Isolation and Screening from Soil Biodiversity for Fungi Involved in the Degradation of Recalcitrant Materials
Summary
Here, we present a protocol for screening soil biodiversity to look for fungal strains involved in the degradation of recalcitrant materials. First, fungal strains able to grow on humic acids or lignocellulose are isolated. Their activity is then tested both in enzymatic assays and on pollutants such as hydrocarbons and plastics.
Abstract
Environmental pollution is an increasing problem, and identifying fungi involved in the bioremediation process is an essential task. Soil hosts an incredible diversity of microbial life and can be a good source of these bioremediative fungi. This work aims to search for soil fungi with bioremediation potential by using different screening tests. Mineral culture media supplemented with recalcitrant substances as the sole carbon source were used as growth tests. First, soil dilutions were plated on Petri dishes with mineral medium amended with humic acids or lignocellulose. The growing fungal colonies were isolated and tested on different substrates, such as complex mixtures of hydrocarbons (petrolatum and used motor oil) and powders of different plastic polymers (PET, PP, PS, PUR, PVC). Qualitative enzymatic tests were associated with the growth tests to investigate the production of esterases, laccases, peroxidases, and proteases. These enzymes are involved in the main degradation processes of recalcitrant material, and their constitutive secretion by the examined fungal strains could have the potential to be exploited for bioremediation. More than 100 strains were isolated and tested, and several isolates with good bioremediation potential were found. In conclusion, the described screening tests are an easy and low-cost method to identify fungal strains with bioremediation potential from the soil. In addition, it is possible to tailor the screening tests for different pollutants, according to requirements, by adding other recalcitrant substances to minimal culture media.
Introduction
Soil is a fundamental component of life on Earth and is the basis of many ecosystems. The minerals, organic matter, and microorganisms in the soil can be considered as one system, with close associations and interactions occurring between them. The interactions of these compounds have an important impact on terrestrial processes, environmental quality, and ecosystem health1. Soil pollution poses serious environmental problems worldwide. The indiscriminate, long-term, and excessive application of recalcitrant and toxic substances, such as pesticides, petroleum products, plastics, and other chemicals, has serious effects on soil ecology and, as a result, can alter soil microbiota. Microbial communities in soils are composed of a wide range of organisms in different physiological states, with the majority being bacteria and fungi. Many of the contaminants in soils have medium- to long-term stability, and their persistence can lead to the development of adaptive mechanisms that allow the microorganisms to utilize recalcitrant substances as nutrients2,3. These microorganisms can, therefore, be considered for bioremediation techniques.
Bioremediation tries to mitigate the effects of pollution by using microorganisms and their enzymes for the degradation or transformation of waste into less toxic or non-toxic compounds. Various species of archaea, bacteria, algae, and fungi possess this bioremediation ability4. As a result of their particular biodegradative actions, fungi are especially promising organisms for bioremediation. They can attack different substrates using their hyphal network, enabling them to penetrate the soil matrix more efficiently than other microorganisms. Additionally, they can reach inaccessible interstices where contaminants are difficult to remove5, and they can also survive low moisture levels6. Moreover, fungi synthesize different cassettes of unspecific enzymes, usually to degrade natural recalcitrant substances such as cellulose, lignin, and humic acids. Those that lack the target substrate can be involved in the degradation of a wide range of recalcitrant pollutants, such as hydrocarbons, plastics, and pesticides7,8,9,10. Therefore, although many fungal species have already been reported as bioremediation agents, there is increasing interest in exploring species that have not yet been studied to select candidates for the bioremediation of recalcitrant contaminating substances. The species already known to have bioremediation properties belong to the phyla Ascomycota11,12,13, Basidiomycota14,15, and Mucoromycota. For example, the genera Penicillium and Aspergillus are well known to be involved in the degradation of aliphatic hydrocarbons13, different plastic polymers16,17,18, heavy metals19, and dyes20. Similarly, studies carried out on basidiomycetes fungi, such as Phanerochaete chrysosporium and Trametes versicolor, have revealed their involvement in the oxidation of recalcitrant materials such as aromatic hydrocarbons13 and plastics21. Another example of fungi involved in the biodegradation processes are the zygomycetes Rhizopus spp., Mucor spp., and Cunninghamella spp.22,23. In particular, Cunninghamella is able to oxidase aromatic hydrocarbons and is considered a model organism for studying the detoxification of products from a wide range of xenobiotics13.
There are several fungal enzymes involved in the major degradative processes of recalcitrant materials24,25, such as esterase, laccase, peroxidase, and protease. Laccases are copper-containing oxidases produced in the cell and subsequently secreted, that allow the oxidation of a variety of phenolic and aromatic compounds. They can degrade ortho and para diphenols, the amino group-containing phenols, lignin, and the aryl group-containing diamines26. Peroxidases use hydrogen peroxide as a mediator to degrade lignin and other aromatic compounds. There are many different peroxidases, but the ones with the greatest potential to degrade toxic substances are lignin peroxidase and manganese peroxidase27.
Esterases and proteases belong to the group of extra- or ecto-cellular enzymes, which act outside their cells of origin but are still bound to them. These enzymes can catalyze the hydrolysis of large recalcitrant molecules into smaller ones. Due to their low substrate specificity, these enzymes can play a key role in the bioremediation of various pollutants, such as textile dyes, effluents released from the pulp and paper industries and leather tanning, petroleum products, plastics, and pesticides28,29,30.
A number of screening methods to select for bioremediative fungal strains have already been published. For example, straw-based agar medium has been used to screen for white-rot fungi with high potential in the polycyclic aromatic hydrocarbons (PAH) degradation31; and small pieces of rotting wood have been placed onto malt extract agar (MEA) to isolate wood-rotting fungi32. However, most of the methods that have already been proposed select very specific fungi for their activity of interest. This research proposes a wider approach for selecting soil fungi with a broader range of action. The method relies on the initial plating of serial dilutions of soil samples onto a medium amended with humic acids or lignocellulose mixed with antibiotics to select fungi with the ability to degrade these natural recalcitrant substances. Humic acids and lignocellulose, in fact, are substances that are extremely resistant to biodegradation since they have very complex molecular structures, and this allows them to be excellent indicators of the degradative ability of the tested fungi33,34. Subsequently, the fungi selected in the first tests are screened to identify those with the potential to degrade specific pollutants such as petrolatum, used engine oil, and plastics. Finally, qualitative enzymatic tests are performed to detect fungal strains able to produce enzymes involved in the biodegradation processes of recalcitrant substances. For this purpose, protease and esterase tests are conducted, while gallic acid and guaiacol are used as indicators of laccase and other ligninolytic enzyme production35,36. These substrates are used because a strong correlation has been found between the ability of fungi to oxidize them to their brown-colored form and the possession of ligninolytic ability37,38,39.
Through these protocols, it is possible to isolate fungal strains with high degradative potential and a broad spectrum of action directly from soil samples. The isolation of these fungal strains could help find new candidates for bioremediation purposes.
Protocol
1. Selection of fungal strains able to degrade recalcitrant materials from soil
- Preparation of antibiotic solution.
- Put penicillin (50 mg/L), streptomycin (40 mg/L), chlortetracycline (40 mg/L), neomycin (100 mg/L), and chloramphenicol (100 mg/L) into 250 mL of deionized sterile water.
- Before adding chloramphenicol to the antibiotic solution, dissolve it into 3 mL of ≥99% ethanol.
- Place the antibiotic solution on a magnetic stirrer (no heat) with a magnetic stirring bar inside the solution for 10 min. Store it at 4 °C.
- Preparation of growth media.
- Put 1 g of commercial humic acid, 3.26 g of Bushnell-Haas broth (BH), and 15 g of agar into 1 L of deionized water and autoclave it for 20 min at 121 °C (humic acid agar).
- Plate the humic acid agar into Petri dishes under a laminar flow cabinet.
- Put 4 g of lignocellulose, 3.26 g of BH, and 15 g of agar into 1 L of deionized water and autoclave it for 20 min at 121 °C (lignocellulose agar).
- Plate the lignocellulose agar into Petri dishes under a laminar flow cabinet.
- Under a laminar flow cabinet, after the media have solidified in the Petri dishes, transfer 1 mL of the antibiotic solution onto the prepared plates using a sterile syringe with a 0.2 µm filter to sterilize it.
- Distribute the antibiotic solution evenly on the plate surfaces using a sterile disposable L-shaped cell spreader and let it air-dry under the laminar flow cabinet.
- Put 20 g of malt extract broth and 15 g of agar into 1 L of deionized water and autoclave it for 20 min at 121 °C (malt extract agar – MEA).
- Plate the MEA into Petri dishes under a laminar flow cabinet.
- Soil sampling and soil dilutions.
- Sample the soil of interest at a depth of 10 cm, removing any plants, grass, and dead leaves from the top layer, and put it into sterile polyethylene bags.
- After reaching the laboratory, sieve the soil using a 2 mm mesh, removing any roots and plant debris.
- If it is not possible to proceed with the downstream analysis right away, store the sieved soil samples at 4 °C.
- Working under a laminar flow cabinet, put 1 g of the sieved soil into a sterile 15 mL tube with 10 mL of sterile deionized water (1 x 10-1 dilution). Shake the tube horizontally for 30 min.
- Under a laminar flow cabinet, before the suspension settles, take 1 mL of the 1 x 10-1 dilution with a sterile pipette and transfer it to a 9 mL of deionized water blank (1 x 10-2 dilution). Vortex it thoroughly.
- Before the suspension settles, take 1 mL of the 1 x 10-2 dilution with a sterile pipette and transfer it to a 9 mL of deionized water blank (1 x 10-3 dilution). Vortex it thoroughly.
- Plating soil dilutions on selective media.
- Working under a laminar flow cabinet, collect 100 µL of the 1 x 10-3 dilution using a micropipette with a sterile point and transfer it onto a humic agar Petri dish. Do this for at least 4 or 5 Petri dishes.
- Distribute the dilution evenly on the Petri dish surface using a sterile disposable L-shaped cell spreader.
- Let it air-dry for 10-15 min under the laminar flow cabinet.
- Repeat Steps 1.4.1.-1.4.3. using lignocellulose Petri dishes.
- Incubate at 25 °C in the dark for a maximum of 15 days.
- Isolation of fungal colonies grown from selective media to growth media.
- Starting from 3 days after preparation, check the prepared selective media Petri dishes daily for any fungal colony growth for a maximum of 15 days.
- Check all the fungal colonies present for similarities. If necessary, prepare a slide to be observed under a light microscope.
NOTE: The aim is to isolate the highest number of fungal colonies, but it is necessary to avoid always isolating the same fungal strain from different plates, so this check is very important. Look for colonies with different macro and micromorphology. - Under a laminar flow cabinet or at a Bunsen burner, isolate each chosen fungal strain by gently removing a small part of the mycelium from the colony with a sterile inoculation needle and transferring it to a new MEA Petri dish.
NOTE: In this phase, it is very important to be delicate and precise because there is a high risk of contamination from the other fungal colonies present in the original Petri dish. If more than one fungal strain grows on the inoculated MEA plate, repeat Step 1.5.3. - Incubate at 25 °C in the dark for 7 days. These are the fungal strains to be tested further on specific recalcitrant substances.
2. Growth tests on recalcitrant substances
- Growth test on petrolatum.
- Transfer 20 mL of liquid BH (3.26 g/L of BH) into 50 mL vials and sterilize them by autoclaving at 121 °C for 20 min.
- Transfer 7 mL of deionized water into 15 mL glass vials and add some pieces of broken glass coverslips into each vial.
- Sterilize them by autoclaving at 121 °C for 20 min.
- Under the laminar flow cabinet, add 1 mL of petrolatum to each 50 mL BH vial using a sterile pipette. Keep some vials with only BH as the negative control.
- Under the laminar flow cabinet, remove the mycelium on the surface of the medium from the MEA plates with the chosen fungal strains (prepared at Step 1.5.3.) using a sterile needle and transfer it to 15 mL glass vials with the broken coverslips.
- Agitate each vial on a vortex mixer for 2 min, allowing the broken coverslips to cut the cube of the fungal colony, making a fungal suspension.
- Inoculate each petrolatum vial with 200 µL of fungal suspension. Make two replicates for each fungal strain.
- For the negative control, put 200 µL of fungal suspension into vials with only liquid BH. Moreover, keep a couple of vials with petrolatum without any fungal inoculation to check for contamination in the medium.
- Incubate the vials at 25 °C in the dark and check them after 15 days and 30 days from the inoculum.
- Growth test on used engine oil.
- Put 3.26 g of BH and 15 g of agar into 1 L of deionized water and autoclave it for 20 min at 121 °C (BHA).
- Plate the BHA into Petri dishes under a laminar flow cabinet.
- When it has solidified, transfer 500 µL of used engine oil onto the BHA Petri dishes and distribute it evenly on the plate surfaces using a sterile disposable L-shaped cell spreader.
- Inoculate each used engine oil BHA plate with each fungal strain, collecting the mycelium from the plates prepared at Step 1.5.3. Work at a Bunsen burner or under a laminar flow hood to avoid contamination. Make two replicates for each fungal strain.
- For the negative control, inoculate the Petri dishes with BHA only. Moreover, keep a couple of used engine oil BHA Petri dishes without any fungal inoculation to check for contamination in the medium.
- Incubate the Petri dishes at 25 °C in the dark and check them after 15 days and 30 days from the inoculum.
- Growth test on plastics.
- Transfer 200 µL of fungal suspension (Steps 2.1.5.-2.1.6.) into a 96-microwell plate. Make three replicates of the fungal strain-plastic combination.
- To each well with the fungal suspension, add 10 mg of a different kind of plastic powder (polyethylene terephthalate – PET, polyvinyl chloride – PVC, high-density polyethylene – HDPE, polystyrene – PS, polyurethane – PUR).
- For the negative control, instead of adding the plastic powder, add 200 µL of sterilized BH solution to the well with the fungal suspension. Moreover, for each type of plastic powder, fill a well with 200 µL of sterilized BH solution and 10 mg of each plastic dust to check for contamination of the medium.
- Incubate the microwell plates at 25 °C in the dark and check them after 15 days and 30 days from the inoculum.
3. Qualitative enzymatic tests
- Qualitative ligninolytic enzymes colorimetric test.
- Put 27 g of potato dextrose broth (PD) and 15 g of agar into 1 L of deionized water and autoclave it for 20 min at 121 °C (PDA).
- When the medium has cooled down a little but is still liquid, add gallic acid (5 g/L) under a laminar flow hood and plate the medium into Petri dishes.
- Inoculate the PDA + gallic acid plates with each fungal strain, collecting the mycelium from the plates prepared at Step 1.5.3. Work at a Bunsen burner or under a laminar flow hood to avoid contamination.
- As negative controls, prepare Petri dishes with only PDA (no gallic acid) and inoculate them with the fungal strains (one for each fungal strain), as well as one Petri dish with PDA + gallic acid and no fungal inoculation.
- Incubate the Petri dishes at 25 °C in the dark and check them after 7 days from the inoculum.
- Qualitative laccases colorimetric test.
- Put 27 g of PD and 15 g of agar into 1 L of deionized water and autoclave it for 20 min at 121 °C (PDA).
- When the medium has cooled down a little but is still liquid, add guaiacol (400 µL/L) under a laminar flow hood and plate it into Petri dishes.
- Inoculate the PDA + guaiacol plates with each fungal strain, collecting the mycelium from the plates prepared at Step 1.5.3. Work at a Bunsen burner or under a laminar flow hood to avoid any contamination.
- As negative controls, prepare Petri dishes with only PDA (no guaiacol) And inoculate them with the fungal strains (one for each fungal strain), as well as one Petri dish with PDA + guaiacol and no fungal inoculation.
- Incubate the Petri dishes at 25 °C in the dark and check them after 7 days from the inoculum.
- Qualitative proteases test.
- Prepare the YES medium by adding K2HPO4 (1 g/L) , KH2PO4 (0.5 g/L), MgSO4·7H2O (0.5 g/L), MnCl2·4H2O (0.001 g/L), CuCl2·2H2O (1.4 x 10-5 g/L), ZnCl2 (1.1 x 10-5 g/L), CoCl2·6H2O (2 x 10-5 g/L), Na2MoO4·2H2O (1.3 x 10-5 g/L), FeCl3·6H2O (7.5 x 10-5 g/L), and gelatin/peptone (0.02 g/L) to deionized water.
- Place the YES medium on a magnetic stirrer (no heat) with a magnetic stirring bar inside the medium for 10 min.
- When all the salts have dissolved into the medium, transfer 5 mL of the solution into 20 mL sterile glass vials and autoclave them for 20 min at 121 °C.
- For the negative control, prepare some 20 mL sterile glass vials with 5 mL of BH solution and autoclave them for 20 min at 121 °C.
- Under a laminar flow cabinet, transfer 200 µL of fungal suspension (Steps 2.1.5.-2.1.6.) for each strain to the YES medium sterilized vials (make at least 2 replicates per fungal strain). For each fungal strain, add 200 µL of the same fungal suspension to BH sterilized vials to have a negative control.
- Incubate the vials at 25 °C in the dark for 21 days and check them every 7 days.
- Qualitative esterases test.
- Prepare the Tween 80 medium by adding peptone (10 g/L), NaCl (5 g/L), CaCl2 (0.1 g/L), and 10 mL of Tween 80 to 1 L of deionized water.
- Place the Tween 80 medium on a magnetic stirrer (no heat) with a magnetic stirring bar inside the medium for 10 min.
- When everything is melted inside the medium, transfer 5 mL of the solution into 20 mL sterile glass vials and autoclave them for 20 min at 121 °C.
- For the negative control, prepare some 20 mL sterile glass vials with 5 mL of BH solution and autoclave them for 20 min at 121 °C.
- Under a laminar flow cabinet, transfer 200 µL of fungal suspension (Steps 2.1.5.-2.1.6.) for each strain to the Tween 80 medium sterilized vials (make at least 2 replicates per fungal strain). For each fungal strain, add 200 µL of the same fungal suspension to BH sterilized vials for negative control.
- Incubate the vials at 25 °C in the dark for 21 days and check them every 7 days.
Representative Results
The selective media methods (Section 1 of the protocol) allowed the rich biodiversity of soil to be screened and the fungi with high bioremediation potential to be selected. With the humic acid and lignocellulose media, more than 100 fungal strains were isolated. These fungi produced enzymes involved in the biodegradation of natural recalcitrant materials, which have a chemical structure resembling many pollutants. However, the fungal strains isolated with the selective media needed further screening. Specifically, the selective tests isolated the strains able to degrade humic acids and lignocellulose, the growth tests screened the isolated fungi for those able to use a specific pollutant as their sole carbon source, and the qualitative enzymatic tests described their metabolic activities.
The growth tests were focused on the fungal ability to degrade hydrocarbons (petrolatum and used engine oil) and plastics (PET, PVC, HDPE, PS, PUR). Each of these substances was used in a test as the sole carbon source for the selected fungal strains. The fungal ability to exploit these substances was evaluated as the difference in colony growth between the samples with the added substance and the control samples with only minimal medium (BH or BHA, depending on the test). A qualitative scale was used to describe the results (Figure 1), ranging from the absence of growth difference with the control (0) to low (+), medium (++), and high (+++) levels of difference in growth.
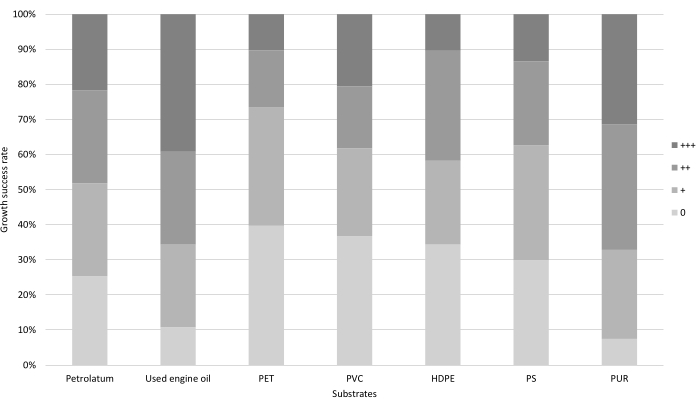
Figure 1: Rate of growth success of fungal strains in the growth tests. The percentage of fungal strains able to grow on petrolatum, used engine oil, and plastic powders (polyethylene terephthalate, PET; polyvinyl chloride, PVC; high-density polyethylene, HDPE; polystyrene, PS; polyurethane, PUR) as their sole carbon source with different degrees of success (no growth, 0; low growth, +; medium growth, ++; high growth, +++). Please click here to view a larger version of this figure.
The petrolatum and used engine oil tests evaluated the hydrocarbon biodegradation potential of the selected fungal strains. Petrolatum, a mixture of long-chain alkanes (>C25), was used as the sole carbon source. The fungal ability to exploit this substance was evaluated as the difference in colony growth between vials with BH + petrolatum and control vials with only BH (Figure 2). Almost 75% of the tested fungal strains were able to use petrolatum as their sole carbon source, and 21% of the strains exhibited maximum growth (+++, Figure 1).
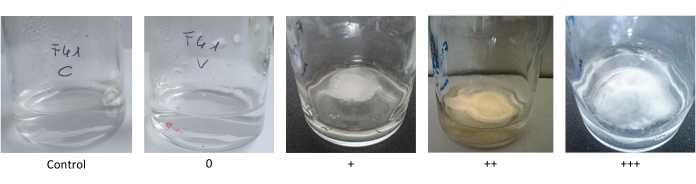

Figure 2: Pictures of the qualitative scale of growth on petrolatum as the sole carbon source. The qualitative scale is based on the growth difference between the treated samples and BH control. It ranges from the absence of growth difference (0), to low (+), medium (++), and high (+++) levels of growth difference. Please click here to view a larger version of this figure.
Used engine oil is a complex mixture of hydrocarbons, engine additives, and metals. It was used in the growth tests to evaluate the fungal ability to utilize a complex and toxic hydrocarbon blend as a carbon source. Like the other growth tests, growth was evaluated by comparing the fungal growth in plates with the engine oil with growth in the control plates containing just BHA (Figure 3). The growth results confirmed the efficacy of the selective media (Section 1 of protocol). Indeed, almost 90% of the isolated fungal strains were able to grow on used engine oil, with 39% showing maximum growth (+++, Figure 1).
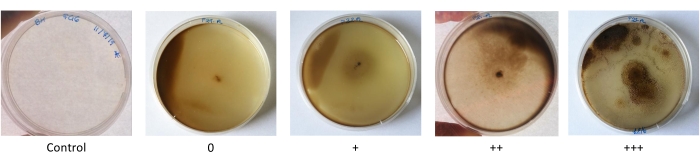
Figure 3: Pictures of the qualitative scale of growth on used engine oil as the sole carbon source. The qualitative scale is based on the growth difference between the treated samples and BHA control. It ranges from absence of growth difference (0), to low (+), medium (++), and high (+++) levels of growth difference. Please click here to view a larger version of this figure.
Growth tests on plastic powders were used to select the fungal strains that could use specific plastic polymers as their primary carbon source; hence, these strains have a high potential for plastic bioremediation. The growth in multiwell plates was observed by stereomicroscope and evaluated using a qualitative scale, ranging from the absence of growth (0) to high production of hyphae (+++). PUR was the plastic powder that had the highest percentage of fungal strains showing growth (92%), with 31% of strains showing maximum growth (+++). Approximately 70% of the strains isolated from humic acid and lignocellulose media grew on PS powder, while almost 60%-65% of them were able to use HDPE, PVC, and PET as their sole carbon source. Therefore, a large percentage of fungi were able to grow on the different plastic powders in multiwell plates, but a very low number of them showed high colonization of plastics. Indeed, only 10% of the strains were able to thrive on PET and HDPE, and 13% were able to thrive on PS. The only plastic where high fungal growth was regularly observed was PUR, on which approximately 30% of fungal strains managed to grow very abundantly.
Qualitative enzymatic tests were performed to detect fungal strains that can produce enzymes involved in the biodegradation processes of recalcitrant substances. Peroxidases and laccases are ligninolytic enzymes whose production is indicated by a brownish halo on the underside of the medium in gallic acid and guaiacol tests (Step 3.1. and Step 3.2. of the protocol). In both tests, the increase in the intensity of the color indicated a higher production of these enzymes (Figure 4).
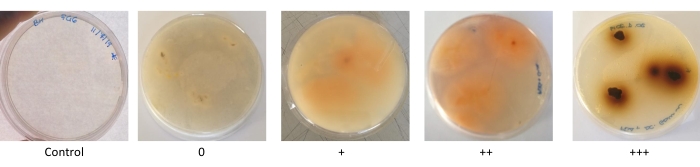
Figure 4: Pictures of the qualitative scale of qualitative ligninolytic enzymes. The qualitative scale is based on the production of a red/brownish halo around the colony. 0 = no activity, + = some activity, ++ = high activity, +++ very high activity. Please click here to view a larger version of this figure.
Approximately 60% of the fungi tested were able to produce ligninolytic enzymes, indicating the success of the selective media. Moreover, 12% of the isolated fungal strains produced an intense dark halo around the colony (+++) in gallic acid medium, with 17% doing so in guaiacol medium (laccases), indicating high fungal secretion of these enzymes (Figure 5).
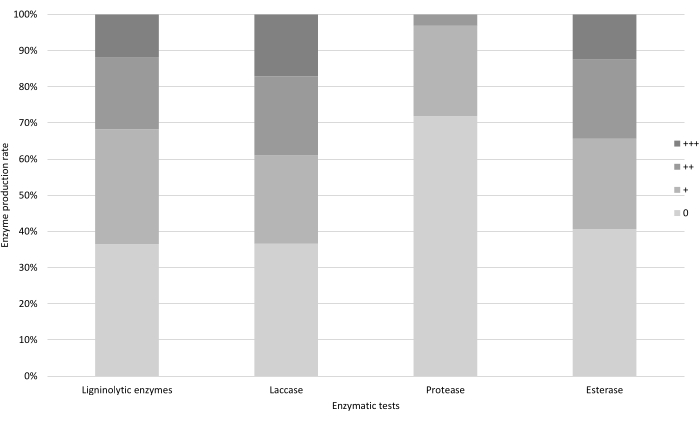
Figure 5: Rate of enzyme activity by fungal strains during qualitative enzymatic tests. The percentage of fungal strains able to produce ligninolytic enzymes, laccases, proteases, and esterases with different levels of success (0 = no activity, + = activity, ++ = high activity, +++ very high activity) during their corresponding qualitative enzymatic tests. Please click here to view a larger version of this figure.
Another screening test used to analyze fungal activity was the proteases test, which was based on the difference in fungal growth between YES medium vials and control vials with BH. However, approximately 70% of the fungal strains in the YES medium showed similar growth to controls, or even decreased growth. No fungal strain reached the maximum level of difference (+++). These results suggest that this test may be unsuitable for screening for protease activity.
Finally, esterases activity was evaluated by the formation of a white precipitate in Tween 80 medium (Figure 5). Almost 60% of the selected fungal strains showed esterase activity, and 13% displayed a high formation of the precipitate (+++), suggesting that the strains in the 13% should be selected to scrutinize their biodegradation potential (Figure 5).
Regarding all the growth and enzymatic tests, the most successful strains (i.e., those that obtained +++ in at least one test) were the most interesting. These fungal strains were able to efficiently use recalcitrant substances as their sole carbon source, or they produced a high level of enzymes involved in the biodegradation of recalcitrant substances. Indeed, 39 of the 115 fungal strains tested were able to thrive (+++) on hydrocarbon sources (petrolatum or used engine oil), and 58 had high growth on at least one plastic polymer. Only 32 showed very high enzymatic activity (+++) in at least one of the enzymatic tests performed. These results show the high efficacy of all the reported screening tests, apart from the proteases test, which did not select any high-performing strains.
Discussion
The rich biodiversity of soil is an abundant source of fungi that possess numerous metabolic abilities, some of which could be potential candidates for bioremediation. Selective media tests (Section 1 of the protocol) are easy-to-perform and effective methods for isolating fungi able to grow on natural complex polymers as their sole carbon source. Fungi can produce extracellular, non-specific hydrolases and oxidoreductases30 such as the ligninolytic enzymes laccases and peroxidases31. These enzymes can degrade lignocellulose and humic acids, but also many different xenobiotics, including hydrocarbons, aromatic compounds, and chlorinated organic compounds32.
The described methods produce results that are easy to interpret and are low-cost to carry out. These characteristics allow them to be used by non-experts. However, certain aspects, such as sterility and morphological identification, require special attention. For example, the 30 min agitation of the soil suspension (Step 1.3.4.) is necessary so that the fungal spores and propagules are freed from the soil microstructure and can be suspended in the solution. In this way, by plating dilutions of this soil suspension, it is possible to isolate the maximum number of fungal strains. Macro and microscopic morphological identification of fungal strains are important in order to differentiate between strains and avoid isolating the same fungal strain multiple times. Moreover, sterility and avoidance of microbial contaminations are crucial because the presence of other active fungal strains could misrepresent the activity of the studied colonies.
After the selective tests, an array of growth tests were performed on different recalcitrant substances, such as petrolatum, used engine oil, and different plastic polymers. This stage can be customized according to the recalcitrant substances of interest. The important step is to add the selected substance to BH or BHA as the sole carbon source and to check the fungal growth against a control without the added recalcitrant substance.
The aim of the growth tests on plastics and hydrocarbons was to assess whether the fungus could use these substances as its sole carbon source by qualitatively observing its growth on the material. Unfortunately, due to the extreme recalcitrance of these materials, it is very difficult to quantify the actual decrease in the weight of the substrate and the increase in the weight of the fungal biomass, especially after a relatively short amount of time. If a fungal strain performs very well in these screening growth tests, further studies should be undertaken to quantify the degradation of the recalcitrant substance.
The used enzymatic tests (Section 3 of the protocol) were chosen because they proved to be efficient and fast to perform, but they produce only qualitative results. They highlight the degradation potential of a fungal strain, but they cannot precisely quantify it. If a strain of particular interest is isolated, further tests, such as spectrophotometric quantitative enzyme essays, could be carried out to describe the metabolic activity more precisely. The only enzymatic test that showed suboptimal efficacy was the qualitative proteases test, with a low number of strains able to grow on the YES medium. Indeed, the high percentage of strains able to grow on PUR suggests that the fungi produced proteases involved in breaking the amide and urethane bonds, with esterase targeting the ester bonds40. The almost complete absence of protease-producing fungi suggests there may have been a problem in the method used. Other tests could be adopted for protease activity, such as casein41, skimmed milk42, or milk powder43 agar plate tests. The gallic acid test proved to be very useful for detecting the production of ligninolytic enzymes, but, unfortunately, it was not very specific and could not distinguish the type of ligninolytic enzyme produced (either peroxidases or laccases).
The results of this work suggest that the use of lignocellulose or humic acids as selective media allows fungal strains with bioremediation potential to be isolated from the rich soil biodiversity. Indeed, more than 70% of the fungal strains isolated by the selective methods were able to grow on petrolatum or used engine oil, and 60% were able to grow on different plastic polymers as the sole carbon source. Moreover, the qualitative enzymatic tests described the metabolic activity of these fungi linked to biodegradation processes. The selection of fungal strains showing +++ activity in one or more tests is recommended because it increases the chances of finding a fungal strain that is very active in bioremediation. Tests using gas mass-chromatography (GC-MS) on the recalcitrant substances after prolonged contact with the fungal strains selected with our screening tests have shown actual biodegradation of the materials9,44. The use of these simple screening tests in different ecosystems will allow the investigation of fungal biodiversity in different soils. Searching for fungi that can degrade recalcitrant materials will increase the number of identified strains that have the potential for bioremediation and help tackle the growing problem of environmental pollution.
Divulgations
The authors have nothing to disclose.
Acknowledgements
We acknowledge Scuola di Alta Formazione Dottorale (SAFD) of the University of Pavia and Professor Solveig Tosi for providing the opportunity for this work.
Materials
| 96 microwell plate | Greiner bio-one | 650185 | |
| Agar | VWR | 84609.05 | |
| Bushnell-Haas Broth | Fluka | B5051 | |
| CaCl2 | Sigma-Aldrich | C5670 | |
| Chloroamphenicol | Eurobio | GABCRL006Z | |
| Chlortetracycline | Sigma-Aldrich | Y0001451 | |
| CoCl2·6H2O | Sigma-Aldrich | C8661 | |
| CuCl2·2H2O | Sigma-Aldrich | C3279 | |
| Ethanol | VWR Chemicals | 20821.296 | |
| FeCl3·6H2O | Sigma-Aldrich | 236489 | |
| Filter 0.2 µm | Whatman | 10462200 | |
| gallic acid | Sigma-Aldrich | G7384 | |
| Glass cover slips | Biosigma | VBS634 | |
| Glass vials 15 mL | SciLabware | P35467 | |
| guaiacol | Sigma-Aldrich | G5502 | |
| High-density polyethylene (HDPE) | Sigma-Aldrich | 434272 | |
| Humic acids | Aldrich Chemistry | 53680 | |
| K2HPO4 | Sigma-Aldrich | P8281 | |
| KH2PO4 | Sigma-Aldrich | P5655 | |
| Lignocellulose | / | / | Sterilized bioethanol production waste |
| L-shaped cell spreader | Laboindustria S.p.a | 21133 | |
| magnetic stirrer | A.C.E.F | 8235 | |
| Malt Extract Broth | Sigma-Aldrich | 70146 | |
| MgSO4·7H2O | Sigma-Aldrich | M2643 | |
| Micropipette 1000 μL | Gilson | FA10006M | |
| Micropipette 200 μL | Gilson | FA10005M | |
| MnCl2·4H2O | Sigma-Aldrich | M5005 | |
| Na2MoO4·2H2O | Sigma-Aldrich | M1651 | |
| NaCl | Sigma-Aldrich | S5886 | |
| Neomycin | Sigma-Aldrich | N0401000 | |
| Penicillin | Sigma-Aldrich | 1504489 | |
| peptone | Sigma-Aldrich | 83059 | |
| Polyethylene terephthalate (PET) | Goodfellow | ES306031 | |
| Petri dishes | Laboindustria S.p.a | 21050 | |
| Petrolatum (Paraffin liquid) | A.C.E.F | 009661 | |
| Potato Dextrose Broth | Sigma-Aldrich | P6685 | |
| Polystyrene (PS) | Sigma-Aldrich | 331651 | |
| Polyurethane (PUR) | Sigma-Aldrich | GF20677923 | |
| Polyvinyl chloride (PVC) | Sigma-Aldrich | 81388 | |
| Sterile falcon tube | Greiner bio-one | 227 261 | |
| Sterile glass vials 20 mL | Sigma-Aldrich | SU860051 | |
| Sterile point 1000 μL | Gilson | F172511 | |
| Sterile point 200 μL | Gilson | F172311 | |
| Sterile polyethylene bags | WHIRL-PAK | B01018 | |
| sterile syringe | Rays | 5523CM25 | |
| Streptomycin | Sigma-Aldrich | S-6501 | |
| Tween 80 | Sigma-Aldrich | P1754 | |
| Used engine oil | / | / | complex mixture of hydrocarbons, engine additives, and metals, provided by an Italian private company |
| Vials 50 mL | Sigma-Aldrich | 33108-U | |
| ZnCl2 | Sigma-Aldrich | Z0152 |
References
- Mohammadi, K., Heidari, G., Khalesro, S., Sohrabi, Y. Soil management, microorganisms and organic matter interactions: A review. African Journal of Biotechnology. 10 (86), 19840-19849 (2011).
- Daccò, C., Girometta, C., Asemoloye, M. D., Carpani, G., Picco, A. M., Tosi, S. Key fungal degradation patterns, enzymes and their applications for the removal of aliphatic hydrocarbons in polluted soils: A review. International Biodeterioration and Biodegradation. 147, (2020).
- Asemoloye, M. D., Ahmad, R., Jonathan, S. G. Synergistic action of rhizospheric fungi with Megathyrsus maximus root speeds up hydrocarbon degradation kinetics in oil polluted soil. Chemosphere. 187, 1-10 (2017).
- Abatenh, E., Gizaw, B., Tsegaye, Z., Wassie, M. The role of microorganisms in bioremediation – A review. Open Journal of Environmental Biology. 2, 038-046 (2017).
- Dix, N. J., Webster, J. . Fungal Ecology. , (1995).
- Magan, N., Esser, K., Lemke, P. A. Fungi in extreme environment. Environmental and Microbial Relationships. The Mycota. 4, 99-114 (2007).
- Aranda, E. Promising approaches towards biotransformation of polycyclic aromatic hydrocarbons with Ascomycota fungi. Current Opinion in Biotechnology. 38, 1-8 (2016).
- Hasan, I. F., AI-Jawhari, V., Kumar, M., Kumar, R., Prasad, Role of Filamentous Fungi to Remove Petroleum Hydrocarbons from the Environment. Microbial Action on Hydrocarbons. , (2018).
- Daccò, C., et al. Trichoderma: evaluation of its degrading abilities for the bioremediation of hydrocarbon complex mixtures. Applied Sciences. 10 (9), 3152 (2020).
- Alarcón, A., Davies, F. T., Autenrieth, R. L., Zuberer, D. A. Arbuscular mycorrhiza and petroleum-degrading microorganisms enhance phytoremediation of petroleum-contaminated soil. International Journal of Phytoremediation. 10, 251-263 (2008).
- Mancera-López, M. E., et al. Bioremediation of an aged hydro-carbon-contaminated soil by a combined system of biostimulation-bioaugmentation with filamentous fungi. International Biodeterioration and Biodegradation. 61, (2008).
- Hatami, E., Abbaspour, A., Dorostkar, V. Phytoremediation of a petroleum-polluted soil by native plant species in Lorestan Province, Iran. Environmental Science and Pollution Research. 26, 24323-24330 (2019).
- Prenafeta-Boldú, F. X., De Hoog, G. S., Summerbell, R. C. Fungal communities in hydrocarbon degradation. Microbial Communities Utilizing Hydrocarbons and Lipids: Members, Metagenomics and Ecophysiology. Handbook of Hydrocarbon and Lipid Microbiology. , 1-36 (2018).
- Gu, J., Ford, T., Mitton, D., Mitchell, R. Microbiological degradation of polymeric materials. Uhlig’s Corrosion Handbook. , 421-438 (2011).
- Tuomela, M., Hatakka, A. Oxidative fungal enzymes for bioremediation. Comprehensive Biotechnology: Environmental and Related Biotechnologies. 6, 224-239 (2019).
- DSouza, G. C., et al. Fungal biodegradation of low-density polyethylene using consortium of Aspergillus species under controlled conditions. Heliyon. 7 (5), 07008 (2021).
- El-Sayed, M. T., Rabie, G. H., Hamed, E. A. Biodegradation of low-density polyethylene (LDPE) using the mixed culture of Aspergillus carbonarius and A. fumigates. Environment, Development, and Sustainability. 23 (10), 14556-14584 (2021).
- Sepperumal, U., Markandan, M., Palraja, I. Micromorphological and chemical changes during biodegradation of polyethylene terephthalate (PET) by Penicillium sp. Journal of Microbiology and Biotechnology Research. 3 (4), 47-53 (2013).
- Leitão, A. L. Potential of Penicillium species in the bioremediation field. International Journal of Environmental Research and Public Health. 6 (4), 1393-1417 (2009).
- Chen, S. H., Ting, A. S. Y. Biosorption and biodegradation potential of triphenylmethane dyes by newly discovered Penicillium simplicissimum isolated from indoor wastewater sample. International Biodeterioration & Biodegradation. 103, 1-7 (2015).
- Orhan, Y., Buyukgungor, H. Enhancement of biodegradability of disposable polyethylene in controlled biological soil. International Biodeterioration and Biodegradation. 45, 49-55 (2000).
- Deshmukh, R., Khardenavis, A. A., Purohit, H. J. Diverse metabolic capacities of fungi for bioremediation. Indian journal of microbiology. 56 (3), 247-264 (2016).
- Viswanath, B., Rajesh, B., Janardhan, A., Kumar, A. P., Narasimha, G. Fungal laccases and their applications in bioremediation. Enzyme research. 2014, 163242 (2014).
- Ali, M., Husain, Q., Ishqi, H. M. Fungal peroxidases mediated bioremediation of industrial pollutants. Fungal Bioremediation. , (2019).
- Nousiainen, P., Kontro, J., Manner, H., Hatakka, A., Sipilä, J. Phenolic mediators enhance the manganese peroxidase catalyzed oxidation of recalcitrant lignin model compounds and synthetic lignin. Fungal Genetics and Biology. 72, 137-149 (2014).
- Srivastava, S., Kumar, M. Biodegradation of polycyclic aromatic hydrocarbons (PAHs): A sustainable approach. Sustainable Green Technologies for Environmental Management. , (2019).
- Wei, R., Zimmermann, W. Microbial enzymes for the recycling of recalcitrant petroleum-based plastics: how far are we. Microbial Biotechnology. 10 (6), 1308-1322 (2017).
- Matsubara, M., Lynch, J. M., De Leij, F. A. A. M. A simple screening procedure for selecting fungi with potential for use in the bioremediation of contaminated land. Enzyme and Microbial Technology. 39 (7), 1365-1372 (2006).
- Mann, J., et al. Screening and selection of fungi for bioremediation of olive mill wastewater. World Journal of Microbiology and Biotechnology. 26 (3), 567-571 (2010).
- Andlar, M., Rezić, T., Marđetko, N., Kracher, D., Ludwig, R., Šantek, B. Lignocellulose degradation: an overview of fungi and fungal enzymes involved in lignocellulose degradation. Engineering in Life Sciences. 18, 768-778 (2018).
- Goméz-Toribio, V., García-Martín, A. B., Martínez, M. J., Martínez, A. T., Guillén, F. Induction of extracellular hydroxyl radical production by white-rot fungi through quinone redox cycling. Applied and Environmental Microbiology. 75, 3944-3953 (2009).
- Belcarz, A., Ginalska, G., Kornillowicz-Kowalska, T. Extracellular enzyme activities of Bjerkandera adusta R59 soil strain, capable of daunomycin and humic acids degradation. Applied Microbiology and Biotechnology. 68 (5), 686-694 (2005).
- Stevenson, F. J. . Humus Chemistry: Genesis, Composition, Reactions. 2nd ed. , (1995).
- Andlar, M., et al. Lignocellulose degradation: An overview of fungi and fungal enzymes involved in lignocellulose degradation. Engineering in Life Sciences. 18 (11), 768-778 (2018).
- Lee, H., et al. Biotechnological procedures to select white rot fungi for the degradation of PAHs. Journal of Microbiological Methods. 97 (1), 56-62 (2014).
- Batista-García, R. A., et al. Simple screening protocol for identification of potential mycoremediation tools for the elimination of polycyclic aromatic hydrocarbons and phenols from hyperalkalophile industrial effluents. Journal of Environmental Management. 198, 1-11 (2017).
- Shleev, S. V., et al. Comparison of physico-chemical characteristics of four laccases from different basidiomycetes. Biochimie. 86 (9-10), (2004).
- Kiiskinen, L. L., Rättö, M., Kruus, K. Screening for novel laccase-producing microbes. Journal of Applied Microbiology. 97, (2004).
- Kumar, V. V., Rapheal, V. S. Induction and purification by three-phase partitioning of aryl alcohol oxidase (AAO) from Pleurotus ostreatus. Applied Biochemistry and Biotechnology. , 163 (2011).
- Loredo-Treviño, A., Gutiérrez-Sánchez, G., Rodríguez-Herrera, R., Aguilar, C. N. Microbial enzymes involved in polyurethane biodegradation: a review. Journal of Polymers and the Environment. 20 (1), 258-265 (2012).
- Garriga, M., et al. Technological and sensorial evaluation of Lactobacillus strains as starter cultures in fermented sausages. International Journal of Food Microbiology. 32 (1-2), 173-183 (1996).
- Zerdani, I., Faid, M., Malki, A. Feather wastes digestion by new isolated strains Bacillus sp. In microcco. African Journal of Biotechnology. 3 (1), 67-70 (2004).
- Nygren, C. M., Edqvist, J., Elfstrand, M., Heller, G., Taylor, A. F. Detection of extracellular protease activity in different species and genera of ectomycorrhizal fungi. Mycorrhiza. 17 (3), 241-248 (2007).
- Asemoloye, M. D., et al. Hydrocarbon degradation and enzyme activities of Aspergillus oryzae and Mucor irregularis isolated from Nigerian crude oil-polluted sites. Microorganisms. 8 (12), 1912 (2020).

