Epithelial Cell Infection Analyses with Shigella
PRÉPARATION DE L'INSTRUCTEUR
concepts
PROTOCOLE ETUDIANT
1. Preparation of reagents and materials
NOTE: All volumes are consistent with an assay using two 6-well plates.
- TSB medium: Add 0.5 L of deionized (DI) water to 15 g of Tryptic Soy Broth (TSB, see Table of Materials) medium and autoclave. Store at room temperature.
- Bile salts medium (TSB + BS): To prepare TSB containing 0.4% (w/v) bile salts, resuspend 0.06 g of bile salts (BS, see Table of Materials) in 15 mL of autoclaved TSB. Filter sterilize using a 0.22 µm PES filter.
NOTE: The bile salts consist of a 1:1 mixture of sodium cholate and sodium deoxycholate. Prepare fresh media immediately before use. - DMEM + 10% (v/v) FBS: Add 5 mL of fetal bovine serum (FBS) to 45 mL of Dulbecco's Modified Eagle Medium (DMEM). Store at 4 °C.
- DMEM + gentamicin: To a 50 mL tube, add 50 mL of DMEM and 50 µL of 50 mg/mL gentamicin (see Table of Materials).
NOTE: Make fresh aliquot and warm in a 37 °C water bath prior to each experiment. - PBS + 1% (v/v) Triton X-100: Add 150 µL of Triton X-100 to 15 mL of Phosphate-buffered saline (PBS).
NOTE: Make fresh aliquot and warm in a 37 °C water bath prior to each experiment. - TSB + Congo red indicator plates: Add 15 g of TSB, 7.5 g of select agar, and 0.125 g of Congo red dye (see Table of Materials) to a 1 L bottle. Add 0.5 L of DI water and autoclave. Pour 10-20 mL of media into individual sterile Petri dishes (100 mm x 15 mm) and let solidify.
CAUTION: Congo red is carcinogenic and a reproductive toxin. Ensure that handling of Congo red is performed using the appropriate personal protective equipment. Consult the product safety data sheet for additional information.
NOTE: Approximately 20 plates are made from 0.5 L Congo red media. Plates can be prepared 2-3 days in advance and left inverted at room temperature until use. For long-term storage, place inverted plates in plastic sleeves at 4 °C for up to 3 months. - DMEM + 10% (v/v) FBS and 5% (v/v) dimethyl sulfoxide (DMSO): Add 42.5 mL of DMEM, 5 mL of FBS, and 2.5 mL of DMSO to a 50 mL tube. Store at 4 °C.
2. Preparation of bacteria
NOTE: All Shigella laboratory cultivation and storage protocols are adapted from Payne, S. M.43.
CAUTION: Shigella spp. are Risk Group 2 pathogens44. Perform all laboratory work in a BSL-2 environment, with additional safety measures undertaken to limit accidental exposures due to the low infectious dose of Shigella spp.
- Growth of Shigella from frozen stocks
- Transfer a small amount of frozen culture from the cryogenic vial to a TSB + Congo red agar plate using a sterile applicator.
- Flame sterilize an inoculating loop and allow it to cool. Streak inoculum back and forth across one quadrant of the plate. Flame the loop, allow it to cool, then streak from the first quadrant onto the second quadrant of the plate. Repeat to streak inoculum into the third and fourth quadrants of the plate.
NOTE: Alternatively, streak inoculum using a fresh sterile applicator between each quadrant. - Invert the plate and incubate at 37 °C overnight.
NOTE: Incubation at temperatures ≥37 °C is required for expression of Shigella virulence factors necessary for observation of Congo red-positive (CR+) phenotype45. Avirulent colonies will have a white appearance and will not be invasive. - Seal the plate with paraffin film and store refrigerated at 4 °C.
NOTE: Bacterial colonies will remain viable on agar plates for 1-2 weeks.
- Overnight growth of Shigella in liquid culture
- Aliquot 3 mL of TSB media into sterile 14 mL culture tubes.
- Pick a single, well-isolated red (CR+) colony using a sterile applicator and resuspend in liquid media.
- Incubate cultures overnight (16-18 h) at 37 °C with shaking at 250 rotations per minute (rpm).
3. Preparation of HT-29 eukaryotic cells
NOTE: All volumes are consistent with an assay using two 6-well plates. HT-29 cell lines were acquired from the American Type Culture Collection (ATCC). HT-29 maintenance protocols are adapted from ATCC recommendations46. All media should be pre-warmed in a water bath at 37 °C prior to use. All HT-29 maintenance protocols should be performed in a biosafety cabinet. Refrain from producing bubbles when mixing/working with HT-29 cells in media to avoid dramatic changes in pH.
- Thawing HT-29 cells from frozen stock
- Thaw the vial of HT-29 cells in a 37 °C water bath.
NOTE: Ensure the cap stays fully above the water to avoid contamination. Thawing should take less than 2 min. - Remove the vial from the water immediately after the culture is fully thawed and decontaminate with 70% ethanol. Ensure that all steps from this point are performed using aseptic techniques.
- Add all the contents of the vial to a 15 mL centrifuge tube containing 9 mL of DMEM + 10% FBS. Centrifuge at 125 x g for 5 min at room temperature.
- Decant the supernatant into a waste container and resuspend the pellet in 10 mL of warm DMEM + 10% FBS. Transfer resuspended cells to a 75 cm2 tissue culture flask (T75) containing 10 mL of warm DMEM + 10% FBS (total volume of 20 mL).
- Incubate cells at 37 °C with 5% CO2 until the cells reach 90% confluency (approximately 6-7 days).
NOTE: Confluency is estimated through visual approximation.
- Thaw the vial of HT-29 cells in a 37 °C water bath.
- Seeding HT-29 cells
- Pre-warm 20 mL of PBS and 50 mL of DMEM + 10% FBS in a 37 °C water bath and pre-warm 3 mL of 0.25% (w/v) Trypsin-EDTA to room temperature.
- Once HT-29 cells (from step 3.1) reach 90% confluency, decant HT-29 cell culture media from the T75 flask into a waste container. Pour ~10 mL of warm PBS into the flask and swirl gently to wash. Decant the PBS into a waste container. Wash with warm PBS again and decant.
- Add 2-3 mL of 0.25% (w/v) Trypsin-EDTA and gently swirl across the entire surface area. Incubate at 37 °C with 5% CO2 for 4 min.
- Remove the flask from the incubator and gently swirl the Trypsin-EDTA, visually ensuring that all cells detach from the surface.
- Immediately add 6 mL of warm DMEM + 10% FBS to deactivate the Trypsin. Pipette up and down to thoroughly mix.
- Transfer all the contents to a 15 mL centrifuge tube and spin it at 500 x g for 5 min at room temperature.
- Gently decant supernatant into a waste container and resuspend the pellet in 6 mL of warm DMEM + 10% FBS.
- Immediately after resuspension, transfer 10 µL of suspended HT-29 cells from the middle of the culture to a 0.2 mL PCR tube. Add 10 µL of Trypan blue dye to the PCR tube and mix.
- Add 10 µL of HT-29 cell/Trypan blue mix to a disposable Countess cell counter chamber slide (see Table of Materials). Enumerate the number of live cells and calculate cell viability.
NOTE: When documenting the number of cells in the sample, read the number under the "live" cell count, not the total cell count. Alternatively, cell enumeration can be performed manually using a hemocytometer. - Seed resuspended HT-29 cells into a fresh T75 flask or 6-well plate.
- For T75 flask:
- Pipet gently to mix, then transfer 2.5 x 106 cells to a fresh T75 flask according to the equation below:
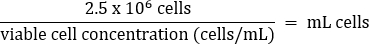
- Add warm DMEM + 10% FBS media to a final volume of 20 mL (final concentration of 1.25 x 105 cells/mL).
- Disperse cells evenly across the flask by gently rocking back and forth.
- Incubate at 37 °C with 5% CO2 until cells reach 80% confluency.
NOTE: For optimal growth, replace the DMEM + 10% FBS media in the T75 flask every ~3 days. Decant media into a waste container and add 10 mL of warm PBS to the flask. Swirl the PBS around gently and decant it into the waste container. Then add 20 mL of fresh, warm DMEM + 10% FBS to the flask and return to the 37 °C, 5% CO2 incubator.
- Pipet gently to mix, then transfer 2.5 x 106 cells to a fresh T75 flask according to the equation below:
- For 6-well plate:
- Pipet gently to mix, then transfer 5.85 x 106 cells to a fresh 50 mL conical tube according to the equation below:
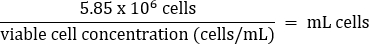
- Add warm DMEM + 10% FBS media to a final volume of 26 mL (final concentration of 2.25 x 105 cells/mL).
- Pipet gently to mix, then dispense 2 mL (4.5 x 105 cells) into individual wells of 6-well plates.
- Disperse cells evenly across the well by gently rocking up/down and left/right 2-3x.
- Incubate at 37 °C with 5% CO2 until cells reach 80%-95% confluency (approximately 3-4 days).
NOTE: 85% confluency is recommended for invasion and intracellular replication assays, while 90%-95% confluency is recommended for adherence assays. Cells should reach ~85% confluency after 48 h incubation with a final concentration of approximately 1 x 106 cells/well. Adjustments to the number of cells seeded and length of incubation may be required.
- Pipet gently to mix, then transfer 5.85 x 106 cells to a fresh 50 mL conical tube according to the equation below:
- For T75 flask:
- Making frozen HT-29 stocks
- Aliquot 1 mL of DMEM + 10% FBS + 5% DMSO media into individual cryogenic vials.
- Add 1 x 106 HT-29 cells from step 3.2.7 to each vial. Calculate the volume of cells according to the formula below:
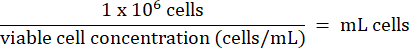
- Store HT-29 cells long-term below -130 °C in liquid nitrogen vapor storage freezer.
4. Adherence assay
NOTE: All volumes are consistent with an assay using two 6-well plates.
- Subculture overnight Shigella cultures via 1:50 dilution into fresh media.
- Vortex, then add 100 µL of each overnight culture to 5 mL of fresh TSB or TSB + BS in a suitably sized culture tube.
NOTE: Limit culture volume to <20% of culture flask or tube volume to ensure proper aeration. - Incubate at 37 °C with shaking at 250 rpm until cells reach an optical density (OD600) of 0.7 (mid-log phase of Shigella growth); about 2-2.5 h.
NOTE: During the subculture, aliquot 50 mL of DMEM and a sufficient volume of PBS for all washing steps and place in a 37 °C water bath. Allow media to reach 37 °C before use.
- Vortex, then add 100 µL of each overnight culture to 5 mL of fresh TSB or TSB + BS in a suitably sized culture tube.
- Transfer 2 x 108 colony forming units (CFUs) subcultured Shigella to individual 2 mL microcentrifuge tubes.
NOTE: 2 x 108 CFUs corresponds to approximately 1 mL of bacterial cells at an OD600 of 0.7. Use OD600 readings to approximate CFU/mL according to the calibration of each individual spectrophotometer. - Wash each Shigella sample 2x with PBS.
- Pellet cells by centrifugation at 17,000 x g for 2 min at room temperature. Aspirate the supernatant, then add 1 mL of warm PBS and resuspend the pellet well, gently pipetting the sample up and down until the mixture is fully homogeneous (8-10x).
- Repeat step 4.3.1 1x additional time.
- Pellet cells by centrifugation at 17,000 x g for 2 min at room temperature, aspirate the supernatant, and resuspend the pellets in 2 mL of warm DMEM.
NOTE: The final concentration of resuspended bacteria will be 1 x 108 CFU/mL.
- Vortex, then add 1 mL (1 x 108 CFUs) of resuspended Shigella to each well of the prepared HT-29 colonic epithelial monolayers in 6-well plates (from step 3.2.10.2).
NOTE: Infections are normally performed at a multiplicity of infection (MOI; ratio of bacterial to epithelial cells) of 100. To test different MOIs, dilute resuspended Shigella in warm DMEM to the desired concentration, then add 1 mL of diluted bacteria to HT-29 monolayers. For example, to test an MOI of 10, dilute bacteria 1:10 by adding 150 µL of 1 x 108 CFU/mL bacteria to 1.35 mL of warm DMEM, then apply 1 mL (1 x 107 CFUs) to HT-29 cells. - Incubate the 6-well plates at 37 °C with 5% CO2 for 3 h.
- During the incubation, determine the bacterial infection titer.
- Prepare 10-fold serial dilutions of resuspended Shigella cells (from step 4.3.3) into PBS.
- Plate 100 µL of the 1 x 10-5 and 1 x 10-6 dilutions onto TSB + Congo red plates and incubate overnight at 37 °C.
NOTE: Plating 100 µL from the 1 x 10-5 and 1 x 10-6 dilutions corresponds to a final dilution factor of 1 x 10-6 and 1 x 10-7, respectively.
- After incubation, wash the monolayers 4-5x with PBS.
- Aspirate media from each well.
NOTE: When aspirating media from 6-well plates, guide the tip of the aspirator along the bottom side of the wells, trying to avoid contact with the HT-29 cells. - Add 1 mL of warm PBS to each well and wash gently.
NOTE: To gently wash 6-well monolayers with PBS, move the plate up and down and side to side on the benchtop. Washing plates in a circular motion and/or removing the plate from the benchtop surface can cause the mechanical removal of cells from plastic. - Repeat steps 4.7.1 and 4.7.2 4x additional times.
- Aspirate media from each well.
- Remove PBS by aspiration and lyse HT-29 cells by adding 1 mL of PBS + 1% Triton X-100 to each well.
- Incubate 6-well plates at 37 °C for 5 min.
- Use a cell scraper or a bent pipette tip to scrape the lysed cells from the bottom of the well and transfer the full 1 mL into a fresh 1.7 mL microcentrifuge tube.
- Determine the number of cell-associated bacteria.
- Vortex each tube (from step 4.10) for at least 30 s to further displace Shigella from the lysed eukaryotic cells.
- Prepare 10-fold serial dilutions of lysates into PBS.
- Plate 100 µL of the 1 x 10-2, 1 x 10-3, and 1 x 10-4 dilutions onto TSB + Congo red plates and incubate overnight at 37 °C.
NOTE: Plating 100 µL from the 1 x 10-2, 1 x 10-3, and 1 x 10-4 dilutions corresponds to a final dilution factor of 1 x 10-3, 1 x 10-4, and 1 x 10-5, respectively.
5. Invasion assay
NOTE: All volumes are consistent with an assay using two 6-well plates.
- Subculture overnight Shigella cultures via 1:50 dilution into fresh media.
- Vortex, then add 100 µL of each overnight culture to 5 mL of fresh TSB or TSB + BS in a suitably sized culture tube.
NOTE: Limit culture volume to <20% of culture flask or tube volume to ensure proper aeration. - Incubate at 37 °C with shaking at 250 rpm until cells reach an OD600 of 0.7 (mid-log phase of Shigella growth); about 2-2.5 h.
NOTE: During the subculture, aliquot 50 mL of DMEM + 50 mg/mL gentamicin and a sufficient volume of PBS for all washing steps and place in a 37 °C water bath. Allow media to reach 37 °C before use.
- Vortex, then add 100 µL of each overnight culture to 5 mL of fresh TSB or TSB + BS in a suitably sized culture tube.
- Transfer 2 x 108 CFUs subcultured Shigella to individual 2 mL microcentrifuge tubes.
NOTE: 2 x 108 CFUs corresponds to approximately 1 mL of bacterial cells at an OD600 of 0.7. Use OD600 readings to approximate CFU/mL according to the calibration of each individual spectrophotometer. - Wash Shigella samples 1x with PBS.
- Pellet cells by centrifugation at 17,000 x g for 2 min at room temperature. Aspirate the supernatant, then add 1 mL of warm PBS and resuspend the pellet well, gently pipetting the sample up and down until the mixture is fully homogeneous (8-10x).
- Repeat step 5.3.1. 1x additional time.
- Pellet cells by centrifugation at 17,000 x g for 2 min at room temperature, aspirate the supernatant, and resuspend the pellets in 2 mL of warm DMEM.
NOTE: The final concentration of resuspended bacteria will be 1 x 108 CFU/mL.
- Vortex, then add 1 mL (1 x 108 CFUs) of resuspended Shigella plus 1 mL of DMEM to each well of the prepared HT-29 colonic epithelial monolayers in 6-well plates (from step 3.2.10.2).
NOTE: Infections are normally performed at a multiplicity of infection (MOI; ratio of bacterial to epithelial cells) of 100. To test different MOIs, dilute resuspended Shigella in DMEM to the desired concentration, then add 1 mL of diluted bacteria to HT-29 monolayers. For example, to test an MOI of 10, dilute bacteria 1:10 by adding 150 µL of 1 x 108 CFU/mL bacteria to 1.35 mL of DMEM, then add 1 mL (1 x 107 CFUs) to HT-29 cells. - To promote bacterial contact with the HT-29 cells, centrifuge the 6-well plates at 2,000 x g for 10 min at room temperature or 37 °C if the temperature setting can be adjusted.
NOTE: Centrifugation promotes bacterial contact with the HT-29 cells, which bypasses the need for adherence factors and allows the bacteria to quickly invade the cells. - Incubate 6-well plates at 37 °C with 5% CO2 for 45 min.
- During the incubation, determine the bacterial infection titer.
- Prepare 10-fold serial dilutions of resuspended Shigella cells (from step 5.3.3) into PBS.
- Plate 100 µL of the 1 x 10-5 and 1 x 10-6 dilutions onto TSB + Congo red plates and incubate overnight at 37 °C.
NOTE: Plating 100 µL from the 1 x 10-5 and 1 x 10-6 dilutions corresponds to a final dilution factor of 1 x 10-6 and 1 x 10-7, respectively.
- Thoroughly wash infected HT-29 cells 3x with 1 mL of PBS.
- Aspirate media from each well.
NOTE: When aspirating media from 6-well plates, guide the tip of the aspirator along the bottom side of the wells, trying to avoid contact with the HT-29 cells. - Add 1 mL of warm PBS to each well and wash gently.
NOTE: To gently wash 6-well monolayers with PBS, move the plate up and down and side to side on the benchtop. Washing plates in a circular motion and/or removing the plate from the benchtop surface can cause the mechanical removal of cells from plastic. - Repeat steps 5.8.1 and 5.8.2 2x additional times.
- Aspirate media from each well.
- Remove PBS by aspiration, then add 2 mL of warm DMEM supplemented with 50 µg/mL gentamicin to each well and incubate for 30 min at 37 °C with 5% CO2.
- Thoroughly wash infected HT-29 cells 3x with 1 mL of PBS.
- Repeat washing step 5.8.
- Remove PBS by aspiration, then add 2 mL of warm DMEM supplemented with 50 µg/mL gentamicin to each well and incubate for 60 min at 37 °C with 5% CO2.
- Thoroughly wash infected HT-29 cells 3x with 1 mL of PBS.
- Repeat washing step 5.8.
- Remove PBS by aspiration and lyse HT-29 cells by adding 1 mL of PBS + 1% Triton X-100 to each well.
- Incubate 6-well plates at 37 °C for 5 min.
- Use a cell scraper or a bent pipette tip to scrape the lysed cells from the bottom of the well and transfer the full 1 mL into a fresh 1.7 mL microcentrifuge tube.
- Determine the number of intracellular bacteria.
- Vortex each tube (from step 5.15) for at least 30 s to further displace Shigella from the lysed eukaryotic cells.
- Prepare 10-fold serial dilutions of lysates into PBS.
- Plate 100 µL of the 1 x 10-2 and 1 x 10-3 dilutions onto TSB + Congo red plates and incubate overnight at 37 °C.
NOTE: Plating 100 µL from the 1 x 10-2 and 1 x 10-3 dilutions corresponds to a final dilution factor of 1 x 10-3 and 1 x 10-4, respectively.
6. Intracellular replication assay
NOTE: All volumes are consistent with an assay using two 6-well plates.
- Subculture overnight Shigella cultures via 1:50 dilution into fresh media.
- Vortex, then add 100 µL of each overnight culture to 5 mL of fresh TSB or TSB + BS in a suitably sized culture tube.
NOTE: Limit culture volume to <20% of culture flask or tube volume to ensure proper aeration. - Incubate at 37 °C with shaking at 250 rpm until cells reach an OD600 of 0.7 (mid-log phase of Shigella growth); about 2-2.5 h.
NOTE: During the subculture, aliquot 50 mL of DMEM + 50 mg/mL gentamicin and a sufficient volume of PBS for all washing steps and place in a 37 °C water bath. Allow media to reach 37 °C before use.
- Vortex, then add 100 µL of each overnight culture to 5 mL of fresh TSB or TSB + BS in a suitably sized culture tube.
- Transfer 2 x 108 CFUs subcultured Shigella to individual 2 mL microcentrifuge tubes.
NOTE: 2 x 108 CFUs corresponds to approximately 1 mL of bacterial cells at an OD600 of 0.7. Use OD600 readings to approximate CFU/mL according to the calibration of each individual spectrophotometer. - Wash Shigella samples 1x with PBS.
- Pellet cells by centrifugation at 17,000 x g for 2 min at room temperature. Aspirate the supernatant, then add 1 mL of warm PBS and resuspend the pellet well, gently pipetting the sample up and down until the mixture is fully homogeneous (8-10x).
- Repeat step 6.3.1. 1x additional time.
- Pellet cells by centrifugation at 17,000 x g for 2 min at room temperature, aspirate the supernatant, and resuspend the pellets in 2 mL of warm DMEM.
NOTE: The final concentration of resuspended bacteria will be 1 x 108 CFU/mL.
- Vortex, then add 1 mL (1 x 108 CFUs) of resuspended Shigella plus 1 mL of DMEM to each well of prepared HT-29 colonic epithelial monolayers in 6-well plates (from step 3.2.10.2).
NOTE: Infections are normally performed at a multiplicity of infection (MOI; ratio of bacterial to epithelial cells) of 100. To test different MOIs, dilute resuspended Shigella in DMEM to the desired concentration, then add 1 mL of diluted bacteria to HT-29 monolayers. For example, to test an MOI of 10, dilute bacteria 1:10 by adding 150 µL of 1 x 108 CFU/mL bacteria to 1.35 mL of DMEM, then apply 1 mL (1 x 107 CFUs) to HT-29 cells. - To promote bacterial contact with the HT-29 cells, centrifuge the 6-well plates at 2,000 x g for 10 min at room temperature or 37 °C if the temperature setting can be adjusted.
NOTE: Centrifugation promotes bacterial contact with the HT-29 cells, which bypasses the need for adherence factors and allows the bacteria to quickly invade the cells. - Incubate 6-well plates at 37 °C with 5% CO2 for 45 min.
- During the incubation, determine the bacterial infection titer.
- Prepare 10-fold serial dilutions of resuspended Shigella cells (from step 6.3.3) into PBS.
- Plate 100 µL of the 1 x 10-5 and 1 x 10-6 dilutions onto TSB + Congo red plates and incubate overnight at 37 °C.
NOTE: Plating 100 µL from the 1 x 10-5 and 1 x 10-6 dilutions corresponds to a final dilution factor of 1 x 10-6 and 1 x 10-7, respectively.
- Thoroughly wash infected HT-29 cells 3x with 1 mL of PBS.
- Aspirate media from each well.
NOTE: When aspirating media from 6-well plates, guide the tip of the aspirator along the bottom side of the wells, trying to avoid contact with the HT-29 cells. - Add 1 mL of warm PBS to each well and wash gently.
NOTE: To gently wash 6-well monolayers with PBS, move the plate up and down and side to side on the benchtop. Washing plates in a circular motion and/or removing the plate from the benchtop surface can cause the mechanical removal of cells from plastic. - Repeat steps 6.8.1 and 6.8.2 2x additional times.
- Aspirate media from each well.
- Remove PBS by aspiration, then add 2 mL of warm DMEM supplemented with 50 µg/mL gentamicin to each well and incubate for 30 min at 37 °C with 5% CO2.
- Thoroughly wash infected HT-29 cells 3x with 1 mL of PBS.
- Repeat washing step 6.8.
- Remove PBS by aspiration, then add 2 mL of warm DMEM with 50 µg/mL gentamicin to each well of the 6-well plates and incubate at 37 °C with 5% CO2 for the desired length of time to allow for intracellular replication (up to 24 h).
- Thoroughly wash cells 2x with 1 mL of PBS.
- Repeat washing step 6.8.
- Remove PBS by aspiration and lyse HT-29 cells by adding 1 mL of PBS + 1% Triton X-100 to each well.
- Incubate 6-well plates at 37 °C for 5 min.
- Use a cell scraper or bent pipette tip to scrape the lysed cells from the bottom of the well and transfer the full 1 mL into a fresh 1.7 mL microcentrifuge tube.
- Determine the number of intracellular bacteria.
- Vortex each tube (from step 6.15) for at least 30 s to further displace Shigella from the lysed eukaryotic cells.
- Prepare 10-fold serial dilutions of lysates into PBS.
- Plate 100 µL of the 1 x 10-2, 1 x 10-3, and 1 x 10-4 dilutions onto TSB + Congo Red plates and incubate overnight at 37 °C.
NOTE: Plating 100 µL from the 1 x 10-2, 1 x 10-3, and 1 x 10-4 dilutions corresponds to a final dilution factor of 1 x 10-3, 1 x 10-4, and 1 x 10-5, respectively.
Epithelial Cell Infection Analyses with Shigella
Learning Objectives
Adherence, invasion, and intracellular replication assays were performed comparing S. flexneri 2457T wild type (WT) to S. flexneri ΔVF (ΔVF), a mutant hypothesized to negatively regulate Shigella virulence. Since Shigella uses bile salts as a signal to regulate virulence17,18,47, experiments were performed after bacterial subculture in TSB media as well as TSB supplemented with 0.4% (w/v) bile salts18. Bile salts exposure during the subculture step acts as a pre-treatment to replicate small intestinal transit prior to colonic infection17,18,47. Figure 1 analyzes the effect of the ΔVF mutation on the ability of S. flexneri to adhere to HT-29 colonic epithelial cells. Percent adherence is plotted on the y-axis and represents the ratio of recovered bacteria following HT-29 lysis standardized to the number of input bacteria. As expected, both S. flexneri WT and ΔVF strains had a significant increase in adherence when subcultured with bile salts supplementation in comparison to TSB without bile salts supplementation18. However, there was no difference in adherence to HT-29 cells between WT and ΔVF mutant strains within each subculture condition. These data indicate that the ΔVF mutation has no effect on the ability of S. flexneri to adhere to HT-29 epithelial cells with or without the bile salts pre-treatment.
In Figure 2, the effect of the ΔVF mutation on the ability of S. flexneri to invade (Figure 2A) and replicate (Figure 2B) inside of HT-29 colonic epithelial cells with or without bile salts pre-treatment was analyzed. Percent recovery is plotted on the y-axis and represents the ratio of recovered bacterial cells following HT-29 cell lysis standardized to the number of input bacteria. In Figure 2A, there was an expected significant increase in the ability of WT S. flexneri 2457T to invade HT-29 cells after pre-exposure to bile salts48, while the S. flexneri ΔVF mutant displayed a smaller increase in invasion following bile salts pre-exposure compared to the WT strain. The ΔVF mutant had increased invasion rates of HT-29 cells compared to the WT subcultured in TSB, but had similar invasion rates as WT when subcultured in TSB supplemented with bile salts (Figure 2A). These results suggest that the ΔVF mutation enhances the ability of S. flexneri to invade HT-29 cells, which lessens the effect of the bile salts pre-exposure even though the invasion ability of the ΔVF mutant did increase further following bile salts subculture.
Overall, 10-fold more bacteria were recovered following overnight incubation (Figure 2B) compared to the 90 min incubation (Figure 2A), which demonstrates the differences in monitoring intracellular growth versus invasion, respectively. When infected HT-29 cells were incubated for 18 h to allow for intracellular replication of the bacteria (Figure 2B), the impact of the bile salts pre-treatment decreased for both the WT and ΔVF strains. However, the reduced effect of bile salts pre-treatment during intracellular replication was more dramatic for the ΔVF mutant. Since the increase in intracellular replication of both strains when pre-exposed to bile salts was smaller than the increase in invasion rates in the same conditions, we hypothesize that bile salts have a greater impact on the early steps in S. flexneri pathogenesis. The ΔVF mutant displayed an increase in percent recovery from overnight replication inside HT-29 cells compared to WT (Figure 2B) following both subculture conditions. However, the percent recoveries of the ΔVF mutant were similar regardless of bile salts pre-exposure. These data trends suggest that the ΔVF mutant replicates more efficiently inside HT-29 cells compared to WT, and that bile salts pre-exposure does not impact the ability of the ΔVF mutant to replicate intracellularly, as observed for WT (Figure 2B). Since the difference between the mutant and WT strains in the bile salts pre-exposure condition was not observed during the 90 min invasion assay, we hypothesize that the product encoded by the deleted VF gene may also regulate S. flexneri replication inside HT-29 cells. Combined, both analyses demonstrate that the ΔVF mutant is more virulent relative to WT, which suggests that the VF gene product is a negative regulator of S. flexneri virulence.
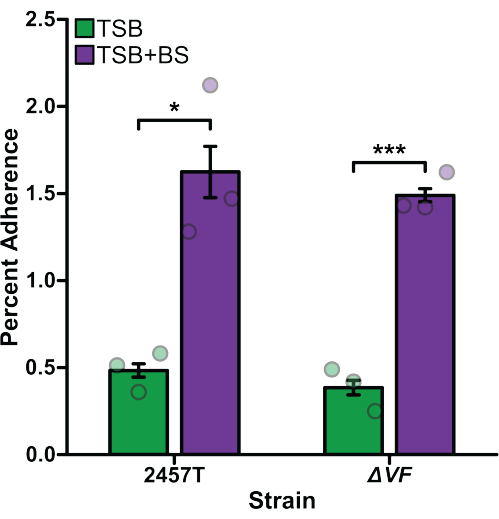
Figure 1: Bile salts pre-exposure induced adherence of S. flexneri to HT-29 cells. S. flexneri 2457T WT and ΔVF mutant cells were subcultured in either TSB or TSB supplemented with 0.4% (w/v) bile salts (TSB+BS) media. The bacteria were then applied to HT-29 cells at a multiplicity of infection (MOI) of 100 and incubated for 3 h to examine adherence. After incubation, infected HT-29 cells were washed and lysed, and serial dilutions of recovered bacteria were plated to enumerate colony-forming units per mL (CFU/mL). The number of adherent bacteria is plotted relative to the input bacteria titers to establish the percent adherence. Data are representative of one biological replicate with three technical replicates (individual dots). Error bars indicate the standard error of the mean (SEM). Statistical significance was determined by a Student's t-test (*p < 0.05; ***p < 0.001). Please click here to view a larger version of this figure.
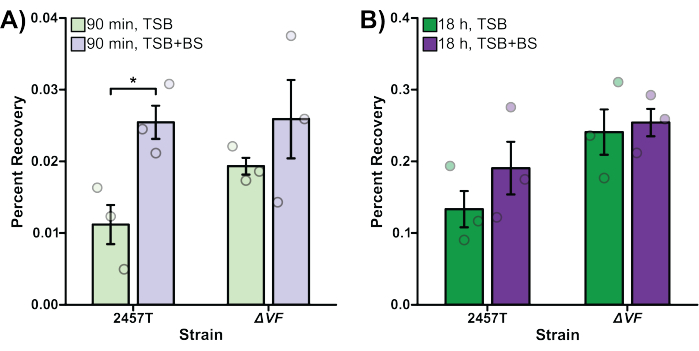
Figure 2: Bile salts pre-exposure increased WT S. flexneri invasion and intracellular replication. S. flexneri 2457T WT and ΔVF mutant cells were subcultured in either TSB or TSB supplemented with bile salts (TSB+BS) media. The bacteria were then applied to HT-29 cells at an MOI of 100, centrifuged onto the cells, and incubated at 37 °C with 5% CO2 for 45 min. Cells were washed with PBS, and extracellular bacteria were lysed by the addition of gentamicin to the DMEM to exclusively recover intracellular bacteria. After 90 min (A, bacterial invasion) or 18 h (B, intracellular replication) incubations, infected HT-29 cells were washed and lysed, and serial dilutions of recovered bacteria were plated to enumerate colony-forming units per mL (CFU/mL). The number of intracellular bacteria is plotted relative to the input bacterial titers to establish the percent recovery. Data are representative of one biological replicate, each with three technical replicates (individual dots). Error bars indicate SEM. Statistical significance was determined by a Student's t-test (*p < 0.05). Please note the differences in the y-axis scales between panels (A) and (B). Please click here to view a larger version of this figure.
List of Materials
| 0.22 μm PES filter | Millipore-Sigma | SCGP00525 | Sterile, polyethersulfone filter for sterilizing up to 50 mL media |
| 14 mL culture tubes | Corning | 352059 | 17 mm x 100 mm polypropylene test tubes with cap |
| 50 mL conical tubes | Corning | 430829 | 50 mL clear polypropylene conical bottom centrifuge tubes with leak-proof cap |
| 6-well tissue culture plates | Corning | 3516 | Plates are treated for optimal cell attachment |
| Bile salts | Sigma-Aldrich | B8756 | 1:1 ratio of cholate to deoxycholate |
| Congo red dye | Sigma-Aldrich | C6277 | A benzidine-based anionic diazo dye, >85% purity |
| Countess cell counting chamber slide | Invitrogen | C10283 | To be used with the Countess Automated Cell Counter |
| Dimethyl sulfoxide (DMSO) | Sigma-Aldrich | D8418 | A a highly polar organic reagent |
| Dulbecco’s Modified Eagle Medium (DMEM) | Gibco | 10569-010 | DMEM is supplemented with high glucose, sodium pyruvate, GlutaMAX, and Phenol Red |
| Fetal Bovine Serum (FBS) | Sigma-Aldrich | F4135 | Heat-inactivated, sterile |
| Gentamicin | Sigma-Aldrich | G3632 | Stock concentration is 50 mg/mL |
| HT-29 cell line | ATCC | HTB-38 | Adenocarcinoma cell line; colorectal in origin |
| Paraffin film | Bemis | PM999 | Laboratory sealing film |
| Petri dishes | Thermo Fisher Scientific | FB0875713 | 100 mm x 15 mm Petri dishes for solid media |
| Phosphate-buffered saline (PBS) | Thermo Fisher Scientific | 10010049 | 1x concentration; pH 7.4 |
| Select agar | Invitrogen | 30391023 | A mixture of polysaccharides extracted from red seaweed cell walls to make bacterial plating media |
| T75 flasks | Corning | 430641U | Tissue culture flasks |
| Triton X-100 | Sigma-Aldrich | T8787 | A common non-ionic surfactant and emulsifier |
| Trypan blue stain | Invitrogen | T10282 | A dye to detect dead tissue culture cells; only live cells can exclude the dye |
| Trypsin-EDTA | Gibco | 25200-056 | Reagent for cell dissociation for cell line maintenance and passaging |
| Tryptic Soy Broth (TSB) | Sigma-Aldrich | T8907 | Bacterial growth media |
Lab Prep
The human-adapted enteric bacterial pathogen Shigella causes millions of infections each year, creates long-term growth effects among pediatric patients, and is a leading cause of diarrheal deaths worldwide. Infection induces watery or bloody diarrhea as a result of the pathogen transiting the gastrointestinal tract and infecting the epithelial cells lining the colon. With staggering increases in antibiotic resistance and the current lack of approved vaccines, standardized research protocols are critical to studying this formidable pathogen. Here, methodologies are presented to examine the molecular pathogenesis of Shigella using in vitro analyses of bacterial adherence, invasion, and intracellular replication in colonic epithelial cells. Prior to infection analyses, the virulence phenotype of Shigella colonies was verified by the uptake of the Congo red dye on agar plates. Supplemented laboratory media can also be considered during bacterial culturing to mimic in vivo conditions. Bacterial cells are then used in a standardized protocol to infect colonic epithelial cells in tissue culture plates at an established multiplicity of infection with adaptations to analyze each stage of infection. For adherence assays, Shigella cells are incubated with reduced media levels to promote bacterial contact with epithelial cells. For both invasion and intracellular replication assays, gentamicin is applied for various time intervals to eliminate extracellular bacteria and enable assessment of invasion and/or the quantification of intracellular replication rates. All infection protocols enumerate adherent, invaded, and/or intracellular bacteria by serially diluting infected epithelial cell lysates and plating bacterial colony forming units relative to infecting titers on Congo red agar plates. Together, these protocols enable independent characterization and comparisons for each stage of Shigella infection of epithelial cells to study this pathogen successfully.
The human-adapted enteric bacterial pathogen Shigella causes millions of infections each year, creates long-term growth effects among pediatric patients, and is a leading cause of diarrheal deaths worldwide. Infection induces watery or bloody diarrhea as a result of the pathogen transiting the gastrointestinal tract and infecting the epithelial cells lining the colon. With staggering increases in antibiotic resistance and the current lack of approved vaccines, standardized research protocols are critical to studying this formidable pathogen. Here, methodologies are presented to examine the molecular pathogenesis of Shigella using in vitro analyses of bacterial adherence, invasion, and intracellular replication in colonic epithelial cells. Prior to infection analyses, the virulence phenotype of Shigella colonies was verified by the uptake of the Congo red dye on agar plates. Supplemented laboratory media can also be considered during bacterial culturing to mimic in vivo conditions. Bacterial cells are then used in a standardized protocol to infect colonic epithelial cells in tissue culture plates at an established multiplicity of infection with adaptations to analyze each stage of infection. For adherence assays, Shigella cells are incubated with reduced media levels to promote bacterial contact with epithelial cells. For both invasion and intracellular replication assays, gentamicin is applied for various time intervals to eliminate extracellular bacteria and enable assessment of invasion and/or the quantification of intracellular replication rates. All infection protocols enumerate adherent, invaded, and/or intracellular bacteria by serially diluting infected epithelial cell lysates and plating bacterial colony forming units relative to infecting titers on Congo red agar plates. Together, these protocols enable independent characterization and comparisons for each stage of Shigella infection of epithelial cells to study this pathogen successfully.
Procédure
The human-adapted enteric bacterial pathogen Shigella causes millions of infections each year, creates long-term growth effects among pediatric patients, and is a leading cause of diarrheal deaths worldwide. Infection induces watery or bloody diarrhea as a result of the pathogen transiting the gastrointestinal tract and infecting the epithelial cells lining the colon. With staggering increases in antibiotic resistance and the current lack of approved vaccines, standardized research protocols are critical to studying this formidable pathogen. Here, methodologies are presented to examine the molecular pathogenesis of Shigella using in vitro analyses of bacterial adherence, invasion, and intracellular replication in colonic epithelial cells. Prior to infection analyses, the virulence phenotype of Shigella colonies was verified by the uptake of the Congo red dye on agar plates. Supplemented laboratory media can also be considered during bacterial culturing to mimic in vivo conditions. Bacterial cells are then used in a standardized protocol to infect colonic epithelial cells in tissue culture plates at an established multiplicity of infection with adaptations to analyze each stage of infection. For adherence assays, Shigella cells are incubated with reduced media levels to promote bacterial contact with epithelial cells. For both invasion and intracellular replication assays, gentamicin is applied for various time intervals to eliminate extracellular bacteria and enable assessment of invasion and/or the quantification of intracellular replication rates. All infection protocols enumerate adherent, invaded, and/or intracellular bacteria by serially diluting infected epithelial cell lysates and plating bacterial colony forming units relative to infecting titers on Congo red agar plates. Together, these protocols enable independent characterization and comparisons for each stage of Shigella infection of epithelial cells to study this pathogen successfully.
