An Improved Chemotaxis Assay for the Rapid Identification of Rhizobacterial Chemoattractants in Root Exudates
Summary
Here, we present an improved chemotaxis assay protocol. The goal of this protocol is to reduce the steps and costs of traditional bacterial chemotaxis methods and to serve as a valuable resource for understanding plant-microbe interactions.
Abstract
Chemotaxis identification is very important for the research and application of rhizosphere growth-promoting bacteria. We established a straightforward method to quickly identify the chemoattractants that could induce the chemotactic movement of rhizosphere growth-promoting bacteria on sterile glass slides via simple steps. Bacteria solution (OD600 = 0.5) and sterile chemoattractant aqueous solution were added dropwise on the glass slide at an interval of 1 cm. An inoculating loop was used to connect the chemoattractant aqueous solution to the bacterial solution. The slide was kept at room temperature for 20 min on the clean bench. Finally, the chemoattractant aqueous solution was collected for bacterial counting and microscopic observation. In this study, through multiple comparisons of experimental results, the method overcame multiple shortcomings of traditional bacterial chemotaxis methods. The method reduced the error of plate counting and shortened the experimental cycle. For the identification of chemoattractant substances, this new method can save 2-3 days compared to the traditional method. Additionally, this method allows any researcher to systematically complete a bacterial chemotaxis experiment within 1-2 days. The protocol can be considered a valuable resource for understanding plant-microbe interactions.
Introduction
Chemotaxis is important for colonization of plant growth-promoting rhizobacterial (PGPR) on roots and for understanding plant-microbe interactions1. A class of low molecular weight compounds (chemoattractants) in plant root exudates induce the chemotactic movement of PGPR to the rhizosphere2. Malic acid, citric acid, and other components in the root exudates stimulate chemotaxis of Bacillus strains3. For example, glucose, citric acid, and fumaric acid in maize root exudates recruit bacteria to the root surface4. D-galactose, which is derived from root exudates,induces the chemotaxis of Bacillus velezensis SQR95. Organic acids, including fumarate, malic acid, and succinate, influence chemotaxis and colonization of various PGPR in the Cajanus cajan – Zea mays intercropping system6. Oleanolic acid in rice root exudates, acts as a chemoattractant for the strain FP357. Other plant exudates (including histidine, arginine, and aspartate) can play a crucial role in the chemotactic response of bacteria8. Plant exudates function as a signal to direct the movement of bacteria, which is the first step during rhizosphere colonization. Plant colonization by PGPR is a process of enormous relevance, as PGPR are beneficial for the plant host.
Many methods have been used for analyzing bacterial chemotaxis. The swimming plate method is one of the methods described previously9. In this method, plates were made with a semisolid medium. A chemotactic buffer containing agar (1.0%, w/v) was added to the plate. The buffer is heated, and then mixed with the chemoattractant. Then, 8 µL of bacteria suspension was added dropwise to the middle of the plate and the plate was placed in an incubator at 28 °C. The plate was regularly observed and photographed. However, the experimental cycle of the swimming plate method was very long. In the capillary-like method10, a pipette tip serves as a chamber for holding 100 µL of bacterial suspension. 1 mL syringe needle was used as a capillary. A syringe needle containing chemoattractants with different concentration gradients was inserted into the 100 µL pipette tip. After incubation at room temperature for 3 h, the syringe needle was removed, the content was diluted and plated on the medium. The bacterial accumulation in the syringe was represented by colony-forming units (CFUs) in the plates. However, the experimental error within replicates for the capillary-like method was large. Another method used a microfluidic SlipChip device11. Briefly, bovine serum albumin (BSA) solution was injected into all channels and removed using a vacuum. The solutions containing different chemoattractants (1 mM concentration for qualitative detection only), bacterial cells suspended in phosphate-buffered saline and phosphate buffered saline buffer (negative control) were added to the top, middle, and bottom microwells, respectively. Incubation was then performed in a dark environment at room temperature for 30 min. The bacterial cells were then detected in the microwells. The microfluidic SlipChip device, however, was expensive. Therefore, each of the methods described above had advantages and disadvantages.
We established an improved chemotaxis assay for the rapid identification of rhizobacterial chemoattractants in root exudates using sterile glass slides without complicated steps. In this study, through multiple comparisons of experimental results, the method overcame multiple shortcomings of traditional bacterial chemotaxis methods. The method reduced the error of plate counting and shortened the experimental cycle. Therefore, if used to identify a chemoattractant substance, this new method can save 2-3 days and reduce the cost of experimental materials.
Protocol
1. Materials and equipment
NOTE: Bacillus altitudinis LZP02 (CP075052) was isolated from the rhizosphere of rice in northeast China12,13 for this study.
- Culture B. altitudinis LZP02 in Luria-Bertani (LB) medium (peptone, 10 g L-1; NaCl, 8 g L-1 and yeast extract, 5 g L-1) for 10 h. Collect cells by centrifugation at 9,569 x g for 2 min at 4 °C. and store with 15% glycerol at -80 °C.
NOTE: For this experiment, rice seeds (Oryza sativa Longgeng 46) were provided by the Rice Research Institute of Heilongjiang Academy of Agricultural Sciences.
2. Collection of root exudates
- Randomly distribute rice seeds in a growth chamber.
NOTE: Rice seeds were sterilized with 30% H2O2 for 30 min and soaked in water overnight. The conditions were as follows: the controlled light (16/8 h light/dark cycle), temperature (22 ± 2 °C) and relative humidity (῀70%). - Culture rice seeds for a week and add sterile water twice.
- Select rice seedlings of similar size and plant in 50 mL of Murashige and Skoog (MS) liquid medium. Incubate for 48 h at 22 °C under aseptic conditions.
NOTE: Rice root exudates will be released into the MS medium14,15,16.
3. Liquid chromatography-mass spectrometry analysis of root exudates
- Collect 100 µL of the sample (MS medium containing root exudates) in a 1.5 mL centrifuge tube. Add 20 µL of the extraction solvent (acetonitrile-methanol-water, 2:2:1, including internal standard). Homogenize the sample at 45 Hz for 4 min, followed by ultrasonication on ice for 5 min in a water bath.
- Repeat the homogenization and ultrasonic cycle three times. Incubate the sample at -20 °C for 1 h, followed by centrifugation at 133,778 x g and 4 °C for 15 min.
- Transfer the resulting supernatant to LC-MS vials and store at -80 °C until UHPLC-QE analysis. Prepare the quality control (QC) samples by mixing equal portions of the supernatant of all samples.
NOTE: Each sample volume was 600 µL (six replicates per experiment) in the presented experiment. - Perform LC-MS/MS analysis using UHPLC system, UPLC HSS T3 column (2.1 mm x 100 mm, 1.8 µm) and Q Exactive12.
- Use 0.1% formic acid aqueous solution and 5 mmol/L ammonium acetate aqueous solution as mobile phase A and acetonitrile as mobile phase B. Use formic acid and ammonium acetate as positive and negative ion modes, respectively.
- Set the elution gradient as follows: 0 min, 1% B; 1 min, 1% B; 8 min, 99% B. 10 min, 99% B. 10.1 min, 1% B; 12 min, 1% B. Set the flow rate and injection volume to 0.5 mL/min and 2 µL, respectively.
NOTE: The macromolecule (>1,000 Daltons) cannot be detected.
4. Chemotaxis assay
- Prepare the chemoattractant aqueous solution. Ensure that it is sterile. Filter the chemoattractant solution with a 0.22 µm bacteria filter.
NOTE: The chemoattractant aqueous solution was the single substance obtained from the LC-MS studies dissolved in water. The concentration and volume can be adjusted appropriately according to different studies. Citric acid solution was used as an example. All actions must be performed by the side of a lamp. - Mark the middle position of the glass slide at an interval of 1 cm. Ensure that the glass slide is sterilized several times on the flame.
- Add the 30 µL of chemoattractant solution on the left of the glass slide. Ensure that bacteria were cultured to the logarithmic stage (2 x 108 CFU/mL) in LB medium. Add 30 µL of the bacterial solution on the right of the glass slide.
NOTE: Prepare a negative control group with an equal volume of sterile water. The saline solution (0.9% NaCI) was used as the positive control in order to eliminate the changes caused by intermolecular force on the experiment. - Sterilize an inoculating loop several times on the flame. Use the inoculating loop to connect the chemoattractant aqueous solution to the bacterial solution and keep it at room temperature for 20 min on a clean bench.
NOTE: The experiment must be carried out in a windless environment. Adjust the time before disconnecting the connection line for strains of different athletic abilities appropriately. - After 20 min, disconnect the connecting line with filter paper.
- Ensure that the 1.5 mL centrifuge tube is sterile. Collect the chemoattractant aqueous solution on the left of the glass slide. Transfer the solution to the 1.5 mL sterile centrifuge tube.
- Add the appropriate volume of safranin to the centrifuge tube. After 2 min, collect the miscible liquids for counting bacteria and microscopic observation with a blood counting chamber.
5. Results analysis
- Determine the number of viable microorganisms attracted using the following equation:
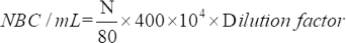
where NBC: Total number of bacteria cells; N: Number of bacteria in 80 grids.
NOTE: Statistical analysis software was used for data analysis. The error was based on three different replicated experimental values and was calculated using one-way ANOVA followed by Turkey's post-hoc analysis. P ≤ 0.05 was considered significant.
Representative Results
A total of 584 and 937 known metabolites were detected in the positive and negative ion indices, respectively. Previous studies have shown that chemoattractants are typically organic acids, amino acids, and carbohydrates17,18.
In this study, 16 kinds of chemoattractants from the LC-MS studies in the rice rhizosphere exudates were selected for subsequent experiments (Table 1). Using the swimming plate method, we screened the low molecular weight compounds secreted by the root for an attractant of the rhizospheric B. altitudinis LZP02. To compare the strengths of the different chemoattractants, the chemotaxis index (RCI), which shows the ratio of average treatment bacteria to the corresponding control, was introduced19. A higher RCI indicates a stronger reaction to the chemoattractant. The results indicated that, 100 µM citric acid (RCI = 2.34) and 100 µM caffeic acid (RCI = 1.85) had the highest chemotaxis indices (Figure 2). Representative images showed that the swarming areas of citric acid and caffeic acid was more compared to CK (Figure 3).
Using the new chemotaxis assay method (Figure 1), the results showed that the bacterial concentration of the citric acid, caffeic acid, and galactose experiments (100 µM) were significantly higher than those of the negative and positive control groups. Citric acid was the strongest chemoattractant. The experimental results (Figure 4 and Table 2) were in agreement with the results of the traditional swimming plate method (Figure 2). Representative microscopic images of the new chemotaxis assay are shown in Figure 5.
Compared with the swimming plate method, our results confirmed the feasibility of the new chemotaxis assay. This technique can save considerable amount of time, reduce the cost of reagents and reduce errors in the result. The results of the new method were intuitive, and the method showed the motility of bacteria.
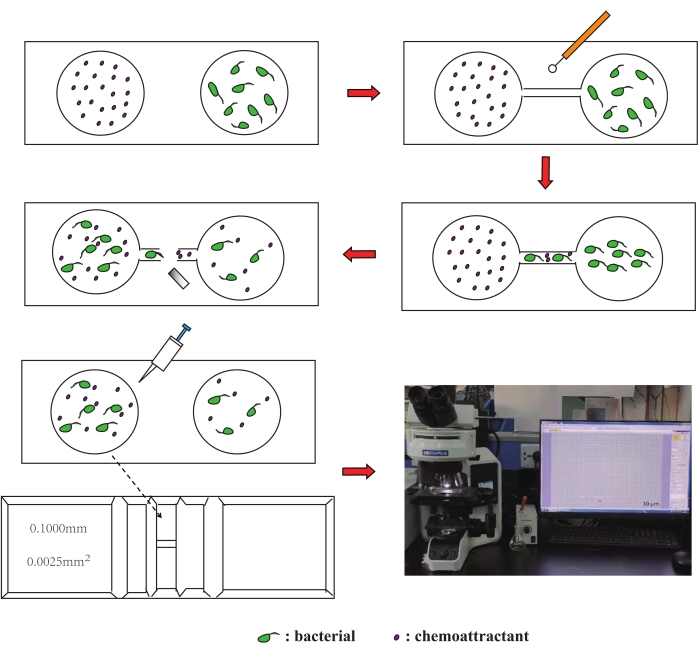
Figure 1: Schematic diagram of the new chemotaxis assay. Please click here to view a larger version of this figure.
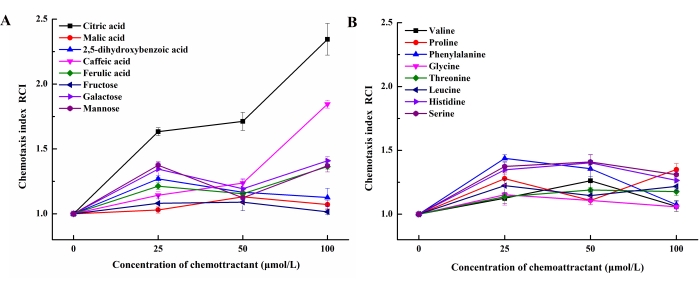
Figure 2: Results of the swimming plate method. (A) Chemotactic response of B. altitudinis LZP02 to increasing concentrations of organic acids, phenolic acids, and carbohydrates. (B) Chemotactic response of B. altitudinis LZP02 to increasing concentrations of amino acids. Please click here to view a larger version of this figure.
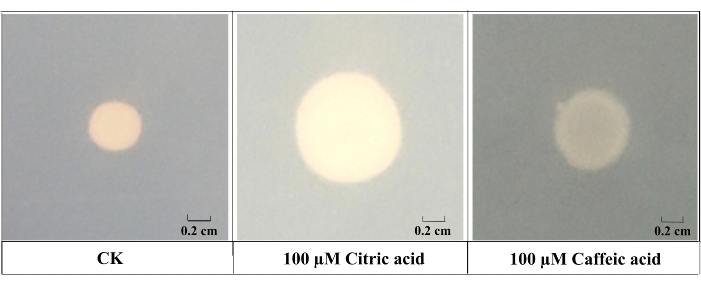

Figure 3: Results of the swimming plate method. Comparison of the chemotactic response of B. altitudinis LZP02 toward organic acids. Please click here to view a larger version of this figure.
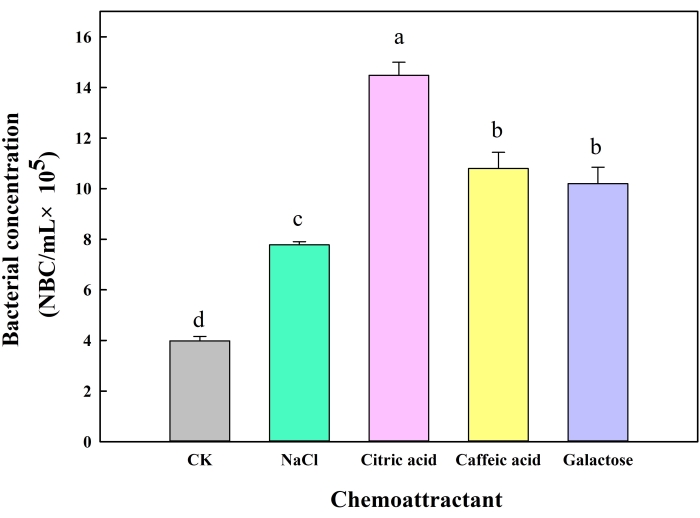
Figure 4: Results of the new chemotaxis assay method. NBC: Total number of bacteria cells. Chemotactic response of B. altitudinis LZP02 to increasing concentrations of citric acid, caffeic acid, and galactose. The results are expressed as the mean of at least three assays for each determination. Error bars indicate the SDs based on three different replicated experimental values. Columns identified by different letters are significantly different at P ≤ 0.05 according to one-way ANOVA followed by Turkey's post-hoc analysis. Please click here to view a larger version of this figure.
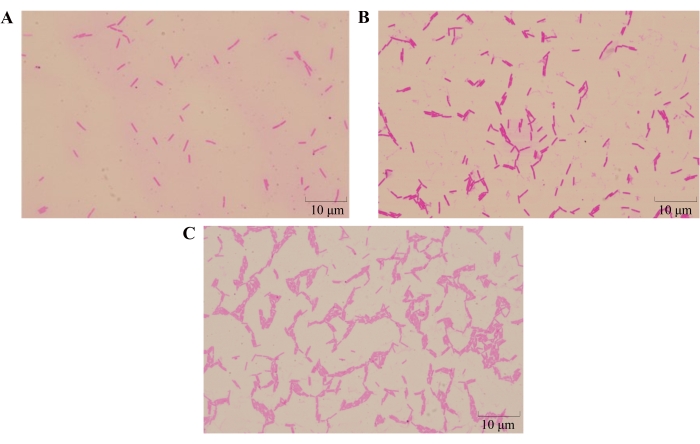
Figure 5: Results of the new chemotaxis assay method. The images were obtained under a light microscope. (A) CK, (B) saline solution, (C) citric acid. Please click here to view a larger version of this figure.
| Index | Name | Score | Abundance |
| POS01062 | Citric acid | 0 | 200471.6 |
| NEG00016 | Malic acid | 0.9944492 | 2093440.97 |
| NEG00038 | 2,5-dihydroxybenzoic acid | 0.8886685 | 114774.98 |
| POS01763 | Caffeic acid | 0 | 6195.5 |
| POS00154 | Ferulic acid | 0.4725294 | 12723.95 |
| POS02156 | Fructose | 0 | 6767.81 |
| POS01289 | Galactose | 0 | 871476.14 |
| NEG00021 | Mannose | 0.9898174 | 683176.34 |
| POS00037 | Valine | 0.9660872 | 5985865.24 |
| POS00018 | Proline | 0.9887274 | 2739147.17 |
| POS00124 | Phenylalanine | 0.6106034 | 11940645.8 |
| POS00092 | Glycine | 0.7418478 | 22787114.1 |
| POS00079 | Threonine | 0.8086754 | 1039040.18 |
| NEG00010 | Leucine | 0.9962673 | 3546561.95 |
| POS00266 | Histidine | 0 | 3010220.41 |
| POS01993 | Serine | 0 | 645602.04 |
| Note: POS and NEG in first column indicate positive and negative ion index, respectively. Third column indicate, which the second-level spectrum was used to score database matching and the ungraded chemoattractants were matched by the first order spectrum. The score ranges from 0 to 1. | |||
Table 1: 16 kinds of chemoattractants from the LC-MS studies in the rice rhizosphere exudates.
| Chemoattractant | RCI |
| NaCl | 1.95±0.03 |
| Citric acid | 3.63±0.12 |
| Caffeic acid | 2.71±0.26 |
| Galactose | 2.56±0.16 |
Table 2: Chemotaxis index (RCI) of the new chemotaxis assay method.
Discussion
Increasing research indicates that plant-bacteria interactions mainly occur in the rhizosphere and are influenced by root exudates20,21,22,23,24. Plant root exudates include a diverse array of primary metabolites, including phenolic acids, organic acids and amino acids as well as more complex secondary compounds25,26,27. For each bacterium, specific substances have been shown to have recruiting effects in rhizosphere exudates. In this research field, many of the chemotaxis methods reported in the literature are highly assay-dependent. There are reports on a variety of approaches, which all have specific advantages and drawbacks.
In the present study, we focused on introducing an improved chemotaxis assay for the rapid identification of rhizobacterial chemoattractants in root exudates. In this protocol, all experiments were performed under aseptic conditions. In step 4, we used the inoculating loop to connect the chemoattractant aqueous solution to the bacteria solution, and we kept the slide at room temperature for 20 min. It is worth noting that the width of the connection line was controlled to approximately 2 mm, and the experiment must be carried out in a windless environment.
Appropriate modifications should be made according to the experimental strain being used. The logarithmic culture time should be selected according to the growth cycle of different strains. For strains of different swarming abilities, the time before disconnecting the connection line can be adjusted appropriately. Small sized bacterium species that cannot be observed under a light microscope are not suitable for this method. For example, nanobacteria can be as small as 80 nm in diameter and can only be seen through an electron microscope. This method is suitable for strains with flagella.
The new method reduces the error of plate counting and shortens the experimental cycle. For the identification of chemoattractant substances, the new method can save 2-3 days. The new method allows any researcher to systematically complete a bacterial chemotaxis experiment within 1-2 days. Compared to the microfluidic device, the new method is less expensive. The protocol also overcomes some shortcomings of traditional bacterial chemotaxis methods. Furthermore, the new method can be used in various other ways. For instance, we should utilize this method to quickly identify bacteria in soil stimulated by signals from specific plant root exudates. The new method is simple in design, easy to perform, and has great application value. This protocol can be presented as a valuable resource for the identification and understanding of plant-microbe interactions.
Divulgations
The authors have nothing to disclose.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (Nos. 31870493), the Key Research and Development Projects in Heilongjiang, China (GA21B007), and the Basic Research Fees of Universities in Heilongjiang Province, China (No. 135409103).
Materials
| 2,5-dihydroxybenzoic acid | Beijing InnoChem Science & Technology C.,Ltd. | 490-79-9 | |
| Acetonitrile | CNW Technologies | 75-05-8 | |
| Ammonium acetate | CNW Technologies | 631-61-8 | |
| Caffeic acid | Beijing InnoChem Science & Technology C.,Ltd. | 331-39-5 | |
| Centrifuge | Thermo Fisher Scientific | Heraeus Fresco17 | |
| Citric acid | Beijing InnoChem Science & Technology C.,Ltd. | 77-92-9 | |
| Clean bench | Shanghai Boxun Industrial Co., Ltd. | BJ-CD | |
| Ferulic acid | Beijing InnoChem Science & Technology C.,Ltd. | 1135-24-6 | |
| Formic acid | CNW Technologies | 64-18-6 | |
| Fructose | Beijing InnoChem Science & Technology C.,Ltd. | 57-48-7 | |
| Galactose | Beijing InnoChem Science & Technology C.,Ltd. | 59-23-4 | |
| Glycine | Beijing InnoChem Science & Technology C.,Ltd. | 56-40-6 | |
| Grinding Mill | Shanghai Jingxin Industrial Development Co., Ltd. |
JXFSTPRP-24 | |
| Histidine | Beijing InnoChem Science & Technology C.,Ltd. | 71-00-1 | |
| Internal standard: 2-Chloro-L-phenylalanine | Shanghai Hengbai Biotech C.,Ltd. | 103616-89-3 | |
| Leucine | Beijing InnoChem Science & Technology C.,Ltd. | 61-90-5 | |
| Malic acid | Beijing InnoChem Science & Technology C.,Ltd. | 6915-15-7 | |
| Mannose | Beijing InnoChem Science & Technology C.,Ltd. | 3458-28-4 | |
| Mass Spectrometer | Thermo Fisher Scientific | Q Exactive Focus | |
| Methanol | CNW Technologies | 67-56-1 | |
| Optical Microscope | Olympus | BX43 | |
| Phenylalanine | Beijing InnoChem Science & Technology C.,Ltd. | 63-91-2 | |
| Proline | Beijing InnoChem Science & Technology C.,Ltd. | 147-85-3 | |
| Scales | Sartorius | BSA124S-CW | |
| Serine | Beijing InnoChem Science & Technology C.,Ltd. | 56-45-1 | |
| Threonine | Beijing InnoChem Science & Technology C.,Ltd. | 72-19-5 | |
| UHPLC | Agilent | 1290 UHPLC | |
| Ultrasound Instrument | Shenzhen Leidebang Electronics Co., Ltd. |
PS-60AL | |
| Valine | Beijing InnoChem Science & Technology C.,Ltd. | 7004-03-7 |
References
- Belas, R. Biofilms, flagella, and mechanosensing of surfaces by bacteria. Trends in Microbiology. 22 (9), 517-527 (2014).
- Haichar, Z., Santaella, C., Heulin, T., Achouak, W. Root exudates mediated interactions belowground. Soil Biology and Biochemistry. 77 (7), 69-80 (2014).
- Zhang, N., et al. Effects of different plant root exudates and their organic acid components on chemotaxis, biofilm formation and colonization by beneficial rhizosphere-associated bacterial strains. Plant and Soil. 374 (1-2), 689-700 (2014).
- Zhang, N., et al. Whole transcriptomic analysis of the plant-beneficial rhizobacterium Bacillus amyloliquefaciens SQR9 during enhanced biofilm formation regulated by maize root exudates. BMC Genomics. 16 (1), 685 (2015).
- Lui, Y., et al. Induced root-secreted D-galactose functions as a chemoattractant and enhances the biofilm formation of Bacillus velezensis SQR9 in an mcpa-dependent manner. Applied Microbiology and Biotechnology. 104 (17), 785-797 (2020).
- Vora, S. M., Joshi, P., Belwalkar, M., Archana, G. Root exudates influence chemotaxis and colonization of diverse plant growth-promoting rhizobacteria in the Cajanus cajan – Zea mays intercropping system. Rhizosphere. 18 (12), 100331 (2021).
- Sampedro, I., et al. Effects of halophyte root exudates and their components on chemotaxis, biofilm formation and colonization of the halophilic bacterium halomonas anticariensis FP35T. Microorganisms. 8 (4), 575 (2020).
- Liu, X. L., Raza, W., Ma, J. H., Huang, Q. W., Shen, Q. R. A dual role amino acid from sesbania rostrata seed exudates in the chemotaxis response of Azorhizobium caulinodans ORS571. Molecular Plant-Microbe Interactions. 32 (9), 1134-1147 (2019).
- Ling, N., Raza, W., Ma, J. H., Huang, Q. W., Shen, Q. R. Identification and role of organic acids in watermelon root exudates for recruiting Paenibacillus polymyxa SQR-21 in the rhizosphere. European Journal of Soil Biology. 47 (6), 374-379 (2011).
- Rudrappa, T., Czymmek, K. J., Pare, P. W., Bais, H. P. Root-secreted malic acid recruits beneficial soil bacteria. Plant Physiology. 148 (3), 1547-1556 (2008).
- Shen, C., et al. Bacterial chemotaxis on Slipchip. Lab on a Chip. 14 (16), 3074-3080 (2014).
- Liu, H., et al. Bacillus pumilus LZP02 promotes rice root growth by improving carbohydrate metabolism and phenylpropanoid biosynthesis. Molecular Plant-Microbe Interactions. 33 (10), 1222-1231 (2020).
- Goswami, M., Deka, S. Isolation of a novel rhizobacteria having multiple plant growth promoting traits and antifungal activity against certain phytopathogens. Microbiological Research. 240, 126516 (2020).
- Kaiira, M., Chemining’Wa, G., Ayuke, F., Baguma, Y., Nganga, F. Profiles of compounds in root exudates of rice, cymbopogon, desmodium, mucuna and maize. Journal of Agricultural Sciences Belgrade. 64 (4), 399-412 (2019).
- Shi, Y., et al. Effect of rice root exudates and strain combination on biofilm formation of Paenibacillus polymyxa and Paenibacillus macerans. African Journal of Microbiology Research. 6 (13), 3343-3347 (2012).
- Lee, H. W., Ghimire, S. R., Shin, D. H., Lee, I. J., Kim, K. U. Allelopathic effect of the root exudates of K21, a potent allelopathic rice. Weed Biology and Management. 8 (2), 85-90 (2008).
- Belimov, A. A., et al. Rhizobacteria that produce auxins and contain 1-amino-cyclopropane-1-carboxylic acid deaminase decrease amino acid concentrations in the rhizosphere and improve growth and yield of well-watered and water-limited potato (Solanum tuberosum). Annals of Applied Biology. 167 (1), 11-25 (2015).
- Ankati, S., Podile, A. R. Metabolites in the root exudates of groundnut change during interaction with plant growth promoting rhizobacteria in a strain-specific manner. Journal of Plant Physiology. 243, 153057 (2019).
- Gordillo, F., Chavez, F., Jerez, C. A. Motility and chemotaxis of Pseudomonas sp. B4 towards polychlorobiphenyls and chlorobenzoates. FEMS Microbiology Ecology. 60 (2), 322-328 (2007).
- Bais, H. P., Weir, T. L., Perry, L. G., Gilroy, S., Vivanco, J. M. The role of root exudates in rhizosphere interactions with plants and other organisms. Annual Review of Plant Biology. 57, 233-266 (2006).
- Badri, D. V., Vivanco, J. M. Regulation and function of root exudates. Plant Cell Environment. 32 (6), 666-681 (2009).
- Badri, D. V., Weir, T. L., Lelie, D., Vivanco, J. M. Rhizosphere chemical dialogues: plant-microbe interactions. Curr Opin Biotech. Current Opinion in Biotechnology. 20 (6), 642-650 (2009).
- Kamilova, F., Kravchenko, L. V., Shaposhnikov, A. I., Makarova, N., Lugtenberg, B. Effects of the tomato pathogen Fusarium oxysporum f. sp. radicis-lycopersici and of the biocontrol bacterium Pseudomonas fluorescens WCS365. Molecular Plant-Microbe Interactions. 19 (10), 1121-1126 (2006).
- Kamilova, F., et al. Organic acids, sugars, and L-tryptophane in exudates of vegetables growing on stonewool and their effects on activities of rhizosphere bacteria. Molecular Plant-Microbe Interactions. 19 (3), 250-256 (2006).
- Badri, D. V., Vivanco, J. M. Regulation and function of root exudates. Plant Cell Environment. 32 (6), 666-681 (2009).
- Hao, W. Y., Ren, L. X., Ran, W., Shen, Q. R. Allelopathic effects of root exudates from watermelon and rice plants on Fusarium oxysporum f.sp. Niveum. Plant and Soil. 336 (1-2), 485-497 (2010).
- Hao, Z. P., Wang, Q., Christie, P., Li, X. L. Allelopathic potential of watermelon tissues and root exudates. Scientia Horticulturae. 112 (3), 315-320 (2007).

