Automated Dissection Protocol for Tumor Enrichment in Low Tumor Content Tissues
Summary
Digital annotation with automated tissue dissection provides an innovative approach to enriching tumor in low tumor content cases and is adaptable to both paraffin and frozen tissue types. The described workflow improves accuracy, reproducibility and throughput and could be applied to both research and clinical settings.
Abstract
Tumor enrichment in low tumor content tissues, those below 20% tumor content depending on the method, is required to generate quality data reproducibly with many downstream assays such as next generation sequencing. Automated tissue dissection is a new methodology that automates and improves tumor enrichment in these common, low tumor content tissues by decreasing the user-dependent imprecision of traditional macro-dissection and time, cost, and expertise limitations of laser capture microdissection by using digital image annotation overlay onto unstained slides. Here, digital hematoxylin and eosin (H&E) annotations are used to target small tumor areas using a blade that is 250 µm2 in diameter in unstained formalin fixed paraffin embedded (FFPE) or fresh frozen sections up to 20 µm in thickness for automated tumor enrichment prior to nucleic acid extraction and whole exome sequencing (WES). Automated dissection can harvest annotated regions in low tumor content tissues from single or multiple sections for nucleic acid extraction. It also allows for capture of extensive pre- and post-harvest collection metrics while improving accuracy, reproducibility, and increasing throughput with utilization of fewer slides. The described protocol enables digital annotation with automated dissection on animal and/or human FFPE or fresh frozen tissues with low tumor content and could also be used for any region of interest enrichment to boost adequacy for downstream sequencing applications in clinical or research workflows.
Introduction
Next generation sequencing (NGS) is increasingly utilized for both patient care and in cancer research to help guide treatments and facilitate scientific discovery. Tissue is often limited and small specimens with variable tumor content are routinely used. Tumor adequacy and integrity, therefore, remain a barrier to obtaining meaningful data. Samples with lower tumor percentages may cause difficulty in distinguishing true variants from sequencing artifacts and are often ineligible for NGS1. Tumor enrichment of low tumor content cases, those below 20%, has been shown to help yield sufficient material in order to generate reproducible sequencing data and ensure low frequency variants are not missed2,3. However, limits will vary depending on the platforms utilized and planned use of the data generated.
Traditionally, enrichment of tumor regions for extraction is performed by manual macrodissection or laser capture microdissection (LCM) of formalin fixed paraffin embedded (FFPE) slides. Manual macrodissection, or scraping specified tissue areas from slides, allows tumor regions to be removed for use in downstream assays with relatively low cost, but with low accuracy and low precision2,4. Minimal technical accuracy can be very effective with higher tumor content cases where large swaths of tumor are present and/or minimal tissue loss does not significantly impact results, but low tumor content cases or cases with more dispersed tumor require increased precision. LCM was therefore invented in the 1990s and became a valuable way to precisely remove small, defined, microscopic regions of tissue from formalin fixed paraffin embedded (FFPE) slides5,6,7,8. LCM can be utilized to collect single cell populations when complex heterogeneity of the sample exists9 allowing for collection of previously difficult to separate cell populations. However, LCM requires costly machinery that requires extensive technical expertise and hands-on time10,11,12,13,14.
The instrument used for automated tissue dissection has precision in between that of LCM (~10 µm) and macrodissections (~1 mm)15. Additionally, it exhibits both cost and technical expertise requirements between that of macrodissection and LCM and is designed to perform rapid tissue enrichment from sequential FFPE slides to alleviate the disadvantages of previous methods15. Automated dissection in this fashion utilizes digital annotations or on-stage slide reference image overlays onto serially sectioned unstained tissue slides for dissecting and enriching regions of interest. The instrument uses plastic spinning blade milling tips, 1.5 mL collection tubes and can be used with a number of different fluids for dissection to collect regions of interest for downstream assays inclusive of nucleic extraction and sequencing. The spinning plastic milling tip utilizes inner and outer syringe barrel reservoirs and a plunger to collect buffer, then mills and collects tissue16. The variable milling tip size diameter (250 µm, 525 µm, 725 µm) can allow for dissection of separate tissue areas for comparison, multifocal regions that can be pooled or individual small areas from single or multiple FFPE slides. Section thicknesses used for harvest can be adjusted based on individual experiment needs and users can ensure regions of interest have not been depleted by performing an additional H&E on one serial section immediately after the last section used for harvest.
Automated dissection was identified as a way to enrich tumor content in low tumor content cases and we tested and expanded the intended functionality of an automated tissue dissection instrument, which is currently marketed for use on FFPE clinical specimens up to 10 µm in thickness. The work shows that automated dissection can be applied to both FFPE and fresh frozen human or animal tissue sections up to 20 µm in thickness for research purposes. The protocol also demonstrates an approach to digitally annotate and automate dissection for tumor enrichment in tissues with low tumor content and/or cases with nested, dispersed tumor where meaningful macrodissection is challenging or not feasible and show both quality and yield of nucleic acid sufficient for NGS. Automated dissection can therefore provide mid-level precision and increased throughput for tumor enrichment and could also be applied to enrich other regions of interest or combined with other platforms to answer research or clinical questions.
Protocol
Prior to initiation, obtain appropriate tissue specimens according to Institutional Review Board (IRB) protocols. All methods described here have been approved by the Institutional Animal Care and Use Committee (IACUC) of Genentech, Inc.
1. Tissue and slide preparation
- Select FFPE or fresh frozen tissue blocks and utilize the corresponding processing method below.
- Cut tissue block sections onto positively charged glass slides at the desired thickness. Serially section the FFPE tissue in ribbons with the first reference section cut at a thickness appropriate for H&E staining (i.e., 4 µm) followed by 1-4 sections at a thickness ranging from 4-20 µm based on the need and tissue availability. Collect the tissue sections onto positively charged glass microscope slides.
NOTE: Fresh frozen reference tissue sections should be stained immediately with Hematoxylin and Eosin (H&E) using routine protocols for frozen sections and the unstained frozen sections held at -20 °C until they are needed for harvest. - Allow all the FFPE sections to dry at room temperature overnight.
- Bake the FFPE reference slides at 60 °C for 30 min and then stain with H&E using routine protocols.
- Scan the H&E stained slides on a whole slide imager at 20x magnification or greater.
- Annotate the scanned slide images for tumor regions of interest using a vendor provided viewing platform or open-source viewer. Export these annotations as either a low-magnification screenshot or save them as a metadata file containing X-Y pixel coordinates corresponding to polygon vertices.
NOTE: The former is less technically challenging to work with, but the latter offers advantages in process automation. - Create digital masks of the annotated regions of interest in line with the approach used and export the manual annotations.
NOTE: If a screenshot/image of the annotations is used, simple image processing software can be used to select a region and to fill in the entire selection. Using X-Y coordinates for each ROI requires the use of a programming language to read both the image data and polygon coordinates to create a low-mag image with filled in regions of interest. The user should work with the automated dissection instrument vendor to establish a process based on their individual software availability and needs. If scanning, digital slide annotation and/or digital mask creation is unavailable, careful on-slide annotation using a marker can be performed and used in place of a digital mask as a reference image. Pseudocode for digital mask creation has been provided in Supplementary File 1.
2. Automated tissue dissection
- Place the unstained sample tissue slides onto the stage in the first through fourth slide positions when using digital slide reference. When using on-slide annotation rather than a digital option, place the unstained sample tissue slides onto the stage in the second through fourth slide positions with a reference slide in the first position.
- Create a milling job using the automated tissue dissection software: Job Selection > Create New Job > Case ID > Name the milling job; go to Thickness > Section Thickness using the up or down arrow tab; then go to Tissue Preparation > Paraffinized or Deparaffinized, Reference Image > From File > Import Image > File to import from the dropdown as the digital reference, if applicable. Select From Stage for on stage slide reference. When fields are complete, scan the stage by selecting the Scan Stage button in the bottom-right corner to capture each sample tissue slide in the first through fourth position.
- Select the tissue area for image capture.
- If using an on-stage reference, drag the box from one corner to the opposite to create a rectangular area over tissue. Select the circular bubble under the rectangular area to capture the stage reference image. If using a digital reference image, overlay the image on the rectangular area selected. Resize and align the digital reference grossly in course zoom to best match the size and position over the sample tissue.
- Copy this rectangle field onto remaining sample tissue slides in the second through fourth slide positions by selecting the copy option in the upper-right corner of the reference image. Align and resize grossly as necessary.
NOTE: When using an on-slide reference image rather than a digital mask, select which slide on the stage should be used as the reference.
- Align the reference and sample slides
- When the reference image is grossly aligned on tissue sample slides in all slide positions, select the Scan Stage button in the lower-right corner of the screen to move into the fine adjustment step. Select the first stage position and Transform tool icon (the third icon down in the right-hand toolbar) to make fine alignment and zoom adjustments of the reference to best match the sample slide overlay. Use the Reference to Sample sliding bar at the bottom of the screen toggling between reference image and sample image along with the Zoom In and Zoom Out feature to adjust and achieve alignment of each slide position. Replicate this process in the second through fourth sample slide positions.
- Select the milling area of region of interest
- Once optimal sample overlay of each of the four slide positions is achieved, draw milling path designations using the Color Picker tool icon (the tenth icon down in the right-hand toolbar) on the colored portion of masked reference image. Select the Extend to Similar box if multiple slides or areas are annotated for dissection and then select the Get Annotation(s) button in lower right to draw milling paths onto sample slides.
- Select the milling path in the first slide position.
NOTE: When the milling path is selected in the first slide position, it will be copied onto the remaining slide positions and milling tip usage will be calculated. The milling tip usage in the upper-left corner is calculated based on the area covered and the tip size selected. If more than four tips are calculated, a larger tip size can be selected to capture the annotated ROI. Tip size can be selected or changed on the left side of the screen under the Milling Tip arrow and tip usage will be recalculated. - When the milling path is calculated, collect the annotated ROI with four or less milling tips. Select the Setup Stage button in the lower right of the screen to prompt loading of milling tips from placed collection tubes in their proper designation on stage.
- Fill the reservoir with 3.0 mL of the dissection buffer most appropriate for the tissue type (FFPE or fresh frozen) and downstream nucleic acid extraction kit needs and select the Dissect button in the lower-right corner of the screen. Use molecular grade mineral oil or an appropriate buffer from commercially available nucleic acid extraction kits.
NOTE: Automated dissection of slides and selected regions of interest then begins and samples are collected by the instrument. The unit head will pick up milling tips from the back of the stage and fill with dissection fluid from the reservoir. Tips then spin along the milling path aspirating sample tissue from slides until complete or full. The collected sample with dissection fluid is then dispensed into collection tubes located at the back of the stage. - When automated dissection is complete, remove the collection tubes and the dissected sample slides from the stage and place them in a tube rack and slide rack, respectively.
NOTE: Fresh frozen harvests should be taken directly into nucleic acid extraction as per the manufacturer's instructions and post-dissection fresh frozen sections should be H&E stained immediately using routine protocols for frozen sections. - Bake the post dissected tissue slides at 60 °C for 30 min and then stain with H&E using routine protocols.
- Scan the post dissected H&E stained slides on a whole slide imager at 20x magnification and/or archive for a reference of what tissue was not collected and remains on the slide.
NOTE: See step 1.5 above for alternative scanning options.
3. Nucleic acid extraction
- Pool and pellet the tissue. Perform the nucleic acid extraction using a commercially available kit and following the manufacturer's instructions.
Representative Results
FFPE and FF mouse liver sections containing metastatic colorectal cancer in xenografts were selected. Sections were H&E stained (Figure 1A,E,I) and scanned on a whole slide imager at 20x magnification. A pathologist digitally annotated tumor regions of interest and a mask was generated using commercial software and formatted as a digital png reference image (Figure 1B,F,J). Serial 10 µm and 20 µm thick unstained sample slides were placed on the stage and automated dissection was performed as described above. Fresh frozen tissues were collected in a lysis buffer from a commercially available kit and carried directly into nucleic acid extraction following the manufacturer's instructions. FFPE samples were collected using molecular grade mineral oil and dissected samples were pooled together and centrifuged at 25,000 x g for 20 min at 4 °C. The supernatant was removed and the minimal mineral oil required was used to resuspend, transfer, and collect the dissected tissue into a single collection tube for each sample appropriately. Samples were shipped at room temperature to a vendor for nucleic acid extraction, RNA and DNA sizing, quantity, integrity, and purity determination following the manufacturer's instructions. Sequencing libraries were created and used for hybridization and capture with commercial options following the manufacturer's instructions. Post dissected sample slides were H&E stained using routine staining protocols to confirm dissection areas in 10 µm (Figure 1C,G,K) and 20 µm (Figure 1D,H,L) slides and dissection metrics were captured (Supplementary Table 1). Exome sequencing generated approximately 75 million 100 bp paired-end reads, yielding an average depth of coverage (before removing duplicate reads) of 150x per sample, with 99.9% reads aligned and a 78% on-target rate. RNA sequencing metrics demonstrated just over 55 million bp paired-end reads, a 98% alignment rate, and a 19.4% duplication rate with 77% concordant reads.
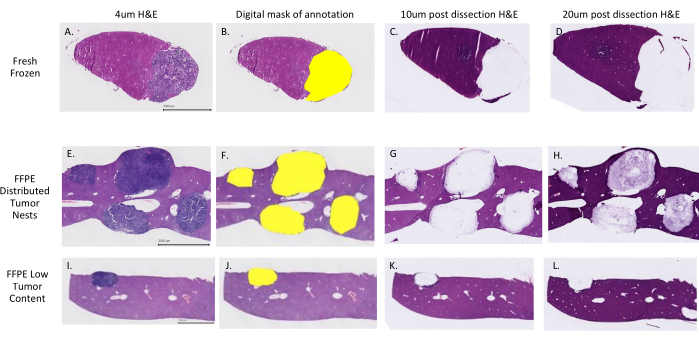
Figure 1; Successful dissection of tumor nests from fresh frozen and FFPE tissue. H&E stained mouse fresh frozen (A–D) and FFPE (E–L) liver tissue with colorectal cancer metastasis. 4 µm reference slides used for digital annotation (A, E, I) demonstrate examples with a low tumor percent for the total tissue area (I) and distributed tumor nests (E) that classically present challenges to tumor enrichment. Annotated and digitally masked H&E reference slides (B, F, J) were generated and post-dissection 10 µm (C, G, K) and 20 µm (D, H,L) H&E stained slides demonstrate successful harvest of selected areas. Please click here to view a larger version of this figure.
Supplementary File 1: Pseudocode utilized to create digital masks from annotations to use in automated dissection. Please click here to download this File.
Supplementary Table 1: Example captured metrics from automated dissection and nucleic acid extraction. Please click here to download this Table.
Discussion
Presented here is a protocol for the application of digital annotation and automated dissection to dissect tumor regions from low tumor content FFPE or fresh frozen tissues for tumor enrichment and use in WES. Combining digital annotation and mask creation with automated dissection significantly reduces the required hands-on time and expertise common to classical methods of tumor enrichment inclusive of manual macrodissection and LCM. The protocol demonstrates a potentially important mid-range tumor enrichment option that allows for not only low tumor content enrichment but also enrichment in cases where it is challenging to dissect distributed tumor nests away from the tumor adjacent normal tissue for meaningful tumor enrichment with high throughput and a moderate level of precision. While the use of our workflow for low tumor content xenograft tissues is demonstrated here, it was also found that this protocol works across tissue types, including human, murine, and xenograft tissues for a variety of normal tissues and cancer indications.
Therefore, it could also apply to a broad range of applications where enriching for specific regions of interest without significant contamination of background tissue would be beneficial (i.e., to enrich for a specific brain region) or even for removal of areas of tissue prior to nucleic acid extraction using classical macrodissection.
Many platforms exist on the market for slide scanning and digital annotation. It is therefore important to remain aware that platform compatibility may present limitations and specified platforms within any protocol may not be broadly available in all laboratories. Therefore, significant efforts were made to provide alternative options within the described protocol that will guide users in making any necessary modifications based on their available resources. An option for removal of the digital annotation component has also been noted to allow for careful manual on-slide annotation. The options provided for modifications will maximize the ability of users to find an option that works with their current platform and software availability.
While digital annotation and automated dissection have been demonstrated to be broadly applied to both FFPE and fresh frozen tissue, it is important to note that the boundaries of the automated tissue dissection instrument have been pushed beyond its intended use with FFPE specimens and the protocol is meant for research use only. Here, successful tumor enrichment was demonstrated through automated tumor dissection of low tumor content FFPE as well as fresh frozen tissues for nucleic acid extraction, WES and RNA sequencing. The protocol shows that xenograft and human tissue regions of interest could be enriched prior to WES and RNA sequencing in basic and translational research settings and also note that other downstream molecular applications, including PCR, from both tissue types would be possible. The protocol expands FFPE automated dissection options and lays the groundwork for fresh frozen tissue automated dissection that could be developed and validated further for use in clinical settings.
Divulgations
The authors have nothing to disclose.
Acknowledgements
The authors would like to thank Carmina Espiritu and Robin E. Taylor for their support in automated dissection development as well as the Genentech Pathology Core Laboratory staff that supported this work.
Materials
| Agilent SureSelectXT | Agilent | G9611A | |
| AVENIO Millisect Fill Station | Roche | 8106533001 | |
| AVENIO Millisect Instrument, Base | Roche | 8106568001 | |
| AVENIO Millisect Instrument, Head | Roche | 8106550001 | |
| AVENIO Millisect Milling Tips Small | Roche | 8106509001 | |
| AVENIO Millisect PC | Roche | 8106495001 | |
| BioAnalyzer | Agilent | G2939BA | |
| Eppendorf 5427R | Eppendorf | 22620700 | Micro-centrifuge |
| Incubation Buffer | Promega | D920D | |
| Leica Autostainer XL | Leica | ST5010 | Automated stainer |
| Molecular Grade Mineral Oil | Sigma | M5904-500ML | |
| Proteinase K | Promega | V302B | Digestion buffer |
| Qiagen AllPrep DNA/RNA Mini Kit | Qiagen | 80284 | |
| RLT Plus buffer | Qiagen | 80204 | |
| Superfrost Plus positively charged microscope slides | Thermo Scientific | 6776214 |
References
- Cho, M., et al. Tissue recommendations for precision cancer therapy using next generation sequencing: a comprehensive single cancer center’s experiences. Oncotarget. 8 (26), 42478-42486 (2017).
- Smits, A. J. J., et al. The estimation of tumor cell percentage for molecular testing by pathologists is not accurate. Modern Pathology: An Official Journal of the United States and Canadian Academy of Pathology, Inc. 27 (2), 168-174 (2014).
- Poole-Wilson, P. A., Langer, G. A. Effect of pH on ionic exchange and function in rat and rabbit myocardium. The American Journal of Physiology. 229 (3), 570-581 (1975).
- Viray, H., et al. A prospective, multi-institutional diagnostic trial to determine pathologist accuracy in estimation of percentage of malignant cells. Archives of Pathology & Laboratory Medicine. 137 (11), 1545-1549 (2013).
- El-Serag, H. B., et al. Gene Expression in Barrett’s Esophagus: Laser capture versus whole tissue. Scandinavian Journal of Gastroenterology. 44 (7), 787-795 (2009).
- Harrell, J. C., Dye, W. W., Harvell, D. M. E., Sartorius, C. A., Horwitz, K. B. Contaminating cells alter gene signatures in whole organ versus laser capture microdissected tumors: a comparison of experimental breast cancers and their lymph node metastases. Clinical & Experimental Metastasis. 25 (1), 81-88 (2008).
- Kim, H. K., et al. Distinctions in gastric cancer gene expression signatures derived from laser capture microdissection versus histologic macrodissection. BMC Medical Genomics. 4, 48 (2011).
- Klee, E. W., et al. Impact of sample acquisition and linear amplification on gene expression profiling of lung adenocarcinoma: laser capture micro-dissection cell-sampling versus bulk tissue-sampling. BMC Medical Genomics. 2, 13 (2009).
- Civita, P., et al. Laser capture microdissection and RNA-seq analysis: High sensitivity approaches to explain histopathological heterogeneity in human glioblastoma FFPE archived tissues. Frontiers in Oncology. 9, 482 (2019).
- Emmert-Buck, M. R., et al. Laser capture microdissection. Science. 274 (5289), 998-1001 (1996).
- Bonner, R. F., et al. Laser capture microdissection: molecular analysis of tissue. Science. 278 (5342), 1481-1483 (1997).
- Hunt, J. L., Finkelstein, S. D. Microdissection techniques for molecular testing in surgical pathology. Archives of Pathology & Laboratory Medicine. 128 (12), 1372-1378 (2004).
- Espina, V., et al. Laser-capture microdissection. Nature Protocols. 1, 586-603 (2006).
- Grafen, M., et al. Optimized expression-based microdissection of formalin-fixed lung cancer tissue. Laboratory Investigation; A Journal of Technical Methods and Pathology. 97 (7), 863-872 (2017).
- Javey, M., et al. innovative tumor tissue dissection tool for molecular oncology diagnostics. The Journal of Molecular Diagnnostics: JMD. (21), 1525-1578 (2021).
- Adey, N., et al. A mill based instrument and software system for dissecting slide-mounted tissue that provides digital guidance and documentation. BMC Clinical Pathology. 13 (1), 29 (2013).

