Novel Atomic Force Microscopy Based Biopanning for Isolation of Morphology Specific Reagents against TDP-43 Variants in Amyotrophic Lateral Sclerosis
Summary
Using atomic force microscopy in combination with biopanning technology we created a negative and positive biopanning system to acquire antibodies against disease-specific protein variants present in any biological material, even at low concentrations. We were successful in obtaining antibodies to TDP-43 protein variants involved in Amyotrophic Lateral Sclerosis.
Abstract
Because protein variants play critical roles in many diseases including TDP-43 in Amyotrophic Lateral Sclerosis (ALS), alpha-synuclein in Parkinson’s disease and beta-amyloid and tau in Alzheimer’s disease, it is critically important to develop morphology specific reagents that can selectively target these disease-specific protein variants to study the role of these variants in disease pathology and for potential diagnostic and therapeutic applications. We have developed novel atomic force microscopy (AFM) based biopanning techniques that enable isolation of reagents that selectively recognize disease-specific protein variants. There are two key phases involved in the process, the negative and positive panning phases. During the negative panning phase, phages that are reactive to off-target antigens are eliminated through multiple rounds of subtractive panning utilizing a series of carefully selected off-target antigens. A key feature in the negative panning phase is utilizing AFM imaging to monitor the process and confirm that all undesired phage particles are removed. For the positive panning phase, the target antigen of interest is fixed on a mica surface and bound phages are eluted and screened to identify phages that selectively bind the target antigen. The target protein variant does not need to be purified providing the appropriate negative panning controls have been used. Even target protein variants that are only present at very low concentrations in complex biological material can be utilized in the positive panning step. Through application of this technology, we acquired antibodies to protein variants of TDP-43 that are selectively found in human ALS brain tissue. We expect that this protocol should be applicable to generating reagents that selectively bind protein variants present in a wide variety of different biological processes and diseases.
Introduction
The presence of protein variants has been implicated as a factor in the progression of many diseases including neurodegenerative diseases such as Alzheimer’s, Parkinson’s, ALS and Frontotemporal Dementia (FTD)1,2,3,4,5,6,7,8,9,10,11. Oligomeric forms of the proteins beta-amyloid and alpha-synuclein are thought to be the toxic species responsible for Alzheimer’s and Parkinson’s, respectively2,3,4,5. Aggregates of the TAR DNA-binding protein 43 (TDP-43) have been linked to ALS and FTD12,13,14. Therefore reagents such as antibodies that can selectively target the different protein variants can be powerful tools to serve as diagnostic markers and potential therapeutics. In this study, we focused on developing reagents that selectively bind variants of the TDP-43 protein implicated in ALS, however the technique outlined in this paper should be applicable to the isolation of reagents against a wide range of protein variants.
Cytoplasmic aggregation of TDP-43 has been identified as a pathological feature in ALS15,16,17,18,19. Typically TDP-43 is found in the nucleus of all cells from a normal individual, although it tends to move between the cytosol and nucleus15,17. However, in ALS aggregated forms of TDP-43 are detected in the cytoplasm of select neurons and glia with lower concentrations found in the nucleus suggesting the movement of TDP-43 from the nucleus to the cytoplasm during disease progression16,20. While aggregation of TDP-43 is found in the majority of ALS cases, it does not account for all cases since 1%-2% of total ALS cases (or 15%-20% of familial ALS cases ) are linked to mutations in the superoxide dismutase 1 (SOD1) gene15,17. Because of the important role of TDP-43 in the vast majority of ALS cases, here we focus on developing antibody based reagents that can selectively bind to TDP-43 variants that are present in human ALS brain tissue utilizing our novel AFM based biopanning techniques.
Initially we need a diverse repertoire of antibody binding domains. We combined three different phage display single chain variable domain antibody fragment (scFvs) libraries, (Tomlinson I and J and Sheets libraries21). The panning process is divided into negative and positive panning phases. Phages from the libraries are first subjected to the negative panning process during which phages reactive to multiple off-target antigens are excluded. After the completion of each round of negative panning against each off-target antigen, the process is monitored by AFM imaging to ensure that all phage binding the off-target antigens have been removed. Only after verifying by AFM imaging that all reactive phages are removed do we proceed to the next target. To isolate reagents against TDP-43 variants implicated in ALS we utilized the following negative panning antigens: 1) BSA to remove phage that bind weakly or non-specifically to proteins; 2) aggregated alpha-synuclein to remove phage that bind to generic structural elements of aggregated proteins; 3) human brain tissue homogenates to remove phage that bind to any proteins or other components present in post-mortem samples of healthy human brain tissue; 4) immunoprecipitated TDP-43 from healthy human brain to remove phage that bind to all TDP-43 forms associated with healthy human brain; and 5) immunoprecipitated TDP-43 isolated from FTD brain homogenates to remove phage that bind TDP-43 variants associated with non-ALS pathology. After removal of all phage reactive to all the off-target antigens, we then proceeded to the positive panning phase during which antibody fragments that bind the antigen of interest are isolated, in this case TDP-43 immunoprecipitated from human ALS brain tissue. These isolated antibodies may be reactive to aggregated or modified forms of TDP-43.
Conventional phage biopanning focuses mainly on the positive panning phase22,23. Usually the target of interest is immobilized, the phage library added and bound phages eluted. The phages are then amplified and added to the target again. This amplification and incubation process is usually repeated several times to increase the percentage of positive binding phage. While variations of this process have been used extensively to isolate antibody reagents against a wide range of target antigens, they generally requires large amounts of purified target antigen24,25,26,27, whereas our process requires only trace amounts of the target antigen. The protocol described here can be used to isolate reagents that selectively bind target antigens that are present at very low concentrations, without the need for purification and the panning can be performed directly against antigen present in complex tissue samples. The use of exhaustive negative panning protocols as verified by AFM ensures that clones isolated against the positive antigen should selectively bind the target even when not purified or enriched.
Kasturirangan and colleagues (2003) have carried out a similar negative and positive biopanning process to isolate antibodies reactive to oligomeric beta-amyloid using nanogram concentration of the target5. Here we expand on this process to enable the generation of reagents that selectively bind disease-specific protein variants directly from human tissue samples. In future studies we intend to further investigate not only the diagnostic value of the reagents isolated here but also assess their therapeutic relevance for treating ALS.
Overall, our novel AFM based biopanning technology should be applicable to the isolation of any disease-specific protein variant in any biological material without the need for protein purification or modification, even when the target antigen concentrations are extremely low.
Protocol
1. Phage Production
- Perform all phage production and biopanning processes in a biosafety cabinet. Produce phages particles from the different libraries (Tomlinson I and J libraries and Sheets library21) for the biopanning process using the manufacturer’s instructions (http://www.lifesciences.sourcebioscience.com/media/143421/tomlinsonij.pdf).
NOTE: We use multiple libraries in our panning process to increase the diversity of the antibodies available. - Briefly, culture bacteria stocks of the three libraries in 200 ml of 2xYT containing 1% glucose and 100 μg/ml of ampicillin on a shaker at 37 °C until OD600 is 0.4.
- Add 8 x 1011 of KM13 helper phage to 200 ml of the Tomlinson I and J libraries and 8 x 1011 of VCSM13 helper phage to 200 ml of the Sheets library. Incubate library samples with helper phage for 30 min at 37 °C without shaking.
- Centrifuge the cultures at 3,000 x g for 10 min and resuspend the cell pellet in 200 ml of 2xYT with 0.1% glucose, 100 μg/ml of ampicillin and 50 μg/ml of kanamycin.
- Incubate the cultures O/N at 30 °C with shaking and then spin for 30 min at 3,300 x g the following day.
- Add 50 ml of Polyethylene glycol (PEG) 8000 in 2.5 M NaCl to the 200 ml of supernatant from each of the libraries and incubate on ice for 1 hr. After PEG precipitation, spin the solutions again for 30 min at 3,300 x g.
- Add 1-2 ml of phosphate buffered saline (PBS) to the cell pellet (depending on the size of the pellet) and transfer to microcentrifuge tubes.
- Incubate phage particles on ice for 1 hr at 37 °C with occasional mixing before centrifuging for 10 min at 11,600 x g. Collect the supernatant and store at -80 °C.
- Before freezing, transfer a small volume of the phage particles from the three libraries to another microcentrifuge tube and titer each library individually using TG1 cells to determine concentration. Below is a description of the titer process for one library.
- Grow TG1 cells to OD600 of 0.4. Add 990 μl of PBS to 6 microcentrifuge tubes.
- To the first tube add 10 μl of the phage particles. After mixing, remove 10 μl from tube one and add to tube 2. Repeat this step for the remaining 4 tubes.
- Add 200 μl of TG1 cells to 6 tubes and add 10 μl of each of the six phage dilutions to these tubes. Allow the phage particles to infect for 30 min with no shaking at 37 °C.
- Plate 100 μl of the cells unto Luria-Bertani agar plates with 100 μg/ml of ampicillin (LBA). Incubate O/N at 37 °C and then determine the pfu/ml using the number of visible colonies.
2. Negative Biopanning Process
NOTE: Avoid too many freeze thaw of the phages throughout the biopanning process. Ideally, it is best to carry out as many of the negative panning steps as possible in a day. After the completion of the rounds of negative or positive panning against each target it is important to save some of the phage in case of contamination at any step.
- 12 Rounds of Negative Panning against Bovine Serum Albumin (BSA)
- Add 4 ml of 1 mg/ml BSA solution in PBS to 12 immunotubes and incubate O/N at 4 °C.
- Combine 1 x 1012 phage particles from each of the three libraries (set aside a small quantity of this phage mixture for future verification of the negative panning process).
- Discard the BSA solution from the first tube and wash the tube 2-3 times with PBS to remove any unbound BSA.
- Add the combined phage libraries to this first tube, incubate for 30 min at RT on the rotator and collect the unbound phages.
- Repeat step 2.1.3 with the second immunotube and add the phage collected from step 2.1.4 to this immunotube. Repeat step 2.1.4.
- Repeat steps 2.1.3-2.1.4 with the remaining 10 tubes of BSA solution and the unbound phage collected after each incubation.
- To verify removal of all BSA phage binders use AFM. Add 10 μl of the phage from step 2.1.2 and 10 μl of the remaining phage after step 2.1.7 separately to 10 μg/ml of BSA bound to cleaved mica. A brief version of the AFM process is outlined in section 8.
- It is important to save a small quantity of phage after finishing the numerous rounds of negative panning against each target to verify success of the panning process.
- 10 Rounds of Negative Panning against Various Forms of Alpha-synuclein
NOTE: In this project, we want to remove phage that could bind other aggregated forms of proteins such as alpha-synuclein (monomer, dimer, trimer, etc.) and so we will eliminate any of these phages (the goal is to eliminate as many cross-reactive phages as possible). Allow purified alpha-synuclein to aggregate for at least 7 days so that both monomeric and oligomeric forms are present for negative panning.- Prepare 4 immunotubes with 500 μg/ml of aggregated alpha-synuclein in PBS and 4 immunotubes with 100 μg/ml of aggregated alpha-synuclein and incubate O/N at 4 °C.
- Discard the solution from one of the tubes with 500 μg/ml of aggregated alpha-synuclein, wash with PBS and add the phage library collected after the 12 rounds of negative panning with BSA.
- Incubate the tube for 30 min at RT on the rotator and collect the unbound phages.
- Repeat steps 2.2.3 and 2.2.4 for the remaining tubes with 500 μg/ml of aggregated alpha-synuclein and then the tubes with 100 μg/ml of aggregated alpha-synuclein using the unbound phage collected each time after step 2.2.4.
- Repeat step 2.1.8 using the phage after the first of negative panning against 500 μg/ml of aggregated alpha-synuclein and the phage after the eighth round of negative panning against 100 μg/ml of alpha-synuclein.
- To eliminate all negative binders against alpha-synuclein prepare two extra immunotubes with 500 μg/ml of aggregated alpha-synuclein and repeat steps 2.2.2-2.2.4.
- 10 Rounds of Negative Panning against Healthy Human Brain Tissue Homogenate
- Add 1x STEN buffer (50 mM Tris (pH 7.6), 150 mM NaCl, 2 mM EDTA, 0.2% NP-40, 0.2% BSA, 20 mM PMSF and protease cocktail inhibitor) to human brain tissue28. Homogenize at 24,000 rpm for 1 min, centrifuge samples at 3,000 rpm and collect supernatant.
- Prepare 2 immunotubes with 2 ml of 50 μg/ml of human brain tissue in PBS and 8 immunotubes with 10 μg/ml of human brain tissue and incubate O/N at 4 °C.
- Repeat steps 2.2.3-2.2.5 first with the tubes containing 50 μg/ml of human brain tissue and then with those containing 10 μg/ml of human brain tissue using the unbound phage collected after the negative panning against alpha-synuclein.
- Repeat step 2.1.8 using the phage after the first of negative panning against 50 μg/ml of human brain tissue and the phage after the tenth round of negative panning against human brain tissue.
- 8 Rounds of Negative Panning against Healthy Immunoprecipitated TDP-43
- Immunoprecipitate healthy TDP-43 protein from healthy human brain tissue using the protocol described in section 9. Due to lower quantity of immunoprecipitated protein, carry out these rounds of negative panning using mica.
- Cleave eight mica surfaces.
- To the first mica add ~100 μl of 5 μg/ml of the immunoprecipitated protein and incubate for 5-10 min at RT.
- Wash the mica 5 times with 0.1% PBS with Tween 20 (PBST).
- Add ~100 μl of the phage after the negative panning against healthy brain tissue and incubate for 5-10 min at RT.
- At midway incubation of step 2.4.5, carry out step 2.4.3 using the second mica.
- Collect the phage from step 2.4.5 and wash the mica 5 times with 0.1% PBST.
- Wash mica from step 2.4.6 and add the phage from 2.4.7.
- Repeat steps 2.4.4 to 2.4.9 until the phage is negatively panned against all eight mica with the immunoprecipitated protein. Repeat step 2.1.7 using the phage after the first and eight round of negative panning.
- 2 Rounds of Negative Panning Against FTD Immunoprecipitated TDP-43
- Immunoprecipitate TDP-43 protein from the brains of individuals with FTD.
- Cleave 2 mica surfaces.
- Add 50 μl of 10 μg/ml of FTD TDP-43 to one mica. Incubate for 5 min at RT.
- Wash the mica 5 times with 0.1% PBST.
- Add 50 μl of the phage collected after negative panning against healthy TDP-43. Incubate for 5 min.
- Collect the unbound phages.
- Repeats steps 2.5.3 to 2.5.6 with the second mica and the unbound phage collected in 2.5.6.
3. Positive Panning against Immunoprecipitated TDP-43 from Individuals with ALS
- Immunoprecipitate TDP-43 protein from the brains of individuals with ALS. Cleave 3 mica surfaces. Add 50 μl of 10 μg/ml of ALS TDP-43 to one mica. Incubate for 5 min at RT. Wash the mica 5 times with 0.1% PBST.
- Add 50 μl of the phage collected after negative panning against healthy TDP-43 to the mica. Incubate for 5 min.
- Collect the unbound phages. Also collect the bound phages using three elution methods. It is important to keep all collected phages in these panning steps separate.
- First, add 50 μl of trypsin (1 mg/ml in PBS) to each mica and incubate for no more than 10 min. Collect the solution and save. Next, add 50 μl of 100 mM triethylamine (TEA) to each mica and incubate for no more than 10 min. Collect the solution, add equal volume of 1 M Tris-HCl buffer pH 7.4 and save.
- Third, add 50 μl of TG1 at OD600 of 0.4 directly to the mica and incubate for 30 min at 37 °C. Also add the phages collected from the two elution methods in 3.3.1 to 100 μl of TG1 at OD600 of 0.4 and incubate for 30 min at 37 °C.
- Repeat steps 3.1 with a second mica.
- Take the unbound phage collected in 3.3.2. using the first mica with ALS TDP-43 and add it to the second mica with ALS TDP-43. Incubate for 5 min.
- Repeat all the steps in 3.3 for these second sets of mica.
- Combine the TG1 cells infected with the phages eluted with trypsin from the two ALS TDP-43 mica and plate on LBA plates. Repeat the same process for the TEA and TG1 eluted phages from the two ALS TDP-43 mica for a total of three plates.
NOTE: Because of TDP-43’s involvement in both ALS and FTD, some of the clones isolated in the two rounds of positive panning against ALS TDP-43 may be specific to this target, but some may also bind FTD. To better isolate ALS specific clones, take the unbound phage collected after 2 rounds of negative panning against FTD TDP-43 (since these two rounds of negative panning should have removed any target that is reactive to FTD TDP-43) and use these phages in one more round of positive panning against the ALS TDP-43 antigen on mica (prepare the ALS TDP-43 mica as described above in 3.3 and 3.4). - Repeat all the steps in 3.3 with this third ALS mica and plate the infected TG1 cells on three LBA plates O/N at 37 °C. The next day count the number of colonies on the 6 LBA plates with the potential ALS clones. There should be fewer colonies on the three LBA plates with the potential ALS clones after the third round of positive panning compared to the three LBA plates after the two rounds of positive panning.
4. Culturing and Phage Production of Potential Positive Clones against ALS Specific TDP-43
- Grow each colony from the 6 plates in 96-well plates containing 100 μl of 2xYT with 1% glucose and 100 μg/ml of ampicillin on a shaker at 37 °C O/N.
- The next day transfer 10 μl from the wells containing the clones from the 3 LBA plates after the third round of positive panning with the ALS TDP-43 protein to microcentrifuge tubes containing 1 ml of 2xYT with 1% glucose and 100 μg/ml of ampicillin.
- Add glycerol to all the cultures in the 96-well plates to a final concentration of 15% and freeze the plates at -80 °C. Allow the 1 ml cultures to grow for 2-3 hr shaking at 37 °C.
- Add 109 of KM13 helper phage and allow the helper phage to infect for 1 hr at 37 °C with shaking. Centrifuge the cells for 10 min at 3,000 x g, discard the supernatant and add 1 ml of 2xYT with 0.1% glucose, 100 μg/ml of ampicillin and 50 μg/ml of kanamycin.
- Culture the cells O/N at 30 °C with shaking. Centrifuge the tubes at 3,300 x g for 30 min, transfer the supernatant to new microcentrifuge tubes and add 250 μl PEG/NaCl to each tube.
- Incubate the tubes on ice for 1 hr. Centrifuge the tubes again at 3,300 x g for 30 min, discard the supernatant and add 100 μl of PBS to each tube.
- Incubate the tubes on ice for 1 hr. Spin the tubes for 10 min at 11,600 x g and transfer the supernatant to new tubes. Store the phages at -80 °C.
5. Sequencing of Potential ALS Clones
- Repeat step 4.2 and allow to cultures to grow O/N at 37 °C with shaking. The next day carry out DNA plasmid preps of the clones according to the manufacturer’s instructions. Sequence the clones using the appropriate forward and reverse primers.
6. Screening Potential Positive Clones against ALS Specific TDP-43 Using Indirect ELISA
- Add 100 μl of 2.5 μg/ml of ALS, FTD and healthy human brain tissue to the wells of a high binding ELISA plate for each potential ALS clone. Incubate the plates at 37 °C for 1 hr with slow shaking.
- Wash the plates 3 times with 0.1% PBST. Add 200 μl of 2% milk powder in PBS to the well and incubate the plates at 37 °C for 1 hr with slow shaking.
- Repeat the wash step and then add 100 μl of each phage diluted at least 1/100 in PBS to the wells. After the incubation and wash steps, add 100 μl of 1/1,000 anti-M13 HRP to each well. Wash the plates and visualize using the ELISA femto chemiluminescence substrate kit.
7. scFv Production and Screening Using Indirect ELISA
- Grow HB 2151 cells to OD600 of 0.4. Add 5 μl of the different phages to 20 μl of the HB 2151 cells. Allow the phages 30 min to infect at 37 °C without shaking. Divide LBA plates into grids so that approximately 20 clones can be plated on each.
- Add 5-10 μl of each phage infected bacteria to the plate and incubate O/N at 37 °C. The next day add individual colonies to 1 ml of 2xYT with 0.1% glucose and 100 μg/ml of ampicillin and shake at 37 °C for 3-4 hr until OD600 is 0.6.
- Add 1 μl of 1 M isopropylthiogalactoside (IPTG) to each tube and incubate O/N at 30 °C with shaking. Spin the tubes at 11,600 x g for 10 min and harvest the supernatant for ELISA testing.
- For the ELISA testing follow the same procedures in section 6, except instead of phage in 6.5 add the scFv undiluted and for 6.6 add 100 μl of 1/1,000 9e10 HRP.
8. AFM Process
- Add 10-20 μl of the target antigen to cleaved mica and incubate for 5-10 min. Wash the mica 5 times with water. Add 10-20 μl of the phages to the mica and incubate for 5-10 min.
- Wash the mica 5 times again with water and allow the mica to air dry. Visualize phage binding using AFM1. Execute the AFM imaging process using AFM tapping mode with a Nanoscope IIIa controller in air at RT29. The silicon AFM probes have a resonant frequency of 300 kHz and a spring constant of 40 N/m. Use a scan rate of 3.05 Hz and a scan resolution of 512 samples/line.
- Process the images by basic flattening of the background to ensure that all particle sizes are comparable within each image. Phage width (not length) may vary from image to image due to other particle sizes and the scanning area, but it will never compromise the accuracy of whether or not phage binds to the antigen.
- If during AFM verification for each set of negative panning phages are visible on the mica, perform addition rounds until no phages are detected before proceeding to the next set of negative panning.
9. Immunoprecipitation of Target Antigen
- To isolate the target antigen (TDP-43), add 250 μg of homogenize brain tissue (from motor cortex), 100 μl of 1/100 dilution of the anti-TDP 43 polyclonal antibody and 1x STEN buffer to 700 μl total volume to an Eppendorf tube28.
- Incubate O/N at 4 °C on a rotator. Wash A/G agarose beads with the 1x STEN buffer 2-3 times and add 25 ml to the tissue mixture. Incubate at 4 °C for 1.5 hr on a rotator and then centrifuge at 2,000-4,000 rpm for 1 min.
- Remove the supernatant and wash the pellet once with 1x STEN buffer and 5 times with PBS. Add 40-60 μl of 1x STEN buffer to the pellet and boil for 5 min. After the samples are cool, centrifuge and collect the supernatant. Use SDS-PAGE and Western blotting to verify success of the immunoprecipitation process30.
10. Dot Blot Analysis
- Cut nitrocellulose paper into small rectangles for each clone that will be tested. Dot 2 μl of ALS, FTD and healthy human brain tissue samples unto the nitrocellulose paper and let it dry completely.
- Add 1-2 ml of 5% milk powder in PBS and let it shake for 1 hr at RT. Add 1 ml of 1/100 dilution of each phage and let it shake O/N at 4 °C. Wash the membrane 3 times for 10 min each with 0.5% PBST.
- Add 1 ml of 1/1,000 dilution of anti-M13 HRP and let it shake for 1 hr at RT. Repeat wash procedure and visualized using Western chemiluminescence solution.
Representative Results
In Figure 1, the schematic demonstrates the negative panning process by which we removed phage binding off-target antigens from our library using immunotubes. We initially started with BSA since this is a common blocking agent and any phage that would react nonspecifically with this target would be problematic in future immunoassays. Next, we removed binders to aggregated alpha-synuclein to eliminate phage that are reactive with generic structures of aggregated proteins (i.e., an antibody that would be cross-reactive to aggregated alpha-synuclein, TDP-43, abeta, etc.). AFM results showed phage binding after 1 round of negative panning against the aggregated alpha-synuclein (Figure 2A) and no binding after 8 rounds (Figure 2B). We then negatively panned against healthy human brain tissue to remove phage binding the many antigens present in healthy human brain homogenate. Figure 2C showed phage binding to healthy tissue after 1 round of negative panning and no phage left binding after 10 rounds of negative panning (Figure 2D). Since the amount of healthy and FTD immunoprecipitated TDP-43 protein available for panning was low, using mica rather than immunotubes utilized less volume and therefore lowered total protein consumed (Figure 3). After 8 rounds of negative panning against healthy immunoprecipitated TDP-43, the phage was divided. Half the phage was expended in two rounds of negative panning against FTD TDP-43. The objective was to eliminate any FTD TDP-43 reactive clones (due to TDP-43’s involvement in FTD), while retaining any potential ALS TDP-43 specific clones.
For the positive panning portion (Figure 4), we also employed mica as a substrate to minimize use of material. We used the unbound phage after the FTD TDP-43 negative panning against ALS TDP-43 to acquire any ALS specific clones. We also positively panned against ALS TDP-43 twice in the event that we did not obtain clones after using the unbound phage from the negative panning against FTD TDP-43. After the entire panning process, our three elution methods yielded 154 clones from the two positive panning against ALS TDP-43 (clones that may be reactive to ALS and FTD) and 45 potential ALS specific clones (using the unbound phage from the FTD TDP-43 negative panning).
To further reduce the 45 potential ALS specific clones, the clones are sequenced and only the clones without any stop codons were considered further, except for one clone which showed up twice. This left us with 23 potential ALS specific clones. After preparing both phage and soluble scFv with these clones, they are tested in indirect ELISAs for specificity to ALS tissue. Almost all of the phages showed a preference for ALS tissue over healthy tissue (Figure 5). Similar results are obtained when comparing phage binding to ALS to FTD tissue (Figure 6). For future studies it is essential that these clones express high yields of functional scFvs, so we produced small batches of the different scFvs and carried out similar indirect ELISAs. Comparison of scFv binding to ALS tissue to both healthy (Figure 7) and FTD tissue (Figure 8) again showed selective binding to ALS tissue in almost all clones.
We used another immunoassay (dot blots) to further verify binding to ALS tissue and also to ascertain which clones are reactive in other immunological applications. Clone 2A binding to ALS tissue is shown (Figure 9). All of these clones showed promising results in either or both of the phage and scFv ELISAs. These results confirm that our AFM based biopanning process is a very powerful technique that can be used to generate reagents that selectively bind disease-specific protein variants directly from complex sources.
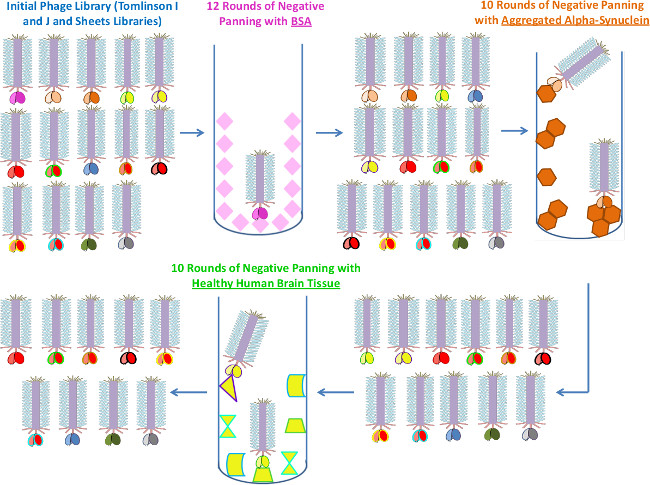
Figure 1. Negative Biopanning Process Utilizing Immunotubes. Schematic demonstrating the negative biopanning process. Phages that are reactive to proteins such as BSA, aggregated alpha-synuclein and healthy brain tissue are removed using immunotubes. Please click here to view a larger version of this figure.
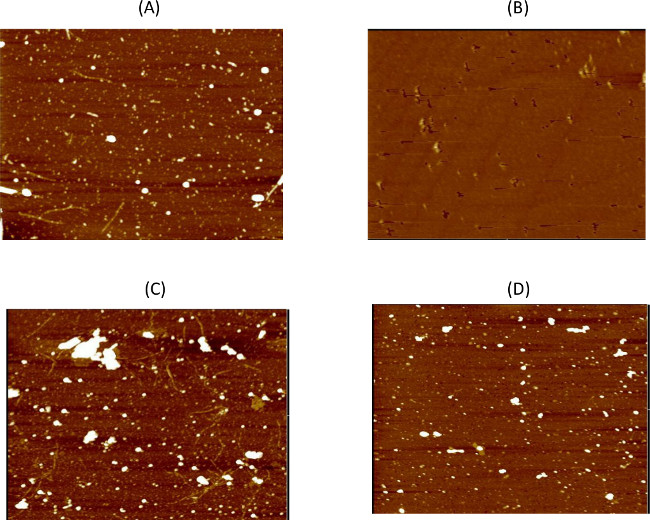

Figure 2. Confirmation of Negative Panning Results. AFM imaging is used to monitor the negative panning steps against the various targets to ensure all reactive phages are removed. Here we show some of the AFM results demonstrating the level of phage binding before and after the negative panning. (A) Phage binding can be detected to aggregated alpha-synuclein particles after the first round of negative panning, but (B) no phage are visible after 8 rounds of negative panning. (C) After the first round of negative panning against healthy tissue phage binding is evident, but (D) no binding is detected after 10 rounds of negative panning. All images are 5 μm. The large white structures on the AFM images are usually due to salts present in the buffers or residual PEG from the PEG precipitation step during phage production. Please click here to view a larger version of this figure.
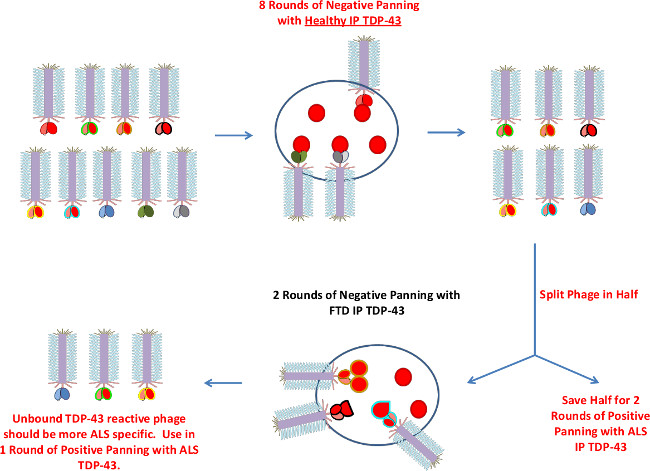
Figure 3. Negative Biopanning Process Utilizing Mica. Schematic demonstrating additional negative biopanning using mica. Due to limited sample mica surface is employed to first eliminate binders to healthy and then FTD immunoprecipitated TDP-43. Before proceeding to the two rounds of negative panning against FTD TDP-43, the phage is split in half (in the event of unsuccessful isolation of ALS TDP-43 exclusive phages during the positive panning phase). Please click here to view a larger version of this figure.
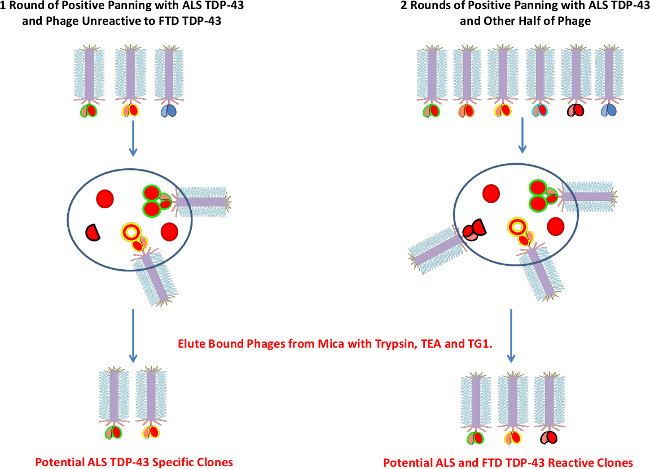
Figure 4. ALS Positive Biopanning Process. Schematic demonstrating the ALS positive biopanning process. The unbound phages after the negative panning against FTD TDP-43 are used in a round of positive panning against ALS TDP-43 to elute more ALS TDP-43 specific clones. Also, two rounds of positive panning against ALS TDP-43 are carried out using mica surface and the unbound phages after negative panning against healthy immunoprecipitated TDP-43 (the bound phages are eluted since these phages should not bind healthy TDP-43, however some may be cross-reactive with both ALS and FTD). Three elution methods are used (trypsin, TEA and TG1 cells) to ensure all bound phages are removed. Please click here to view a larger version of this figure.
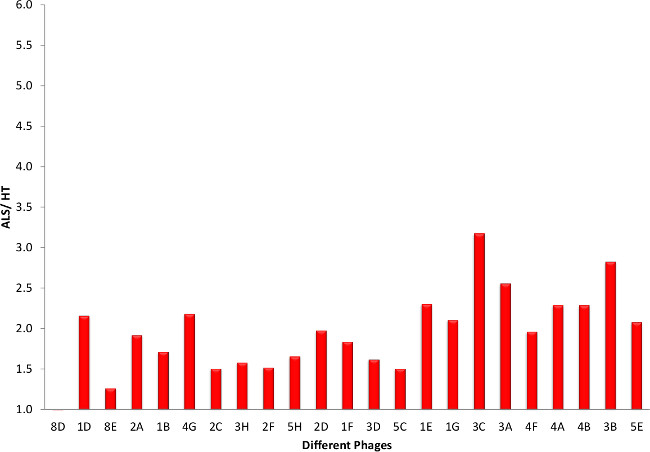
Figure 5. Phage ELISA Screening of Potential ALS Clones (ALS versus HT). Using homogenized human ALS, FTD and healthy brain tissue (HT) samples we performed indirect ELISA using phage produced from the different clones. Results are represented as the ratio to healthy tissue samples. Results showed that some clones distinguish between TDP-43 found in ALS patients and those in healthy. The ALS, FTD and HT tissues are a mix of brain samples (motor cortex) from three individuals. Please click here to view a larger version of this figure.
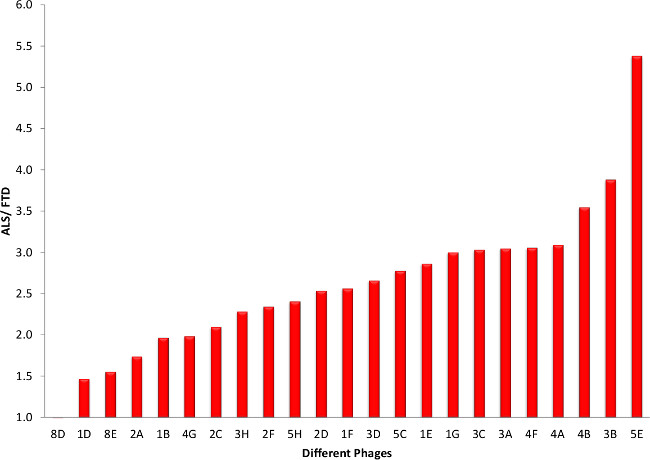
Figure 6. Phage ELISA Screening of Potential ALS Clones (ALS versus FTD). Here we show the phage ELISA results of ALS versus FTD patients. Most of the clones have a preference for ALS over FTD. Please click here to view a larger version of this figure.
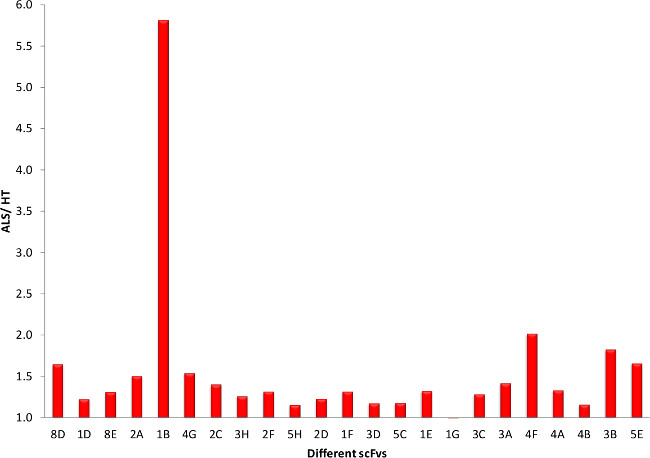
Figure 7. scFv ELISA Screening of Potential ALS Clones (ALS versus HT). Using the scFvs produced from each clone we can observe clones that have a preference for the TDP-43 from ALS patients over healthy. Please click here to view a larger version of this figure.
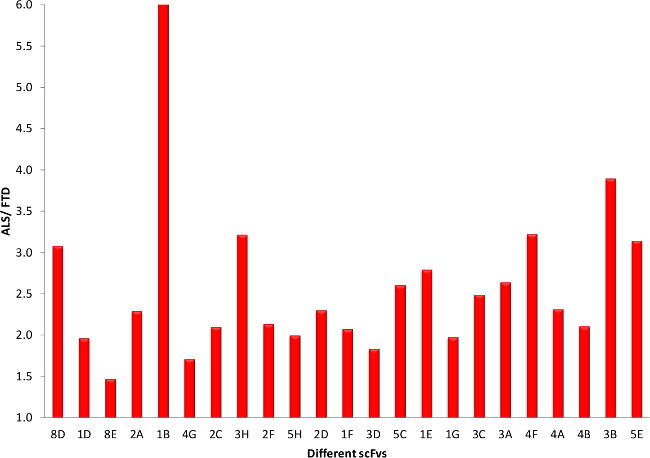
Figure 8. scFv ELISA Screening of Potential ALS Clones (ALS versus FTD). The success of our panning process is further demonstrated when comparing ALS to FTD for the different scFvs in the indirect ELISAs. Please click here to view a larger version of this figure.
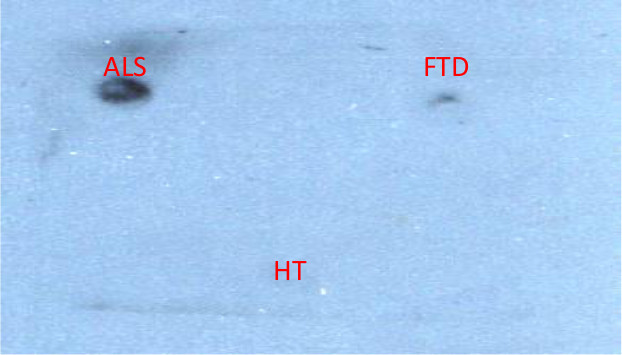
Figure 9. Dot Blot Analysis of Potential ALS Clones. Using dot blot instead of ELISA is another technique to show specificity of the clones for ALS over FTD or HT. Clone 2A is shown. Please click here to view a larger version of this figure.
Discussion
Protein variants have been shown to be involved in the progression of many neurodegenerative diseases such as Alzheimer’s, Parkinson’s, ALS and FTD1,2,3,4,5,6,7,8,9,10,11. Isolation of antibodies that can selectively recognize these different protein variant targets can be effective reagents to study, diagnose and potentially treat such ailments. To generate such variant specific antibodies we have developed a novel biopanning process that utilizes atomic force microscopy to monitor the progress of each step. Previous work using a similar negative and positive biopanning combination resulted in a clone that binds oligomeric beta-amyloid5. Here we extend these studies to demonstrate that we can efficiently generate reagents that selectively bind disease-specific protein variants present in human brain tissue samples. We isolated morphology specific reagents that selectively recognize TDP-43 variants uniquely present in human ALS brain tissue13,14. Future studies necessitate characterization of the TDP-43 variants being targeted (whether aggregated or modified).
The key steps to ensuring that we can obtain phages specific to any protein variant target during the positive panning phase is to confirm that the negative panning steps have removed all potential cross-reacting phage clones. We utilized AFM imaging to verify that essentially all phage clones binding off-target antigens have been removed during each of the rounds of negative panning. Since a high percentage of the phage recovered from the positive binding are then selective for the desired target, only minimal amounts of the target antigen is needed for screening.
It is essential to perform any of the panning steps where the phages will be exposed to the external environment in a biosafety cabinet to prevent contamination by other phage. To guard against this event, it is important to save a small aliquot of phage after each panning round. If contamination is discovered, a phage aliquot from a previous round can then be used to continue the process. Freezing and thawing the phages several times may be harmful to stability and so it is beneficial to perform many of the negative panning steps continuously31. Because the panning process does not utilize phage amplification steps after each round of positive panning32 the process is less likely to isolate empty or incomplete phage sequences.
The AFM based panning process generates a number of potential protein variant specific clones after the positive panning step. To reduce the number of potential clones, we tested each clone for its ability to distinguish ALS from other diseased and healthy human brain tissue samples. The AFM based biopanning process described here represents a powerful technique that can be used to isolate reagents that selectively recognize disease-specific protein variants directly from complex human samples. The target protein variants can be present at very low concentrations, and there is no need to purify, modify or concentrate the target antigen.
Divulgations
The authors have nothing to disclose.
Acknowledgements
This research was supported by a grant from NIH: R21AG042066. We would like to thank Philip Schulz for his contributions in creating the screen capture videos.
Materials
| Name of Reagent/ Equipment | Company | Catalog Number | Comments/Description |
| Tomlinson I and J Libraries | MRC (Cambridge, England) | ||
| Sheets Library | MRC (Cambridge, England) | ||
| 2xYT | BD Sciences | 244020 | |
| Glucose | Amresco | 0188-2.5KG | |
| Ampicillin | Amresco | 0339-25G | Irritant |
| KM13 Helper Phage | MRC (Cambridge, England) | ||
| Kanamycin | OmniPur | 5880 | Irritant |
| Polyethylene Glycol 8000 | OmniPur | 6510 | Irritant |
| Sodium Chloride | Macron | 7647-14-5 | |
| Sodium Phosphate Dibasic | Amresco | 0404-1KG | Irritant |
| Potassium Chloride | EMD | PX1405-1 | Irritant |
| Potassium Phosphate Monobasic | Amresco | 0781-500G | Irritant |
| TG1 Cells | MRC (Cambridge, England) | ||
| Luria-Bertani Agar | EMD | 1.10283.0500 | |
| Bovine Serum Albumin | Amresco | 0332-100G | |
| STEN buffer | Crystalgen Inc. | 33429775 | |
| Immunotubes | Thermo Scientific | 470319 | |
| Mica | Spruce Pine Mica | 24365 | |
| Tween 20 | EMD | ||
| Trypsin | Sigma | T-0303 | Irritant |
| Triethylamine | Sigma | T-0886 | Flammable |
| Glycerol | Amresco | 0854-1L | Irritant |
| DNA Plasmid Prep Kit | qiagen | 27106 | Irritant |
| Non-Fat Milk Powder | Carnation | ||
| 96-Well High Binding ELISA Plate | Costar | 3590 | |
| Anti-M13 HRP | GE Healthcare Life Sciences | 27-9421-01 | |
| ELISA Femto Chemiluminescence Substrate Kit | Thermo Scientific | 37074 | |
| Anti-TDP 43 Polyclonal Antibody | ProteinTech | 10782-2-AP | |
| A/G Agarose Beads | Santa Cruz Biotechnology | sc-2003 | |
| HB 2151 Cells | MRC (Cambridge, England) | ||
| Isopropylthiogalactoside | Teknova | 13325 | |
| 9e10 HRP | Santa Cruz Biotechnology | sc-40 | |
| Nitrocellulose Membrane | Biorad | 162-0115 | Flammable |
| Centrifuge | Thermo Scientific | Sorvall RC 6+ | |
| Nanoscope IIIa Atomic Force Microscope | Veeco | ||
| AFM Probes | VistaProbes | T300R-10 |
References
- Hedieh, B., Sharareh, E., Philip, S., Michael, R. S. Isolating recombinant antibodies against specific protein morphologies using atomic force microscopy and phage display technologies. Protein Engineering Design and Selection. 19, 497-502 (2006).
- Emadi, S., Barkhordarian, H., Wang, M. S., Schulz, P., Sierks, M. R. Isolation of a Human Single Chain Antibody Fragment Against Oligomeric α-Synuclein that Inhibits Aggregation and Prevents α-Synuclein-induced Toxicity. Journal of Molecular Biology. 368, 1132-1144 (2007).
- Emadi, S., Kasturirangan, S., Wang, M. S., Schulz, P., Sierks, M. R. Detecting Morphologically Distinct Oligomeric Forms of α-Synuclein. Journal of Biological Chemistry. 284, 11048-11058 (2009).
- Kasturirangan, S., et al. Nanobody specific for oligomeric beta-amyloid stabilizes nontoxic form. Neurobiology of Aging. 33, 1320-1328 (2012).
- Kasturirangan, S., et al. Isolation and characterization of antibody fragments selective for specific protein morphologies from nanogram antigen samples. Biotechnology Progress. 29, 463-471 (2013).
- Zameer, A., Kasturirangan, S., Emadi, S., Nimmagadda, S. V., Sierks, M. R. Anti-oligomeric Aβ Single-chain Variable Domain Antibody Blocks Aβ-induced Toxicity Against Human Neuroblastoma Cells. Journal of Molecular Biology. 384, 917-928 (2008).
- Boddapati, S., Levites, Y., Sierks, M. R. Inhibiting β-Secretase Activity in Alzheimer’s Disease Cell Models with Single-Chain Antibodies Specifically Targeting APP. Journal of Molecular Biology. 405, 436-447 (2011).
- Boddapati, S., Levites, Y., Suryadi, V., Kasturirangan, S., Sierks, M. R. Bispecific Tandem Single Chain Antibody Simultaneously Inhibits β-Secretase and Promotes α-Secretase Processing of AβPP. Journal of Alzheimer’s Disease. 28, 961-969 (2012).
- Zhou, C., Emadi, S., Sierks, M. R., Messer, A. A Human Single-Chain Fv Intrabody Blocks Aberrant Cellular Effects of Overexpressed [alpha]-Synuclein. Mol Ther. 10, 1023-1031 (2004).
- Vanden Broeck, L., Callaerts, P., Dermaut, B. TDP-43-mediated neurodegeneration: towards a loss-of-function hypothesis. Trends in Molecular Medicine. 20, 66-71 (2014).
- Akamatsu, M., et al. A unique mouse model for investigating the properties of amyotrophic lateral sclerosis-associated protein TDP-43, by in utero electroporation. Neuroscience Research. 77, 234-241 (2013).
- Keage, H. A., et al. TDP-43 in the Population: Prevalence and Associations with Dementia and Age. Journal of Alzheimer’s Disease. 42, 641-650 (2014).
- Honda, D., et al. The ALS/FTLD-related RNA-binding proteins TDP-43 and FUS have common downstream RNA targets in cortical neurons. FEBS Open Bio. 4, 1-10 (2014).
- Baloh, R. H. TDP-43: the relationship between protein aggregation and neurodegeneration in amyotrophic lateral sclerosis and frontotemporal lobar degeneration. FEBS Journal. 278, 3539-3549 (2011).
- Ling, S. -. C., Polymenidou, M., Cleveland, D. W. . Converging Mechanisms in ALS and FTD: Disrupted RNA and Protein. 79, 416-438 (2013).
- Sasaki, S., Takeda, T., Shibata, N., Kobayashi, M. Alterations in subcellular localization of TDP-43 immunoreactivity in the anterior horns in sporadic amyotrophic lateral sclerosis. Neuroscience Letters. 478, 72-76 (2010).
- Robertson, J., et al. Lack of TDP-43 abnormalities in mutant SOD1 transgenic mice shows disparity with ALS. Neuroscience Letters. 420, 128-132 (2007).
- Shan, X., Vocadlo, D., Krieger, C. Mislocalization of TDP-43 in the G93A mutant SOD1 transgenic mouse model of ALS. Neuroscience Letters. 458, 70-74 (2009).
- Yamashita, T., Hideyama, T., Teramoto, S., Kwak, S. The abnormal processing of TDP-43 is not an upstream event of reduced ADAR2 activity in ALS motor neurons. Neuroscience Research. 73, 153-160 (2012).
- Dong, H., et al. Curcumin abolishes mutant TDP-43 induced excitability in a motoneuron-like cellular model of ALS. Neurosciences. 272, 141-153 (2014).
- Sheets, M. D., et al. Efficient construction of a large nonimmune phage antibody library: The production of high-affinity human single-chain antibodies to protein antigens. Proceedings of the National Academy of Sciences. 95, 6157-6162 (1998).
- Hairul Bahara, N. H., et al. Phage display antibodies for diagnostic applications. Biologicals. 41, 209-216 (2013).
- Azzazy, H. M. E., Highsmith, W. E. Phage display technology: clinical applications and recent innovations. Clinical Biochemistry. 35, 425-445 (2002).
- Zhang, X., et al. Rapid isolation of single-chain antibodies from a human synthetic phage display library for detection of Bacillus thuringiensis (Bt). Cry1B toxin. Ecotoxicology and Environmental Safety. 81, 84-90 (2012).
- Liu, H., et al. Selection and characterization of single-chain recombinant antibodies against spring viraemia of carp virus from mouse phage display library. Journal of Virological Methods. 194, 178-184 (2013).
- Cukkemane, N., Bikker, F. J., Nazmi, K., Brand, H. S., Veerman, E. C. I. Identification and characterization of a salivary-pellicle-binding peptide by phage display. Archives of Oral Biology. 59, 448-454 (2014).
- Adamson, C. S., et al. Novel single chain antibodies to the prion protein identified by phage display. Virology. 358, 166-177 (2007).
- Hebron, M. L., et al. Parkin Ubiquitinates Tar-DNA Binding Protein-43 (TDP-43) and Promotes Its Cytosolic Accumulation via Interaction with Histone Deacetylase 6 (HDAC6). Journal of Biological Chemistry. 288, 4103-4115 (2013).
- Wang, M. S., Zameer, A., Emadi, S., Sierks, M. R. Characterizing Antibody Specificity to Different Protein Morphologies by AFM. Langmuir. 25, 912-918 (2008).
- Williams, S., Sakic, B., Hoffman, S. A. Circulating brain-reactive autoantibodies and behavioral deficits in the MRL model of CNS lupus. Journal of Neuroimmunology. 218, 73-82 (2010).
- Jończyk, E., Kłak, M., Międzybrodzki, R., Górski, A. The influence of external factors on bacteriophages—review. Folia Microbiol. 56, 191-200 (2011).
- Hammers, C. M., Stanley, J. R. Antibody Phage Display: Technique and Applications. J Invest Dermatol. 134, e17 (2014).

