Lung CT Segmentation to Identify Consolidations and Ground Glass Areas for Quantitative Assesment of SARS-CoV Pneumonia
Summary
The aim of this protocol is to provide a time efficient way to segment volumes of interest on high-resolution CT scans to use for further radiomics analysis.
Abstract
Segmentation is a complex task, faced by radiologists and researchers as radiomics and machine learning grow in potentiality. The process can either be automatic, semi-automatic, or manual, the first often not being sufficiently precise or easily reproducible, and the last being excessively time consuming when involving large districts with high-resolution acquisitions.
A high-resolution CT of the chest is composed of hundreds of images, and this makes the manual approach excessively time consuming. Furthermore, the parenchymal alterations require an expert evaluation to be discerned from the normal appearance; thus, a semi-automatic approach to the segmentation process is, to the best of our knowledge, the most suitable when segmenting pneumonias, especially when their features are still unknown.
For the studies conducted in our institute on the imaging of COVID-19, we adopted 3D Slicer, a freeware software produced by the Harvard University, and combined the threshold with the paint brush instruments to achieve fast and precise segmentation of aerated lung, ground glass opacities, and consolidations. When facing complex cases, this method still requires a considerable amount of time for proper manual adjustments, but provides an extremely efficient mean to define segments to use for further analysis, such as the calculation of the percentage of the affected lung parenchyma or texture analysis of the ground glass areas.
Introduction
In the current year, the world is facing a health emergency, the pandemic caused by the novel Coronavirus, Sars-CoV2. Even if, up to the present date, many aspects regarding the physiopathology of the COVID-19 infection are still unclear, it shares several characteristics with its “ancestors” SARS1 and MERS. In particular, it has been proven that the virion spike proteins interact with the Angiotensin Converting Enzyme Type-2, a receptor well represented on the alveolar endothelial cells, but ubiquitous in the human organism, thus having the potentiality to give systemic symptoms1.
For diagnosis, the current standard is the real-time reverse transcriptase-polymerase chain reaction (rt-PCR), a test performed on pharyngeal swabs. Although radiological imaging is not officially recognized in the diagnostic path for the detection of the disease, high resolution computed tomography (HRCT) proved to be a valuable aid to the clinical and epidemiological management of affected patients, due to the relatively low sensitivity of the rt-PCR, the current scarcity of specialized laboratories and of the necessary reagents, and the high operator-dependency.
The Radiological Society of North America (RSNA) released a consensus statement, endorsed by the society of Thoracic Radiology and the American College of Radiology (ACR), that classifies the CT appearance of COVID19 into four categories in order to standardize the reporting, dividing the interstitial pneumonia patterns into “typical”, “atypical”, “indetermined”, and “negative”2.
The “typical” pattern is characterized by the presence of round-shaped Ground Glass Opacities (GGO), usually with a sub-pleural location on the dorsal basal segments. The GGO can be associated with “Crazy Paving” areas of thickened septa, or other signs of organizing pneumonia. The “indetermined” pattern is characterized by the absence of the “typical” pattern findings, with diffuse GGO areas with a perihilar distribution, with or without consolidative areas. The “atypical” pattern is characterized by either the absence of the “typical” or “undetermined” signs, and the presence of lobar consolidations, “tree in bud”, smooth thickening of the septa and pleural effusion; in this presentation no GGO are detectable. The “negative” pattern is characterized by the absence of the aforementioned pathological findings.
According to literature, some patients may present with a high clinical suspect of COVID-19 supported by epidemiological criteria and imaging finding with negative rt-PCR3,4. On the other hand, it has been reported that patients with a positive rt-PCR and suggestive clinical findings, do not present pathological findings on HRCT5.
Nowadays, it is of paramount interest for the scientific community to apply image analysis techniques when quantitatively studying the characteristics of this disease. A recent study has applied automated segmentation technique of the lung parenchyma to identify the percentage of aerated lung in patients affected by COVID-19, correlating this value with the prognosis, and demonstrating that patients with a more severe lung involvement presented more risk of being admitted in the Intensive Care Unit (ICU), and having worse outcomes6.
The segmentation is the contouring of regions of interest (ROIs) inside a volume acquired through an imaging technique, such as HRCT. This activity may be carried out though three methods: manual, semi-automatic and automatic. Manual segmentation, thanks to the experience of a trained radiologist, consists of labelling voxels belonging to the pathological area. The main disadvantages of this method are the large amount of time required and the fact that it is operator-dependent.
Semi-automatic methods allow to speed up the segmentation as the operator can modify a segmentation mask obtained through the classic methods of image processing (e.g., threshold on pixel intensity, clustering, etc.). However, these techniques are not easy to implement in clinical practice as they require extensive manual intervention in the most complicated cases18.
Automatic segmentation methods, currently of limited use, employ artificial intelligence to obtain ROIs. In particular, a recent study aims to use automatic segmentation in the quantification of ground glass areas in patients suffering from COVID-19 interstitial pneumonia19. The definition of a segmentation protocol for the pathological areas on the HRCT images is the real first step toward the subsequent radiomics analysis, in order to identify features that could help to further understand the physiopathology of the disease, and serve as an accurate prognostic factor potentially influencing the treatment.
This paper offers a guide to obtain accurate and efficient segments representing the pathological findings of COVID-19 pneumonia using “3D Slicer”7,8,9,10.
Protocol
This protocol follows the guidelines of the institutional human research ethics committee.
1. Downloading the DICOM images
- Download the DICOM images and transfer them in the workstation dedicated to the segmentation, with the 3D software installed. If planning to work on a personal computer, anonymize the DICOM data.
2. Importing the HRCT study on the 3D Slicer software
- In the software opening screen (corresponding to the Welcome to Slicer section in the drop-down menu) select Load DICOM Data. Alternatively, select the DCM icon in the upper left corner of the toolbar.
- In the upper left corner of the DICOM Browser panel select Import, then select the location of the HRCT study. Select the folder with the DCM images, and then select Import.
- Now that the HRCT study has been successfully imported, click on the Load button.
3. Creating the Segments
- Create segments in the Segmentations section of the drop-down menu or directly in the sub-section Segment Editor found in the drop-down menu’s Segmentation section, or again as a dedicated icon in the toolbar.
- In the drop-down menu next to Master Volume, select the HRCT study.
- Select Add and create three new segments, which will be automatically named Segment 1, 2 and 3. Double click on each one and rename them respectively “tlv” (Total Lung Parenchyma), “ggo” (Ground Glass Opacities) and “cons” (Consolidations). If in the HRCT study additional pathological findings coexist, such as pleural effusion, lung cancer, fibrotic areas and so on, create additional segments. The same applies to artifacts.
4. Definition of the TLP segment
NOTE: An accurate definition of the TLP segment is fundamental, as it will be used for masking the HRCT during the definition of the GGO and CDs segments.
- As a first step, in the Segment Editor section, after selecting the TLP segment, choose the Threshold instrument.
- Set a threshold large enough to include both the healthy lung parenchyma and the ground glass opacities. Using the values reported in scientific literature with the necessary adjustments to comply with the textbook definition of GGO, we found that setting the threshold between -1000 HU and -250 HU works well6,14,15,16,17. Then select Apply.
- The segment so defined will include both the air inside the lungs and the air outside the chest (i.e., the air outside the patient). In order to isolate the lung parenchyma, use the Islands tool, found in the Segment Editor, and choose Keep selected island. Left click inside the chest so that anything outside the chest will be excluded from the segment.
- Next, include any area of consolidation inside the TLP segment. The consolidations usually have attenuation values (Hounsfield Units: HU) similar to those of soft tissues that constitute the chest wall and the mediastinum. This is the reason why the Threshold instrument cannot be used for this purpose and the consolidations will need to be added manually, using the Scissors and Paint tools found in the segment editor.
- Use the Scissors tool to add portions of lung parenchyma to the segment. Choose the Fill Inside operation, select Free Form as the shape and Symmetric as the slice cut. Choose a thickness appropriate to the size of the consolidation (e.g., from 3 to 20 mm); and then, start progressively adding small portions of TLP, until the consolidation has been completely included.
- The Paint tool is easier and sometimes faster to use, but might be less precise, depending on the area to include. When the Sphere Brush option is active, this tool can add 3D portions of lung parenchyma to the segment. The size of the brush can be easily modified.
- While adding the consolidations to the TLP segment, do not include portions of thoracic wall or mediastinum. This could prove difficult as in patients affected by COVID19 consolidations adjacent to the posterior thoracic wall are quite common. Use the Erase tool to correct the segmentation or select Undo to discard the last action.
- At this stage, use the smoothing algorithm once more to delete small imperfections that might have occurred during the manual part of the segmentation. Now the TLP segment definition is concluded.
5. Definition of the GGO segment
- To define the GGO segment, use the threshold tool.
- Select the GGO segment.
- Set the threshold between -750 HU and -150 HU.
- Before selecting Apply, go to the Masking section right underneath and select TLP in the Editable Area drop down menu. In the Overwrite Other Segment drop-down menu, select None. This is extremely important as, if skipped, once defined the GGO segment, also the TLP segment would be modified. This must be avoided, as the TLP segment is the mask on which all the other segments are defined.
- Now select Apply.
- At this point it will probably be necessary to use the smoothing algorithm once again, in order to exclude all the small vessels and the physiological interstitial elements that have a density similar to that of ground glass from the segment. Start with a Kernel size of 3 mm and, if needed, increase it progressively, to a maximum of 6-7 mm. Note that increasing the Kernel size too much might determine the loss of small areas of ground glass, which would be left outside the GGO segment. Because of this, the smoothing algorithm should be used with the proper caution.
- In this case too, apply needed corrections with the Paint, Scissors and Erase tool. During this phase remember to keep the TLP mask active at all times to avoid including portions of chest wall adjacent to the “ground glass” areas inside the segment. This is because, if erroneously segmented, they would automatically be excluded.
- When obtaining this segment, pay close attention to the possible presence of artifacts generated by movements of hearth and diaphragm (this happens if the patient could not hold his breath during the exam). If present and previously included in the TLP segment, eliminate these artifacts from the “ground glass” segment, for example through the Islands tool with the Remove Selected Island option or using the Scissors and Erase tools. Depending on the study’s goal, the artifacts might be segmented separately, using a dedicated segment or included in the TLP segment and excluded from other segments. Now the GGO segment has been defined.
6. Definition of the CDs segment
- To define the CDs segment, proceed the same way as to define the GGO segment.
- Select the CDs segment. Be sure to work with the TLP masking always active.
- Set an adequate threshold. For consolidations the range varies, roughly, from -150 HU to 100 HU.
- Apply the smoothing algorithm varying the Kernel size as needed.
- Use a combination of the tools Scissors, Erase, Paint and Islands in order to keep inside the segment only the real consolidations, excluding large vessels, pleural effusions, artifacts, disventilatory bands and other non COVID-related lesions.
- If needed, apply a final smoothing might be applied.
7. Saving the segments
- Save segments as an “.nrrd” file, or convert to binary label maps from the "Data" module.
8. Extracting volumes from the defined segments.
- From the Segment Statistics module, obtain a table with details on volume and surfaces of the segments.
Representative Results
The proposed method has been refined through trials and errors, testing it on 117 patients affected by COVID-19 pneumonia with a positive rt-PCR test.
After a short learning curve, the time needed to obtain the segments can vary from 5 to 15 minutes, depending on the presentation pattern.
As shown on Figure 1, the method yields precise segments: this can be observed by noticing the exact correspondence with the HRCT. The 3D rendering helps to assess the correspondence and quickly review the segmentation results. A quantitative assessment of the amount of lung parenchyma affected can be obtained, to replicate the results reported from Colombi et al.6 and Lanza et al.13.
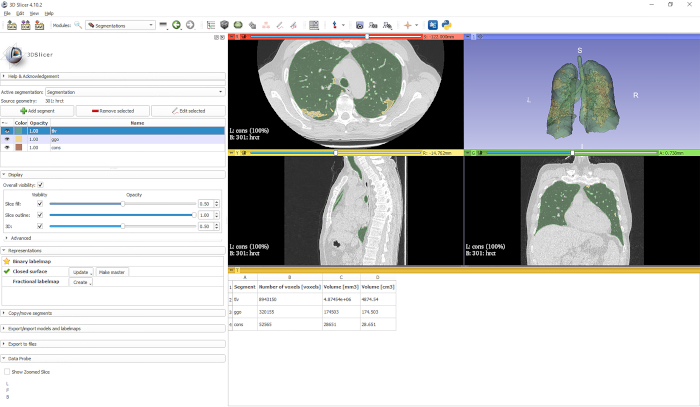
Figure 1: Representative results volume and surface analysis. The screenshot from the 3D Slicer interface represents the results obtained from the “Segment Statistics” module, which can be used to quantitatively evaluate the volume of the affected lung parenchyma. Please click here to view a larger version of this figure.
Discussion
Segmentation represents a fundamental step for performing modern quantitative radiology studies, and is necessary to apply radiomics or texture analysis techniques. Pathological findings in the lungs represent one of the most challenging to segment, for the lack of defined anatomical borders and a small difference in attenuation value when compared to the healthy areas.
The source images must present with a minimum of artifacts if possible, especially on the pathological areas, and this is sometimes hard to achieve when studying a disease that compromises breath holding; therefore, researchers can consider excluding compromised HRCTs or defining a segment dedicated to artifacts to be eliminated from further analysis.
It is possible to install an extension to 3D Slicer, called the “chest imaging platform”11, which allows faster, more automated operations on lung segments, with particular interest on aerated lung. It was decided not to adopt this method, since it requires extensive manual intervention when GGO and consolidations have a sub-pleural distribution, which is the case of the pathology explored in this paper.
An automatic segmentation method for interstitial lung diseases has been reported12; even so, this method required having previous knowledge of the affected areas’ features. The technique proposed in this study represents an easy to learn and reproducible approach to segmentation of pathological findings of the lung, the features extracted from which could provide future means of automatically segment viral interstitial lung diseases, and represent accurate prognostic factors.
The segmentation method proposed has some limitations.
First of all, downloading the DICOM images from the workstations is a process that might require a variable amount of time, thus doing so for vast numbers of patients might be a troublesome work. Any potential application of this segmentation method to the clinic practice must consider this critical issue, until the point when segmentation plugins become widely available on PACS platforms.
Secondly, the segmentation of the COVID-19 related pathological areas might be complicated in patients with concurrent chronic lung disease (e.g., lung cancer, lung fibrosis, etc.) whose radiological findings consist in areas with the same densities of those typical of COVID-19 patterns. The same concern should be considered in CT scans with respiratory artifacts. These artifacts are quite common in patients with COVID-19 infection, being commonly related to dyspnea and respiratory failure, especially in old/middle aged patients.
Moreover, patients with a severe interstitial pneumonia (characterized by many lung consolidations and crazy paving opacities) require a more extensive manual segmentation and consequently, a vast amount of time. In general, the higher the severity of the interstitial pneumonia, the more extensive the manual segmentation required, the longer the segmentation time.
However, the degree of precision offered by advanced evaluation techniques such as the ones proposed by Lanza and Colombi could add limited information on patients with already severe lung conditions compared to the standard clinical and radiological evaluations20.
Finally, it should be noted that any radiologist who has no experience with 3D slicer needs an adequate training time, as it is not an intuitive software and requires some time to be mastered even in its basilar functions.
Divulgations
The authors have nothing to disclose.
Acknowledgements
This work was supported by funding from the Department of Radiology of the University of Bologna.
Materials
| CT Scanner | General Electrics Healthcare | 64-MDCT VCT lightSpeed | The CT scanner used for HRCT acquisitions |
| Desktop Computer | ThinkCentre | The computer used to download the DICOM files and run 3D Slicer |
References
- Zheng, Y., et al. COVID-19 and the cardiovascular system. Nature Reviews Cardiology. 17, 259-260 (2020).
- Simpson, S., et al. Radiological Society of North America Expert Consensus Statement on Reporting Chest CT Findings Related to COVID-19. Endorsed by the Society of Thoracic Radiology, the American College of Radiology, and RSNA. Radiology: Cardiothoracic Imaging. 2, 2 (2020).
- Xie, X., et al. Chest CT for Typical 2019-nCoV Pneumonia: Relationship to Negative RT-PCR Testing. Radiology. , 200343 (2020).
- Huang, P., et al. Use of Chest CT in Combination with Negative RT-PCR Assay for the 2019 Novel Coronavirus but High Clinical Suspicion. Radiology. 295 (1), 22-23 (2020).
- Fang, Y., et al. Sensitivity of Chest CT for COVID-19: Comparison to RT-PCR. Radiology. , 200432 (2020).
- Colombi, D., et al. Well-aerated Lung on Admitting Chest CT to Predict Adverse Outcome in COVID-19 Pneumonia. Radiology. , 201433 (2020).
- . 3D Slicer [software] Available from: https://www.slicer.org (2020)
- Kikinis, R., Pieper, S. D., Vosburgh, K., Jolesz, F. A. 3D Slicer: a platform for subject-specific image analysis, visualization, and clinical support. Intraoperative Imaging Image-Guided Therapy. 3 (19), 277-289 (2014).
- Kapur, T., et al. Increasing the impact of medical image computing using community-based open-access hackathons: The NA-MIC and 3D Slicer experience. Medical Image Analysis. 33, 176-180 (2016).
- Fedorov, A., et al. 3D Slicer as an Image Computing Platform for the Quantitative Imaging Network. Magnetic Resonance Imaging. 30 (9), 1323-1341 (2012).
- . Chest Imaging Platform [software] Available from: https://chestimagingplatform.org (2020)
- Wang, J., Li, F., Li, Q. Automated segmentation of lungs with severe interstitial lung disease in CT. Medical Physics. 36 (10), 4592-4599 (2009).
- Lanza, E., et al. Quantitative chest CT analysis in COVID-19 to predict the need for oxygenation support and intubation. European Radiology. , (2020).
- Yao, G. Value of window technique in diagnosis of the ground glass opacities in patients with non-small cell pulmonary cancer. Oncology Letters. 12 (5), 3933-3935 (2016).
- Kauczor, H. U., et al. Automatic detection and quantification of ground-glass opacities on high-resolution CT using multiple neural networks: comparison with a density mask. American Journal of Roentgenology. 175 (5), 1329-1334 (2000).
- Funama, Y., et al. Detection of nodules showing ground-glass opacity in the lungs at low-dose multidetector computed tomography: phantom and clinical study. Journal of Computed Assisted Tomography. 33 (1), 49-53 (2009).
- Hansell, D. M., et al. Fleischner Society: glossary of terms for thoracic imaging. Radiology. 246, 697-722 (2008).
- Heye, T., et al. Reproducibility of Dynamic Part II. Comparison of Intra- and Interobserver Variability with Manual Region of Interest Placement versus semiautomatic lesion segmentation and histogram analysis. Radiology. 266, 812-821 (2013).
- Zhang, H. T., et al. Automated detection and quantification of COVID-19 pneumonia: CT imaging analysis by a deep learning-based software. Nuclear Medicine and Molecular Imaging. 14, 1-8 (2020).
- Alhazzani, W., et al. Surviving Sepsis Campaign: guidelines on the management of critically ill adults with Coronavirus Disease 2019(COVID-19). Intensive Care Medicine. 46 (5), 854-887 (2020).

