Production, Characterization and Potential Uses of a 3D Tissue-engineered Human Esophageal Mucosal Model
Summary
This manuscript describes the production, characterization and potential uses of a tissue engineered 3D esophageal construct prepared from normal primary human esophageal fibroblast and squamous epithelial cells seeded within a de-cellularized porcine scaffold. The results demonstrate the formation of a mature stratified epithelium similar to the normal human esophagus.
Abstract
The incidence of both esophageal adenocarcinoma and its precursor, Barrett’s Metaplasia, are rising rapidly in the western world. Furthermore esophageal adenocarcinoma generally has a poor prognosis, with little improvement in survival rates in recent years. These are difficult conditions to study and there has been a lack of suitable experimental platforms to investigate disorders of the esophageal mucosa.
A model of the human esophageal mucosa has been developed in the MacNeil laboratory which, unlike conventional 2D cell culture systems, recapitulates the cell-cell and cell-matrix interactions present in vivo and produces a mature, stratified epithelium similar to that of the normal human esophagus. Briefly, the model utilizes non-transformed normal primary human esophageal fibroblasts and epithelial cells grown within a porcine-derived acellular esophageal scaffold. Immunohistochemical characterization of this model by CK4, CK14, Ki67 and involucrin staining demonstrates appropriate recapitulation of the histology of the normal human esophageal mucosa.
This model provides a robust, biologically relevant experimental model of the human esophageal mucosa. It can easily be manipulated to investigate a number of research questions including the effectiveness of pharmacological agents and the impact of exposure to environmental factors such as alcohol, toxins, high temperature or gastro-esophageal refluxate components. The model also facilitates extended culture periods not achievable with conventional 2D cell culture, enabling, inter alia, the study of the impact of repeated exposure of a mature epithelium to the agent of interest for up to 20 days. Furthermore, a variety of cell lines, such as those derived from esophageal tumors or Barrett’s Metaplasia, can be incorporated into the model to investigate processes such as tumor invasion and drug responsiveness in a more biologically relevant environment.
Introduction
The esophageal mucosa comprises a stratified, squamous epithelium above a layer of connective tissue, the lamina propria, and is one of the first sites to encounter ingested environmental stressors. Exposure to dietary toxins is implicated in the development of esophageal squamous carcinoma, while duodenogastro-esophageal reflux is a critical factor in the pathogenesis of Barrett’s Metaplasia, which is associated with increased risk of progression to esophageal adenocarcinoma. Esophageal carcinomas are the 8th most common malignant tumor in UK males and esophageal adenocarcinoma is rapidly increasing in the Western world1. Furthermore, there has been little improvement in disease prognosis, with an overall 5-year survival rate of around 15%. Consequently there is a need for experimental platforms to investigate the impact of exposure to environmental stressors on this esophageal epithelium and their potential involvement in the development of metaplasia or neoplasia.
Although immortalized or tumor cell lines allow researchers to study the response of epithelial cells to these stressors in vitro, they remain proliferative and fail to differentiate into the mature epithelial cells found on the uppermost layers of the esophageal mucosa. Furthermore, cells lines that have already undergone tumorigenesis may provide only limited information regarding the initial responses of normal cells within the epithelium to environmental factors; and this is the stage when the potential for therapeutic intervention may be highest. Finally, conventional cell culture systems fail to capture the potentially important interactions between epithelial and mesenchymal cells and between these cells and the surrounding matrix that occur within tissues in vivo.
Animal models provide a more realistic microenvironment for studying the responses of the esophageal epithelium and can incorporate the artificial induction of gastro-esophageal reflux disease2. However it can be more challenging to manipulate the environmental stressors in these models and they may not fully represent the response within the human esophagus.
Other experimental human esophageal models have been developed that utilize primary cells, immortalized cells or tumor cell lines on a collagen, or combined collagen/Matrigel, scaffold containing fibroblasts3,4. It is less labor intensive to generate these scaffolds than the acellular esophageal scaffold described in this manuscript, and these organotypic models provide a useful tool, particularly in the study of tumor invasion5,6, where tumor cell infiltration into the collagen gel can be readily observed. However these collagen gels have non-native mechanical properties and lack certain features of the original tissue, including a specific basement membrane and the appropriate surface topography. This can influence the behavior of cells resulting in, for example, poorer adhesion between the epithelium and scaffold when using a collagen gel scaffold7. As a consequence the acellular porcine esophageal scaffold was developed, with the advantage of being a more biologically realistic scaffold and thus more appropriate for use as an experimental platform. It has also been shown that it is better to incorporate primary cells into the esophageal constructs than immortalized esophageal epithelial cell lines, such as Het-1A, since these cells form a multi-layered epithelium but fail to stratify or differentiate4,7,8.
Consequently, this protocol has been adapted from a method already in use in the MacNeil laboratory for making tissue engineered skin and oral mucosa9,10 and incorporates a de-cellularized porcine esophageal scaffold combined with primary human esophageal epithelial cells and fibroblasts. This protocol produces a mature, stratified epithelium, similar to that of the normal human esophagus as demonstrated by CK4, CK14, Ki67 and involucrin staining. The resulting model provides an experimental platform to study responses to environmental stressors, and has been used effectively to investigate changes in gene expression in the esophageal epithelium in response to refluxate components11.
Protocol
Human esophageal cells are obtained from patients undergoing gastric or esophageal surgery. Informed consent is obtained for the tissue to be used for research purposes, and the tissue used anonymously under the appropriate ethical approvals (SSREC 165/03, Human Research Tissue Bank Licence 12179).
1. Isolation of Human Esophageal Epithelial Cells
- Working in a laminar flow tissue culture hood and using sterile technique, prepare the epithelial culture medium by mixing Dulbecco’s Modified Eagle’s medium (DMEM) and Ham’s F12 in a 3:1 ratio (v:v). Add 10% (v/v) FCS, 10 ng/ml epidermal growth factor, 0.4 µg/ml hydrocortisone, 1.8 x 10-4 M adenine, 5 µg/ml insulin, 5 µg/ml transferrin, 2 x 10-7 M triiodothyronine, 1 x 10-10 M cholera toxin, 2 x 10-3 M glutamine, 100 IU/ml penicillin, 100 µg/ml streptomycin, and 0.625 µg/ml amphotericin B.
- Using standard sterile techniques, put 11 ml of DMEM, 10% FCS, 2 x 10-3 M L-glutamine, 100 IU/ml penicillin, 100 µg/ml streptomycin, and 0.625 µg/ml amphotericin B into a T75 tissue culture flask. Add 1 x 106 lethally irradiated mouse 3T3 fibroblasts12 in 1 ml of the same medium and incubate overnight at 37 °C in a 5% CO2, humidified atmosphere. This will produce a feeder layer for the subsequent culture of epithelial cells.
- Using a scalpel and following the recommendations of a histopathologist, dissect an approximately 2 cm2 sample of tissue from the disease-free background region of esophageal squamous mucosa obtained from patients undergoing gastric or esophageal surgery. Transport to the laboratory in a 50 ml container of sterile transport medium (PBS 100 IU/ml penicillin, 100 µg/ml streptomycin and 0.625 µg/ml amphotericin B) at room temperature.
- At this point, store the tissue in the transport medium for up to 12 hr (overnight) at 4 °C, for convenience; however do not use longer storage periods as they result in reduced cell viability.
- Working in a laminar flow tissue culture hood and using sterile technique, sterilize a scalpel blade by soaking in 70% ethanol for 5 min. Rinse in sterile PBS.
- Place the tissue in a sterile Petri dish and use the scalpel to cut it into 0.5 cm strips. Add 5 ml of 0.1% (w/v) trypsin in PBS to the Petri dish, place the lid on the dish and incubate for 1 hr at 37 °C.
- Add 5 ml of FCS to the tissue in the Petri dish to inhibit the trypsin. Holding the strip of tissue with sterile forceps, gently scrape the epithelial surface with the scalpel blade for 1-2 min per strip to remove the epithelial cells into the surrounding medium. Remove the scraped tissue and set aside. Repeat the process for all the tissue strips.
- Collect the medium containing the detached epithelial cells into a 15 ml centrifuge tube and centrifuge (5 min, 200 x g). Pour off the supernatant and resuspend the pellet in 12 ml of epithelial medium (described in step 1.1).
- Pour off and discard the medium from the feeder cell layer in the T75 flask prepared in step 1.2. Use a sterile pipette to add the cell suspension containing the freshly isolated epithelial cells to the flask and incubate at 37 °C in a 5% CO2, humidified atmosphere.
- After 24 hr pour off and discard the culture medium, which will also contain cell debris, and replace with 12 ml of fresh epithelial medium. Continue incubating at 37 °C in a 5% CO2, humidified atmosphere. Colonies will start to appear over the next 24 to 48 hr and can be viewed by placing the culture flask under a standard light microscope. Pour off the spent culture medium and replace with an equal volume of fresh medium every 2 to 3 days.
- Once the flask has reached 80% confluency, passage the cells13. Split the cells in a ratio of 1:5 to increase cell numbers for use in the model. Follow step 1.2 to establish feeder layers of irradiated 3T3 cells in each flask that will be receiving epithelial cells 24 hr prior to passaging the cells.
Note: Use the epithelial cells in the esophageal model described below between passages 1 and 4 only.
2. Isolation of Human Esophageal Fibroblasts
- Working in a laminar flow cabinet and using sterile technique, take the tissue set aside in step 1.7. Place in a sterile Petri dish and mince finely by chopping with a sterile scalpel blade. Add 10 ml of 0.5% (w/v) collagenase A solution and place the lid on the Petri dish and incubate at 37 °C overnight in a 5% CO2, humidified atmosphere.
- Transfer the digest to a 15 ml centrifuge tube and centrifuge (10 min, 200 x g), carefully pour off and discard the supernatant and resuspend the pellet in 10 ml of fibroblast culture medium (DMEM 10% FCS, 2 x 10-3 M L-glutamine, 100 IU/ml penicillin, 100 µg/ml streptomycin, and 0.625 µg/ml amphotericin B).
- Place the suspension into a T75 flask and incubate at 37 °C in a 5% CO2, humidified atmosphere. After 24 hr pour off the medium, which will contain debris and replace with 12 ml of fresh fibroblast medium. Replace the culture medium every 2 to 3 days and passage cells once they have reached 80% confluency13, splitting the cells in a 1:5 ratio to increase cell numbers.
Note: Use the fibroblast cells in the model described below between passages 4 and 10 only.
3. Preparation of the De-cellularized Esophageal Scaffold
- Obtain intact porcine esophagi at an abattoir from freshly slaughtered Landrace pigs that are used for food production. One single esophagus will provide sufficient scaffolds for approximately 30 separate constructs.
Note: Due to the extensive and time consuming sterilization protocol required, it is normally worth obtaining and processing a minimum of 10 esophagi at one time. - Immediately place the freshly excised esophagus into a 180 ml sterile pot containing approximately 150 ml of 10% povidone-iodine solution for 5 min to reduce any contaminating microbial load. Transfer the esophagus to a second pot containing approximately 100 ml of sterile PBS containing 200 IU/ml penicillin, 200 µg/ml streptomycin, and 1.25 µg/ml amphotericin B.
- Replace the lid on the container and transport the esophagi back to the laboratory at room temperature. Two or three esophagi can be transported in each container, depending upon their size.
- Once at the laboratory, handle the tissue in a laminar flow hood using aseptic technique to minimize microbial contamination. Sterilize tweezers, scissors and scalpel blades by soaking for 5 min in 70% ethanol, and rinsing in sterile PBS.
- Handle the esophagus using sterile tweezers. Using the scissors, cut open the esophagus longitudinally. Rinse the tissue by placing the esophagus into a 180 ml sterile pot containing approximately 100 ml of sterile PBS and shaking gently to remove any debris.
- Clean a cork dissection board by spraying liberally with 70% ethanol and allow to dry.
- Remove the esophagus from the PBS, and using sterile needles, pin out the esophagus onto the board, mucosal surface uppermost. Grasp the mucosal surface at one end of the esophagus with sterile tweezers and pull away from the board. This will separate the mucosa from the underlying submucosa.
- Then, use a sterile scalpel blade to dissect the esophagus longitudinally along the lamina propria, removing pins as the dissection progresses to allow the mucosa to be lifted away.
- Retain the dissected mucosa and discard the underlying submucosa.
- Cut off and discard the most proximal and distal 2 cm of the mucosa as there is a greater number of submucosal glands and a thicker lamina propria in those areas, respectively7.
- Use a scalpel to cut the remaining tissue into 5 cm2. Place all the cut tissue into a 180 ml sterile pot containing approximately 150 ml of sterile PBS and agitate gently to rinse the tissue. If processing more than five esophagi at one time it may be necessary to use multiple pots for this and the three subsequent steps.
- Using sterile tweezers, transfer the cut tissue into a 180 ml pot containing 150 ml of sterile 1 M NaCl, 200 IU/ml penicillin, 200 µg/ml streptomycin and 1.25 µg/ml amphotericin B. Incubate for 72 hr at 37 °C.
- Place the tissue into a fresh 180 ml pot containing 150 ml sterile PBS. The epithelium will have begun to visibly separate from the underlying tissue. Gently peel the detaching epithelium from the pig esophagus using sterile forceps and discard.
- Place the remaining tissue into a fresh 180 ml pot and add 150 ml of sterile PBS. Agitate gently for 5 min to wash. Pour off the PBS and repeat a further two times with fresh PBS.
- Place all the tissue into a 500 ml glass bottle containing 400 ml of sterile 80% (v/v) glycerol solution for 24 hr at room temperature. Transfer to 400 ml of sterile 90% (v/v) glycerol solution for a further 24 hr.
- Store in 400 ml of sterile 100% glycerol solution at room temperature for a minimum of 4 months to dehydrate and sterilize the tissue while preserving the integrity of the basement membrane.
4. Production of the Culture Media
- Prepare chelated newborn calf serum for use in the culture medium.
- Pour 100 g of Chelex 100 into a glass 1 L bottle and add de-ionized water. The exact volume is not critical but sufficient water must be added to cover the Chelex powder. Add a magnetic stirrer and sterilize by autoclaving (121 °C for 15 min).
- Allow the bottle to cool and the Chelex to settle. In a laminar flow cabinet pour off the excess water, add 500 ml of newborn calf serum (NCS) and leave to stir overnight at 4 °C.
- Stop stirring and allow the Chelex to settle, this can take up to 30 min. In a laminar flow cabinet decant the chelated serum using a sterile pipette, taking care not to disturb the Chelex, and store in 10 ml aliquots at -20 °C.
- Prepare three different media in a laminar flow cabinet, commencing with pro-proliferative and ending with pro-differentiative formulations.
- Prepare Composite Medium I comprised of DMEM and Hams F12 in a 3:1 ratio (v:v), 4 x 10-3 M L-glutamine, 0.5 µg/ml hydrocortisone, 1 x 10-4 M O-phosphorylethanolamine, 2 x 10-12 M triiodothyronine, 1.8 x 10-4 M adenine, 1.88 x 10-3 M CaCl2, 4 x 10-12 M progesterone, 10 µg/ml insulin, 10 µg/ml transferrin, 10 ng/ml selenium, 1 x 10-3 M ethanolamine and 0.1% (v/v) chelated NCS.
- Prepare Composite Medium II as Composite Medium I except replace chelated serum with 0.1% (v/v) non-chelated NCS.
- Prepare Composite Medium III as Composite Medium II except omit progesterone and increase serum to 2% (v/v) non-chelated NCS.
5. Production of the Human Esophageal Mucosa Model
- Perform all work in a laminar flow hood, using aseptic technique to maintain a sterile environment. Sterilize all tools by soaking in 70% ethanol for 5 min, before rinsing in sterile PBS.
- The day before it is required, rehydrate the acellular porcine esophageal scaffold. One piece of scaffold is required for each construct to be prepared.
- To rehydrate the tissue place sufficient pieces of porcine scaffold into a pot containing 100 ml of sterile PBS, agitate and soak for 10 min. Decant the PBS and replace with fresh PBS. Repeat the washing process five times to ensure removal of all glycerol traces.
- Test the scaffold sterility by incubating the rehydrated tissue overnight in 100 ml of DMEM at 37 °C. If there is no change to the color or turbidity of the culture medium at the end of the incubation period the scaffold is assumed to be sterile. Once sterility has been confirmed, place the 5 cm2 acellular scaffold into a 6 well plate, submucosal side uppermost.
- Place a sterile medical grade stainless steel ring (internal diameter 10 mm, external diameter 20 mm) onto the center of each scaffold and gently press down using sterile forceps, to ensure an adequate seal with the tissue.
- Harvest the fibroblasts using Trypsin-EDTA13 to produce a cell suspension. Count the cells using a haemocytometer14. Centrifuge the cell suspension (10 min, 200 x g). Resuspend the cell pellet into an appropriate volume of fibroblast medium to produce a final cell count of 2.5 x 106 cells per ml.
- Add 0.2 ml of fibroblast medium, containing 5 x 105 human esophageal fibroblasts, within each ring. Flood the area around the outside of the ring with approximately 2 ml of fibroblast medium. Incubate at 37 °C in a 5% CO2, humidified atmosphere.
- After 24 hr remove the rings and fibroblast medium and add at least 5 ml of fresh fibroblast medium to each well, ensuring that the scaffold is totally immersed in medium. Culture for 1 week, at 37 °C in a 5% CO2, humidified atmosphere replacing the medium every 2 to 3 days.
- After 1 week remove the 6 well plates containing the scaffolds to a laminar flood hood, remove the medium and invert each scaffold using sterile forceps, placing the mucosal surface uppermost. This time point is referred to as Day 0.
- Place the steel rings on the mucosal surface in the center of the tissue as in step 5.4.
- Prepare the T75 flasks of epithelial cells for trypsinization13. After the PBS wash and prior to the addition of trypsin-EDTA solution, add 2 ml of sterile 0.02% (w/v) EDTA solution to each flask. Incubate for 2 min at 37 °C. Tap the flask gently to selectively detach the i3T3 feeder layer while leaving the epithelial cells attached.
Note: It is generally not practical or necessary to patient match the epithelial cells to the fibroblasts already incorporated in the model the previous week. - Pour off the EDTA solution containing the feeder layer and add trypsin-EDTA solution to harvest the epithelial cells13. Count the cells using a haemocytometer14. Centrifuge the cell suspension (10 min, 200 x g). Resuspend the cell pellet into an appropriate volume of Composite Medium 1 to produce a final cell count of 5 x 106 cells per ml.
- Add 0.2 ml of Composite Medium I, containing 1 x 106 epithelial cells, within the ring. Flood the area around the outside of the ring with approximately 2 ml of Composite Medium I and place the constructs into the incubator at 37 °C in a 5% CO2, humidified atmosphere.
- After 24 hr (Day 1) remove the rings and medium and add at least 5 ml of fresh Composite Medium I, ensuring the scaffolds are fully submerged and return to the incubator. After a further 24 hr (Day 2) remove the medium and replaced with at least 5 ml of Composite Medium II, again ensuring the scaffolds are fully submerged and return to the incubator.
- On Day 4 place sterile medical grade stainless steel mesh grids (2 cm wide x 2 cm long x 0.5 cm high), into fresh 6 well plates, one per construct. Use sterile tweezers to transfer the constructs onto the top of the steel grids, mucosal surface uppermost.
- Add sufficient Composite Medium III so that the liquid reaches the underside of the composite, but the surface is exposed to the air, this ensures that the sample is maintained at an air-liquid interface. The exact volume of medium required will vary depending upon the thickness of the scaffold, the volume of the well and the precise height of the stainless steel grid.
- Maintain the composites at an air-liquid interface for between 10 and 20 days (Day 14 to Day 24), depending upon the requirements of the experiment. Replace the Composite Medium III with fresh medium every 2 to 3 days (Monday, Wednesday, Friday is a user-friendly regime).
Note: After 5 days at an air-liquid interface (Day 9) the constructs have a sufficiently mature esophageal epithelium to be used in experiments of the impact of environmental factors on the esophageal epithelium. - At the end of the experiment, fix the constructs for histological analysis and immunohistochemical staining7, or extract protein15 or RNA11,16 from the epithelium for further analysis. If required, retain the conditioned culture medium for analysis of extracellular signaling molecules17.
Representative Results
This manuscript describes the process required, shown in schematic form in Figure 1, to culture 3D models of the human esophageal epithelium successfully. To confirm the suitability of the model as an experimental platform histological and immunohistochemical studies have been undertaken comparing the cultured tissues with normal human esophageal squamous mucosa.
Histological assessment of the epithelium produced by the method described shows a mature, multi-layered, stratified squamous epithelium (Figure 2B) which is comparable to that observed with the normal human esophagus (Figure 2A), albeit thinner (5 to 10 layers of cells compared with 10 to 20 for the normal esophagus), with the cells becoming progressively flatter and ultimately anuclear as they migrate towards the surface.
Immunohistochemical characterization of key markers of proliferation and differentiation demonstrate that the microanatomy of the model epithelium is similar to the normal human esophageal epithelium. Comparable Ki67 expression is observed in both the native esophagus and the model epithelium, with staining restricted to a subset of cells within the basal and immediately suprabasal layers (Figures 3A and 3B). This is analogous to studies which report that less than 10% of cells generally show expression of the proliferation marker, Ki67, in the normal esophageal epithelium18. CK4 is normally expressed in stratified and columnar epithelia but is generally absent from the basal layers while CK14 is only positive in the basal layer19. In both the normal and model esophageal epithelia, CK14 is observed in all cells in the basal layer (Figures 3D and 3E) while CK4 is observed throughout the epithelium except for the two most basal layers (Figures 3G and 3H). Involucrin expression is an early marker of differentiation, expressed in the suprabasal layers of the epithelium20. Again both the normal human esophageal tissue and the model epithelium show staining that reflects this (Figures 3J and 3K).
Attempts to replace the primary esophageal epithelial cells with immortalized esophageal epithelial cells, such as Het-1A, were less successful and did not produce a valid model of the normal esophageal epithelium. A multi-layered epithelium was formed; however the proliferative marker Ki67 was detected throughout the epithelium (Figure 3C) with no expression detected for any markers of differentiation (Figures 3F, 3I and 3L), indicating that the Het-1A cells produced a hyperproliferative epithelium with no evidence of normal stratification or maturation.
However, the model has been successfully modified to incorporate tumor cells by the replacement of primary epithelial cells with either esophageal adenocarcinoma (OE33) or squamous carcinoma (OE21) cells. This demonstrates the flexibility of the model, enabling its use in investigating a range of esophageal disorders at different stages of progression through the inclusion of a number of different cell lines. It can be seen that there is a marked difference in the responses from these two cell lines. OE21 squamous carcinoma cells produces an epithelium visible on the construct as a defined yellow region (Figure 4A) with large clefts within the epithelium (Figure 4B), likely to reflect dysfunctional cell adhesion molecules. Including OE33 adenocarcinoma cells within the model results in a large amount of scaffold degradation, visible by eye as a thinning of the scaffold after 2 weeks growth at the air/liquid interface (Figure 4C) and confirmed in H+E analysis as an obvious reduction in the thickness of the scaffold in the region below the cells (Figure 4D). This is a likely to be a result of an interaction between the tumor cells and fibroblasts, since the scaffold degeneration is not observed in the absence of fibroblasts (Figures 4E and 4F). We have observed similar impacts on tumor invasion in the presence and absence of fibroblasts in equivalent melanoma models21.
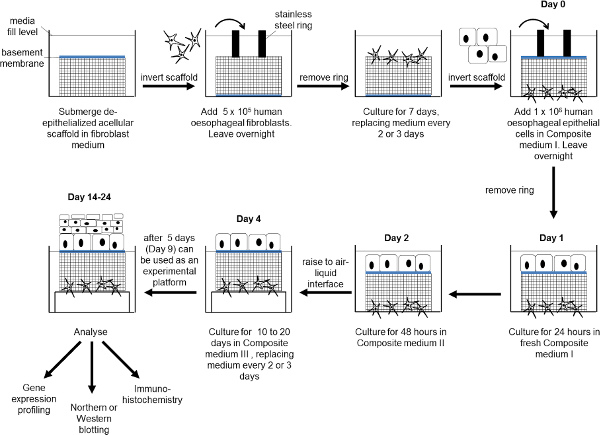
Figure 1: Schematic diagram showing the production of the esophageal mucosa model. Human esophageal fibroblasts are seeded onto the submucosal surface of the scaffold and cultured for 7 days. The scaffold is inverted, human esophageal epithelial cells added and cultured submerged for 4 days. The construct is raised to an air-liquid interface for between 10 and 20 days. At the end of the experiment the construct can undergo further analysis as required. Please click here to view a larger version of the figure.
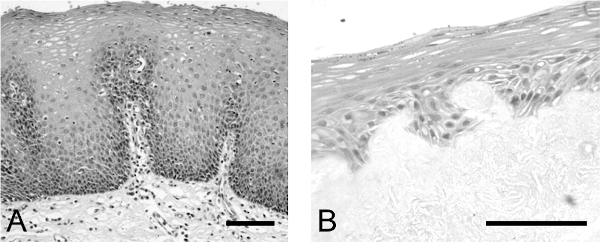
Figure 2: Comparison of epithelium produced in esophageal model with normal human esophageal epithelium. H+E analysis of (A) normal human esophageal epithelium and (B) esophageal epithelium formed in the model of the human esophageal mucosa. Scale bar is 500 µm.Please click here to view a larger version of the figure.
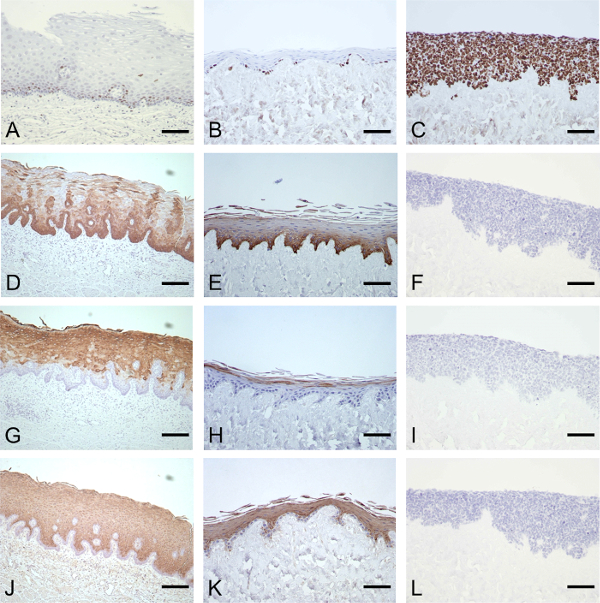
Figure 3: Immunohistochemical characterization of the esophageal epithelium produced in the model and comparison with the normal human esophagus. IHC analysis of normal esophagus (column 1) and the model esophagus produced using primary human esophageal epithelial cells (column 2) or immortalized esophageal epithelial cells (column 3). Characterization was by Ki67 (A–C), CK14 (D, E), CK4 (G, H) and involucrin staining (J, K). Scale bar is 200 µm. (This figure has been modified7.) Please click here to view a larger version of the figure.
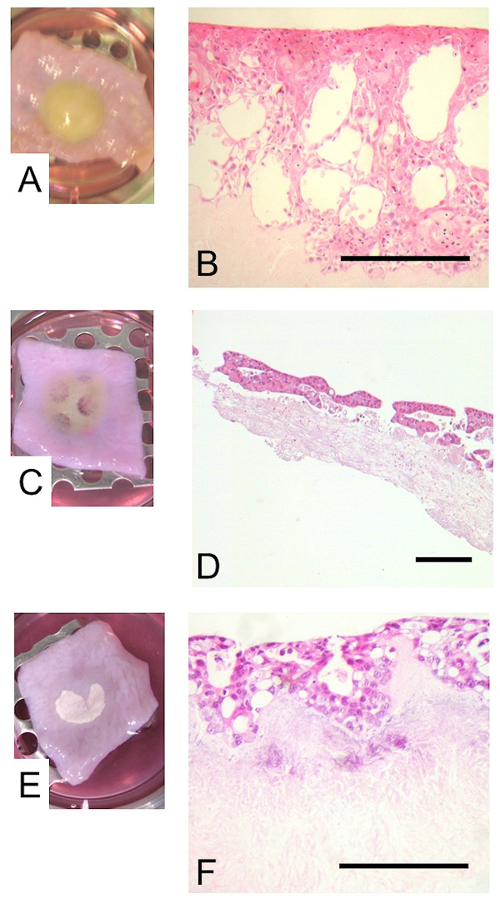
Figure 4: The inclusion of tumor cell lines within the esophageal model. The primary epithelial cells were replaced by the squamous carcinoma cell line, OE21, or the adenocarcinoma cell line, OE33. Images show the construct with the OE21 cells (A) or OE33 cells either in conjunction with (C) or in the absence of (E) fibroblasts, prior to fixing at the end of the culture period. The epithelia including the OE21 (B) or OE33 cells with (D) or without (F) fibroblasts are visualized by H+E staining. Scale bar is 500 µm. Please click here to view a larger version of the figure.
Discussion
This manuscript describes the production and characterization of a biologically relevant human esophageal mucosal model suitable for use as an experimental platform to study the impact of exposure to environmental stressors upon the esophageal epithelium.
The most critical steps for the successful production of a human esophageal mucosal model are: ensuring that the majority of the epithelial cells remain proliferative and have not already begun to differentiate prior to seeding them on the scaffold; maintaining an air-liquid interface for the composite to ensure correct maturation of the epithelium; maintaining sterility throughout the extended culture period.
Due to the limited amounts of human tissue available for cell isolation it is not generally possible to obtain sufficient cells from a single tissue sample to seed freshly isolated cells directly onto the scaffold and consequently a cell expansion phase is required where the cells are grown in standard 2D cell culture conditions. It is important to ensure that cells remain proliferative during this period. This is achieved by ensuring that the cultures are never allowed to reach full confluency, using cells in the model only up to passage 4 and carefully monitoring cell morphology. Thus cells should retain their distinctive proliferative polygonal morphology and the tight “cobblestone” pavement-like appearance characteristic of proliferative epithelial cells rather than the more diffuse appearance observed in cultures which have begun to differentiate. The second critical step is controlled by ensuring that the cell culture medium covers the top surface of the stainless steel grids used to lift the cultures to air-liquid interface but the uppermost surface of the construct itself is not submerged. The third step is dependent upon strict adherence to the extended sterilization protocol when producing the de-cellularized porcine scaffold and the use of good sterile technique throughout the process.
Our results show that primary human esophageal cells are required to produce a normal, mature, stratified epithelium. If an immortalized cell line was used, albeit derived from the human esophageal epithelium, the cells remained proliferative and failed to differentiate to form a mature stratified epithelium. The resulting epithelium could not be used as a satisfactory model for the normal epithelium, demonstrating the unsuitability of the immortalized cell line Het-1A for use in the model. However other cell lines such as those derived from esophageal tumors or Barrett’s Metaplasia can be incorporated into the model either in place of or in addition to the primary epithelial cells to produce models for the study of tumor progression, invasion and the response to pharmacological agents.
Experiments can be performed using the esophageal model to simulate the exposure of the esophagus to discrete, pulsatile events during swallowing or reflux. This is achieved by repeatedly submerging the constructs in culture medium containing the compound(s) to be tested and rinsing in PBS before returning the construct to an air-liquid interface. The exposure times and frequency can be modified to reflect the process being simulated, ensuring that the epithelial layer is exposed to the environmental factor(s) while allowing the culture to continue at an air-liquid interface. This method has been used in our laboratory to investigate the impact of reflux on the normal esophageal epithelium, where the model was exposed to specific gastro-esophageal reflux components for 10 minutes twice a day for 11 days11. Alternatively the constructs can be exposed to continuous levels of environmental stressors by including these compounds in the culture medium. However, since the epithelium must be exposed to an air-liquid interface for the epithelium to mature and differentiate properly7, it is not possible to continuously submerge the constructs in the culture medium. Consequently, exposure to the stressor will only be via the submucosal surface in these experiments, which may limit the relevance of results obtained.
The technique is relatively labor intensive and consequently is better suited to the screening of relatively limited numbers of compounds. As a result, when using the model as an experimental platform, it is appropriate to first perform preliminary studies using conventional cell culture techniques to identify a limited number of conditions prior to further analysis using the more physiologically relevant esophageal model described here11. In addition to immunohistochemical analysis it has also been possible to examine changes in the epithelial gene expression profile following exposure to environmental stressors. This was achieved by manually stripping away the epithelium, extracting the RNA and converting it to cDNA to isolate the genes being expressed and analyzing by microarray to obtain gene expression profiles11. In this way it has been possible to increase the information available from the model as an experimental platform. Early changes in the epithelial gene expression profile following simulated exposure to gastro-esophageal reflux have been proposed and from these results new areas have been suggested for further investigation in the prevention of the development of Barrett’s Metaplasia, the metaplastic precursor to OAC11. The model could also be used to study protein expression using methods such as Western blot analysis15 and gene expression using Northern blot analysis16. Moreover studies of extracellular signaling molecules could be performed on the culture medium17.
Divulgations
The authors have nothing to disclose.
Acknowledgements
We are grateful to Mr Roger Ackroyd, Mr Andrew Wyman and Mr Chris Stoddard, Consultant Surgeons at Sheffield Teaching Hospitals NHS Foundation Trust, for their help in acquiring esophageal tissue samples and their support of our work. We thank Ashraful Haque for his help incorporating tumor cell lines into the model. We gratefully acknowledge financial support for this study by grants from the Bardhan Research and Education Trust (BRET) and Yorkshire Cancer Research (YCR).
Materials
| Trypsin | BD Biosciences | 215240 | Prepare 0.1% w/v solution in PBS and filter sterilise. Warm in 37°C water bath before use |
| DMEM | Labtech | LM-D1112 | Warm in 37°C water bath before use |
| Ham's F12 | Labtech | LM-H1236 | Warm in 37°C water bath before use |
| Foetal Calf Serum | Labtech | FB-1090 | |
| Epidermal Growth Factor | R+D Systems | 236-EG-200 | Prepare 200 µg/ml stock solution in 10 mM acetic acid, 1% FCS |
| Hydorcortisone | Sigma-Aldrich | H0396 | Prepare stock solution in PBS and filter sterilise before use |
| Adenine | Sigma-Aldrich | A2786 | Prepare stock solution in PBS and filter sterilise before use |
| Insulin | Sigma-Aldrich | I2767 | Prepare 10 mg/ml solution in 0.01M HCl, dilute 1:10 in distilled water and filter sterilise before use |
| Transferrin | Sigma-Aldrich | T2036 | Prepare stock solution in distilled water and filter sterilise before use |
| Triiodothyronine | Sigma-Aldrich | T2752 | Prepare stock solution in distilled water and filter sterilise before use |
| Cholera toxin | Sigma-Aldrich | C8052 | Prepare stock solution in water |
| L-Glutamine | Sigma-Aldrich | G7513 | |
| Penicillin-Streptomycin | Sigma-Aldrich | P0781 | |
| Amphotericin B | Gibco | 15290-026 | Brand name Fungizone |
| PBS | Oxoid | BR0014 | Dissolve 1 tablet in 100 ml water and autoclave to sterilise |
| Collagenase A | Roche | 10103578001 | |
| Povidone-iodine solution | Ecolab | 10830E | Brand name Videne |
| Ethanol | Sigma-Aldrich | E7023 | |
| NaCl | Sigma-Aldrich | 433209 | Prepare 1M solution and autoclave to sterilise before use (121 ˚C for 15 min) |
| Glycerol | Sigma-Aldrich | G2025 | Autoclave to sterilise before use (121 ˚C for 15 min) |
| Chelex 100 | Sigma-Aldrich | C7901 | |
| Newborn calf serum | Gibco | 26010074 | |
| Progesterone | Sigma-Aldrich | P8783 | Prepare stock solution in DMEM and filter sterilise before use |
| Ethanolamine | Sigma-Aldrich | E9508 | Prepare stock solution in DMEM and filter sterilise before use |
| Hydrocortisone | Sigma-Aldrich | H0888 | Prepare stock solution in DMEM and filter sterilise before use use |
| O-phosphorylethanolamine | Sigma-Aldrich | P0503 | Prepare stock solution in DMEM and filter sterilise before use |
| ITS (insulin, transferrin, selenium) | Lonza | 17-838Z | Used for composite media preparation |
| Trypsin-EDTA | Sigma-Aldrich | T3924 | Warm in 37°C water bath before use |
| EDTA 0.02% solution | Sigma-Aldrich | E8008 | Warm in 37°C water bath before use |
| T75 culture flask | VWR | 734-2313 | |
| 50 ml centrifuge tube | Fisher | 11819650 | |
| 15 ml universal tube | SLS | SLS7504 | |
| 180 ml pot | VWR | 216-2603 | |
| Petri dish | SLS | 150350 | |
| 6 well plate | VWR | 734-2323 | |
| stainless steel rings | Manufactured in house – medical grade stainless steel, internal diameter 10 mm, external diameter 20 mm | ||
| steel mesh grids | Manufactured in house – sheets have 0.3 cm diameter holes, bent to produce grid 2cm (w) x2 cm (d) x 0.5 cm (h) | ||
| ki67 | Novocastra | KI67-MM1-L-CE | Clone MM1 Use at 1:100 |
| CK4 | Abcam | ab9004 | Clone 6B10 Use at 1:200 |
| CK14 | Novocastra | LL002-L-CE | Clone LL002 Use at 1:200 |
| Involucrin | Novocastra | INV | Clone SY5 Use at 1:100 |
| OE21 | Sigma-Aldrich | 96062201 | |
| OE33 | Sigma-Aldrich | 96070808 | |
| Het-1A | ATCC-LGC | CRL-2692 | |
| Mouse 3T3 fibroblasts | ATCC-LGC | CRL-1658 | previously growth arrested by irradiation (60 Gy) |
References
- Pera, M., Manterola, C., Vidal, O., Grande, L. Epidemiology of esophageal adenocarcinoma. J. Surg. Oncol. 92 (3), 151-159 (2005).
- Manea, G. S., Lupusoru, C., Caruntu, I. D. Experimental study on the effect of bile, and bile and hydrochloric acid mixture on the esophageal mucosa. Rom. J. Morphol. Embryol. 50 (1), 61-66 (2009).
- Andl, C. D., et al. Epidermal growth factor receptor mediates increased cell proliferation, migration, and aggregation in esophageal keratinocytes in vitro and in vivo. J. Biol. Chem. 278 (3), 1824-1830 (2003).
- Underwood, T. J., et al. A comparison of primary oesophageal squamous epithelial cells with HET-1A in organotypic culture. Biol. Cell. 102 (12), 635-644 (2010).
- Nystrom, M. L., et al. Development of a quantitative method to analyse tumour cell invasion in organotypic culture. J. Pathol. 205 (4), 468-475 (2005).
- Okawa, T., et al. The functional interplay between EGFR overexpression, hTERT activation, and p53 mutation in esophageal epithelial cells with activation of stromal fibroblasts induces tumor development, invasion, and differentiation. Genes & Development. 21 (21), 2788-2803 (2007).
- Green, N., et al. The development and characterization of an organotypic tissue-engineered human esophageal mucosal model. Tissue Eng Part A. 16 (3), 1053-1064 (2010).
- Harada, H., et al. Telomerase induces immortalization of human esophageal keratinocytes without p16INK4a inactivation. Mol. Cancer Res. 1 (10), 729-738 (2003).
- Bhargava, S., Chapple, C. R., Bullock, A. J., Layton, C., MacNeil, S. Tissue-engineered buccal mucosa for substitution urethroplasty. BJU. Int. 93 (6), 807-811 (2004).
- Ralston, D. R., et al. Keratinocytes contract human dermal extracellular matrix and reduce soluble fibronectin production by fibroblasts in a skin composite model. Br. J. Plast. Surg. 50 (6), 408-415 (1997).
- Green, N. H., et al. Pulsatile exposure to simulated reflux leads to changes in gene expression in a 3D model of oesophageal mucosa. Int. J. Exp. Pathol. 95 (3), 216-228 (2014).
- Wang, C. S., et al. Selective culture of epithelial cells from primary breast carcinomas using irradiated 3T3 cells as feeder layer. Pathol. Res. Pract. 197 (3), 175-181 (2001).
- Ricardo, R., Phelan, K. Trypsinizing and subculturing mammalian cells. J. Vis. Exp. (16), e755 (2008).
- . Using a Hemacytometer to Count Cells. Basic Methods in Cellular and Molecular Biology. , (2014).
- Mahmood, T., Yang, P. C. Western blot: technique, theory, and trouble shooting. N. Am. J. Med. Sci. 4 (9), 429-434 (2012).
- Streit, S., Michalski, C. W., Erkan, M., Kleeff, J., Friess, H. Northern blot analysis for detection and quantification of RNA in pancreatic cancer cells and tissues. Nat. Protoc. 4 (1), 37-43 (2009).
- Morgan, E., et al. Cytometric bead array: a multiplexed assay platform with applications in various areas of biology. Clin. Immunol. 110 (3), 252-266 (2004).
- Xu, M., et al. The abnormal expression of retinoic acid receptor-beta, p 53 and Ki67 protein in normal, premalignant and malignant esophageal tissues. World J. Gastroenterol. 8 (2), 200-202 (2002).
- Moll, R., Divo, M., Langbein, L. The human keratins: biology and pathology. Histochem. Cell Biol. 129 (6), 705-733 (2008).
- Rice, R. H., Green, H. Presence in human epidermal cells of a soluble protein precursor of the cross-linked envelope: activation of the cross-linking by calcium ions. Cell. 18 (3), 681-694 (1979).
- Eves, P., et al. Melanoma invasion in reconstructed human skin is influenced by skin cells – investigation of the role of proteolytic enzymes. Clin. Exp. Metastasis. 20 (8), 685-700 (2003).

