action potential |
| APD | action potential duration |
| CFU | colony forming unit |
| ChR | channelrhodopsin |
| CM | cardiomyocyte |
| eGFP | enhanced green fluorescent protein |
| ESD | end systolic cell deformation |
| EU | endotoxin units |
| F | force |
| FFT | fast Fourier transform |
| GtACR | Guillardia theta anion channelrhodopsin |
| GUI | graphical user interface |
| I-clamp | current-clamp |
| IU | international units |
| MOI | multiplicity of infection |
| poly-HEMA | poly(2-hydroxyethyl methacrylate) |
| V-clamp | voltage-clamp |
Table 5: List of abbreviations.
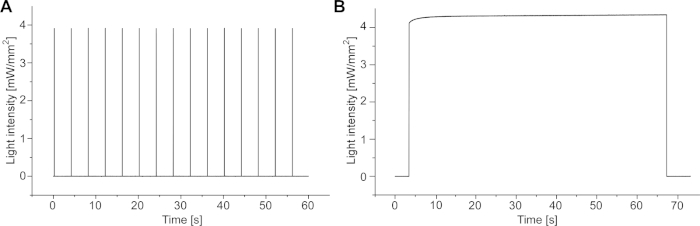
Supplemental Figure 1: Light intensity measurements with optical power meter. (A) Measurement of 10 ms light pulses at 4 mW/mm2. (B) Measurement of sustained illumination of 64 s at 4 mW/mm2. Please click here to view a larger version of this figure.
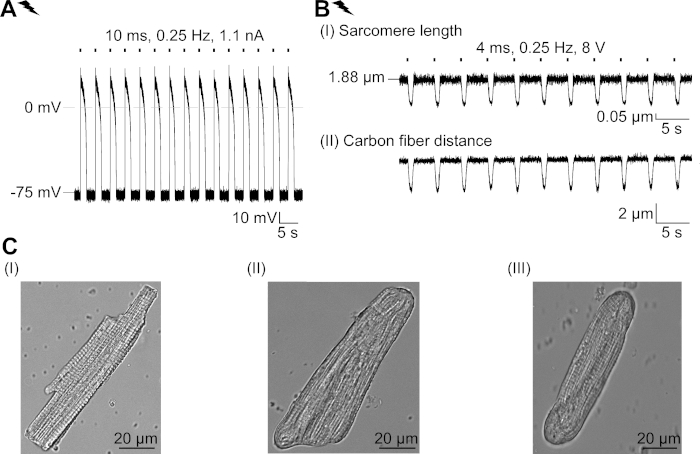
Supplemental Figure 2: Properties of freshly isolated CM and their structural adaptation in culture. (A) AP recording of a freshly isolated CM (APD 20 of 1.11 ± 0.34 s, APD 90 of 1.96 ± 0.32 s, n = 7, N = 2). Mean resting membrane potential of -79.3 ± 0.8 mV (n = 7, N = 2). (B) Carbon fiber recording of an electrically paced freshly isolated CM. Mean peak force of 205 ± 78 µN/mm2 (n = 7, N = 2). (C) Confocal images of a freshly isolated CM (I); untransduced (II) and transduced (III) CM after 48 hours in culture. Please click here to view a larger version of this figure.
Supplemental Material: MatLab script to determine APD and resting membrane potential. Please click here to download this file.
Discussion
Whereas optogenetic tools enable modulation of excitable cell electrophysiology in a non-invasive manner, they need thorough characterization in different cell types (e.g., CM) to allow one to choose the best available tool for a specific experimental design. The patch-clamp technique is a standard method for assessing cellular electrophysiology. In the whole-cell configuration, it allows one to record photo-activated currents across the plasma membrane or temporal changes in membrane voltage following light stimulation/inhibition. Optogenetic manipulation of electrical excitation also affects CM contractions. We use sarcomere tracking and carbon fiber-assisted force measurements to quantify the effects of optical interrogation on the mechanical activity of myocytes.
We describe a protocol to characterize the basic effects of a light-gated chloride channel, GtACR1, in CM. As model system, we chose rabbit CM, as their electrophysiological characteristics (e.g., AP shape and refractory period) resemble those observed in human CM more closely than rodent CM. Moreover, rabbit CM can be cultured for several days, long enough for adenoviral delivery and expression of GtACR1-eGFP. Notably, isolated CM change their structural properties in culture over time, including rounding of cell endings and gradual loss of cross-striation, T-tubular system and caveolae23,24. In line with this, functional alterations have been reported in cultured CM: depolarization of the resting membrane potential, prolongation of the AP and changes in cellular Ca2+ handling. For review of cellular adaptations in culture, please see Louch et al.25. Supplemental Figure 2 shows exemplary AP and contraction measurements of freshly isolated CM for comparison with those observed in cultured CM (Figure 6, Figure 7) using the here presented protocol.
Whole-cell patch-clamp recordings enable direct measurements of photocurrent properties (e.g., amplitudes and kinetics) and light-induced changes in membrane potential or AP characteristics at high temporal resolution. However, such recordings have several limitations: Firstly, the cytosol is replaced by the pipette solution in whole-cell recordings, which is advantageous to control ionic electrochemical gradients, but has the intrinsic disadvantage of washing-out cellular organelles, proteins and other compounds, thus potentially affecting cellular electrical responses. Secondly, side effects like activation of additional ion channels resulting from non-physiologically long depolarization (e.g., slow time constants of light-gated ion channels) are difficult to assess as our method only allows one to detect changes in APD, but not to conduct direct measurements of ionic concentrations in electrophysiologically relevant cell compartments. This could be done with fluorescent indicators (e.g., Ca2+ sensors) or ion-selective electrodes. Further characterization may include light intensity titrations, determination of pH-dependency, photocurrent kinetics at different membrane potentials, and recovery kinetics during repetitive light stimulation.
In contrast to patch-clamp recordings, single-cell force measurements enable analysis of cellular contractions of intact myocytes without affecting their intracellular milieu. Secondary effects on ion concentrations (e.g., Ca2+) can be indirectly assessed by determining generated force amplitude and dynamics (e.g., maximum velocity of contraction and relaxation; here not analyzed). Force measurements with the carbon fiber technique have an advantage over freely contracting cells as they provide direct information on passive and active forces in pre-loaded cells (i.e., in conditions that are more similar to the in situ or in vivo settings). Mechanical preloading is especially important when analyzing cellular contractility, as stretch affects force production and relaxation26,27.
Optogenetic approaches allow for spatiotemporally precise manipulation of the cellular membrane potential, both in single CM and intact cardiac tissue. Classically, ChR2, a light-gated cation non-selective channel, has been used for depolarization of the membrane potential, whereas light-driven proton and/or chloride pumps were used for membrane hyperpolarization. Both groups of optogenetic actuators require high expression levels, as ChR2 is characterized by an intrinsically low single-channel conductance28 and light-driven pumps maximally transport one ion per absorbed photon. Furthermore, prolonged activation of ChR2 in CM may lead to Na+ and/or Ca2+ overload, and light-driven pumps may change trans-sarcolemmal H+ or Cl– gradients29,30. In search for alternative tools for optogenetic control of CM activity, we recently tested the natural anion channelrhodopsin GtACR1, characterized by a superior single-channel conductance and higher light sensitivity compared to cation ChR such as ChR2. We found that GtACR1 activation depolarizes CM and can be used for optical pacing and inhibition, depending on the light pulse timing and duration. An additional advantage of using ACR instead of cation ChR might be the more negative reversal potential of Cl– compared to Na+, reducing artificially introduced ion currents. As we have previously shown, optical pacing with GtACR1 may lead to AP prolongation as a result of the slow component of GtACR1 channel closure, which could be overcome by using faster GtACR1 mutants19. However, AP prolongation is much less pronounced when using a lower, more physiological intracellular Cl– concentration (see Figure 6). Moreover, GtACR1-mediated inhibition by prolonged illumination results in profound membrane depolarization, which again could activate secondary Na+ and Ca2+ influx, thereby altering the activity of voltage-gated channels. In our measurements, we find that AP and contraction parameters recover to baseline within 40 s after a light-induced inhibition for 1 min (see Kopton et al. 2018, Figure 6, Figure 7). Light-gated K+ channels offer a potent alternative for silencing CM without affecting the CM resting membrane potential31.
In future we would like to quantitatively compare different optogenetic tools for their potential to inhibit cardiac activity. To this end, we test a variety of light-gated ion channels including ACR, ChR2 and red-shifted ChR variants32, as well as hyperpolarizing actuators such as halorhodopsin or the light gated adenylyl cyclase bPAC in combination with the potassium channel SthK (PAC-K)31.
The here presented protocol can be used for in-depth characterization of the electromechanical properties of CM. It is principally applicable also to CM from other species, and to CM isolated from diseased myocardium. Optical stimulation allows one to pace CM at different frequencies, and different preloads can be tested during carbon fiber contraction experiments. An interesting experiment would be to use low-intensity illumination for subthreshold depolarization, to mimic gradual increase in the resting membrane potential, as can be observed during the development of cardiac tissue remodeling during disease progression. Finally, functional measurements could be combined with Ca2+ imaging for further insight into excitation-contraction coupling, or with pharmacological interventions to evaluate the effects of different drugs on CM activity.
Divulgations
The authors have nothing to disclose.
Acknowledgements
We thank Stefanie Perez-Feliz for excellent technical assistance, Dr. Jonas Wietek (Humboldt-University, Berlin, Germany) for providing the pUC57-GtACR1 plasmid, Prof. Dr. Michael Schupp (Charité- Universitätsmedizin Berlin, Institut für Pharmakologie, Berlin) for the adenovirus production and Dr. Anastasia Khokhlova (Ural Federal University) for sharing her expertise to improve the cell isolation protocol and to re-design the pipette bending setup. The project was funded by the German Research Foundation (SPP1926: SCHN 1486/1-1; Emmy-Noether fellowship: SCHN1486/2-1) and the ERC Advanced Grant CardioNECT.
Materials
| Equipment – Cell isolation/Culturing/Transduction | |||
| Adeno-X Adenoviral System 3 CMV | TaKaRa, Clontech Laboratories, Inc., Mountain View, California, USA | ||
| Aortic cannula | Radnoti | 4.8 OD x 3.6 ID x 8-9 L mm | |
| Coverslips ø 16 mm, Thickness No. 0 | VWR International GmbH, Leuven, Belgium | 631-0151 | Borosilicate Glass |
| Griffin Silk, Black, 2 m Length, Size 3, 0.5 mm | Samuel Findings, London, UK | TSGBL3 | |
| Incubator | New Brunswick, Eppendorf, Schönenbuch, Switzerland | Galaxy 170S | |
| Langendorff-perfusion set-up | Zitt-Thoma Laborbedarf Glasbläserei, Freiburg, Germany | Custom-made | |
| Langendorff-pump | Ismatec, Labortechnik-Analytik, Glattbrugg-Zürich, Switzerland | ISM444 | |
| Mesh: Nylon Monodur filter cloth | Cadisch Precision Meshes Ltd | 800 µm holes, 1 m wide | |
| Neubauer chamber | VWR International GmbH, Leuven, Belgium | 717806 | |
| Rabbit, New Zealand White | Charles River | Strain Code: 052 | |
| Scissors | Aesculap AG, Tuttlingen, Germany | BC774R | Bauchdeckenschere ger. 18cm |
| Sterile filter, 0.22 µm | Merck, Darmstadt, Germany | SLGP033RB | |
| Equipment – Patch-clamp | |||
| Amplfier | AxonInstruments, Union City, CA, United States | Axopatch 200B | |
| Coverslip ø 50 mm, Thickness No. 1 | VWR International GmbH, Leuven, Belgium | 631-0178 | Borosilicate Glass |
| Digitizer Axon Digidata | Molecular Devices, San José, CA, United States | 1550A | |
| Filter (530/20) | Leica Microsystems, Wetzlar, Germany | 11513878 | BZ:00 |
| Filter (630/20) | Chroma Technology, Bellows Falls, Vermont, United States | 227155 | |
| Headstage | AxonInstruments, Union City, CA, United States | CV203BU | |
| Interface | Scientifica, Uckfield, UK | 1U Rack, 352036 | |
| LED 525 nm | Luminus Devices, Sunnyvale, CA, United States | PT-120-G | |
| LED control software | Essel Research and Development, Toronto, Canada | ||
| LED control system | custom-made | ||
| Micropipette Puller | Narishige Co., Tokyo, Japan | PP-830 | |
| Microscope inverted | Leica Microsystems, Wetzlar, Germany | DMI4000B | |
| Motorised Micromanipulator | Scientifica, Uckfield, UK | PatchStar | |
| Optical power meter | Thorlabs, Newton, NJ, United States | PM100D | |
| Silicone Grease | RS Components, Corby, UK | 494-124 | |
| Silver wire | A-M Systems, Sequim, WA, United States | 787500 | Silver, Bare 0.015'', Coated 0.0190'', Length 25 Feet |
| Soda lime glass capillaries | Vitrex Medical A/S, Vasekaer, Denmark | 160213 BRIS, ISO12772 | 1.55 OD x 1.15 ID x 75 L mm |
| Software Axon pClamp | Molecular Devices, San José, CA, United States | Version 10.5 | |
| Software MatLab2017 | The MathWorks, Inc. | ||
| Stage micrometer | Graticules Optics LTD, Tonbridge, UK | 1 mm | |
| Equipment – Carbon fiber | |||
| Carbon fibers | provided from Prof. Jean-Yves Le Guennec | BZ:00 | |
| Digitizer Axon Digidata | Molecular Devices, Sunnyvale, CA, United States | 1550B | |
| Filter (530/20) | Leica Microsystems, Wetzlar, Germany | 11513878 | |
| Filter (630/20) | Chroma Technology, Bellows Falls, Vermont, United States | 227155 | |
| Fluorescence System Interface | IonOptix, Milton, MA United States | FSI-800 | 2.0 OD x 1.16 ID x 100 L mm |
| Force Transducer System | Aurora Scientific Inc., Ontario, Canada | 406A | |
| Glass capillaries for force measurements | Harvard Apparatus, Holliston, Massachusetts, United States | GC200F-10 | |
| Interface National Instruments | National Instruments, Budapest, Hungary | BNC-2110 | |
| LED 525 nm | Luminus Devices, Sunnyvale, CA, United States | PT-120-G | |
| LED control box | Essel Research and Development, Toronto, Canada | ||
| LED control system | custom-made | ||
| Microcontroller | Parallax Inc., Rocklin, California, United States | Propeller | |
| Micropipette Puller | Narishige Co., Tokyo, Japan | PC-10 | |
| Microscope inverted | Leica Microsystems, Wetzlar, Germany | DMI4000B | |
| MyoCam-S camera | IonOptix, Dublin, Ireland | ||
| MyoCam-S camera Power | IonOptix, Milton, MA, United States | MCS-100 | |
| MyoPacer Field Stimulator | IonOptix Cooperation, Milton, MA, United States | MYP100 | |
| Piezo Motor | Physik Instrumente (PI) GmbH & Co. KG, Karlsruhe, Germany | E-501.00 | |
| Silicone Grease | RS Components, Corby, UK | 494-124 | |
| Software Axon pClamp | Molecular Devices, San José, CA, United States | Version 10.5 | |
| Software IonWizard | IonOptix, Dublin, Ireland | Version 6.6.10.125 | |
| Software MatLab2017 | The MathWorks, Inc. | ||
| Stage micrometer | Graticules Optics LTD, Tonbridge, UK | 1 mm | |
| Chemicals | |||
| Adenosine | Sigma-Aldrich, St. Louis, Missouri, United States | A9251-100G | |
| Bovine serum albumin | Sigma-Aldrich, St. Louis, Missouri, United States | A7030-50G | |
| CaCl2 | Honeywell Fluka, Muskegon, MI, USA | 21114-1L | |
| L-Carnitine hydrochloride | Sigma-Aldrich, St. Louis, Missouri, United States | C9500-25G | |
| Collagenase type 2, 315 U/mg | Worthington, Lakewood, NJ, USA | LS004177 | |
| Creatine | Sigma-Aldrich, St. Louis, Missouri, United States | C0780-50G | |
| Cytosine-β-D-arabinofuranoside | Sigma-Aldrich, St. Louis, Missouri, United States | C1768-100MG | |
| EGTA | Carl Roth GmbH + Co. KG, Karlsruhe, Germany | 3054.3 | |
| Esketamine hydrochloride, Ketanest S 25 mg/mL | Pfizer Pharma PFE GmbH, Berlin, Germany | PZN-07829486 | |
| Fetal Bovine Serum | Sigma-Aldrich, St. Louis, Missouri, United States | F9665 | |
| Gentamycin 50 mg/mL | Gibco, Life Technologies, Waltham, MA, USA | 15750-037 | |
| Glucose | Sigma-Aldrich, St. Louis, Missouri, United States | G7021-1KG | |
| Heparin-Sodium, 5,000 IU/mL | Braun Melsungen AG, Melsungen, Germany | PZN-03029843 | |
| HEPES | Sigma-Aldrich, St. Louis, Missouri, United States | H3375-1KG | |
| Insulin (bovine pancreas) | Sigma-Aldrich, St. Louis, Missouri, United States | I6634-50MG | |
| K-aspartate | Sigma-Aldrich, St. Louis, Missouri, United States | A6558-25G | |
| KCl | VWR International GmbH, Leuven, Belgium | 26764.260 | 1 mg/mL |
| KOH | Honeywell Fluka, Muskegon, MI, USA | 35113-1L | |
| Laminin from Engelbreth-Holm-Swarm murine sarcoma basement membrane | Sigma-Aldrich, St. Louis, Missouri, United States | L2020-1MG | |
| M199-Medium | Sigma-Aldirch, St. Louis, Missouri, United States | M4530 | |
| Mg-ATP | Sigma-Aldrich, St. Louis, Missouri, United States | A9187-1G | |
| MgCl2 | Sigma-Aldrich, St. Louis, Missouri, United States | 63069-500ML | |
| NaCl | Fisher Scientific, Loughborough, Leics., UK | 10428420 | |
| NaCl-Solution 0.9%, Isotone Kochsalz-Lösung 0.9% | Braun Melsungen AG, Melsungen, Germany | 3200950 | |
| NaOH | AppliChem GmbH, Darmstadt, Germany | A6579 | without Ca2+/Mg2+ |
| Na-pyruvat | Sigma-Aldrich, St. Louis, Missouri, United States | P2256-100MG | |
| Phosphate Buffered Saline | Sigma-Aldrich, St. Louis, Missouri, United States | D1408-500ML | |
| Poly(2-hydroxyethyl methacrylate) | Sigma, Poole, UK | 192066 | |
| Protease XIV from Streptomyces griseus | Sigma-Aldrich, St. Louis, Missouri, United States | P5147-1G | |
| Taurine | Sigma-Aldrich, St. Louis, Missouri, United States | T0625-500G | |
| Thiopental Inresa 0.5 g | Inresa Arzneimittel GmbH, Freiburg, Germany | PZN-11852249 | |
| Xylazine hydrochloride, Rompun 2% | Bayer Vital GmbH, Leverkusen, Germany | PZN-01320422 |
References
- Harz, H., Hegemann, P. Rhodopsin-regulated calcium currents in Chlamydomonas. Nature. 351, 489-491 (1991).
- Litvin, F. F., Sineshchekov, O. A., Sineshchekov, V. A. Photoreceptor electric potential in the phototaxis of the alga Haematococcus pluvialis. Nature. 271, 476-478 (1978).
- Nagel, G., et al. Channelrhodopsin-1: A Light-Gated Proton Channel in Green Algae. Science. 296 (5577), 2395-2398 (2002).
- Nagel, G., et al. Channelrhodopsin-2, a directly light-gated cation-selective membrane channel. Proceedings of the National Academy of Sciences of the United States of America. 100 (24), 13940-13945 (2003).
- Boyden, E. S., Zhang, F., Bamberg, E., Nagel, G., Deisseroth, K. Millisecond-timescale, genetically targeted optical control of neural activity. Nature Neuroscience. 8 (9), 1263-1268 (2005).
- Bruegmann, T., et al. Optogenetic control of heart muscle in vitro and in vivo. Nature Methods. 7 (11), 897-900 (2010).
- Lozier, R. H., Bogomolni, R. A., Stoeckenius, W. Bacteriorhodopsin: a light-driven proton pump in Halobacterium Halobium. Biophysical journal. 15 (9), 955-962 (1975).
- Schobert, B., Lanyi, J. K. Halorhodopsin is a light-driven chloride pump. Journal of Biological Chemistry. 257 (17), 10306-10313 (1982).
- Inoue, K., et al. A light-driven sodium ion pump in marine bacteria. Nature Communications. 4, 1678 (2013).
- Han, X., et al. A High-Light Sensitivity Optical Neural Silencer: Development and Application to Optogenetic Control of Non-Human Primate Cortex. Frontiers in Systems Neuroscience. 5, 18 (2011).
- Zhang, F., et al. Multimodal fast optical interrogation of neural circuitry. Nature. 446 (7136), 633-639 (2007).
- Grimm, C., Silapetere, A., Vogt, A., Bernal Sierra, Y. A., Hegemann, P. Electrical properties, substrate specificity and optogenetic potential of the engineered light-driven sodium pump eKR2. Scientific Reports. 8, 9316 (2018).
- Wietek, J., et al. Conversion of channelrhodopsin into a light-gated chloride channel. Science. 344 (6182), 409-412 (2014).
- Berndt, A. Structure-Guided Transformation. Science. 344 (6182), 420-424 (2014).
- Govorunova, E. G., Sineshchekov, O. A., Janz, R., Liu, X., Spudich, J. L. Natural light-gated anion channels: A family of microbial rhodopsins for advanced optogenetics. Science. 349 (6248), 647-650 (2015).
- Mohamed, G. A., et al. Optical inhibition of larval zebrafish behaviour with anion channelrhodopsins. BMC Biology. 15 (1), 103 (2017).
- Mauss, A. S., Busch, C., Borst, A. Optogenetic Neuronal Silencing in Drosophila during Visual Processing. Scientific Reports. 7, 13823 (2017).
- Govorunova, E. G., Cunha, S. R., Sineshchekov, O. A., Spudich, J. L. Anion channelrhodopsins for inhibitory cardiac optogenetics. Scientific Reports. 6, 33530 (2016).
- Kopton, R. A., et al. Cardiac Electrophysiological Effects of Light-Activated Chloride Channels. Frontiers in Physiology. 9, 1806 (2018).
- Peyronnet, R., et al. Load-dependent effects of apelin on murine cardiomyocytes. Progress in Biophysics and Molecular Biology. 130, 333-343 (2017).
- Wang, K., et al. Cardiac tissue slices: preparation, handling, and successful optical mapping. American Journal of Physiology-Heart and Circulatory Physiology. 308 (9), 1112-1125 (2015).
- Nishimura, S., et al. Single cell mechanics of rat cardiomyocytes under isometric, unloaded, and physiologically loaded conditions. American Journal of Physiology-Heart and Circulatory Physiology. 287 (1), 196-202 (2004).
- Mitcheson, J. S., Hancox, J. C., Levi, A. J. Action potentials, ion channel currents and transverse tubule density in adult rabbit ventricular myocytes maintained for 6 days in cell culture. Pflugers Archiv European Journal of Physiology. 43 (6), 814-827 (1996).
- Burton, R. A. B., et al. Caveolae in Rabbit Ventricular Myocytes: Distribution and Dynamic Diminution after Cell Isolation. Biophysical Journal. 113 (5), 1047-1059 (2017).
- Louch, W. E., Sheehan, K. A., Wolska, B. M. Methods in cardiomyocyte isolation, culture, and gene transfer. Journal of Molecular and Cellular Cardiology. 51 (3), 288-298 (2011).
- Janssen, P. M., Hunter, W. C. Force, not sarcomere length, correlates with prolongation of isosarcometric contraction. The American Journal of Physiology. 269 (2), 676-685 (1995).
- Monasky, M. M., Varian, K. D., Davis, J. P., Janssen, P. M. L. Dissociation of force decline from calcium decline by preload in isolated rabbit myocardium. Pflugers Archiv European Journal of Physiology. 456 (2), 267-276 (2008).
- Kleinlogel, S., et al. Ultra light-sensitive and fast neuronal activation with the Ca 2+-permeable channelrhodopsin CatCh. Nature Neuroscience. 14 (4), 513-518 (2011).
- Schneider-Warme, F., Ravens, U. Using light to fight atrial fibrillation. Cardiovascular Research. 114 (5), 635-637 (2018).
- Chow, B. Y., et al. High-performance genetically targetable optical neural silencing by light-driven proton pumps. Nature. 463 (7277), 98-102 (2010).
- Bernal Sierra, Y. A., et al. Potassium channel-based optogenetic silencing. Nature Communications. 9 (1), 4611 (2018).
- Oda, K., et al. Crystal structure of the red light-activated channelrhodopsin Chrimson. Nature Communications. 9 (1), 3949 (2018).

