A Modified Precipitation Method to Isolate Urinary Exosomes
Summary
This manuscript describes a protocol for isolating exosomes from human urine using a modified precipitation technique.
Abstract
Identification of biomarkers that allow early detection of kidney diseases in urine and plasma has been an area of active interest for several years. Urinary exosome vesicles, 40-100 nm in size, are released into the urine under normal conditions by cells from all nephron segments and may contain protein, mRNA and microRNA representative of their cell type of origin. Under conditions of renal dysfunction or injury, exosomes may contain altered proportions of these components, which may serve as biomarkers for disease. There are currently several methods available for isolation of urinary exosomes, and we have previously conducted an experimental comparison of each of these approaches, including three based on ultracentrifugation, one using a nanomembrane ultrafiltration concentrator, one using a commercial precipitation reagent and one using a modification of the precipitation technique using ExoQuick reagent that we developed in our laboratory. We found the modified precipitation method produced the highest yield of exosome particles, miRNA, and mRNA, making this approach suitable for the isolation of exosomes for subsequent RNA profiling. We conclude that the modified exosome precipitation method offers a quick, scalable, and effective alternative for the isolation of exosomes from urine. In this report, we describe our modified precipitation technique using ExoQuick reagent for isolating exosomes from human urine.
Introduction
Urinary exosomes are small internal vesicles of 100 nm of multivesicular bodies (MVB) derived from all cell types under normal and pathological conditions in urinary space including glomerular podocytes, renal tubule cells and cells lining the urinary drainage systems that appear to be enriched for miRNAs1-2. Many researchers have detected distinct proteins in exosomes isolated from urine of healthy individuals as well as those with renal and/or systematic diseases3-5. Exosomes can be isolated using different methods such as two-step differential ultracentrifugation with or without sucrose6-7, combining nanomembrane filter with ultracentrifugation8-9, or precipitation10-11. We have recently developed a simple, fast, scalable and effective method to isolate urinary exosomes and detect miRNA and protein biomarkers11. Although biomarkers can be detected in a variety of different biological fluids, urine is the most convenient choice for patients with diseases of the kidney and urinary tract because it can be obtained in large amounts in a relatively non-invasive manner.
Although there are several methods to isolate urinary exosomes6-11, the most commonly used procedure depends on differential ultracentrifugation with a 30% sucrose cushion. While this method is efficient and the quality of the resulting exosomes isolated with this procedure is high, the technique itself requires skilled personnel and is time-consuming, requiring several ultracentrifugation steps. Other methods, such as filtration and precipitation, have been developed for the isolation of exosomes, including one that uses a proprietary precipitation reagent (i.e., ExoQuick-TC: System Biosciences; Mountain View, CA). Although precipitation using this reagent is fast and relatively easy, the purity of the exosomes isolated is low compared to the ultracentrifugation method. We recently compared six methods for the isolation of urinary exosomes, including a modification of the precipitation method using ExoQuick-TC that we developed in our laboratory11. We found that this modified precipitation method yielded higher quantities of protein, miRNA, and mRNAs and the procedure itself was faster and easier to implement than ultracentrifugation. Here, we describe the experimental protocol of our modified precipitation method in a stepwise fashion, and include a discussion of the advantages and disadvantages of this technique compared with other methods of exosome isolation. The modified precipitation method involves an initial precipitation of exosome particles, followed by removal of Tamm-Horsfall protein, which is correlated with exosomal proteins, and known to reduce exosome yield from urine. We found that these modifications increased the yield and purity of the exosomal preparation from urine.
Protocol
NOTE: The steps involved in the isolation of the urinary exosomes using the modified precipitation method are illustrated in the Figure 1. The initial steps involve removal of large dead cells and cell debris by centrifugation. The final pellet obtained from the supernatant collected from each subsequent step corresponds to the exosome, which is dissolved in isolation solution for further extraction of protein, RNA, and DNA.
1. Collection of Urine
- Prior to urine collection, prepare protease inhibitor cocktail according to the manufacturer’s instructions and add 3.125 ml of this cocktail to an empty sterile receptacle. The tube containing the protease inhibitor cocktail can then be distributed to volunteers for urine collection.
NOTE: The reason for adding protease inhibitor is to minimize protein degradation in the urine sample. Although we specifically used a protease inhibitor cocktail from Sigma, other commercially available protease inhibitors can be substituted. - Collect approximately 10 ml of first-void urine. Process the urine immediately after collection without storage or refrigeration. Isolate the urinary exosomes in duplicate using the modified precipitation method; one of the duplicates is for DNA, total RNA and protein extraction, while the other is for the quantification of urinary exosome particles.
NOTE: A 24 hr urine sample can be collected for exosome isolation by storing at 4 °C.
2. Removal of Cell Debris and Tamm-Horsfall Protein (THP)
- Transfer urine samples to 15 ml centrifuge tubes. Centrifuge 10 ml of urine at 17,000 x g for 10 min at 37 °C to eliminate cells and cell debris (Figure 1).
- Using a pipette, transfer the supernatant to a fresh tube for further processing. Resuspend the pellet in 500 µl of isolation solution (250 mM sucrose, 10 mM triethanolamine, pH 7.6) followed by 10 min incubation with DL-dithiothreitol (DTT: final concentration 200 mg/ml) at 37 °C.
NOTE: DTT is added to degrade THP, which traps exosomes, leading to decreased yields. Take care to leave the pellet undisturbed during transfer of the supernatant, and confirm that the supernatant is clear of pellet. - Vortex pellet samples every 2 min until the pellet is completely dissolved. Then add 500 µl of isolation solution to the dissolved pellet.
- Immediately following the addition of isolation solution to the pellet, centrifuge at 17,000 x g for 10 min at 37 °C. Pipet the supernatant and combine with the supernatant collected earlier.
- Resuspend the pellet in 50 µl of isolation solution for SDS-PAGE analysis.
NOTE: PBS or TBS can be used instead of isolation solution to resuspend the pellet.
3. Isolation of Exosome Pellet
- To 11 ml of the collected supernatant, add 3.3 ml of ExoQuick-TC reagent and incubate at 4 °C for at least 12 hr.
NOTE: The ratio of volume of supernatant to ExoQuick-TC reagent can be adjusted to increase or decrease the exosomal yield. We observed that lesser volumes of added reagent produce smaller yields of exosomes. - Following incubation, centrifuge the sample/ExoQuick-TC mix at 10,000 x g for 30 min at 25 °C.
- Transfer the supernatant to a fresh tube for SDS-PAGE analysis.
- The exosome pellet can be stored at -80 °C for DNA, RNA and protein extraction and quantification of exosome particles using CD9 ELISA11.
NOTE: The exosome pellet can be stored at -80 °C for 1 year with limited freezing and thawing.
4. Biomarker Detection and Analysis of Exosome Pellet
- For downstream analyses, extract DNA, RNA and protein from the exosome pellet using the DNA/RNA/Protein isolation kit and an RNA isolation kit according to the manufacturer’s instructions. Determine concentrations using a Nanodrop spectrophotometer by measuring absorbance at 260 and 280nm with 2 µl of sample.
- Perform quantitative real time RT-PCR (qPCR) using TaqMan microRNA assays (in our case, we used human miR-192- and miR-1207-5p-specific assays) or miRNA-specific primers.
- For SDS-PAGE analysis, use 4-12% Bis-Tris gel and separate protein samples by one-dimensional electrophoresis. Visualize gels using the silver staining kit according to the manufacturer’s instructions.
- Perform western blot analysis using PVDF membranes according to the instructions provided by the manufacturer. Use rabbit polyclonal antibody to apoptosis-linked gene-2 interacting protein X or ALIX and mouse monoclonal antibody to tumor susceptibility gene 101 protein or TSG101 in the western blot detection. Quantify the density of the bands by open access NIH ImageJ software (http://rsbweb.nih.gov/ij/) or a similar program.
- Quantify the number of urinary exosome particles obtained using a CD9 ELISA kit as per manufacturer’s instructions.
Representative Results
Urinary exosomes carry protein, miRNA and mRNA biomarkers secreted from renal epithelial cells. The isolation and detection of these exosomes may provide a useful diagnostic tool for early detection of kidney diseases such as diabetic nephropathy. There are several methods for isolation of exosomes from urine samples collected from human subjects. We previously compared six methods of urine exosome isolation and investigated resulting protein, mRNA, and miRNA levels11. Our aim here was to describe in detail the modified precipitation method we have developed for efficient isolation of urinary exosomes.
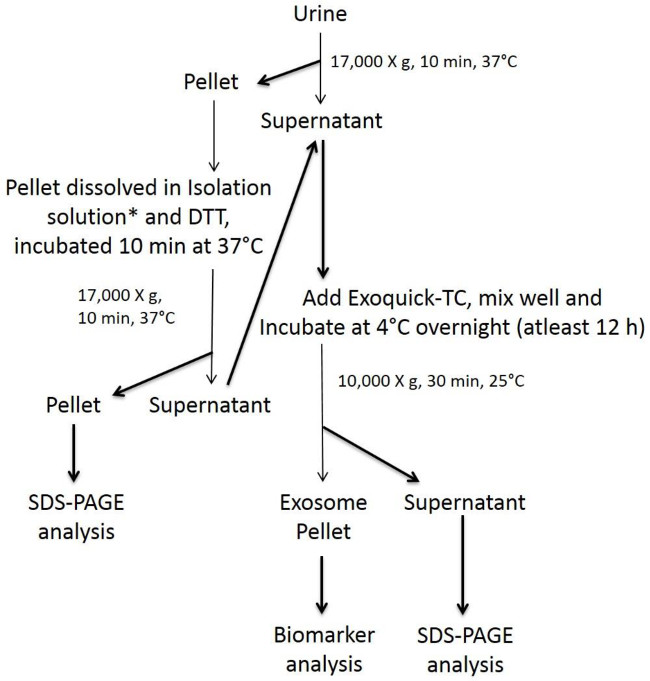
Figure 1 describes the schematic representative flow chart of various steps involved in the isolation of urinary exosomes using the modified precipitation method. One of the duplicate exosome pellet obtained is measured for CD9 using the CD9 ELISA kit, which provides us with the relative quantification of exosome particles. The other pellet processed using Qiagen DNA/RNA/Protein mini kit yields DNA, RNA and protein, which were quantified using Nanodrop spectrophotometer by measuring the absorbance at 260 and 280 nm.
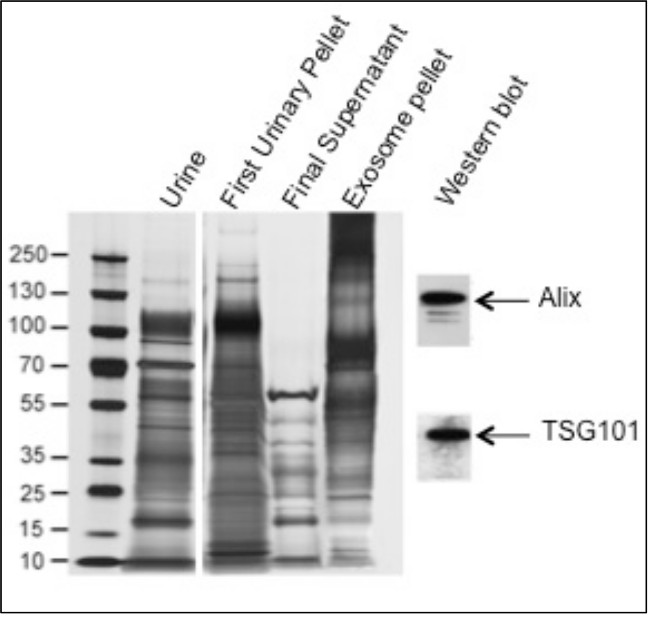
Figure 2 is the modified reproduction of the results on SDS-PAGE analysis and biomarker detection using western blot analysis (adapted from reference 11). In lanes with unprocessed urine and first urinary pellet, a protein band corresponding to 90 to 100 kDa was observed indicating the presence and removal of THP protein respectively. While in lanes with final supernatant and exosome pellet, the protein band corresponding to THP was not observed, indicating the removal of THP in the earlier steps, thereby increasing the yield of exosome pellet. To access the purity and sensitivity of the exosome pellets, western blot detection of biomarkers such as Alix and TSG101 was employed. We detected Alix and TSG101 even when low quantities of sample were used indicating significantly pure exosome pellet for biomarker analysis and absence of abundant soluble proteins including THP. Densitometric analysis of the western blots was done to quantify and measure the level of detection (data not shown).
Discussion
Most methods involving precipitation of exosomes from urine do not address removal of the polymeric network formed by Tamm-Horsfall protein (THP). This protein is present in large amounts in urine, where it putatively traps exosomes during initial low speed centrifugation. In our modified method we reduced THP using DTT prior to exosome precipitation. As shown in Figure 2, a higher amount of THP was observed in the unprocessed urine and pellet, with no THP in the final supernatant and exosome pellet, indicating removal of the protein. Quantification of protein in the pellet indicated six times higher yield, and on CD9 ELISA quantification of the pellet, it was observed that yield of exosome per ml of urine was five times higher than the unmodified precipitation method (results not shown11). The biomarkers Alix6 and TSG1018 were detected in the exosome pellet; we could only detect these proteins using concentrated samples in the unmodified precipitation method. We also observed the highest levels of miRNA (i.e., miR-192 and miR-1207-5p) and mRNA expression (i.e., TSG101, PDCD6IP and AQP212) using the modified precipitation method11. The advantages and disadvantages of this modified precipitation method are shown in Table 1.
| Advantages | Disadvantages |
| Easy simple steps and requires relative little expertise to perform | Low purity of the proteins obtained from Urinary exosomes |
| Does not require an ultracentrifugation step | Due to absence of the Ultracentrifugation steps, the Urinary exosome pellet obtained is not as pure as compared to the exosome pellet obtained from Ultracentrifugation methods |
| Uses small volume of Urine sample for isolation of exosomes | To obtain pure protein for specific analysis, protein purification steps such as affinity chromatography would be needed |
| Obtained good amount of miRNA, mRNA and proteins from Urinary exosomes compared to non-modified precipitation method | Few extra steps needed with addition of DTT to reduce entrapment by Tamm-Horsfall protein compared to non-modified precipitation method |
| Specificity and sensitivity of the biomarkers analyzed were similar to that obtained and analyzed using Ultracentrifugation method | |
| Urinary exosomes can be isolated from large number of samples at a time, suitable for clinical trails | |
| Less hands-on time compared to Ultracentrifugation methods |
Table 1. Advantages and disadvantages of modified precipitation method for urinary exosome isolation.
We conclude that our modification of the precipitation method is a simple, fast, efficient way to isolate urinary exosomes and is a suitable alternative to the ultracentrifugation method, which is labor-intensive and requires expensive equipment. Our method requires no ultracentrifugation step, is easy to perform, and requires less steps, while still providing a high yield or pure exosomes for subsequent molecular analyses. We also show that the yields of miRNA and mRNA using this method are high, and hence can be directly used in subsequent downstream RNA profiling applications. However, we note that a limitation of this method is the quality of the protein obtained, which is not as pure as that obtained using the sucrose gradient ultracentrifugation methods, and requires additional purification steps prior to utilization in downstream proteomic analyses. We are currently assessing the utility of this method using urine samples obtained from patients with diabetic nephropathy.
Divulgations
The authors have nothing to disclose.
Acknowledgements
We are grateful to the Roney Family Foundation for the support of this study. We acknowledge the contribution of Dr. M. Lucrecia Alvarez to the planning and execution of the experiments described in this work.
Materials
| Name of Material/ Equipment | Company | Catalog Number | Comments/Description |
| Corning 15ml Centrifuge Tube | Corning (Sigma Aldrich) | CLS430791 | |
| Protease Inhibitor Cocktail | Sigma Aldrich | P8340 | |
| Centrifuge | Sorvall | ||
| DL-dithiothreitol | Sigma Aldrich | D9779 | used at final concentration of 200 mg/ml |
| Novex NuPAGE SDS-PAGE system (4-12 % Bis-Tris Gel 1.0 mm, 10 well | Invitrogen | NP0321BOX | For SDS-PAGE analysis |
| ECL Advance Western blotting detection kit | GE Healthcare Life Sciences | RPN2135 | For western blot detection of biomarkers |
| Exoquick-TC Reagent | System Biosciences (SBI) | EXOTC50A-1 | For exosome isolation |
| Exosome CD9 ELISA Complete Kit | System Biosciences (SBI) | EXOEL-CD9-A-1 | For exosome quantification |
| AllPrep DNA/RNA/Protein mini kit | Qiagen | 80004 | For DNA, RNA, Protein isolation |
| Rneasy MinElute Cleanup kit | Qiagen | 74204 | |
| NanoDrop 2000 UV-Vis Spectrophotometer | Thermo Scientific | For Protein Quantification | |
| ProteoSilver Sliver Stain Kit | Sigma Aldrich | PROTSIL1-1KT | For staining SDS-PAGE gels |
| Xcell SureLock Mini-Cell Electrophoresis System | Life Technologies | For running the SDS-PAGE gels | |
| TaqMan microRNA assays | Life Technologies | For RT-PCR assays | |
| qBasePlus v1.5 software | Biogazelle NV | For analysis of RT-PCR assay data | |
| Rabbit polyclonal antibody to ALIX | Millipore | For western blot detection of biomarkers | |
| Mouse monoclonal antibody to TSG101 | Abcam | For western blot detection of biomarkers |
References
- Miranda, K. C., Bond, D. T., McKee, M. Nucleic acids within urinary exosomes/microvesicles are potential biomarkers for renal disease. Kidney Int. 78, 191-199 (2010).
- Pisitkun, T., Shen, R. F., Knepper, M. A. Identification and proteomic profiling of exosomes in human urine. Proc Natl Acad Sci USA. 101, 13368-13373 (2004).
- Raj, D. A., Fiume, I., Capasso, G., Pocsfalvi, G. A multiplex quantitative proteomics strategy for protein biomarker studies in urinary exosomes. Kidney Int. 81 (12), 1263-1273 (2012).
- Pisitkun, T., Gandolfo, M. T., Das, S., Knepper, M. A., Bagnasco, S. M. Application of systems biology principles to protein biomarker discovery: urinary exosomal proteome in renal transplantation. Proteomics Clin Appl. 6 (5-6), 268-278 (2012).
- Gonzales, P. A., Zhou, H., Pisitkun, T. Isolation and purification of exosomes in urine. Methods Mol Biol. 641, 89-99 (2010).
- Zhou, H., Yuen, P. S., Pisitkun, T. Collection, storage, preservation, and normalization of human urinary exosomes for biomarker discovery. Kidney Int. 69, 1471-1476 (2006).
- Thery, C., Amigorena, S., Clayton, A. Isolation and characterization of exosomes from cell culture supernatants and biological fluids. Curr Prot Cell Biol. 43, 3.22.1-3.22.29 (2006).
- Cheruvanky, A., Zhou, H., Pisitkun, T. Rapid isolation of urinary exosomal biomarkers using a nanomembrane ultrafiltration concentrator. Am J Physiol Renal Physiol. 292, 1657-1661 (2007).
- Gonzales, P. A., Pisitkun, T., Hoffert, J. D. Large-scale proteomics and phosphoproteomics of urinary exosomes. J Am Soc Nephrol. 20, 363-379 (2009).
- Rood, I. M., Deegens, J. K., Merchant, M. L. Comparison of three methods for isolation of urinary microvesicles to identify biomarkers of nephrotic syndrome. Kidney Int. 78, 810-816 (2010).
- Alvarez, M. L., Khosroheidari, M., Kanchi Ravi, R., DiStefano, J. K. Comparison of protein, microRNA, and mRNA yields using different methods of urinary exosome isolation for the discovery of kidney disease biomarkers. Kidney Int. Nov. 82 (9), 1024-1032 (2012).
- Kang, S. W., Kim, Y. W., Kim, Y. H. Study of the association of -667 aquaporin-2 (AQP-2) A/G promoter polymorphism with the incidence and clinical course of chronic kidney disease in Korea. Renal Fail. 29, 693-698 (2007).

