Detection of a Circulating MicroRNA Custom Panel in Patients with Metastatic Colorectal Cancer
Summary
We present a protocol to evaluate the expression levels of circulating microRNA (miRNAs) in plasma samples from cancer patients. In particular, we used a commercially available kit for circulating miRNA extraction and reverse transcription. Finally, we analyzed a panel of 24 selected miRNAs using real-time pre-spotted probe custom plates.
Abstract
There is increasing interest in liquid biopsy for cancer diagnosis, prognosis and therapeutic monitoring, creating the need for reliable and useful biomarkers for clinical practice. Here, we present a protocol to extract, reverse transcribe and evaluate the expression levels of circulating miRNAs from plasma samples of patients with colorectal cancer (CRC). microRNAs (miRNAs) are a class of non-coding RNAs of 18-25 nucleotides in length that regulate the expression of target genes at the translational level and play an important role, including that of pro- and anti-angiogenic function, in the physiopathology of various organs. miRNAs are stable in biological fluids such as serum and plasma, which renders them ideal circulating biomarkers for cancer diagnosis, prognosis and treatment decision-making and monitoring. Circulating miRNA extraction was performed using a rapid and effective method that involves both organic and column-based methodologies. For miRNA retrotranscription, we used a multi-step procedure that considers polyadenylation at 3' and the ligation of an adapter at 5' of the mature miRNAs, followed by random miRNA pre-amplification. We selected a 24-miRNA custom panel to be tested by quantitative real-time polymerase chain reaction (qRT-PCR) and spotted the miRNA probes on array custom plates. We performed qRT-PCR plate runs on a real-time PCR System. Housekeeping miRNAs for normalization were selected using GeNorm software (v. 3.2). Data were analyzed using Expression suite software (v 1.1) and statistical analyses were performed. The method proved to be reliable and technically robust and could be useful to evaluate biomarker levels in liquid samples such as plasma and/or serum.
Introduction
CRC represents the third most frequently diagnosed malignancy and the fourth cause of cancer-related death worldwide. To date, bevacizumab (B), a monoclonal antibody directed against vascular-endothelial growth factor (VEGF), and cetuximab (C) or panitumumab (P), monoclonal antibodies directed against epidermal growth factor receptor (EGFR), have been approved for first-line treatment in combination with chemotherapy (CT) regimens.
Mutations in K-RAS and N-RAS genes are the only clinically useful biomarkers capable of identifying patients who are least likely to benefit from anti-EGFR-based CT. Although several studies have been conducted to search for biomarkers that are predictive of response to B-based CT, there is still a substantial lack of reliable and effective drugs for use in clinical practice1.
miRNAs are small RNAs (18-25 nucleotides in length) that regulate the translation of target genes and play a crucial role in numerous physiopathological processes, including embryogenesis and carcinogenesis. These molecules are highly stable in biological fluids such as plasma/serum, urine and sputum, which renders them robust biomarkers to use for non-invasive sampling2. Using a panel of miRNAs involved in the angiogenic pathway, we aimed to identify new circulating biomarkers capable of predicting clinical outcome in patients with metastatic CRC (mCRC) treated with a B-based CT regimen.
We analyzed a series of 52 mCRC patients treated with B-based CT within the prospective multicenter randomized phase III trial "Italian Trial in Advanced Colorectal Cancer" (ITACa). The present protocol was approved by the Local Ethics Committee (Comitato Etico Area Vasta e Istituto Scientifico Romagnolo per lo Studio e la Cura dei Tumori (IRST) IRCCS, no. 674) on 19th September 2007. All patients gave informed consent before blood sample collection. For each patient, venous blood samples were collected before treatment and at the first clinical evaluation (after 8 weeks) to evaluate baseline miRNA expression and its modulation during treatment in relation to patient outcome.
Through a search of the literature, we selected 21 miRNAs correlated with the angiogenic process that are known to be detectable in human plasma: hsa-miR-107, hsa-miR-126-3p, hsa-miR-145-5p, hsa-miR-194-5p, hsa-miR-199a-5p, hsa-miR-200b-3p, hsa-miR-20b-5p, hsa-miR-21-5p, hsa-miR-210-3p, hsa-miR-221-3p, hsa-miR-24-3p, hsa-miR-27a-3p, hsa-miR-29b-3p, hsa-miR-335-5p, hsa-miR-424-5p, hsa-miR-497-5p, hsa-miR-520d-3p, hsa-miR-92a-3p, hsa-miR-17-5p and hsa-miR-155-5p. We also selected hsa-miR-223-3p and hsa-mir-484 for endogenous normalization3,4,5,6, and cel-miR-39 purified from C. elegans as a spike-in for exogenous normalization. All data normalizations were performed using the 2 ̂ ̄(ΔΔCt) method.
The detection of circulating miRNAs presents some technical difficulties because the molecules are present at very low levels in plasma and their amplification is chemically challenging given their short sequence. For these reasons, we selected a procedure that considers both organic extraction with phenol and a glass-fiber column-based methodology. Circulating miRNA extraction is a hot topic in the field of liquid biopsy, and we selected a commercial kit that has shown to be one of the most reliable in terms of amount and quality of recovered yields7,8. We selected a protocol to reverse transcribe miRNAs by the addition of an adapter at 5' and a poly(A) tail to 3' of the mature miRNA to enhance the selectivity and specificity of the reaction.
Given the robustness of the method, we designed custom plates with pre-spotted probes for RT-PCR to assess each sample in duplicate and analyzed 2 patients within each plate, as shown in Figure 1.
Protocol
NOTE: Perform all of the following steps under a sterilized fume hood according to Good Laboratory Practice (GLP).
1. Plasma Collection and Storage
- Collect 3 mL of peripheral blood sample in a K3E EDTA tube.
- Centrifuge the tube at room temperature at 1,880 x g for 15 min.
- Recover the supernatant plasma in aliquots of 450 µL.
- Store the samples at -80 °C until use.
NOTE: Process the peripheral blood tube within 2 h of collection. When recovering the plasma from the tube, do not disturb the lymphocyte layer.
2. Extraction of Circulating miRNAs from Plasma Samples (Table of Materials)
- Prepare all the required solutions and thaw the samples for the extraction steps.
- Add 375 µL of 2-mercaptoethanol to the 2x denaturating solution and mix well.
- Add 21 mL of ACS grade 100% ethanol to the miRNA wash solution I and mix well.
- Add 40 mL of ACS grade 100% ethanol to the wash solution 2/3 and mix well.
- Allow a plasma aliquot to thaw at room temperature and then begin the extraction steps.
- Organic extraction
- Transfer 400 µL of the plasma sample into a RNase-free 2 mL tube.
- Add 400 µL of 2x denaturating solution, mix by vortexing and briefly centrifuge.
- Add 4 µL of 1 nM spike-in, mix by vortexing and briefly centrifuge.
- Add 800 µL of acid phenol-chloroform.
- Vortex for 60 s and centrifuge at maximum speed (≥10,000 x g) for 15 min.
- Transfer the upper aqueous phase to a fresh tube. Do not disturb the interphase. Note the volume recovered and discard the tube containing the non-acqueous phase.
NOTE: The 2x denaturing solution is stored at 4 °C and may solidify at this temperature. Before use, warm the solution to 37 °C, agitating occasionally for 10-15 min. Be careful to withdraw the bottom phase containing the acid phenol-chloroform, not the acqueous buffer that lies on top of the mixture. Preheat the elution solution or nuclease-free water to 95 °C. Be careful to heat an adequate volume of solution (100 µL per sample + 10% excess).
- Small RNA enrichment
- Add 1/3 volume of ethanol 100% to the aqueous phase from step 2.2.6 and mix thoroughly.
- Place a filter cartridge into a fresh collection tube for each sample.
- Transfer up to 700 µL of lysate from step 2.3.1 and ethanol into the filter cartridge and centrifuge at 10,000 x g for 30 s.
- If there is >700 µL of mixture left, place the filtrate in a fresh tube and repeat step 2.3.3; repeat until all the mixture has passed through the filter cartridge.
- Measure the total volume of the flow-through.
- Add 2/3 volume of ethanol 100% to filtrate and mix thoroughly.
- Place a second filter cartridge in a fresh collection tube.
- Transfer up to 700 µL of sample volume into the cartridge and centrifuge at 10,000 x g for 30 s. Discard the flow-through.
- Repeat step 2.3.8 until all the mixture has passed through the filter cartridge.
- Apply 700 µL of miRNA wash solution 1 to the filter cartridge and centrifuge the filter cartridge with the collection tube at 10,000 x g for 15 s. Discard the flow-through. Place the filter cartridge in the same collection tube.
- Apply 500 µL of wash solution 2/3 to the filter cartridge and centrifuge the filter cartridge with the collection tube at 10,000 x g for 15 s. Discard the flow-through. Place the filter cartridge in a fresh collection tube.
- Place the filter cartridge in a fresh collection tube and repeat step 2.3.11 with 500 µL of wash solution 2/3. Discard the flow-through and transfer the filter cartridge to a fresh collection tube.
- Centrifuge the tubes with the filter cartridges at 10,000 x g for 1 min.
- Transfer the filter cartridge into a fresh 1.5 mL collection tube and add 100 µL of preheated (95 °C) elution solution or nuclease-free water.
- Centrifuge at 10,000 x g for 30 s to recover miRNAs and store at -80 °C.
NOTE: A white precipitate may form in the wash solution 2/3. This is excess EDTA released from the solution. Avoid drawing up these crystals during the pipetting steps.
3. Perform Reverse Transcription and miRNA Pre-amplification (Table of Materials)
- Perform polyadenylation reaction
- Thaw all reagents on ice, then briefly vortex and centrifuge to spin down the contents. In addition, thaw 50% PEG 8000, allowing it to reach room temperature.
- Prepare the reaction mix in a 0.5 mL centrifuge tube, obtaining a final volume of 3 µL of reaction cocktail for each sample. Use the following volumes, plus 10% excess volume for each reagent.
- Add 1.7 µL of RNase-free water, 0.5 µL of 10x Poly(A) buffer, 0.5 µL of 10 mM ATP, and finally 0.3 µL 5 U/µL PolyA enzyme.
- Briefly vortex and centrifuge the cocktail mix to spin down the contents.
- Transfer 2 µL of the sample to each well of the reaction plate or reaction tube and add 3 µL of the reaction cocktail. Briefly vortex and centrifuge to spin down the contents.
- Place the plate/tubes into a thermal cycler and perform the following steps.
- Incubate at 37 °C for 45 min (polyadenylation step).
- Incubate at 65 °C for 10 min (stop reaction), with a following hold at 4 °C.
- Perform the ligation reaction.
- Prepare the reaction mix in a centrifuge tube with the following volumes for each sample and 10% of volume in excess.
- Add 0.4 µL of RNase-free water, 3 µL of 5x DNA ligase buffer, 0.6 µL of 25x ligation adaptor, 4.5 µL of 50% PEG 8000, and finally 1.5 µL of RNA ligase.
- Briefly vortex and centrifuge the cocktail mix to spin down the content.
- Add 10 µL of the mix to each well/reaction tube containing the poly(A) tailing product. Mix by a plate shaker at 1,900 rpm for 1 min or by briefly vortexing and spin down the contents.
- Place the plate/reaction tubes in a thermal cycler with the following settings: incubation at 60 °C for 60 min, with a following hold at 4 °C.
NOTE: 50% PEG 8000 is highly viscous. Use at room temperature, aspirating and dispensing slowly. After each aspiration and dispensing step, hold the plugger for 10 s to allow the reagent to flow up and down.
- Perform the reverse transcription reaction.
- Prepare the reaction mix in a centrifuge tube, with the following volumes for each sample and 10% of volume in excess.
- Add 3.3 µL of RNase-free water, 6 µL of 5x RT buffer, 1.2 µL of dNTP mix (25 mM each), 1.5 µL of 20x Universal RT primers, and finally 3 µL of 10x RT enzyme mix.
- Briefly vortex and centrifuge the cocktail mix to spin down the contents.
- Add 15 µL of the mix to each well/reaction tube containing the ligation product. Mix by a plate shaker at 1,900 rpm for 1 min or by briefly vortexing and spin down the contents.
- Place the plate/reaction tubes in a thermal cycler with the following settings.
- Incubate at 42 °C for 15 min (reverse transcription).
- Incubate at 85 °C for 5 min (stop reaction), with a following hold at 4 °C.
NOTE: The RT product can now be stored at -20 °C and will remain stable for up to 2 months.
- Perform the miR-Amp reaction (Table of Materials).
- Prepare the reaction mix in a centrifuge tube with the following volumes for each sample and 10% of volume in excess.
- Add 17.5 µL of RNase-free water, 25 µL of the 2x miR-Amp master mix, and 2.5 µL of 20x miR-Amp primer mix.
- Briefly vortex and centrifuge the cocktail mix to spin down the contents.
- For each sample, transfer 45 µL of the reaction mix to each well of a new reaction plate or a reaction tube. Add 5 µL of the RT product to each well or reaction tube. Mix by a plate shaker at 1,900 rpm for 1 min or by briefly vortexing and spin down the contents.
- Place the plate/reaction tubes in a thermal cycler and run as shown in Table 1.
4. Perform Real-time PCR
- Dilute each cDNA sample 1:10 in 0.1x TE buffer.
- Calculate the number of wells/replicates for each sample. Prepare the reaction mix in a centrifuge tube with the following volumes for each sample, adding 10% of volume in excess.
- Add 4 µL of RNase-free water, 10 µL of 2x fast advanced master mix (Table of Materials), and finally 5 µL of diluted cDNA.
- Mix by vortexing and briefly centrifuge to spin down the contents.
- Transfer 19 µL of reaction mix to each well, seal the plate and briefly centrifuge.
- Perform the thermal protocol (Table 2) on a RT-PCR system.
Representative Results
We analyzed a panel of angiogenesis-related circulating miRNAs in relation to progression-free survival (PFS), overall survival (OS) and objective response rate (ORR) in a 52 mCRC patients treated with B-based CT. The evaluation of such biomarkers in plasma samples is challenging because of technical difficulties. We used established methods for miRNA extraction from patient plasma samples, reverse transcription and pre-amplification. Following the manufacturer's instructions and with only a few modifications required, especially in the extraction steps, we successfully evaluated all our targets in the patient population (Figure 2).
In particular, we analyzed 2 plasma samples/patient, correlating baseline and therapy-modified miRNA expression with patient outcome.
We performed serial dilutions of cel-miR-39 at 0.05 nM, 0.5 nM, 5nM and 5 pM in 800 µL of plasma sample (400 µL of plasma plus 400 µL of 2x denaturating solution) to determine the spike-in concentration to use. As shown in Figure 4, the exogenous control showed threshold cycle (Ct) values that were congruent with the serial dilutions, indicating the robustness of the entire protocol (i.e., extraction, reverse transcription and qRT-PCR evaluation). Moreover, these serial dilutions enabled us to select the spike-in concentration that would not interfere with the reverse-transcription of all target miRNAs.
This method allowed us to correlate baseline miRNAs with clinical pathological features. In particular, we found that hsa-miR-199a-5p, hsa-miR-335-5p and hsa-miR-520d-3p were significantly upregulated in left-sided with respect to right-sided lesions (p = 0.03, p = 0.006 and p = 0.008, respectively). Conversely, hsa-miR-21-5p was significantly downregulated in RAS-mutated patients (K-RAS, p = 0.01 and N-RAS, p = 0.008), whereas hsa-miR-221-3p was upregulated (p = 0.05, p = 0.01, K- and N-RAS, respectively).
Comparing the expression levels of miRNA between baseline and first clinical evaluation, we observed that an increase in hsa-miR-155-5p expression levels was associated with shorter PFS (p = 0.04) and OS (p = 0.02). In particular, in patients with a ≥30% increase in hsa-miR-155-5p, PFS was 9.5 months (95% CI, 6.8-18.7) and OS was 15.9 months (95% CI, 8.4-not reached) compared to 22.3 months (95% CI, 10.2-25.5) and 42.9 months (95% CI, 24.8-not reached) for those with a <30% increase (Figure 3). The median value of variation in the case series (30%) was set as the cutoff point. Circulating basal levels of miRNAs were dichotomized into "high" or "low" according to median values9.
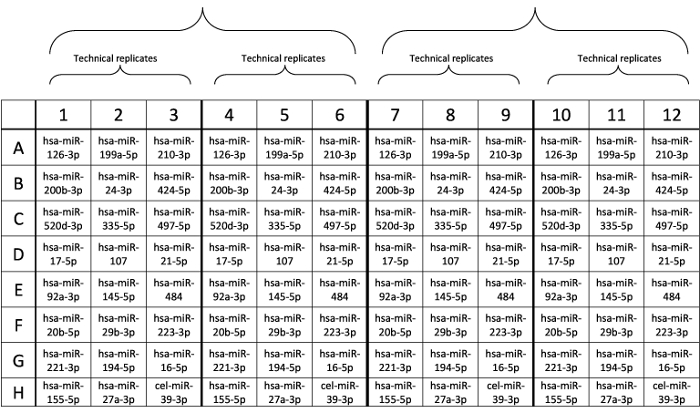
Figure 1: Design of custom plate layout. Probes were pre-spotted in each well. Please click here to view a larger version of this figure.
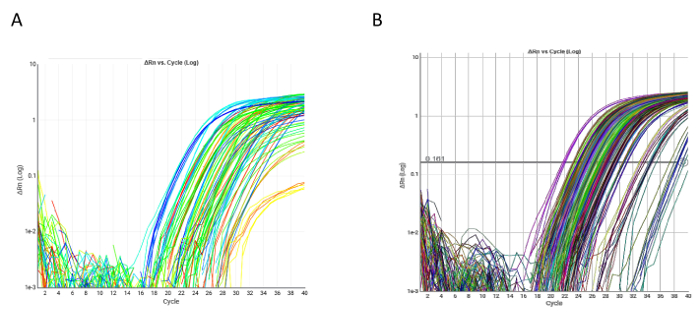
Figure 2: Examples of qRT-PCR amplification plots. (A) Amplification plot of a single qRT-PCR custom plate. All markers were evaluable in both single plate samples. (B) qRT-PCR of hsa-miR-17-5p in the overall case series. Please click here to view a larger version of this figure.
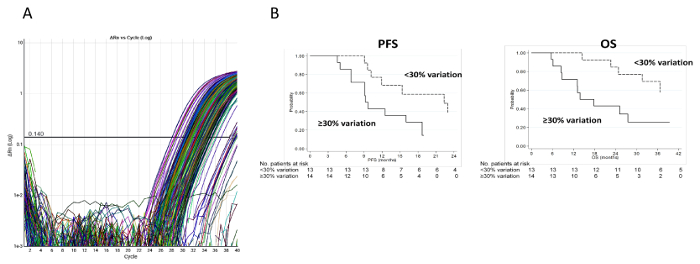
Figure 3: Experimental and clinical results relative to hsa-miR-155-5p. (A) Ct values for hsa-miR-155-5p. Amplification plot of hsa-miR-155-5p in the overall case series. (B) PFS and OS of patients with a ≥30% or <30% increase in circulating hsa-miR-155-5p. PFS was calculated as the time from the date of randomization to the date of the first documented evidence of tumor progression, last tumor assessment or death in the absence of disease progression. OS was calculated as the time from the date of randomization to the date of death from any cause or last follow-up. Please click here to view a larger version of this figure.
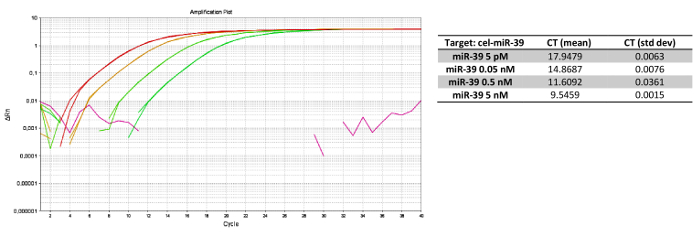
Figure 4: Experimental results forcel-miR-39 serial dilution. (A) qRT-PCR for serial dilutions of cel-miR-39, with (B) relative Cts. Dilutions were performed at 5 nM, 0.5 nM, 0.05 nM and 5 pM in plasma samples from the same patient. Such assay linearity enabled us to select the best concentration to use in the overall case series. Please click here to view a larger version of this figure.
| Step | Temperature | Duration | Cycles |
| Enzyme activation | 95 °C | 5 min | 1 |
| Denature | 95 °C | 3 s | 14 |
| Anneal/extend | 60 °C | 30 s | |
| Stop reaction | 99 °C | 10 min | 1 |
| Hold | 4 °C | Hold | ∞ |
Table 1: Thermal protocol of mir-AMP reaction.
| Step | Temperature | Duration | Cycles |
| Enzyme activation | 95 °C | 20 s | 1 |
| Denature | 95 °C | 3 s | 40 |
| Anneal/extend | 60 °C | 30 s | |
| Hold | 4 °C | Hold | ∞ |
Table 2: Thermal protocol of qRT-PCR.
Discussion
miRNAs are small non-coding RNAs (18-25 nucleotides in length) capable of binding the 3' UTR region of their target messenger RNA and inhibiting and/or degrading it. They can thus be considered gene expression regulators at the translational level. In the last decade, several miRNAs have been correlated with cancer initiation and development, making them useful clinical biomarkers for diagnosis, prognosis and therapy. Protected by RNases, miRNAs are present in a stable form in biological fluids such as blood, plasma, serum, urine and saliva, which has led to a growing interest in their potential usefulness in clinical practice10.
Despite this, the evaluation of circulating miRNAs is a challenging task and difficulties regarding sample collection, target extraction, platform choice and global normalization are hotly debated topics within the scientific community.
For sample collection and storage, pre-analytical variables are closely linked to plasma handling and processing. We focused on plasma rather than serum samples given that there is ample evidence of higher miRNA concentrations in the latter, possibly due to the release of these molecules during the coagulation process10,11,12. To address the issue of the release of cellular nucleic acids deriving from blood cell lysis, we centrifuged the samples within 2 h of blood collection. We also used K3E EDTA tubes because other anti-coagulants are known to interfere with amplification steps (i.e., heparin) and may trigger hemolysis (i.e., citrate). As it is critical to avoid platelet and lymphocyte contamination in the pre-analytical phase, we removed the plasma from the tube very slowly and did not aspirate the last ~50 µL of the sample from the tube.
Various studies have shown the superiority of column-based extraction methods over guanidine/phenol/chloroform-based protocols11,13, but Khoury et al. effectively isolated good quality small RNAs from serum using such reagents without contamination14. The commercial kit we used sequentially performs both extraction methods, permitting suitable miRNA yields in terms of spike-in concentrations. We only performed one modification with respect to the manufacturer's instructions, i.e., fresh collection tubes were always used after each filter cartridge washing step to eliminate/reduce the risk of reagent contamination. In order to identify the best spike-in concentration in our samples and thus avoid interference with subsequent reactions, we first performed the entire protocol without adding the cel-miR-39 during the extraction step to evaluate the median expression levels of our targets. We then performed de novo miRNA extraction from plasma samples of the same patient using a serial dilution of cel-miR-39 to identify the best concentration to add to the plasma samples of our case series (Figure 4). It must, however, be underlined that the optimal cel-miR-39 concentration, identified for only one sample, may not necessarily be the best concentration for the overall case series because of a series of internal variables such as age, gender or life habits, which could influence absolute miRNA expression within blood.
miRNA reverse transcription and amplification represent 2 critical steps. Of the two main techniques for miRNA detection (i.e., qRT-PCR and microarray), we chose qRT-PCR given its high sensitivity and our need to evaluate only a selected panel of miRNAs (a major limitation of this methodology is its low throughput)15. We used reverse transcription chemistry based on 3' polyA tailing and 5' ligation adaptor sequences to extend the mature miRNA and permit annealing of universal RT primers. Based on single base-pair recognition, the 5' ligation adaptor could help to avoid common biases such as those linked to 5' mismatch. This kit also includes a pre-amplification step that is required for low yield samples, such as plasma. Although necessary, this step could introduce some amplification biases before the detection step.
In this protocol, we analyzed a panel of 24 selected miRNAs correlated with angiogenesis. In particular, we chose pre-spotted probe custom plates and followed the manufacturer's instructions for the qRT-PCR runs. The main issues relating to miRNA qRT-PCR are the choice of reference genes and their subsequent data analysis normalization.
Although numerous studies have been conducted to identify the best miRNA to use for normalization, no uniform consensus has been reached. To address this limitation, we carried out a literature search to identify the circulating miRNAs that can be used as reference genes in our case series.
We also selected the 4 miRNAs with strong evidence of stable expression in plasma samples from mCRC patients and tested them in a smaller part of our case series. The 2 most stable miRNAs were identified by GeNorm analysis as reference housekeeping genes.
In conclusion, the evaluation of circulating miRNAs in peripheral blood samples has a number of known difficulties. However, we believe that the methodologies we used are reliable and robust, allowing for an in-depth and accurate study of these biomarkers which have the potential to change the treatment scenario for colorectal cancer.
Divulgations
The authors have nothing to disclose.
Acknowledgements
This study was partially funded by Roche S.p.A. and the Italian Medicines Agency (AIFA).
Materials
| Rnase-free Safe-lock 1.5 mL tubes | Eppendorf | 0030 123.328 | |
| Rnase-free Safe-lock 2 mL tubes | Eppendorf | 0030 123.344 | |
| Rnase-free 20 µL tips | Starlab | S1123-1810 | |
| Rnase-free 200 µL tips | Starlab | S1120-8810 | |
| Rnase-free 1000 µL tips | Starlab | S1122-1830 | |
| mirVana PARIS RNA and Native Protein Purification Kit | Thermo Fisher | AM1556 | |
| TaqMan Advanced miRNA cDNA Synthesis Kit | Thermo Fisher | A28007 | |
| 100% ethanol anidrous ACS grade | Carlo Erba Reagents | 414605 | |
| 2-mercapto-ethanol | Sigma-Aldrich | M3148 | |
| TaqMan Fast Advanced Master Mix | Thermo Fisher | 4444558 | |
| TaqMan Advanced miRNA Assays | Thermo Fisher | A25576 | |
| 7500 Real-Time PCR System | Applied Biosystems | 4406984 | |
| 0.2-2, 1-10, 2-20, 20-200, 100-1000 µL laboratory pipettes | |||
| Benchtop microcentrifuge | |||
| Vortex | |||
| Benchtop heating block | |||
| Fume hood | |||
| 0.2 mL PCR tubes |
References
- Luo, H. -. Y., Xu, R. -. H. Predictive and prognostic biomarkers with therapeutic targets in advanced colorectal cancer. World journal of gastroenterology. 20 (14), 3858-3874 (2014).
- Toiyama, Y., Okugawa, Y., Fleshman, J., Richard, C., Goel, A. MicroRNAs as potential liquid biopsy biomarkers in colorectal Cancer: A systematic review. BBA Reviews on Cancer. 5 (6), (2018).
- Kok, M. G. M., Halliani, A., Moerland, P. D., Meijers, J. C. M., Creemers, E. E., Pinto-Sietsma, S. J. Normalization panels for the reliable quantification of circulating microRNAs by RT-qPCR. FASEB Journal. 29 (9), 3853-3862 (2015).
- Marabita, F., De Candia, P., Torri, A., Tegnér, J., Abrignani, S., Rossi, R. L. Normalization of circulating microRNA expression data obtained by quantitative real-time RT-PCR. Briefings in Bioinformatics. 17 (2), 204-212 (2016).
- Danese, E., et al. Reference miRNAs for colorectal cancer: Analysis and verification of current data. Scientific Reports. 7 (1), 1-12 (2017).
- Zheng, G., et al. Identification and validation of reference genes for qPCR detection of serum microRNAs in colorectal adenocarcinoma patients. PLoS ONE. 8 (12), 1-10 (2013).
- Lv, W., et al. Optimization of the Original TRIzol-Based Technique Improves the Extraction of Circulating MicroRNA from Serum Samples. Clinical laboratory. 61 (12), 1953-1960 (2015).
- Tan, G. W., Khoo, A. S. B., Tan, L. P. Evaluation of extraction kits and RT-qPCR systems adapted to high-throughput platform for circulating miRNAs. Scientific reports. 5, 9430 (2015).
- Altman, D. G., McShane, L. M., Sauerbrei, W., Taube, S. E. Reporting Recommendations for Tumor Marker Prognostic Studies (REMARK): explanation and elaboration. PLoS medicine. 9 (5), (2012).
- Tiberio, P., Callari, M., Angeloni, V., Daidone, M. G., Appierto, V. Challenges in using circulating miRNAs as cancer biomarkers. BioMed Research International. 2015, (2015).
- Kroh, E. M., Parkin, R. K., Mitchell, P. S., Tewari, M. Analysis of circulating microRNA biomarkers in plasma and serum using quantitative reverse transcription-PCR (qRT-PCR). Methods (San Diego, Calif). 50 (4), 298-301 (2010).
- Cheng, H. H., et al. Plasma Processing Conditions Substantially Influence Circulating microRNA Biomarker Levels. PLoS ONE. 8 (6), 1-11 (2013).
- Moret, I., et al. Assessing an improved protocol for plasma microRNA extraction. PloS one. 8 (12), (2013).
- Khoury, S., Ajuyah, P., Tran, N. Isolation of small noncoding RNAs from human serum). Journal of visualized experiments JoVE. (88), e51443 (2014).
- Schwarzenbach, H., Nishida, N., Calin, G. A., Pantel, K. Clinical relevance of circulating cell-free microRNAs in cancer. Nature Reviews Clinical Oncology. 11 (3), 145-156 (2014).

