Preparation of Cell Extracts by Cryogrinding in an Automated Freezer Mill
Summary
We describe a reliable method for the preparation of whole cell extracts from yeast or other cells using a cryogenic freezer mill that minimizes degradation and denaturation of proteins. The cell extracts are suitable for purification of functional protein complexes, proteomic analyses, co-immunoprecipitation studies and detection of labile protein modifications.
Abstract
The ease of genetic manipulation and the strong evolutionary conservation of eukaryotic cellular machinery in the budding yeast Saccharomyces cerevisiae has made it a pre-eminent genetic model organism. However, since efficient protein isolation depends upon optimal disruption of cells, the use of yeast for biochemical analysis of cellular proteins is hampered by its cell wall which is expensive to digest enzymatically (using lyticase or zymolyase), and difficult to disrupt mechanically (using a traditional bead beater, a French press or a coffee grinder) without causing heating of samples, which in turn causes protein denaturation and degradation. Although manual grinding of yeast cells under liquid nitrogen (LN2) using a mortar and pestle avoids overheating of samples, it is labor intensive and subject to variability in cell lysis between operators. For many years, we have been successfully preparing high quality yeast extracts using cryogrinding of cells in an automated freezer mill. The temperature of -196 °C achieved with the use of LN2 protects the biological material from degradation by proteases and nucleases, allowing the retrieval of intact proteins, nucleic acids and other macromolecules. Here we describe this technique in detail for budding yeast cells which involves first freezing a suspension of cells in a lysis buffer through its dropwise addition into LN2 to generate frozen droplets of cells known as “popcorn”. This popcorn is then pulverized under LN2 in a freezer mill to generate a frozen “powdered” extract which is thawed slowly and clarified by centrifugation to remove insoluble debris. The resulting extracts are ready for downstream applications, such as protein or nucleic acid purification, proteomic analyses, or co-immunoprecipitation studies. This technique is widely applicable for cell extract preparation from a variety of microorganisms, plant and animal tissues, marine specimens including corals, as well as isolating DNA/RNA from forensic and permafrost fossil specimens.
Introduction
Yeast is a popular model organism for protein studies, as it is a simple eukaryotic organism with an abundance of genetic and biochemical tools available for researchers1. Because of their sturdy cell wall, one challenge that researchers face is in efficiently lysing the cells without damaging the cellular contents. Different methods are available for obtaining protein extracts through disruption of yeast cells which include enzymatic lysis (zymolyase)2,3, chemical lysis4, physical lysis by freeze-thaw5, pressure-based (French press)6,7, mechanical (glass beads, coffee grinder)8,9, sonication-based10 and cryogenic2,11. The efficiency of cell lysis and the protein yield can vary considerably depending on the technique employed, thus affecting the end result or suitability for the desired downstream application for the lysate. When studying proteins that are unstable, have fleeting posttranslational modifications, or are temperature sensitive, it is particularly important to use a method that will minimize sample loss or degradation during preparation.
| Extract preparation technique | Details | Advantages | Disadvantages | Downstream analysis | Reference |
| French press: High-pressure homogenizer (aka Microfluidizer) with enzymatic pretreatment using Zymolyase | Zymolyase-20T, a Microfluidizer high-pressure homogenizer. The disruptor consists of an air-driven, high-pressure pump (ratio 1:250; required air pressure 0.6-l MPa) and a special disruption chamber with an additional back pressure unit. A minimum sample size of 20 mL is required for processing. | Final total disruption obtained using the combined protocol approached 100 % with 4 passes at a pressure of 95 MPa, as compared to only 32 % disruption with 4 passes at 95 MPa using only homogenization without the Zymolyase. | Not appropriate for small scale applications. The enzymes can get expensive for large scale preparations. | Protein purification | 6 |
| Bead beater: Zymolyase treated cells lysed with glass beads in a fastprep instrument | Roughly an equal volume of cold, dry, acid-washed 0.5 mm glass beads is added to a given volume of cell pellet in lysis buffer and the cells are disrupted by vigorous manual agitation. | It is particularly useful when making extracts from many different small yeast cultures for assaying purposes rather than for protein purification. | During the glass bead procedure, proteins are treated harshly causing extensive foaming leading to protein denaturation. The amount of cell breakage varies, while proteolysis as well as modification of the proteins may result from heating of the extract above 4°C during the mechanical breakage. | Mostly DNA & RNA analyses, but also protein analysis by denaturing gel electropheoresis, either with or without Western blotting. | 8 |
| Zymolyase treatment followed by lysis using a combination of osmotic shock and Dounce homogenization | After enzymatic digestion of cell walls, spheroplasts are lysed with 15 to 20 strokes of a tight-fitting pestle (clearance 1 to 3 µm) in a Dounce homogenizer. | Advantageous to use protease-deficient strains such as BJ926 or EJ101. This is the gentlest way to break yeast cells and hence it is most suitable for preparing extracts that can carry out complex enzymatic functions (e.g., translation, transcription, DNA replication) and in which the integrity of macromolecular structures (e.g., ribosomes, splicesomes) has to be maintained. It is also useful for isolating intact nuclei that can be used for chromatin studies (Bloom and Carbon, 1982) or for nuclear protein extracts (Lue and Kornberg, 1987). | The major disadvantages of the spheroplast lysis procedure are that it is relatively tedious and expensive, especially for large-scale preparations (>10 liters), and the long incubation periods can lead to proteolysis or protein modification. For chromatin preparations, they seem to be of varying or lower quality than those produced by the differential centrifugation (based on nucleosome ladder integrity). | Isolating intact nuclei for chromatin studies, extracts that can carry out complex enzymatic functions, extracts requiring the integrity of macromolecular structures, nuclear protein extracts. |
2 |
| Cell Disruption of flash frozen cells by grinding in Liquid Nitrogen using a mortar/pestle or a blender | Cells are frozen immediately in liquid nitrogen and then lysed by grinding manually in a mortar using a pestle, or using a Waring blender in the presence of liquid nitrogen. | The protocol is quick and easy. It can accommodate varying amounts of yeast cells including very large cultures. Its main advantage is that cells are taken immediately from the actively growing state into liquid nitrogen (−196°C), decreasing degradative enzyme activities such as proteases and nucleases as well as activities that modify proteins (e.g., phosphatases and kinases). It is particularly suited for making whole-cell extracts from a single yeast culture for large-scale protein purification. | A bit messy and potentially dangerous to the careless investigator. Small samples (i.e., 10- to 100-ml yeast cultures) are not easily processed because there is not enough mass of frozen cell clumps to fracture effectively in the blender. It is time-consuming to process individual samples and to clean the equipment between uses. | Whole-cell extracts from a single yeast culture for large-scale protein purification. | 2 |
| Autolysis, Bead mill | pH 5.0, 50 °C, 24 h, 200 rpm / Ø 0.5 mm, 5 × 3 min/3 min | Quick and efficient lysis, especially for small scale extract preparation | Heat generation leads to denaturation and degradation of macromolecules. Bead beating equipment required. | Small scale analyses. | 10 |
| Autolysis, Sonication | pH 5.0, 50 °C, 24 h, 200 rpm, 4 × 5 min/2 min, pulser 80%, power 80% | Sonication equipment is usually available in most institutions. | Heat generation leads to denaturation and degradation of macromolecules. Sonication equipment required. Slow lysis can take more than 24 hours. | Yeast cell wall preparations. | |
| Boiling and freeze-thaw process | No specialized equipment needed other than a standard freezer and a heating block or hot water bath. | Efficient, reproducible, simple and inexpensive. | Heat generation leads to denaturation and degradation of macromolecules. | DNA analyses by PCR. | 5 |
Table 1: Comparison of methods available for the preparation of yeast extracts.
Cryogrinding (aka cryogenic grinding/cryogenic milling) is commonly employed to retrieve nucleic acids, proteins or chemicals from temperature sensitive samples in a reliable manner for quantitative or qualitative analyses. It has been used successfully for multiple applications in diverse fields including biotechnology, toxicology, forensic science12,13, environmental science, plant biology14 and food science. Isolation of intact biological macromolecules is usually critically dependent on the temperature. Extremely low temperatures ensure that the proteases and nucleases stay inactive, resulting in a reliable isolation of intact proteins, nucleic acids and other macromolecules for subsequent analyses. Indeed, a freezer mill typically maintains a sample temperature of -196 °C (the boiling point of LN2), thus minimizing DNA/RNA or protein denaturation and degradation.
The freezer mill employs an electromagnetic grinding chamber that rapidly moves a solid metal bar or cylinder back and forth within a vial containing the sample to be pulverized between stainless steel end plugs. The instrument creates and rapidly reverses a magnetic field within the grinding chamber. As the magnetic field shifts back and forth, the magnet crushes the sample against the plugs thus achieving the 'cryogrinding' and the pulverization of the popcorn. The freezer mill replaces the mortar and pestle and allows the sequential processing of multiple samples (or up to 4 smaller samples simultaneously) with high reproducibility and avoids the user-to-user variability associated with manual grinding. Once the samples are processed, the cell extracts can be used for a variety of downstream applications.
Protocol
1. Preparation of Yeast Popcorn
- Grow yeast cells in 0.5 L of YPD media to a density of 1 x 107 cells/mL. Count cells using a Coulter Counter or any other means.
- Centrifuge cells for 10 min at 2,400 g and 4 °C.
- Wash each sample once with 500 mL of ice-cold deionized 18 mega Ohm Milli-Q water.
- Resuspend pellets in 15 mL of ice-cold Lysis Buffer [20 mM HEPES-KOH pH 7.5, 110 mM KCl, 0.1% Tween, 10% glycerol, with freshly added reducing agent 10 mM β-mercaptoethanol, protease inhibitor cocktail, 10 μM proteasome inhibitor MG-132, 1 mM deacetylase inhibitor sodium butyrate and phosphatase inhibitors (1 mM sodium vanadate, 50 mM sodium fluoride, 50 mM sodium β-glycerophosphate)]. See the Table of Materials for additional details. Keep resuspended cells on ice until the samples are ready for the next step.
NOTE: A wide variety of lysis buffers containing a number of non-ionic detergents can be used with the samples and specific inhibitors included in it based on the downstream application. Inclusion of protease and proteasomal inhibitors are crucial for preventing protein degradation, especially once the extracts are thawed. - Make snap frozen yeast popcorn by slowly adding the cell suspension one drop at a time using a pre-chilled serological pipette into one or more 50 mL centrifuge tubes kept on dry ice and filled with LN2 until just below the rim. Top up liquid nitrogen in the tube frequently to compensate for its loss due to rapid evaporation. Work in a well-ventilated area to avoid the hazards associated with nitrogen asphyxiation.
- Instead of a 50 mL tube, use multiple 50 mL tubes tied together (or any container with a large mouth, such as a large plastic centrifuge bottle that can tolerate cryogenic temperatures) for pop-corn preparation (Figure 1A). The larger the container and its opening, the easier is the popcorn preparation as this prevents clumping of the popcorn to form large aggregates. A size range of 0.3-0.5 cm for the popcorn is ideal for achieving proper grinding/pulverization using the freezer mill (Figure 1B).
- Ensure that the LN2 has evaporated completely from the tubes containing the popcorn prior to storing them at -80 °C. It is possible to stop after preparing the yeast popcorn as they are stable for several years when maintained at -80 °C.
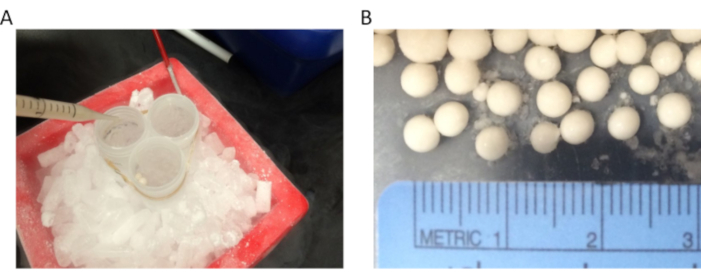

Figure 1: Yeast popcorn preparation. (A) Yeast "popcorn" is made by the dropwise freezing of the cell suspension in LN2. We use one to three 50 mL tubes held together with a rubber band and placed in an ice bucket filled with dry ice. The tubes are filled with LN2 until just below their rims and are topped up with liquid nitrogen frequently to keep them nearly full until all the cell suspension had been made into popcorn (B) The size of the popcorn is an important determinant of optimal grinding efficiency. The size range of the popcorn should be between 0.3 and 0.5 cm in diameter. Please click here to view a larger version of this figure.
2. Cryogrinding
- To start, ensure that there is enough LN2 available to chill the freezer mill, grinding vials, supplies and to operate the machine for all samples. For less than five samples, 30-35 L of LN2 should suffice. Fill the freezer mill chamber with LN2 up to the fill line.
WARNING: All steps involving LN2 should be performed in a well-ventilated area and the freezer mill itself should be located in such an area to avoid risk of asphyxiation. Wear personal protective equipment including proper footwear, lab coat, safety glasses, a base layer of nitrile gloves, then thermal gloves, followed by another pair of nitrile gloves. Use extreme caution when handling LN2. - Close the freezer mill lid slowly, avoiding the splash of LN2 and allow a few min for the machine to cool down. It may be necessary to refill the chamber with LN2 to compensate for the loss due to evaporation before proceeding to the next step.
NOTE: Refill LN2 as needed only up to the fill line. Do not overfill the chamber as this can be both dangerous and detrimental to the machine. Automated refill systems are available for freezer mill that can replenish the evaporating LN2 as needed directly from a storage tank connected to the freezer mill. - Pre-chill the large grinding vials and the magnetic impactor bar by dunking them in LN2 kept in a separate small dewar. Decant all the LN2 when the LN2 stops bubbling. Then add the sample/yeast popcorn to the grinding vial and seal it tightly with the two stainless steel end plugs.
- Do not fill more than one-third of the grinding vial with the sample as that can reduce the efficiency of grinding. Instead, larger samples can be processed by dividing them into two or more vials and grinding them sequentially. Smaller grinding vials can be used for smaller samples (up to 3 mL).
- Place the grinding vial in the freezer mill chamber and lock it in place. Close the lid.
- For budding yeast, grind the samples for a total of three cycles for 2 min/cycle (with 2 min break for cooling between cycles) at a crushing rate of 14. When the machine stops after completing the cycles, open the freezer mill lid slowly and carefully unlock the vial with the powdered frozen cell lysate and remove it from the freezer mill. If multiple small vials are used, work quickly to remove one vial at a time and place them in dry ice.
NOTE: There is usually no need to adjust the grinding parameters when using vials of different sizes with the same type of sample. However, the grinding parameters such as the number of cycles and the grinding rate need to be empirically determined for different sample types, depending on the ease with which they lyse. Most mammalian cells and soft tissues will lyse in 1-2 cycles at a crushing rate of 10. Harder to lyse samples such as bacteria, yeast, fly larvae and adult fruit flies require 3-6 cycles at the maximum rate, while hard tissues such as bone, teeth, etc. may require up to 10 cycles. A small aliquot of the thawed sample can be viewed under the microscope before and after grinding to count the number of intact, unlysed cells to determine the lysis efficiency. An excellent analysis of the impact of cryogrinding parameters on the release of proteins and DNA from budding yeast has been published previously15. - Working quickly so as not to allow the sample to thaw out, carefully unscrew one of the end pieces (pick the one that appears to have become somewhat loose during the grinding) using the opening tool. Then, use a pair of long forceps (that are pre-chilled in a bucket with dry ice) to remove the impactor bar. Collect the pulverized sample by inverting and tapping the grinding vial onto a polystyrene weighing dish pre-chilled with LN2 and kept on dry ice.
- Once all of the powdered frozen cell lysate has been recovered from the grinding vial, pour the powdered lysate from the weighing dish back into the 50 mL tube and proceed immediately to the slow thawing step described next for best results. Alternatively, store the frozen powdered lysate overnight at -80 °C, although this may result in some degradation.
- Prepare an ice bucket with a 50% slurry of ice and water and place it on a stir plate with a magnetic stirrer. Submerge a wire rack or another suitable rack in the ice slurry to hold the samples.
- Then, slowly thaw the samples on an ice slurry bath with constant agitation of the slurry using a magnetic stirrer. Add more ice to replace the melting ice (some water may need to be discarded to prevent the ice bucket from overflowing). Since several inhibitors have short half lives in aqueous buffers, add additional protease inhibitor cocktail (see Table of Materials) and 10 μM proteasome inhibitor MG-132 to the lysate once the samples start thawing (after approximately 30 min).
- Remove ice formed on the outside of the tubes every 5 min to expedite the thawing process. Note that rapid thawing of the samples at room temperature or higher can lead to significant degradation.
- After the samples have thawed completely (it may take well over an hour depending on the amount of sample), centrifuge the lysate at ~3,220 g for 20 min in a refrigerated tabletop centrifuge at 4 °C to remove the bulk of the cell debris from the lysate.
- Transfer the supernatant to 50 mL polycarbonate centrifuge tubes that have been pre-chilled on ice and centrifuge the samples at 16,000 g for 20 min at 4 °C (see Table of Materials).
- Transfer only the clear supernatant consisting of the clarified whole cell extract from the center of the liquid column in the tube very slowly and carefully, without disturbing the cloudy lipid layer (Figure 2), into a chilled 15 mL tube using pre-chilled serological pipettes. Avoid removing all of the lysate in the centrifuge tube to prevent carryover of lipids and debris. The leftover lysate in the centrifuge tube can be centrifuged multiple times to recover small amounts of extract after each spin.
NOTE: Lipids along with any denatured proteins would appear cloudy/milky and are located at the top of the liquid/air interface, and occasionally above the pellet itself that consists of insoluble cellular debris (Figure 2).
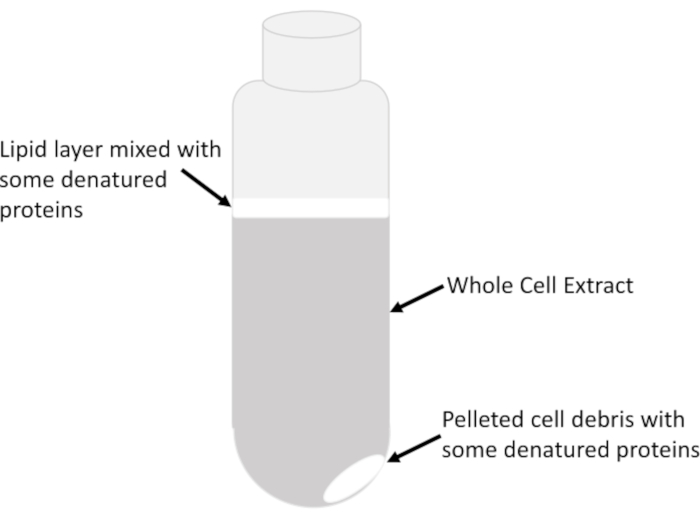
Figure 2: Extract layers in a centrifuge tube. The major visible features of the whole cell extract in a tube following centrifugation at 16,000 g for 20 min are indicated. The relative abundance of each feature depends on the sample type, the growth phase of the cells (exponential versus stationary), the amount of lysis buffer used to resuspend cells and the lysis efficiency. Please click here to view a larger version of this figure.
- Centrifuge any remaining lysate for 5 min to recover more of the extract. Repeat this 5 min centrifugation multiple times if needed to recover as much of the clear extract as possible. Discard the cloudy lysate containing lipids close to the bottom of the tube near the pellet.
- Use these extracts for downstream applications such as protein complex purification, immunoprecipitation and proteomic analysis.
Representative Results
We compared two different methods for yeast cell lysis, namely glass bead milling at 4 °C and an automated cryogrinding method at -196 °C, to assess the relative recovery proteins in the cell extracts prepared with both methods. For this study, we chose to use a budding yeast strain YAG 1177 (MAT a lys2-810 leu2-3,-112 ura3-52 his3-Δ200 trp1-1[am] ubi1-Δ1::TRP1 ubi2-Δ2::ura3 ubi3-Δub-2 ubi4-Δ2::LEU2 [pUB39] [pUB221])16 carrying a high copy plasmid expressing a tandem HIS-MYC tagged Ubiquitin (HIS-MYC-Ub) so we can assess the efficiency of recovery of tagged ubiquitinated proteins from extracts following affinity purification.
We tested a quick protocol that uses glass bead milling to lyse yeast cells9 at 4 °C and the cryogrinding method using frozen yeast cells (yeast popcorn) and grinding them in LN2 in a freezer mill. We report a clear difference with the two methods of extract preparation.
In Figure 3A we demonstrate that we can achieve a decidedly higher total protein yield from whole cell extracts (WCE) prepared using freezer grinding as determined by Ponceau S staining after loading equivalent amounts of WCE prepared from the same number of cells using the two different extract preparation methods. Further, Figure 3B shows higher recovery of HIS-MYC tagged ubiquitylated proteins following pull down using Talon beads from the WCE prepared by cryogrinding, when compared to the WCE prepared using the bead beater, despite using extracts prepared from the same number of cells in both cases.
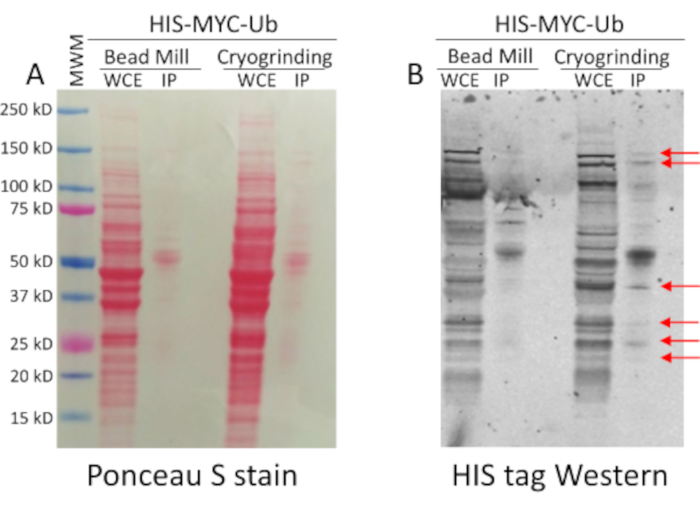
Figure 3: Cryogrinding leads to higher total protein yields in cell extracts and better recovery of tagged proteins following pull down of proteins from these extracts. (A) Whole cell extracts (WCE) were prepared as described in this protocol for the cryogrinding or as previously described for the bead milling method.9 Five hundred mL of the yeast strain carrying HIS-MYC-Ub was grown to density of 107 cells/mL. Then 250 mL of the culture was used in parallel to prepare extracts by each method. Equivalent amounts of WCE was prepared from the same number of cells using the two different methods. HIS tagged proteins were then pulled down using Talon beads (IP) from the same amount of each extract. Finally, these affinity purified proteins were compared by resolving them on a denaturing polyacrylamide gel and transferring to a nitrocellulose membrane followed by staining with Ponceau S. Notice the stronger staining throughout the lane containing WCE prepared using the freezer grinding method. (B) The Ponceau S stained nitrocellulose membrane shown in (A) was then processed for Western blotting using HIS tag antibodies to compare the amounts of HIS-Ub tagged proteins pulled down from equal amounts of extract prepared by the two methods. Several bands were clearly overrepresented in the pulldown using the WCE prepared by cryogrinding (indicated by the red arrows), while the corresponding bands in the pulldown using WCE prepared by bead beating were either considerably weaker in intensity, or undetectable. Please click here to view a larger version of this figure.
Overall, the representative results show that cryogrinding of yeast popcorn leads to higher yields and superior protein isolation based on the stronger Ponceau S stained protein bands in the WCE prepared by cryogrinding (Figure 3A). This is likely due to more efficient lysis and limited protein degradation in the extract prepared by cryogrinding, as indicated by the presence of stronger bands in the Western blot lane showing HIS-tagged proteins pulled down (IP) from the cryogenic extract (Figure 3B). While the glass bead milling protocol is faster and less labor intensive, it may not be suitable for proteins that are temperature sensitive or highly unstable. We also find that the isolation of posttranslational modifications such as polyubiquitinated proteins is more efficient with the cryogrinding protocol16,17,18.
Discussion
A limitation of studying native proteins from yeast is the inefficient lysis of yeast cells due to their tough cell wall. Although several methods have been developed, the most consistent and efficient method in our hands is the cryogrinding of yeast cells flash frozen as popcorn. This method allows the reliable preparation of high-quality whole cell extracts from budding yeast compared to other lysis methods. The representative results demonstrated that cryogrinding is superior to a popular mechanical method for yeast lysis that employs bead milling at 4 °C (Figure 3A and 3B). Cryogrinding efficiently lyses yeast cells while the samples are processed at a temperature of -196 °C in a LN2 bath, therefore maintaining protein integrity by greatly limiting heat induced protein denaturation and degradation that occurs with many other methods of extract preparation. This allows for the analysis of highly labile macromolecules in the samples and permits the purification of functional protein complexes. Further, the use of the freezer mill also replaces the highly labor-intensive traditional manual grinding of yeast popcorn with a mortar, allowing for faster and efficient sample processing. Finally, the achievement of yeast cell lysis without the use of any expensive enzymes provides for significant savings in the long run.
The major downside of a freezer mill is the upfront cost involved in purchasing the equipment, as well as the minimal maintenance and running costs (i.e., large quantities of liquid nitrogen) involved. It also requires careful handling of potentially dangerous LN2 for both the preparation of yeast popcorn and the cryogrinding steps. Larger samples need to be processed individually, which may become time consuming if multiple samples need to be processed. In addition, when processing multiple samples, the grinding vials, caps and impactor bars must be properly washed and wiped clean before re-use (alternatively, several sets of spare accessories must be maintained).
The protocol demonstrated here is valuable for the preparation of yeast protein samples for protein affinity purification, immunoprecipitation studies, sample preparation for Western blotting for labile proteins or their modifications, or for proteomic analyses16,19. In addition, cryogrinding of the yeast popcorn can also be utilized for sheared genomic DNA13,20,21 or RNA22,23 isolation while minimizing nuclease activity, thus ensuring sample preservation24. This versatile method can be used as is for most microorganisms such as bacteria21,22 and yeasts15, and can be quickly adapted for mammalian cells, both plant25 and animal26 tissues, food products27, marine specimens21,28 including corals23, forensic samples20 including hair29 and even fossils30 as well as the remains of extinct lifeforms frozen in permafrost31.
In conclusion, cryogrinding in a freezer mill is an efficient method for protein and other macromolecule isolation from a variety of samples derived from diverse sources for multiple downstream biochemical applications.
Divulgations
The authors have nothing to disclose.
Acknowledgements
Research in the Gunjan lab is supported by funding from the National Institutes of Health, National Science Foundation and the Florida Department of Health. We thank undergraduate student John Parker for technical assistance.
Materials
| 50 mL polycarbonate tubes with screw caps | Beckman | 357002 | Centrifuge tubes |
| BD Bacto Peptone | BD Biosiences | 211677 | Yeast YPD media component |
| BD Bacto Yeast Extract | BD Biosiences | 212750 | Yeast YPD media component |
| Beckman Avanti centrifuge | Beckman | B38624 | High speed centrifuge |
| Beckman JLA-9.1000 | Beckman | 366754 | Rotor |
| D-(+)-Dextrose Anhydrous | MP Biomedicals | 901521 | Yeast YPD media component |
| Eppendorf A-4-44 | Eppendorf | 22637461 | Swinging bucket rotor |
| Eppendorf refrigerated centrifuge 5810 R | Eppendorf | 22625101 | Refrigerated centrifuge |
| Glycerol | SIGMA-ALDRICH | G5150-1GA | Volume excluder and cryoprotectant |
| HEPES | FisherBiotech | BP310-100 | Buffer |
| HIS6 antibody | Novagen | 70796 | Antibody for HIS tag |
| KCl | SIGMA-ALDRICH | P9541-1KG | Salt for maintaining ionic strength |
| MG-132 | CALBIOCHEM | 474790 | Proteasome Inhibitor |
| Phosphatase inhibitor cocktail | ThermoFisher Scientific | A32957 | Phosphatase inhibitor cocktail |
| Ponceau S | SIGMA | P7170-1L | Protein Stain |
| Protease inhibitor cocktail | ThermoFisher Scientific | A32963 | Protease inhibitor cocktail |
| Rotor JLA 25.500 | Beckman | JLA 25.500 | Rotor |
| Sodium Butyrate | EM Science | BX2165-1 | Histone Deacetylase Inhibitor |
| Sodium Fluoride | Sigma-Aldrich | S6521 | Phosphatase Inhibitor |
| Sodium Vanadate | MP Biomedicals | 159664 | Phosphatase Inhibitor |
| Sodium β-glycerophosphate | Alfa Aesar | 13408-09-8 | Phosphatase Inhibitor |
| Spex Certiprep 6850 freezer mill | SPEX Sample Prep | 6850 | Freezer Mill |
| TALON Metal Affinity Resin | BD Biosiences | 635502 | For pulling down HIS tagged proteins |
| Tween 20 | VWR International | VW1521-07 | Non-ionic detergent |
| β-Mercaptoethanol | AMRESCO | M131-250ML | Reducing agent |
References
- Botstein, D., Chervitz, S. A., Cherry, J. M. Yeast as a model organism. Science. 277 (5330), 1259-1260 (1997).
- Dunn, B., Wobbe, C. R. Preparation of protein extracts from yeast. Current Protocols in Molecular Biology. , (2001).
- Holm, C., Meeks-Wagner, D. W., Fangman, W. L., Botstein, D. A rapid, efficient method for isolating DNA from yeast. Gene. 42 (2), 169-173 (1986).
- Nandakumar, M. P., Marten, M. R. Comparison of lysis methods and preparation protocols for one- and two-dimensional electrophoresis of Aspergillus oryzae intracellular proteins. Electrophoresis. 23 (14), 2216-2222 (2002).
- Silva, G. A. D., Bernardi, T. L., Schaker, P. D. C., Menegotto, M., Valente, P. Rapid yeast DNA extraction by boiling and freeze-thawing without using chemical reagents and DNA purification. Brazilian Archives of Biology and Technology. 55, 319-327 (2012).
- Baldwin, C., Robinson, C. W. Disruption of Saccharomyces cerevisiae using enzymatic lysis combined with high-pressure homogenization. Biotechnology Techniques. 4 (5), 329-334 (1990).
- Rigaut, G., et al. A generic protein purification method for protein complex characterization and proteome exploration. Nature Biotechnology. 17 (10), 1030-1032 (1999).
- Hudspeth, M. E., Shumard, D. S., Tatti, K. M., Grossman, L. I. Rapid purification of yeast mitochondrial DNA in high yield. Biochimica et Biophysica Acta. 610 (2), 221-228 (1980).
- Szymanski, E. P., Kerscher, O. Budding yeast protein extraction and purification for the study of function, interactions, and post-translational modifications. Journal of Visualized Experiments. (80), e50921 (2013).
- Bzducha-Wróbel, A., et al. Evaluation of the efficiency of different disruption methods on yeast cell wall preparation for β-glucan isolation. Molecules. 19 (12), 20941-20961 (2014).
- Umen, J. G., Guthrie, C. A novel role for a U5 snRNP protein in 3′ splice site selection. Genes & Development. 9 (7), 855-868 (1995).
- Smith, B. C., Fisher, D. L., Weedn, V. W., Warnock, G. R., Holland, M. M. A systematic approach to the sampling of dental DNA. Journal of Forensic Sciences. 38 (5), 1194-1209 (1993).
- Sweet, D., Hildebrand, D. Recovery of DNA from human teeth by cryogenic grinding. Journal of Forensic Sciences. 43 (6), 1199-1202 (1998).
- Lorenz, W. W., Yu, Y. S., Dean, J. F. An improved method of RNA isolation from loblolly pine (P. taeda L.) and other conifer species. Journal of Visualized Experiments. (36), e1751 (2010).
- Singh, M. R., Roy, S., Bellare, J. R. Influence of Cryogenic Grinding on Release of Protein and DNA from Saccharomyces cerevisiae. International Journal of Food Engineering. 5 (1), (2009).
- Singh, R. K., Kabbaj, M. H., Paik, J., Gunjan, A. Histone levels are regulated by phosphorylation and ubiquitylation-dependent proteolysis. Nature Cell Biology. 11 (8), 925-933 (2009).
- Liang, D., Burkhart, S. L., Singh, R. K., Kabbaj, M. H., Gunjan, A. Histone dosage regulates DNA damage sensitivity in a checkpoint-independent manner by the homologous recombination pathway. Nucleic Acids Research. 40 (19), 9604-9620 (2012).
- Singh, R. K., Gonzalez, M., Kabbaj, M. H., Gunjan, A. Novel E3 ubiquitin ligases that regulate histone protein levels in the budding yeast Saccharomyces cerevisiae. PLoS One. 7 (5), 36295 (2012).
- Gunjan, A., Verreault, A. A Rad53 kinase-dependent surveillance mechanism that regulates histone protein levels in S. cerevisiae. Cell. 115 (5), 537-549 (2003).
- Gill, P., et al. Identification of the remains of the Romanov family by DNA analysis. Nature Genetics. 6 (2), 130-135 (1994).
- Alain, K., et al. DNA extractions from deep subseafloor sediments: novel cryogenic-mill-based procedure and comparison to existing protocols. Journal of Microbiological Methods. 87 (3), 355-362 (2011).
- Mohammad, F., Buskirk, A. Protocol for Ribosome Profiling in Bacteria. Bio-Protocol. 9 (24), 3468 (2019).
- Liew, Y. J., et al. Identification of microRNAs in the coral Stylophora pistillata. PLoS One. 9 (3), 91101 (2014).
- Lopez de Heredia, M., Jansen, R. P. RNA integrity as a quality indicator during the first steps of RNP purifications: a comparison of yeast lysis methods. BMC Biochem. 5 (14), (2004).
- Grant, L. J., et al. Purified plant cell walls with adsorbed polyphenols alter porcine faecal bacterial communities during in vitro fermentation. Food & Function. 11 (1), 834-845 (2020).
- Lolo, M., et al. Cryogenic grinding pre-treatment improves extraction efficiency of fluoroquinolones for HPLC-MS/MS determination in animal tissue. Analytical and Bioanalytical Chemistry. 387 (5), 1933-1937 (2007).
- Santos, D., et al. Determination of Cd and Pb in food slurries by GFAAS using cryogenic grinding for sample preparation. Analytical and Bioanalytical Chemistry. 373 (3), 183-189 (2002).
- da Silva, E. G. P., et al. Fast method for the determination of copper, manganese and iron in seafood samples. Journal of Food Composition and Analysis. 21 (3), 259-263 (2008).
- Kamogawa, M. Y., Nogueira, A. R. A., Costa, L. M., Garcia, E. E., Nobrega, J. A. A new strategy for preparation of hair slurries using cryogenic grinding and water-soluble tertiary-amines medium. Spectrochimica Acta Part B-Atomic Spectroscopy. 56 (10), 1973-1980 (2001).
- Sillen, A., Hall, G., Richardson, S., Armstrong, R. Sr-87/Sr-86 ratios in modern and fossil food-webs of the Sterkfontein Valley: Implications for early hominid habitat preference. Geochimica Et Cosmochimica Acta. 62 (14), 2463-2473 (1998).
- Nielsen-Marsh, C. M., et al. Sequence preservation of osteocalcin protein and mitochondrial DNA in bison bones older than 55 ka. Geology. 30 (12), 1099-1102 (2002).

