Isolation of Endothelial Cells from the Lumen of Mouse Carotid Arteries for Single-Cell Multi-Omics Experiments
Summary
We present a method for isolating endothelial cells and nuclei from the lumen of mouse carotid arteries exposed to stable or disturbed flow conditions to perform single-cell omics experiments.
Abstract
Atherosclerosis is an inflammatory disease of the arterial regions exposed to disturbed blood flow (d-flow). D-flow regulates the expression of genes in the endothelium at the transcriptomic and epigenomic levels, resulting in proatherogenic responses. Recently, single-cell RNA sequencing (scRNAseq) and single-cell Assay for Transposase Accessible Chromatin sequencing (scATACseq) studies were performed to determine the transcriptomic and chromatin accessibility changes at a single-cell resolution using the mouse partial carotid ligation (PCL) model. As endothelial cells (ECs) represent a minor fraction of the total cell populations in the artery wall, a luminal digestion method was used to obtain EC-enriched single-cell preparations. For this study, mice were subjected to PCL surgery to induce d-flow in the left carotid artery (LCA) while using the right carotid artery (RCA) as a control. The carotid arteries were dissected out two days or two weeks post PCL surgery. The lumen of each carotid was subjected to collagenase digestion, and endothelial-enriched single cells or single nuclei were obtained. These single-cell and single-nuclei preparations were subsequently barcoded using a 10x Genomics microfluidic setup. The barcoded single-cells and single-nuclei were then utilized for RNA preparation, library generation, and sequencing on a high-throughput DNA sequencer. Post bioinformatics processing, the scRNAseq and scATACseq datasets identified various cell types from the luminal digestion, primarily consisting of ECs. Smooth muscle cells, fibroblasts, and immune cells were also present. This EC-enrichment method aided in understanding the effect of blood flow on the endothelium, which could have been difficult with the total artery digestion method. The EC-enriched single-cell preparation method can be used to perform single-cell omics studies in EC-knockouts and transgenic mice where the effect of blood flow on these genes has not been studied. Importantly, this technique can be adapted to isolate EC-enriched single cells from human artery explants to perform similar mechanistic studies.
Introduction
This laboratory previously demonstrated that induction of d-flow leads to quick and rugged atherosclerosis development in hyperlipidemic mice1,2. The novel mouse model of d-flow-induced atherosclerosis was possible using partial carotid ligation (PCL) surgery 3. PCL surgery induces low and oscillatory blood flow condition or d-flow in the ligated left carotid artery (LCA). In contrast, the contralateral right carotid artery (RCA) continues to face stable laminar flow (s-flow). Previously, to understand the effect of d-flow on endothelial cells, the carotid arteries were dissected out after partial ligation surgery and flushed with a phenol and guanidine isothiocyanate-based lysing agent (luminal RNA/DNA flushing method)2,4, which provided endothelial-enriched "pooled bulk" RNAs or DNAs. These pooled bulk RNAs or DNAs were then processed for transcriptomic studies or epigenomic DNA methylome studies, respectively4,5,6. These studies helped discover multiple flow-sensitive genes and microRNAs whose roles in endothelial biology and atherosclerosis were extensively investigated4,6,7.
However, despite endothelial enrichment, these bulk RNA/DNA studies could not distinguish the specific role of each cell type in the artery wall in d-flow-induced atherosclerosis. Endothelial-enriched single-cell (sc) isolation and scRNA and scATAC sequencing studies were performed to overcome this limitation8. For this, C57Bl6 mice were subjected to the PCL surgery to induce d-flow in the LCA while using the s-flow-exposed RCA as control. Two days or two weeks after the PCL surgery, the mice were sacrificed, and the carotids were dissected and cleaned up. The lumen of both LCAs and RCAs were infused with collagenase, and the luminal collagenase digests containing ECs as a significant fraction and other arterial cells were collected. The single-cell suspension (scRNAseq) or single-nuclei suspension (scATACseq) were prepared and barcoded with unique identifiers for each cell or nucleus using a 10x Genomics setup. The RNAs were subjected to cDNA library preparation and sequenced.
The scRNAseq and scATACseq datasets were processed using the Cell Ranger Single-Cell Software and further analyzed by Seurat and Signac R packages9,10. Each cell and nucleus was assigned a cell type from these analyses and clustered into the cell type based on the marker genes and unique gene expression patterns. The results of the scRNAseq and scATACseq demonstrated that these single-cell preparations are enriched with ECs and also contain smooth muscle cells (SMCs), fibroblasts, and immune cells.
Further analysis revealed that the EC population in the luminal digestion is highly divergent and plastic (8 different EC clusters) and responsive to blood flow. Most importantly, these results demonstrated that d-flow reprograms ECs from an athero-protected anti-inflammatory phenotype to pro-atherogenic phenotypes, including pro-inflammatory, endothelial-to-mesenchymal transition, endothelial stem/progenitor cell transition, and most surprisingly, endothelial-to-immune cell-like transition. In addition, scATACseq data reveal novel flow-dependent chromatin accessibility changes and transcription factor binding sites in a genome-wide manner, which form the basis of several new hypotheses. The methodology and protocol for preparing single endothelial cells for single-cell multi-omics studies from the mouse carotid arteries are detailed below.
Protocol
All animal procedures described below were approved by the Institutional Animal Care and Use Committee at Emory University. Non-hypercholesterolemic, age- and sex-matched C57BL/6 mice were used to mitigate sex-dependent variation and offset any complication of hypercholesterolemic conditions.
1. Partial carotid ligation (PCL) surgery
NOTE: Partial carotid artery ligation of LCA was carried out as previously described and demonstrated3.
- Anesthetize the mouse by isoflurane inhalation (5% isofluorane in oxygen for induction and 1.5% after that for maintenance) throughout the procedure using an anesthetic vaporizer. Place the mouse supine on a Deltaphase isothermal pad to prevent loss of body heat during surgery.
- Apply an ocular ointment to prevent exposure keratitis, and confirm the depth of anesthesia by the absence of toe pinch response.
- Inject buprenorphine 0.05 mg/kg subcutaneously to manage pain preemptively. Disinfect the epilated area by applying and cleaning with betadine and isopropanol solution three times. Ensure that betadine is the last step to sterilize the surgery area, starting from the center and finishing at the sides.
- Using aseptic techniques, make a ventral midline incision (~1 cm) in the neck region, and expose the LCA branch point by blunt dissection.
- Ligate the left internal carotid, external carotid, and occipital arteries with 6-0 silk suture; leave the superior thyroid artery untouched.
- Approximate the skin and close the incision with tissue glue and/or sutures.
- Transfer the mouse to a pre-warmed recovery cage (maintained at 37 °C) and place it on a clean towel for up to 1 h to avoid post-surgery hypothermia.
NOTE: In our experience, a pre-emptive single dose of Buprenorphine (0.05 mg/kg) is usually sufficient for post-surgery pain management. A repeat dose of Buprenorphine can be administered if the animal is in distress after the first 24 hours. If performed correctly, there is no mortality associated with this procedure, and postoperative stress to the mouse is minimal. The mice are returned to their respective cages and monitored daily following the surgery. - Prepare the perfusion setup.
- Use 0.9% NaCl (normal saline) containing 10 U/mL heparin in an IV bag. Hook the IV bag 244-274 cm (8-9 feet) from the ground or approximately 122-152 cm (4-5 feet) higher than the dissection board. Attach the butterfly needle to the end of the saline line and flush any air bubbles out of the IV line.
2. Isolation of carotid arteries post sacrifice
- Sacrifice the mice by CO2 inhalation following the institutional IACUC protocol.
- Spray 70% ethanol on the mouse skin and place it in the supine position on the dissection board containing adsorbent paper towels. Secure the paws of the mouse to the adsorbent towels on the dissection board using adhesive tape or a 21 G needle.
- Using a pair of sterile scissors, remove the skin of the mouse starting from the abdomen and all the way to the top of the thorax.
- Use a pair of scissors to open the abdominal wall below the ribcage by blunt dissection.
- Using the scissors, carefully make an incision along the length of the diaphragm, and continue through the ribs on both sides of the thorax until the sternum can be lifted away. Gently lift away the sternum with a pair of forceps; after that, remove the ribcage to expose the heart.
- Carefully remove the thymus and any connective tissue over the heart to visualize the major vessels.
- Sever the vena cava with scissors to allow blood to exit from closed circulation.
- Insert a 21 G butterfly needle connected to the IV line through the apex of the heart into the left ventricle and allow retrograde perfusion for 2-3 min with normal saline at room temperature. Ensure that a constant flow rate of normal saline is maintained by keeping the saline bag at the height of 8-9 feet from the ground.
NOTE: The lungs and liver become pale, indicating optimal perfusion. Approximately 20 mL of perfusion buffer is used for each mouse. - Remove the skin from the neck region and remove all the fat, muscles, and connective tissues to expose the carotid arteries (Figure 1A).
NOTE: The rest of the procedure is carried out under the dissecting microscope. - Adjust the field of view to locate the ligation sites. Using the 10 mm micro-dissection scissors, make a small incision below the ligation site in the LCA to allow perfusion.
- Perfuse again for an additional 1 min via the left ventricle and ensure that the LCA is well perfused and free from any visible traces of blood.
- Use fine-tip forceps and small spring scissors to carefully remove the peri-adventitial tissues surrounding the carotids while the carotid is attached to the body.
NOTE: Be extremely careful not to squeeze or stretch the carotid arteries during this cleaning step, as this can increase the number of non-viable cells. If performed correctly, the cell viability in the final cell suspension is usually >93%.
3. Endothelial-cell-enriched single-cell isolation from mice carotids
NOTE: The reagents described below can be prepared in advance and stored at 4 °C until use: 1x and 0.1x single-nuclei lysis buffers with the reagents listed in Table 1; single-nuclei wash buffer with the reagents listed in Table 1; single-nuclei Nuclei Buffer recipe with the reagents listed in Table 1. The working stocks of these buffers are to be prepared following the manufacturer's protocol. Digestion buffer composition: Collagenase Type II 600 U/mL and DNase I 60 U/mL in 0.5% fetal bovine serum (FBS)-containing phosphate-buffered saline (PBS).
- While the carotids are still attached, wash the exterior of the carotid arteries with normal saline solution to flush away any traces of blood.
- Using an insulin syringe fitted with a 29 G needle, inject ~ 50 µL of the digestion buffer into the lumen of the distal end of the left carotid artery. As the digestion buffer starts to fill the carotid artery, clamp the proximal end of the carotid with a micro-clip (Figure 1B). Introduce an additional 15-20 µL of digestion buffer into the lumen and clip the distal end to avoid the release of the digestion buffer.
- Carefully explant the carotid arteries from the mouse (Figure 1B,C) and place them in 35 mm dishes containing warm (at 37 °C) Hepes-buffered saline solution.
NOTE: Make sure that both the clamps are securely placed. If the clamp becomes loose, the digestion solution can leak out of the lumen and affect the obtained cell count. If this happens, add additional enzymatic solution using an insulin syringe fitted with a 29 G needle from the open end of the carotid and secure the clamp again (Figure 1D). If the enzyme buffer solution leaks again, it is likely that the carotid is severed during the explantation process. In this case, discard the carotid and exclude the sample from the study. Repeated rough handling of the carotid will decrease the cell viability and the quality of single-cell preparation. Take care to identify and correctly label the left and right carotid arteries. - Incubate the explanted carotids for 45 min at 37 °C with intermittent rocking.
4. Flushing the carotid arteries
- After completing the luminal enzymatic digestion, remove the carotid artery together with the clamps from the 35 mm dish. Carefully remove each clamp, taking care that the digestion buffer should not leak.
- Gently hold one end of the carotid artery with fine forceps on top of a 1.5 mL microcentrifuge tube. Insert a 29 G needle fitted with an insulin syringe containing 100 µL of warm digestion buffer (37 °C) into the lumen with the other hand (Figure 1D).
- Quickly flush the lumen of the carotid into the microcentrifuge tube. Block the enzymatic reaction by adding 0.3 mL of FBS into the 1.5 mL tube. Place the tube on ice.
NOTE: The flushing contains the cells from the luminal enzymatic digestion. Adding FBS and reducing the temperature stops the enzymatic digestion process. If the number of cells in the preparation is high and contains debris or fat tissue, the use of a 50 or 70 µm cell strainer is highly recommended to filter out unwanted tissues debris. To increase the single-cell count, flushing multiple carotids is recommended. Here, to obtain at least 5,000 cells, 10-12 carotids were flushed and pooled as one sample. - Centrifuge the cells at 500 × g for 5 min at 4 °C using a centrifuge equipped with a fixed-angle rotor (see the Table of Materials).
- Discard the supernatant and resuspend the cell pellet in digestion buffer containing cell dissociation reagent (see the Table of Materials) for 5 min at 37 °C to separate all the cells into single cells.
NOTE: If the number of cells in the preparation is low, increase the centrifuge speed up to 2,000-3,000 × g to spin down the cells. Moreover, using 0.5 mL centrifuge tubes facilitates better visualization of the cell pellet at the bottom of the tube. - Block the enzymatic reaction by adding 0.15 mL of FBS into the 0.5 mL tube.
- Centrifuge the single-cell suspension at 500 × g for 5 min at 4 °C using a centrifuge equipped with a fixed-angle rotor, as in step 4.4.
- Discard the supernatant and resuspend the cells in 100 µL of ice-cold 1% bovine serum albumin (BSA) solution in PBS in a 0.2 mL microcentrifuge tube.
NOTE: Using 0.2 mL centrifuge tubes enhances the visualization of the single-cell-pellet at the bottom of the tube.
5. Single-cell and single-nucleus analyses
- Resuspend and submit the single-cell preparation for scRNAseq.
- Resuspend the single-cell pellet with 100 µL of ice-cold 1% BSA in PBS in a 0.2 mL tube.
- After resuspending the cells for single-cell encapsulation, immediately proceed for single-cell encapsulation and barcoding using a microfluidics-based single-cell partitioning and barcoding system (see the Table of Materials).
- Before submitting the samples to the Genomics Core, count and inspect the single-cell preparations; ensure the absence of cell aggregates.
NOTE: The aggregation of single cells can be further minimized by increasing the BSA amount up to 2%.
- Resuspend and submit the single-cell preparation for scATACseq.
- Resuspend the cell pellet in 100 µL of 0.04% BSA in ice-cold 1x PBS and centrifuge the single-cell suspension at 500 × g at 4 °C.
- Lyse the single-cell suspension with ice-cold 0.1x lysis buffer (Table 1) and incubate for 5 min on ice.
- Mix the lysate 10 times with a P-20 pipette and incubate for an additional 10 min.
- Add 500 µL of chilled wash buffer (Table 1) to the lysed cells and mix 5 times using a pipette.
NOTE: Use a 70 µm cell strainer to filter any debris from the cell suspension and transfer them into a 2 mL tube. - Centrifuge the lysate at 500 × g for 5 min at 4 °C. Discard the supernatant and resuspend the nuclei-pellet in the diluted nuclei buffer (150 µL) (see the Table of Materials and Table 1). Count and inspect the single-nuclei preparations using a hemocytometer11.
- Submit the single-nuclei sample to the Genomics Core for single-nuclei transposition, nuclei partitioning, library preparation, and sequencing.
NOTE: If the initial cell number is low, it is common to observe debris along the nuclei. Therefore, it is recommended to start with a high cell number.
Representative Results
Partial carotid ligation surgeries were performed on 44 mice, and the onset of d-flow in the LCA was validated by performing ultrasonography one day post partial ligation surgery. Successful partial ligation surgery causes reduced blood flow velocity and reverses blood flow (disturbed flow) in the LCA3. The carotid arteries were dissected out either at two days or at two weeks post ligation. The lumen of each carotid was subjected to collagenase digestion, and endothelial-enriched single-cells or single-nuclei suspensions were prepared. Single-cell suspensions were pooled from 10 RCAs and LCAs to increase the cell yield. The cells/nuclei prepared were subsequently barcoded and sequenced. For the sc-RNAseq study, the number of single cells obtained was ~9,700. The distribution of cells from 4 samples is shown in Table 2.
Likewise, for the scATACseq study, single cells were isolated from 12 RCAs and LCAs that were prepared, subjected to transposase treatment, barcoded, and sequenced. Sequencing was performed for 18,324 single nuclei, which were pooled from 1,291 (2-day RCA [2D-R]), 5,351 (2-day LCA [2D-L]), 5,826 (2-week RCA [2W-R]), and 5,856 (2-week LCA [2W-L]) (Table 2). Representative single-cell and single-nuclei preparations as visualized by brightfield microscopy and phase-contrast microscopy are shown in Figure 2A,B. The efficiency of single-nuclei preparation from single-cell suspension is shown in Figure 2C.
The nuclei (~7,000 each) from 2D (RCAs and LCAs) and 2W (RCAs and LCAs) samples were incubated with a Transposition Mix (Tn5 transposase enzyme and buffer) see the Table of Materials) for 60 min at 37 °C following the manufacturer's protocol. Based on the manufacturer's recommendation, a mild detergent condition helped keep the nuclei intact during tagmentation. A master mix, consisting of a Barcoding Reagent, Reducing Agent B, and Barcoding Enzyme, was then loaded onto a microfluidic cell/nuclei encapsulation platform to prepare single-nuclei gel emulsions with barcoding according to the manufacturer's instructions.
Post sequencing, the Cell Ranger Single-Cell Software suite was used for demultiplexing, barcode processing alignment, and initial clustering of the raw scATACseq and scRNaseq profiles. For the scRNAseq study, the distribution of genes per cell, unique molecular identifier (UMI) per cell, mitochondrial reads per cell, and sequencing saturation information are shown in Figure 3A-C. Likewise, for the scATACseq study, quality control metrics showing the insert size distribution (nucleosome banding pattern) and normalized TSS enrichment score are shown in Figure 3D–F. Additionally, the percent fragment reads in the peaks, peak region fragments, TSS enrichment score, ratio of reads in blacklist genomic sites, and nucleosome signal ratio are shown in Figure 3G–K.
Endothelial enrichment was quantitated by comparing this method to that of enzymatic digestion of the whole carotid artery12. The endothelial cell count from the complete carotid artery digestion was 3-5% of the total cells obtained, whereas this method allowed enrichment of endothelial cells to >50% 8. Similarly, another single-cell study that used the whole mouse aorta showed that endothelial fraction was <7% of the total cell count. For an in-depth single-cell RNAseq and single-cell ATACseq analysis, readers are requested to refer to 8.
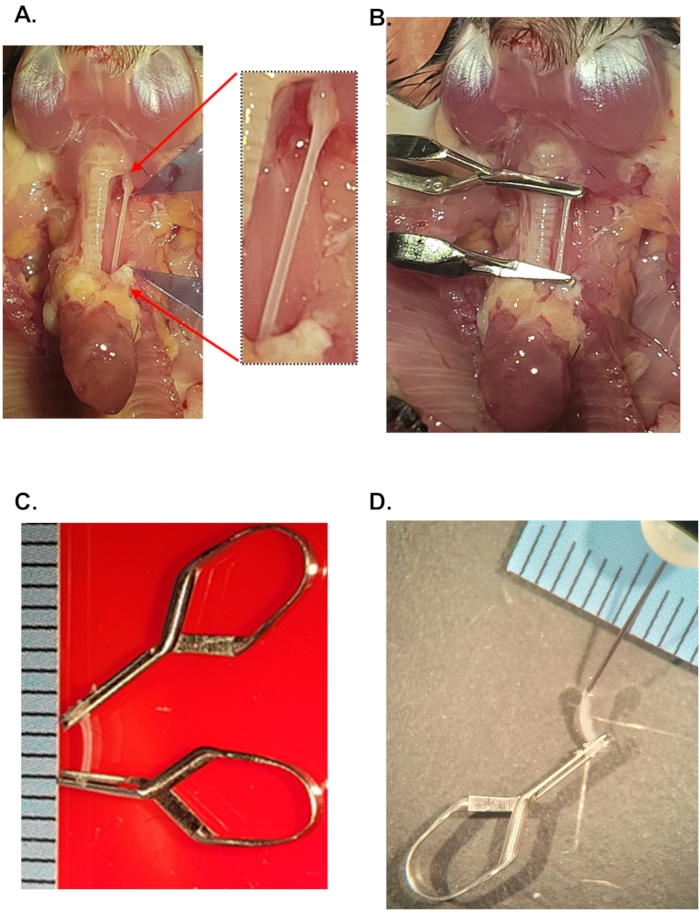
Figure 1: Isolation of carotid arteries for single-cell preparation. (A) Anatomical view of the carotid arteries in mice. Red arrows and inset show the isolated left carotid artery after clean up of periadventitial fat. For a schematic of the carotid anatomy and ligations, refer to Figure 1 in Nam et al3 (B) Image shows the location of micro-clips after filling the carotid artery with the digestion buffer. (C) Explanted carotid artery containing digestion buffer. Scale: distance between black lines = 1 mm. (D) A 29 G needle in the lumen of the mouse carotid artery. This step helps replenish the carotid artery with digestion buffer if needed. Please click here to view a larger version of this figure.
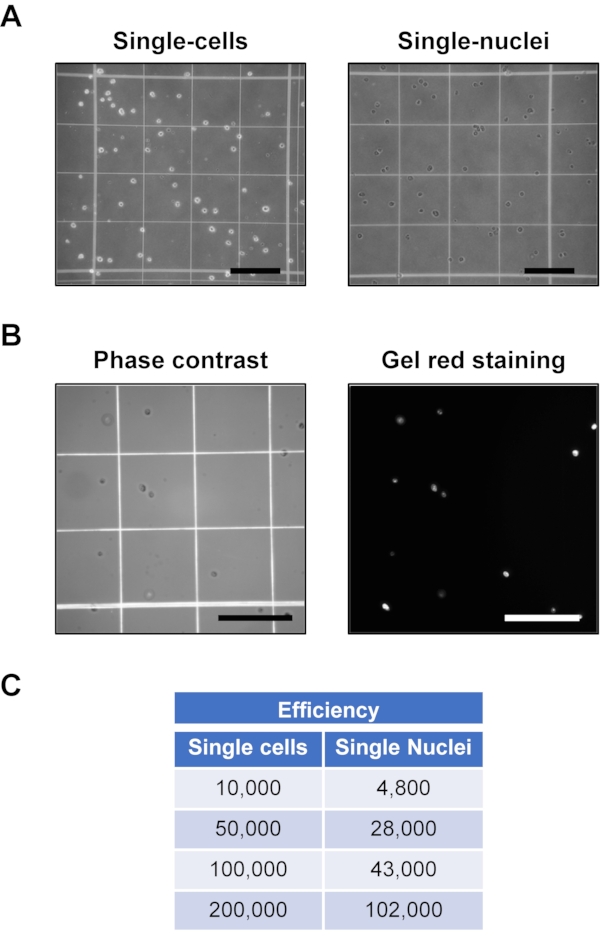
Figure 2: Single-cell and single-nuclei preparation from luminal enzymatic digestion of mouse carotid artery. (A) Representative single-cell and single-nuclei preparations. Scale bars = 0.25 mm (B) Representative phase-contrast and Gel-Red images of single-nuclei preps. Scale bars = 0.25 mm. (C) Efficiency of single-nuclei preparation in the left column shows the number of single cells at the start while the right column shows the number of single nuclei after processing with nuclei isolation buffer in different steps. Please click here to view a larger version of this figure.
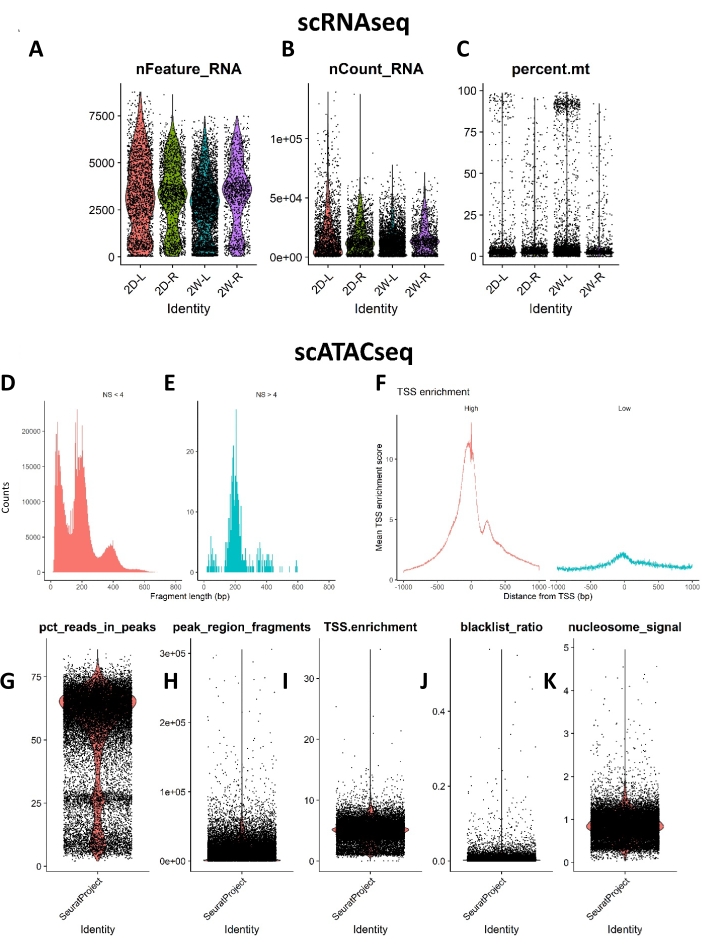
Figure 3: Standard QC metrics for the scRNAseq and scATACseq study. Violin plots show (A) the distribution of genes per cell (nFeature RNA), (B) UMI per cell (nCount_RNA), (C) mitochondrial reads per cell (percent mt) for the scRNAseq data. (D and E) show the nucleosome banding pattern for the scATACseq study. The histogram of DNA fragment sizes exhibits a strong nucleosome banding pattern corresponding to the length of DNA wrapped around a single nucleosome. (F) Normalized TSS enrichment score at each position relative to the TSS. The scatter/violin plots show (G) percent fragment reads in the peaks, a measure of sequencing depth, (H) peak region fragments showing the number of fragments overlapping peaks, (I) TSS enrichment score, a ratio between the aggregate distribution of reads centered on TSSs and that flanking the corresponding TSS, (J) blacklist ratio, a ratio of reads in blacklist genomic sites, and (K) nucleosome signal, a ratio of mononucleosomal to nucleosome-free fragments. Abbreviations: scRNAseq = single-cell RNA sequencing; scATACseq = single-cell Assay for Transposase Accessible Chromatin sequencing; QC = quality control; UMI = unique molecular identifier; TSS = transcription start site. Please click here to view a larger version of this figure.
Table 1: Composition of single-nuclei lysis and wash buffers. Please click here to download this Table.
Table 2: Single-cell and single-nuclei count from mouse carotid artery luminal digestion. The table also shows means reads/cell and number of genes/cell for scRNAseq data. For the scATACseq data, mean fragments/cell and total reads obtained per sample are shown8. Abbreviations: scRNAseq = single-cell RNA sequencing; scATACseq = single-cell Assay for Transposase Accessible Chromatin sequencing; 2D = 2-day; 2W = 2-week; R/RCA = right carotid artery; L/LCA = left carotid artery. Please click here to download this Table.
Discussion
This paper provides a detailed protocol to isolate single-cell preparations from the mouse carotid arteries. The influence of d-flow on the endothelial cells can be accurately studied if the PCL surgery is performed correctly. It is crucial to correctly identify the branches of the common carotid, such as the external carotid, internal carotid, occipital artery, and superior thyroid artery. Validation of flow patterns by ultrasonography further validates the successful onset of d-flow conditions. Although PCL surgery can be performed on mice irrespective of their age, the preferred age is 10 ± 2 weeks. Older mice and mice with hypercholesterolemic backgrounds generally tend to have more periadventitial fat and stiffer skin.
It is essential to carefully dissect the carotid artery endothelial-enriched single-cell preparations free of the surrounding tissue and periadventitial fat. The lumen of the carotids must be perfused thoroughly to avoid contaminating blood cells in the single-cell preparations. Improper tissue identification and labeling can result in high standard deviation. Careful planning and time management are required for this protocol. A skilled surgeon and operator can take 15-20 min to perform one PCL surgery. Sacrificing, carotid isolation, and luminal enzymatic digestion from each mouse would take an additional 35-40 min. The hands-on processing time from endothelial cell flushing to single-cell/single-nuclei prep is an additional 2 h. Meticulous planning and teamwork are highly recommended.
As with any experimental protocol, some disadvantages and limitations should be considered. The current protocol does not incorporate steps to avoid artifactual activation of immediate early response during the luminal tissue dissociation. It has been reported in the literature that enzymatic digestion can lead to stress signaling events, such as inflammation and apoptosis. This can be addressed by incorporating appropriate control groups in the experimental design. In addition, wet laboratory methods that can minimize the artifactual activation of immediate response during enzymatic digestion, such as cold-active proteases, should be considered13,14.
This protocol can be used for any mouse strain irrespective of its genetic background. However, the endothelial-enriched preparations can best answer research questions pertaining to the changes in endothelium and are therefore well suited for EC-specific knockouts and EC-specific transgenic mice. This single-cell isolation method can be adapted to perform single-cell multi-omics studies and other flow cytometry-based assays such as Imaging Flow Cytometry. The luminal digestion approach could be used for freshly obtained mouse aortas, arteries from large animals (rabbits and pigs), as well as for freshly obtained human vascular explants. Collectively, this method would allow us to fully understand the precise role of blood flow on endothelial function and reprogramming at a single-cell resolution.
Divulgations
The authors have nothing to disclose.
Acknowledgements
This work was supported by funding from National Institutes of Health grants HL119798, HL095070, and HL139757 to HJ. HJ is also supported by the Wallace H. Coulter Distinguished Faculty Chair Professorship. The services provided by the Emory Integrated Genomics Core (EIGC) were subsidized by the Emory University School of Medicine and were also partly supported by the Georgia Clinical and Translational Science Alliance of the National Institutes of Health under award no. UL1TR002378. The content provided above is solely the authors' responsibility and does not reflect the official views of the National Institutes of Health.
Materials
| Chemicals, Peptides, and Recombinant Proteins | |||
| 1x PBS (Cell Culture Grade) | Corning | 21040CMX12 | |
| 1.5 mL Protein LoBind Microcentrifuge Tubes | Eppendorf | 022-43-108-1 | |
| 15 mL Centrifuge Tube – Foam Rack, Sterile | Fablab | FL4022 | |
| 50 mL SuperClear Centrifuge Tubes | Labcon | 3191-335-028 | |
| 6-0 Silk Suture Sterile | Covidien | s-1172 c2 | |
| 70 µm Cell Strainer, White, Sterile, Individually Packaged | Thermo Fisher Scientic | 08-771-2 | |
| Accutase solution,sterile-filtered | Sigma-Aldrich | A6964-100ML | or equivalent |
| ATAC Buffer (Component I of Transposition Mix) | 10x Genomics | 2000122 | |
| ATAC Enzyme (Component II of Transposition Mix) | 10x Genomics | 2000123/ 2000138 | |
| Bovine Serum albumin | Sigma-Aldrich | A7906-500G | |
| Buprenorphine | Med-Vet International | RXBUPRENOR5-V | |
| Chromium Controller & Next GEM Accessory Kit | 10X Genomics | 1000204 | |
| Chromium Next GEM Single Cell 3' Reagent Kits v3.1 | 10X Genomics | 1000121 | |
| Chromium Next GEM Single Cell ATAC Reagent Kits v1.1 | 10X Genomics | 1000175 | |
| Collagenase II | MP Biomedicals | 2100502.5 | |
| Digitonin | Sigma-Aldrich | D141-100MG | |
| Dissecting Forceps | Roboz Surgical Instruments Co | RS-5005 | |
| Dnase1 | New England Biolabs Inc | M0303S | |
| Centrifuge (Benchtop-Model # 5425) | Eppendorf | 22620444230VR | |
| Fetal Bovine Serum – Premium Select | R&D systems | S11550 | |
| Fixed Angle Rotor | Eppendorf | FA-45-24-11-Kit Rotor | |
| HEPES buffered saline | Millipore Sigma | 51558 | |
| Insulin syringe (3/10 mL 29 G syringe) | BD | 305932 | |
| Isoflurane | Patterson vet | 789 313 89 | |
| MACs Smart Strainers (30 µm) | Miltenyi Biotec | 130-098-458 | |
| MACS SmartStrainers (100 µm) | Miltenyi Biotec | 130-098-463 | |
| Normal Saline (0.9% sodium chloride) | Baxter International Inc | 2B1323 | |
| Nuclei Buffer (20x) | 10x Genomics | PN 2000153/2000207 | |
| PBS (10x), pH 7.4 | Thermo Fisher Scientic | 70011-044 | |
| Small scissors | Roboz Surgical Instruments Co | RS-5675 | |
| Stainless Steel Micro Clip Applying Forceps With Lock | Roboz Surgical Instruments Co. | RS-5480 | or similar |
| Tissue Mend II | Webster Veteinanry | 07-856-7946 | |
| Type II Collagenase | MP biomedicals | 2100502.1 | |
| Deposited Data | |||
| scATACseq FastQ files | NCBI | www.ncbi.nlm.nih.gov/bioproject Accession # PRJNA646233 | |
| scRNAseq FastQ files | NCBI | www.ncbi.nlm.nih.gov/bioproject Accession # PRJNA646233 | |
| Software and Algorithms | |||
| Cell Ranger 3.1.0 | 10X Genomics | https://support.10xgenomics.com/ single-cell-gene-exp | |
| Cicero | Pliner et al., 2018 | https://cole-trapnell-lab.github.io/cicero-release/ | |
| Ggplot2 v3.2.1 | Hadley Wickham | https://cran.r-project.org | |
| Harmony | Korsunsky et al., 2019 | https://github.com/immunogenomics/harmony | |
| ImageJ | Schneider et al., 2012 | https://imagej.nih.gov | |
| Monocle 2.8.0 | Qiu et al., 2017 | https://github.com/cole-trapnell-lab/ monocle-release | |
| R version 3.6.2 | R Foundation | https://www.r-project.org | |
| Seurat 3.1.3 | Stuart et al., 2019 | https://github.com/satijalab/seurat | |
| Signac 0.2.5 | Stuart et al., 2019 | https://github.com/timoast/signac |
References
- Kumar, S., Kang, D. W., Rezvan, A., Jo, H. Accelerated atherosclerosis development in C57Bl6 mice by overexpressing AAV-mediated PCSK9 and partial carotid ligation. Laboratory Investigation. 97 (8), 935-945 (2017).
- Nam, D., et al. Partial carotid ligation is a model of acutely induced disturbed flow, leading to rapid endothelial dysfunction and atherosclerosis. American Journal of Physiology Heart and Circulation Physiology. 297 (4), 1535-1543 (2009).
- Nam, D., et al. A model of disturbed flow-induced atherosclerosis in mouse carotid artery by partial ligation and a simple method of RNA isolation from carotid endothelium. Journal of Visualized Experiments: JoVE. (40), e1861 (2010).
- Dunn, J., et al. Flow-dependent epigenetic DNA methylation regulates endothelial gene expression and atherosclerosis. Journal of Clinical Investigation. 124 (7), 3187-3199 (2014).
- Kumar, S., Kim, C. W., Son, D. J., Ni, C. W., Jo, H. Flow-dependent regulation of genome-wide mRNA and microRNA expression in endothelial cells in vivo. Scientific Data. 1 (1), 140039 (2014).
- Ni, C. W., et al. Discovery of novel mechanosensitive genes in vivo using mouse carotid artery endothelium exposed to disturbed flow. Blood. 116 (15), 66-73 (2010).
- Son, D. J., et al. The atypical mechanosensitive microRNA-712 derived from pre-ribosomal RNA induces endothelial inflammation and atherosclerosis. Nature Communications. 4, 3000 (2013).
- Andueza, A., et al. Endothelial reprogramming by disturbed flow revealed by single-cell RNA and chromatin accessibility study. Cell Reports. 33 (11), 108491 (2020).
- Stuart, T., Srivastava, A., Lareau, C., Satija, R. Multimodal single-cell chromatin analysis with Signac. bioRxiv. , (2020).
- Stuart, T., et al. Comprehensive integration of single-cell data. Cell. 177 (7), 1888-1902 (2019).
- Crowley, L. C., Marfell, B. J., Christensen, M. E., Waterhouse, N. J. Measuring cell death by trypan blue uptake and light microscopy. Cold Spring Harbor Protocols. 2016 (7), (2016).
- Li, F., et al. Single-cell RNA-seq reveals cellular heterogeneity of mouse carotid artery under disturbed flow. Cell Death and Discovery. 7 (1), 180 (2021).
- Wu, Y. E., Pan, L., Zuo, Y., Li, X., Hong, W. Detecting activated cell populations using single-cell RNA-seq. Neuron. 96 (2), 313-329 (2017).
- O’Flanagan, C. H., et al. Dissociation of solid tumor tissues with cold active protease for single-cell RNA-seq minimizes conserved collagenase-associated stress responses. Genome Biology. 20 (1), 210 (2019).

