Natural Transformation, Protein Expression, and Cryoconservation of the Filamentous Cyanobacterium Phormidium lacuna
Summary
Phormidium lacuna is a filamentous cyanobacterium that was isolated from marine rockpools. This article describes the isolation of filaments from natural sources, DNA extraction, genome sequencing, natural transformation, expression of sfGFP, cryoconservation, and motility methods.
Abstract
Cyanobacteria are the focus of basic research and biotechnological projects in which solar energy is utilized for biomass production. Phormidium lacuna is a newly isolated filamentous cyanobacterium. This paper describes how new filamentous cyanobacteria can be isolated from marine rockpools. It also describes how DNA can be extracted from filaments and how the genomes can be sequenced. Although transformation is established for many single-celled species, it is less frequently reported for filamentous cyanobacteria. A simplified method for the natural transformation of P. lacuna is described here. P. lacuna is the only member of the order Oscillatoriales for which natural transformation is established. This paper also shows how natural transformation is used to express superfolder green fluorescent protein (sfGFP). An endogenous cpcB promoter induced approximately 5 times stronger expression than cpc560, A2813, or psbA2 promoters from Synechocystis sp. PCC6803. Further, a method for the cryopreservation of P. lacuna and Synechocystis sp. CPP 6803 was established, and methods for assessing motility in a liquid medium and on agar and plastic surfaces are described.
Introduction
Cyanobacteria are prokaryotic organisms that utilize photosynthesis as an energy source1,2. Research is increasingly focused on cyanobacterial species. Several cyanobacteria can be transformed with DNA3. Genes can be knocked out or overexpressed in these species. However, transformation is restricted to a few species4,5,6,7,8,9,10,11, and it can be difficult to establish transformation in strains from culture collections or the wild8. Strains of the filamentous species Phormidium lacuna (Figure 1) were isolated from marine rockpools, in which environmental conditions, such as salt concentrations or temperature, fluctuate over time. These filamentous cyanobacteria can be used as model organisms for the order Oscillatoriales12 to which they belong.
During trials testing gene transfer by electroporation13,14 it was found that P. lacuna can be transformed by natural transformation15. In this process, DNA is taken up naturally by some cells. Compared to other methods of transformation16,17, natural transformation has the advantage of not requiring additional tools that could complicate the procedure. For example, electroporation requires proper cuvettes, intact wires, and selection of the proper voltage. P. lacuna is presently the only Oscillatoriales member susceptible to natural transformation. Because the original protocol is based on electroporation protocols, it still included several washing steps that might be unnecessary. Different approaches were tested to simplify the protocol, leading to the transformation protocol presented here.
The genome sequence is essential for further molecular studies based on gene knockout or overexpression. Although genome sequences can be obtained with next-generation sequencing machines within short periods, the extraction of DNA can be difficult and depends on the species. With P. lacuna, several protocols were tested. A modified cetyl trimethyl ammonium bromide (CTAB)-based method was then established, resulting in acceptable purity of DNA and DNA yields of each purification cycle for continued work in the laboratory. The genome of five strains could be sequenced with this protocol. The next logical transformation step was to establish protein expression in P. lacuna.
The sfGFP used as a marker protein in this protocol can be detected with any fluorescence microscope. All promoters that were tested could be used for P. lacuna sfGFP expression. The increasing number of strains arising from transformation has resulted in the need for a method for storing the cultures. Such methods are established for Escherichia coli and many other bacteria18. In standard protocols, glycerol cultures are prepared, transferred in liquid nitrogen, and stored at -80 °C. This method requires only a few steps and is highly reliable for those species for which it is established. The standard protocol was not feasible for P. lacuna because living cells could not be recovered in all cases. However, when glycerol was removed after thawing, cells of all trials survived. Simple methods are presented for the analysis of motility of P. lacuna, which can be combined with knockout mutagenesis to investigate type IV pili or the role of photoreceptors. These assays are different from those of single-celled cyanobacteria19,20,21 and can also be useful for other Oscillatoria.
Protocol
1. Isolation from the natural environment
NOTE: Green algae, diatoms, filamentous cyanobacteria, and other microalgae can be isolated. The protocol can be used for any microalga species from rockpools growing under laboratory conditions. Filamentous cyanobacteria that belong to Oscillatoriales can be easily recognized by their movement and filamentous shape. The species can be identified in a semipure state by genome sequencing or 16S rRNA sequencing.
- Transfer liquid seawater samples from marine rockpools (i.e., cavities in the rocky coast) into 50 mL flasks. For each flask, note the exact place or coordinates of the natural source. If possible, filter the content through 50 µm nets to reduce the amounts of zooplankton. Store the samples at 4 °C until they can be subcultured.
- Transfer 1 mL cultures to 10 cm Petri dishes containing 3% bacto-agar in f/2 medium22,23 (see the Table of Materials). Prepare up to 20 plates. Cultivate under white light of 50 µmol m-2 s-1.
NOTE: Higher light intensities may be used for cultivation. Intensity up to 400 µmol m-2 s-1 can be used for P. lacuna, although other species might be more light-sensitive. - After one week, transfer the desired cells to fresh agar plates using sterile forceps. Isolate the cells under a binocular microscope under sterile conditions. Store the old agar plate at 4 °C until cells appear and grow on the new agar plate.
- Repeat this transfer step every week to eliminate contamination. Use the naked eye for detecting heavy contamination and a microscope with 400x magnification for additional checks for contamination.
- If a sample seems free of contamination, test for bacterial or fungal contamination on agar plates. Transfer a fraction of the culture with an inoculation loop to an LB24 agar plate (10 cm diameter), keep the plate at room temperature, and check for the growth of contaminants over 1-3 days.
- If a sterile filamentous cyanobacterial species is obtained, use it for further culture work. Cultivate P. lacuna in liquid or on bacto-agar plates. Use 250 mL flasks with 50 mL of f/2 medium or f/2+ medium for liquid culture.
2. DNA extraction
NOTE: This method is adopted from 25 26
- Prepare two flasks with 50 mL of f/2 medium. Inoculate each with ~1 mL of P. lacuna filaments from other growing cultures. Keep the cultures for 7 days or longer under agitation (horizontal rotation) at 50 rpm under white light (50 µmol m-2 s-1) at 25 °C.
- Treat the culture with ultrasound (see the Table of Materials) for 2 min with full energy. Measure OD at 750 nm; check to ensure it is ~0.5. Continue to grow the cultures if the OD is too low.
- Collect the filaments by 5,000 × g, 20 min centrifugation. Remove the supernatant. Transfer the filaments with residual liquid to the chamber of a French Press27. Set the pressure of the French Press to 20,000 psi and extract the cells.
NOTE: The French Press will lyse all the cells and release the DNA; strong shear forces will produce 1,500 bp DNA fragments. - Centrifuge the sample for 10 min at 10,000 × g and remove the supernatant.
- Add 400 µL of lysis buffer (4 M urea, 0.1 M Tris/Cl, pH 7.4) and 50 µL of proteinase K (10 mg/mL) to the pellet. Heat the sample to 55 °C for 60 min with shaking at 550 rpm.
- Add 1 mL of DNA extraction buffer (3% CTAB, 1.4 M NaCl, 10 mM EDTA, 0.1 M Tris/Cl, 1% Sarkosyl, 0.1 M DTT, pH 8) and incubate for 60 min at 55 °C and 550 rpm. Transfer the solutions to centrifugation tubes, and add two volumes of chloroform/isoamylalcohol (24/1).
- After shaking, centrifuge the sample for 5 min at 9,000 × g. Transfer the upper, aqueous phase into reaction vials and add 1 mL of ice-cold ethanol and 50 µL of 3 M sodium acetate.
- Vortex the sample and place it at -20 °C for 1 h or longer.
- Centrifuge for 5 min at 10,000 × g (4 °C) and discard the supernatant. Wash the pellet with 70% ethanol.
- Centrifuge the sample again. Remove the supernatant and dry the pellet overnight. Dissolve the DNA in nuclease-free water. Measure the DNA spectrum to check whether the OD 260 nm/OD 280 nm is between 1.6 and 1.9.
- Analyze the size of the DNA on an agarose electrophoresis gel28.
- Sequence the genomic DNA by next-generation sequencing for 300 cycles, with a paired-end setting and read length of 150 bases (see the Tables of Materials).
- Perform the assembly with the appropriate computer program; see the example given in the Table of Materials.
- Submit the draft genome to the RAST server for annotation.
NOTE: Upload DNA sequences to obtain complete annotation within a few minutes.
3. Natural transformation and GFP expression
NOTE: Transformation is based on a plasmid vector propagated in E. coli; pGEM-T or pUC19 may be used as backbone vectors. Cloning techniques are established in many laboratories; see also standard protocols28 and the articles on transformation vectors for P. lacuna15,29. Examples for vectors for sfGFP expression are described in the representative results section. Details of four yet unpublished vectors are provided in Supplemental File 1.
- Perform all steps using sterile material under sterile laboratory conditions (clean bench, sterile glassware).
- Inoculate 2 x 50 mL of f/2 liquid medium in two 250 mL flasks with 2 x 1 mL of P. lacuna filaments from a running culture. Cultivate in white light (50 µmol m-2 s-1) under agitation (horizontal rotation, 50 rpm) for ~5 days at 25 °C.
- Prepare ~200 µg of the transformation vector DNA using a midi prep kit (see the Table of Materials) according to the manufacturer's instructions.
- Homogenize 100 mL of P. lacuna cell suspension (see the Table of Materials) at 10,000 rpm for 3 min. Measure OD at 750 nm (desired value = 0.35).
- Centrifuge the cell suspension for 15 min at 6,000 × g. Remove the supernatant, and suspend the pellet in 800 µL (total volume including residual liquid and filaments) of the remaining liquid and additional f/2+ medium.
- Take eight f/2+ bacto-agar plates (10 cm diameter) containing 120 µg/mL kanamycin. Pipette 10 µg of DNA into the middle of each agar plate. Immediately pipette 100 µL of cell suspension into the middle of each agar plate (on top of the DNA).
- Keep the agar plate without a lid on the clean bench to allow the excess liquid to evaporate. Close the plate and cultivate it in white light at 25 °C for 2 days.
- Distribute the filaments of each agar plate with an inoculation loop onto several fresh f/2+ bacto-agar plates containing 120 µg/mL kanamycin. Cultivate the plates in white light at 25 °C and check the cultures regularly under a microscope.
- Identify living, transformed filaments after 7-28 days under the microscope. Look for healthy, green filaments (Figure 2) that are different from other filaments. If these green filaments can be identified, continue with the next step; otherwise, keep the plate for another 7 days.
- Use forceps to transfer these identified living filaments into 50 mL of liquid f/2+ medium with 250 µg/mL kanamycin. Cultivate in white light at 25 °C on a shaker (horizontal rotation, 50 rpm). Observe growth for up to four weeks.
- Transfer the filaments back to agar medium containing 250 µg/mL kanamycin and wait for the filaments to grow. After several days, transfer single filaments to a fresh agar plate with a higher concentration of kanamycin, e.g., 500 µg/mL. Keep the original plate.
- Ensure that the filaments are propagated in a high concentration of kanamycin in liquid culture or on agar. Increase the kanamycin concentration again to speed up segregation.
NOTE: Transformed P. lacuna grows in up to 10,000 µg/mL kanamycin. Other species might not tolerate such high concentrations. - If resistant cells are grown and distributed broadly over a plate, test the integration of the insert into the genome of P. lacuna by performing PCR with outer and inner primers.
- Use primers that were designed for cloning of the insertion as inner primers.
- For design of the outer primers, select sequences that are 5' and 3' of the proposed insertion site on the genome of P. lacuna but outside the insertion.
- For PCR reactions, use inner primers and outer primers. Use the resistant strain(s) and the wild type.
NOTE: Inner primers indicate that the insert is present; outer primers show that the insert is inserted at the correct locus.
- For each PCR reaction, place ~10 mg of the filaments directly in the PCR tubes and perform PCR according to standard protocols24. If no product is obtained, vary the annealing temperature and wash the filaments with water.
NOTE: Many different polymerases can be used in PCR. Standard polymerases, such as Taq polymerase, have a higher error rate than error-checking polymerases, which are more expensive. This analytical PCR does not require any error-checking polymerase. However, error-checking polymerase should be tested if no PCR product is obtained with a standard polymerase. - Analyze the PCR products of the resistant line on agarose electrophoresis24.
- Compare band positions with marker and compare the wild type and transformant. With inner and outer primers, look for a larger band for the transformant than the wild type (due to the insertion of the resistance cassette) or two bands for the transformant: one with the size of the wild-type band and a larger one. As the latter case indicates incomplete segregation, continue cultivation with high kanamycin concentrations.
NOTE: For more details on PCR and electrophoresis, see15,24 or other standard literature.
- Compare band positions with marker and compare the wild type and transformant. With inner and outer primers, look for a larger band for the transformant than the wild type (due to the insertion of the resistance cassette) or two bands for the transformant: one with the size of the wild-type band and a larger one. As the latter case indicates incomplete segregation, continue cultivation with high kanamycin concentrations.
- For GFP expression: observe single filaments with a fluorescence microscope (see the Table of Materials) at a magnification of the objective set at 40x or 63x. Capture a brightfield transmission image and a fluorescence image. Use the following settings for GFP: 470 nm bandpass for excitation, 525 nm bandpass for emission, and a 495 nm beam splitter, initial exposure time of 500 ms.
- Adjust the exposure time for clear fluorescence signals, avoiding saturating intensities. Try to use the same setting for all samples.
- As the wild-type filaments will also display fluorescence, capture images with the same settings as above for this background fluorescence.
NOTE: The strain expressing GFP must have a higher signal; otherwise, it is not expressing GFP. - Based on exposure times and the pixel intensities of the fluorescence images, calculate and compare the GFP content of the different filaments.
4. Cryoconservation
NOTE: P. lacuna and the single-celled cyanobacterium Synechocystis sp. PCC 6803 are used. The present method works better for P. lacuna.
- Cultivate P. lacuna or Synechocystissp PCC 6803 for at least 10 days in 10 mL of f/2+ or BG-11 medium, respectively, under white light (50 µmol m-2 s-1) at 25 °C under agitation (horizontal rotations, 50 rpm).
- Homogenize the P. lacuna culture (see the Table of Materials) at 10,000 rpm for 3 min or with an ultrasound device (see the Table of Materials) for 2 min at full energy. Determine OD 750 nm of either culture to check whether the value is between 1 and 7.
- Collect the cells by centrifugation at 6,000 × g for 15 min. Remove the supernatant.
- Suspend the cell pellet in 800 µL of f/2+ or BG-11 medium (final volume) and transfer to a 2 mL cryovial. Add 800 µL of a 50% glycerol solution to the cell suspension. Close the vial and mix by repeated inverting.
- Transfer the cryovial to liquid nitrogen and store it in a cryobox in a -80 °C freezer. Note the position of the box within the freezer and the coordinates of the sample within the box.
- For recovery of the cells, take out the cryovial and thaw the contents at room temperature. Transfer the contents to a 2 mL reaction tube.
- Wash the sample twice. For the 1st wash, centrifuge at 6,000 × g for 5 min. Remove the supernatant, and resuspend the pellet in 2 mL of f/2+ or BG-11 medium. For the 2nd wash, recentrifuge at 6,000 × g for 5 min, remove the supernatant, and suspend the pellet in 2 mL of f/2+ or BG-11 medium.
- To check the integrity of these cells that are ready for cultivation, transfer the pellet to 9 mL of medium and cultivate them in white light (50 µmol m-2 s-1) under agitation (55 rpm). Compare the OD 750 nm of the culture on the first day and after 1 week.
5. Motility of Phormidium lacuna
NOTE: Three different assays will be described. The same culture is used in all cases.
- Cultivate P. lacuna in f/2 medium under horizontal agitation (50 rpm) in white light (50 µmol m-2 s-1) for ~5 days until the estimated OD 750 nm is 0.35. Store the sample at 4 °C until use.
- Homogenize the filaments (see the Table of Materials) at 10,000 rpm for 3 min or with ultrasound (see the Table of Materials) for 1 min at maximum power and cycle of 1. Measure OD 750 nm. If above 0.35, dilute the fraction with f/2 medium. Use this solution in motility assays in steps 5.3, 5.4, and 5.5.
- Assay for movement in liquid medium
- For direct observation of motility, transfer 8 mL of medium containing P. lacuna (from step 5.2) into a 6 cm Petri dish. Wait a few minutes until the sample reaches room temperature. Cover the Petri dish with cellophane foil.
- Place a microscope slide on the x-y table of a standard microscope with a camera. Switch on the microscope light. Ideally, always use the same electrical and optical settings for the lighting. Move a 4x or 10x objective into the path of the light.
- Place the Petri dish on top of the slide. Adjust single filaments or filament bundles by x, y, and z movements of the table.
NOTE: Due to the three-dimensional arrangement, only a part of the relevant section can be in focus. The cellophane foil allows adjusting the focus without restriction. - Observe movements of single filaments or bundles. Ensure that the objective lens does not touch the liquid. Record the movements of filaments with a standard microscope camera (see Supplemental Video S1).
- Assay for movement on the surface
- For the observation of filament motility on agar surfaces, prepare 6 cm Petri dishes with f/2 bacto-agar. Ensure that the agar is high enough for the objective lens to get close to the agar surface. Alternatively, prepare a ~3 mm thick agar layer and record the filaments through the agar (keep the plate upside down or use an inverted microscope).
- Pipette 0.5 mL of a solution containing P. lacuna (from step 5.2) on the bacto-agar surface of a 6 cm Petri dish. Allow the liquid to enter the surface. Close the Petri dish and observe the movement of the filaments on the surface using a 4x or 10x objective.
- Ensure that the same electrical and optical settings of the microscope are used throughout the recording and in subsequent recordings.
- Capture time-lapse recordings using an ocular camera and minicomputer system. Ensure that the time interval between subsequent images is 5 s-1 min. Program the Linux script of the minicomputer to control the time-lapse recording. See Supplemental File 2 for an example script and Supplemental Video S2 as an example.
- Assay for phototaxis
- For phototaxis experiments, prepare light-emitting diode (LED) holders (here, with a 3D printer) in which the selected 5 mm LEDs are mounted to irradiate an area of 20 mm2 from below to above (Figure 3). If required, use many LED holders in parallel, connecting each LED electrically through a resistor and potentiometer to an adjustable power supply. Measure and adjust the LED intensities, depending on the experiment. Ensure that the whole setting is in a dark room or a closed dark container.
- Place 8 mL of the medium containing P. lacuna (from step 5.2) into a 6 cm Petri dish. Adjust the light intensity of the LED. Close the Petri dish with the lid and place it on an LED holder so that the LED is in the center of the Petri dish.
- After the desired duration (typically 2 days), capture an image of the Petri dish with a smartphone camera aimed directly at the position of the light treatment. Use a white LED panel for irradiation of the specimen. Use the manual settings of the camera; avoid reflections of light; always adjust to get the same distance between the camera lens and the specimen. Ensure that the exposure settings give an image suitable for later analyses using ImageJ.
- Quantify the diameter of the central circle of filaments using ImageJ software.
- Open ImageJ, click on File | Open, select the desired file, and click Enter.
- Select the Straight button (with a straight line). Press the left mouse button to draw a line from one end of the Petri dish to the opposite end. Ensure that the line passes through the center of the circle of filaments.
- Press Ctrl-K on the keyboard or click Analyze | Plot Profile in the ImageJ menu. Look for an x-y window with pixel intensities plotted versus distance-a 1D profile of the Petri dish. Ensure that the lowest pixel intensity is slightly above 0 and the highest value below 255.
- Estimate an average value for the pixel intensity outside the circle and another average value for the pixel intensity in the circle. At the y- position between these values, estimate the x-values of both sides of the circle by pointing with the mouse on these positions. Note both values and calculate the difference.
- Obtain the highest x-value by pointing the mouse at the y-axis on the right. Note that this value e represents the diameter of the Petri dish. If this diameter is 5 cm, calculate the diameter of the central filament circle as d/e × 5 cm.
Representative Results
Following the above-mentioned methods, 5 different strains of P. lacuna were isolated from rockpools and sequenced (Figure 1 and Table 1). All cultures were sterile after ~1 year of subculturing except P. lacuna HE10JO. This strain is still contaminated with Marivirga atlantica, a marine bacterium. During subsequent Helgoland excursions, other filamentous cyanobacteria were isolated from rock pools, which are different from P. lacuna and need to be characterized.
Several DNA extraction and purification methods were tested for P. lacuna. The best results were obtained with an optimized CTAB method as described above. DNA yields were 310 ± 50 µg/mL, OD 260 nm/OD 280 nm was 1.7 ± 0.03, and OD 260 nm/OD 230 nm was 0.78 ± 0.04 (n = 17). Genome sequencing showed that the DNA of all strains was slightly different, as expected (Table 1). Core protein sequences showed a maximum difference of 0.04% (Table 2). Although all draft genomes were incomplete, one can assume that >98% of the genome of HE10JO30 was sequenced. This estimation is based on the number of incomplete open reading frames. Partial protein sequences could be easily identified after RAST annotation of HE10DO and HE10JO. In HE10JO, 60 proteins out of ~4,500 had a missing N- or C-terminal sequence. The genome sequences can be found in the supplement (Supplemental File 3, Supplemental File 4, Supplemental File 5, Supplemental File 6, and Supplemental File 7).
Interestingly, strains of the same species were isolated from two islands, Helgoland and Giglio. The linear distance between both islands is 1,400 km. There must be a link between both places, e.g., by ships via the sea or, more likely, by migratory birds. Many bird species can be found on both islands, and many of them are migratory birds. The diversity within P. lacuna strains of one island was greater than between the closest Helgoland and Giglio strains (Table 2). This indicates an intense exchange between both places.
The natural transformation was tested with HE10DO as the major strain and with HE10JO. The present protocol is more straightforward than the protocol described earlier12 because of the reduced number of washing steps and fewer transfer steps after transformation. This new method is continuously used in the laboratory; ~15 successful transformations were achieved.
The KanR resistance cassette was usually integrated into the homologous site defined by the adjacent regions, as shown by PCR using inner and outer primers. Like most cyanobacteria, P. lacuna is polyploid. It can have more than 100 chromosomes per cell12. A PCR test with outer primers ~1 week after the transformation typically has 2 bands on the electrophoresis gel, one with the size of the wild-type band and one slower migrating band that indicates the insertion of the resistance cassette (Figure 4). The double band indicates that only a subfraction of the chromosomes contains the insertion. After 4 weeks of selection on kanamycin, segregation is usually complete, and only one large PCR band appears on gels. However, in the case of the transformation with pMH1 (see below), segregation was complete after more than 3 months.
The vectors pAK1, pAK2, pAK3, and pMH1 were constructed for tests on sfGFP expression. In pAK1, pAK2, and pAK3, the sfGFP gene is under the control of the cpc560, A2813, and psbA2 promoters, respectively. These promoters are from Synechocystis sp. PCC 6803 or Synechococcus sp. PCC 700231. For the construction of these vectors, the sfGFP promoter and terminator sequences were taken from vectors used for the transformation of Synechococcus sp. PCC 700231. The relevant sequences were integrated into the homologous chwA (sc_7_37) site of pFN1 (or pFN_7_37_KanR15). The pMH1 expression vector was constructed by DNA synthesis using P. lacuna sequences as templates (Supplemental File 6). The cpcB–cpcA (phycocyanin ß and phycocyanin α) sequences of P. lacuna are serially arranged. A 100 bp intergenic region separates both coding regions. The synthetic sequence contained this endogenous cpcB–cpcA sequence and the cpcB promoter. The sfGFP and KanR cassette is placed just 3' of the cpcB stop codon (5' of cpcA). The entire synthetic sequence with cpcB promoter, cpcB, sfGFP, KanR, cpcA (5' to 3') is cloned into pUC19. A map is shown in Figure 5. More details on the cloning of pAK1, pAK2, and pAK3 and the complete sequence of pMH1 are given in Supplemental File 1.
All 4 transformants (with pAK1, pAK2, pAK3, and pMH1) expressed GFP; all fluorescence levels were above the background fluorescence of wild-type filaments (Figure 6). The pMH1 transformants with incomplete segregation revealed a GFP signal that was very variable between the filaments. The fluorescence signal was evenly distributed when segregation was complete (Figure 6E). The microscope signals of pAK1, pAK2, and pAK3 transformants were similar but ~5x weaker than that of pMH1 (Figure 6E).
The established cryoconservation method is based on a method that was established for E. coli. When 2 washing steps were performed for glycerol removal after thawing, 15 out of 15 P. lacuna samples survived (Table 3). This protocol could also be used for Synechocystis PCC 6803, but only with 2 washing steps and not with 1 (Table 3).
Another feature of Oscillatoriales filaments is their motility: P. lacuna filaments move continuously on surfaces (Figure 7) and in a liquid medium (Figure 8). Both kinds of motion can be studied easily in Petri dishes without or with agar medium. Time-lapse recording is required because movement on agar is slow. Filaments move towards the light cone if a light beam comes from below (Figure 3). The effects of light intensity, wavelength, and time can be easily studied with a simple setup. The photoreceptors of this effect are not yet clear. Possible candidates can be addressed with knockout mutants. The mechanism underlying how the filaments find the light is also unclear. For this question, an infrared system is required to record the filaments during their movement from darkness to light.
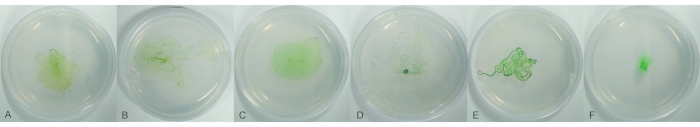
Figure 1: Strains of Phormidium lacuna collected from Helgoland and Giglio. Filaments are propagated for 11 days on f/2 agar in 6 cm Petri dishes. (A) strain GI08AO; (B) strain GI08IO; (C) strain GI09CO; (D) strain HE10DO; (E) strain HE10JO; (F) strain HE15M2G1. Please click here to view a larger version of this figure.
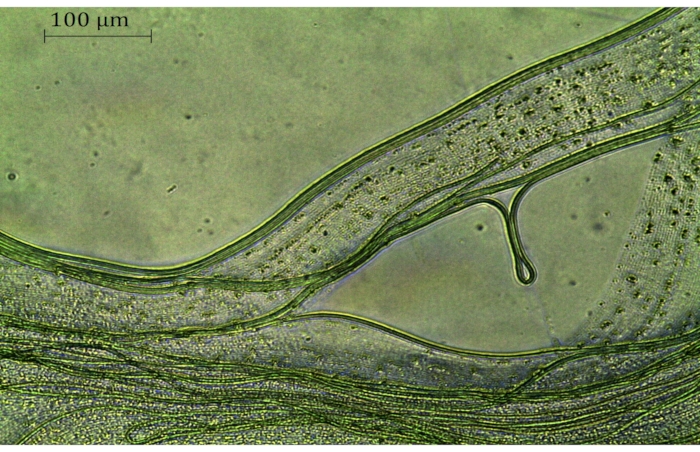
Figure 2: Phormidium lacuna filaments 5 weeks after transformation. The sfGFP expression vector pMH1 was used; selection occurred on f/2+ medium with 120 µg/mL kanamycin. The greenish filaments are resistant and alive; other filaments have died. Scale bar = 100 µm. Please click here to view a larger version of this figure.
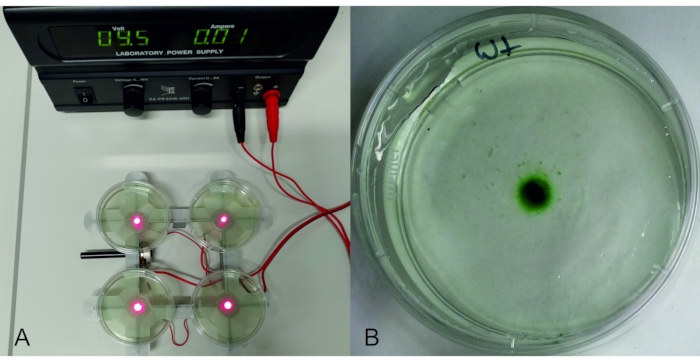
Figure 3: Phototaxis experiment. Left: LED holder with 4 red LEDs, connected to an adjustable power supply. On top of each LED, there is a 6 cm Petri dish with 8 mL of a Phormidium lacuna culture. Right: Petri dish with P. lacuna after 2 days on the red LED (15 µmol m-2 s-1). Abbreviation: LED = light-emitting diode. Please click here to view a larger version of this figure.
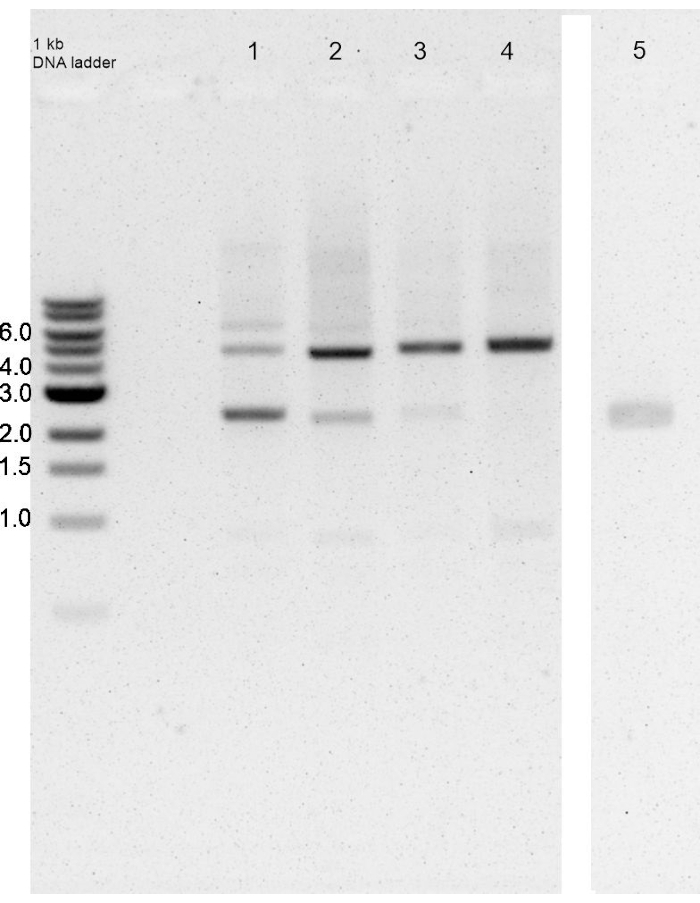
Figure 4: Integration and segregation of insert after transformation of Phormidium lacuna with pAK1. PCR with outer primers. The expected sizes of the product without and with insert are 2371 and 5016 bp, respectively. Left lane: marker, lanes 1, 2, 3, 4: PCR products of filaments 7 days, 11 days, 14 days, and 17 days after the isolation of a resistant filament (4 weeks after transformation), respectively. Lane 5: PCR product of wild-type (from a different gel). In the 7 day sample, the insert is present in a small fraction of the chromosomes. This fraction increases until 17 days, where no wild-type band is visible, i.e., segregation is complete. Please click here to view a larger version of this figure.
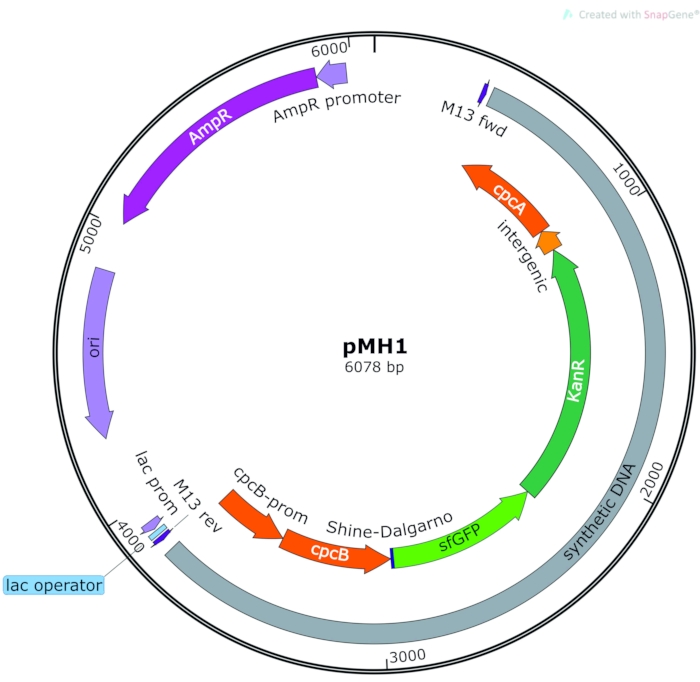
Figure 5: Vector for sfGFP expression under the control of endogenous cpcB promoter. Orange: Phormidium lacuna homologous sequence, violet/blue: pUC-19 vector backbone, green: insert with sfGFP and KanR. Abbreviations: sfGFP = superfolder green fluorescent protein; KanR = kanamycin resistance. Please click here to view a larger version of this figure.
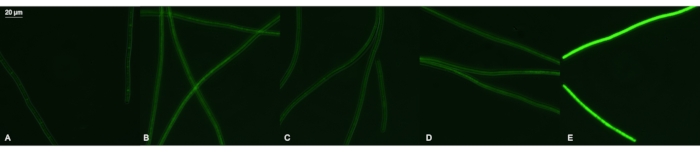
Figure 6: Expression of sfGFP in Phormidum lacuna. Fluorescence images of P. lacuna wild-type filaments (A) and after transformation with pAK1 (B), pAK2 (C), pAK3 (D), and pMH1 (E). In pMH1, the sfGFP gene is placed 3' of the phycocyanin ß gene and therefore driven by the endogenous cpcß promoter; in the other cases, sfGFP is driven by cpc560, A2813, or psbA2s promoters from Synechocystis PCC 6803, respectively. The fluorescence settings are specific for GFP; all images were recorded with the same integration time and optical settings. Please click here to view a larger version of this figure.
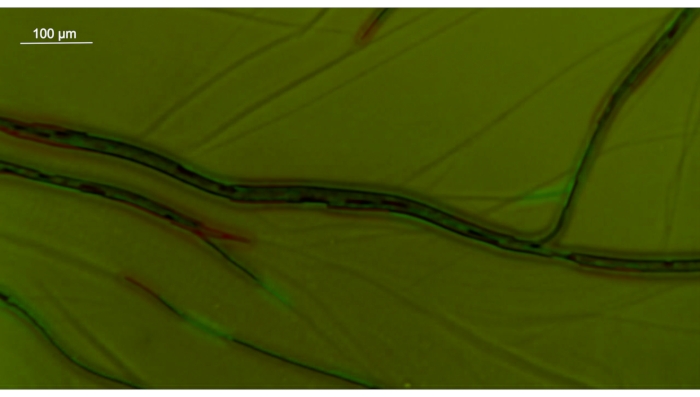
Figure 7: Merged image of Phormidium lacuna on agar surface at 4x magnification. The first image is presented in red, the second (taken 1 min later) in green. Note also the traces on the agar. Scale bar = 100 µm. Please click here to view a larger version of this figure.
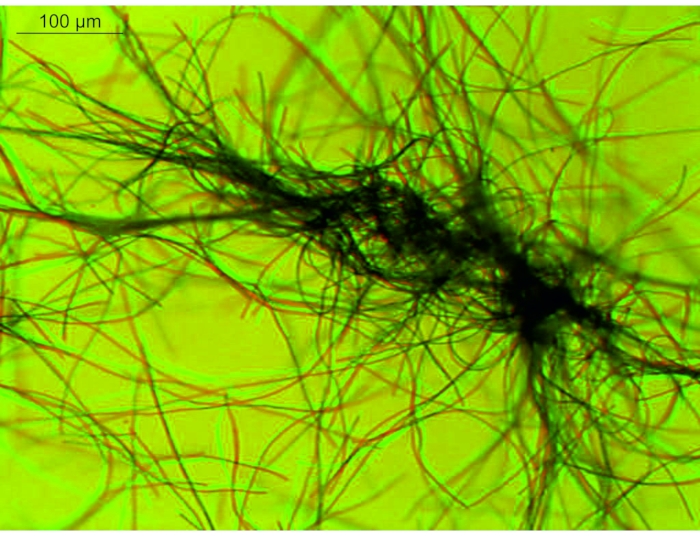

Figure 8: Merged image of Phormidium lacuna in liquid medium. The time interval between both images was 10 s. The first image is printed in red; the second is printed in green. Comparing both colors shows the movement within 10 s. Scale bar = 100 µm. Please click here to view a larger version of this figure.
| strain | HE10JO | HE10DO | GI08AO | GI09CO | HE15M2G1 |
| Contigs | 104 | 174 | 218 | 102 | 154 |
| total bp | 48,19,017 | 47,88,491 | 47,78,775 | 36,69,922 | 45,98,395 |
Table 1: Phormidium lacuna strains.
| Gi09CO | HE10DO | HE10JO | HE15M2G1 | |
| Gi08AO | 42 | 40 | 0 | 42 |
| Gi09CO | 2 | 42 | 0 | |
| HE10DO | 40 | 2 | ||
| HE10JO | 42 |
Table 2. Amino acid differences between strains in sequences of 20 core proteins with 10,876 amino acids.
| Cell density OD 750 nm | 1 | 1 | 3 | 3 | 5 | 5 | 7 | 7 |
| Washes | 1 | 2 | 1 | 2 | 1 | 2 | 1 | 2 |
| Synechocystis PCC 6803 | 2/5 | 5/5 | 1/4 | 4/4 | 0/4 | 4/4 | 0/4 | 3/4 |
| Phormidium lacuna HE10DO | 4/4 | 4/4 | 4/4 | 4/4 | 3/ 4 | 4/4 | 3/3 | 3/3 |
Table 3: Cryoconservation trials with Synechocystis PCC 6803 and Phormidium lacuna HE10DO cyanobacteria. The first number shows the number of cultures that survived after freezing/thawing; the second number shows the total trials.
Supplemental Video S1: Movement of Phormidium lacuna filaments in liquid solution, without time-lapse. Please click here to download this Video.
Supplemental Video S2: Movement of Phormidium lacuna filaments on agar surface, with time-lapse. Please click here to download this Video.
Supplemental File 1: Cloning of vectors for transformation of Phormidium lacuna. List of transformation vectors; list of primers for cloning; sequence of pMH1 in gb format. Please click here to download this File.
Supplemental File 2: Shell scripts (sh) for Raspberry Pi minicomputer. Please click here to download this File.
Supplemental File 3: DNA sequence of HE152G1. Please click here to download this File.
Supplemental File 4: DNA sequence of GI08AO. Please click here to download this File.
Supplemental File 5: DNA sequence of GI09CO. Please click here to download this File.
Supplemental File 6: DNA sequence of HE10DO. Please click here to download this File.
Supplemental File 7: DNA sequence of HE10JO. Please click here to download this File.
Discussion
Although many strains of cyanobacteria are available from culture collections32,33,34,35,36, there is still a demand for new cyanobacteria from the wild because these species are adapted to specific properties. P. lacuna was collected from rockpools and is adapted to variations of salt concentrations and temperature30. Strains of this species were found during excursions in 2008, 2009, and 2010. With the procedure described here, 5 strains of P. lacuna were isolated, and 4 of these strains were sterile. The strain P. lacuna HE10JO is permanently contaminated with the bacterium Marivirga atlantica, a marine bacterium identified by rRNA and genome sequencing. This bacterium could not be separated from the cyanobacterium in spite of the application of mechanical separation, growth at different temperatures, treatments with antibiotics, or chemical treatments. Despite the contamination, P. lacuna HE10JO can be cultivated similar to the other strains. In later excursions, other members of Oscillatoriales were found, which are yet not analyzed in detail. P. lacuna was not found again. It is not clear why P. lacuna was isolated in subsequent years and two different places but not found later. Its abundance is certainly dependent on nonpredictable conditions. Temperature, salt concentrations, and inorganic or organic nutrients are highly variable in rockpools. Therefore, the species composition could fluctuate over time in an unpredictable manner.
Natural transformation is established for different cyanobacteria, mostly single-celled species. The filamentous P. lacuna is the only species of the order Oscillatoriales for which natural transformation has been established. The transformation was almost always successful with the present protocol. In general, the numbers of resistant filaments after a transformation trial vary considerably, and sometimes transformation fails, resulting in the loss of valuable time. It is therefore advisable to perform several transformation projects in parallel. The time for complete segregation, usually 4 weeks after isolation of the resistant strain, can also vary. Because there is no guarantee for complete segregation after growth on kanamycin, it is crucial to perform the PCR tests using outer and inner primers.
Every vector must contain 2 x 500-1,000 bp homologous sequences, e.g., amplified from the host by PCR and interrupted by a resistance cassette (e.g., the kanamycin cassette KanR used here)37,38. For expression, a promoter, coding sequence (e.g., for sfGFP39), terminator, and resistance cassette must be cloned between the homologous sequences. The cloning strategies are species-specific and depend on the aim of the experiment.
This transformation method could be possible with other Oscillatoriales strains or other cyanobacteria as well: natural transformation is based on type IV pili, which are present in almost every other cyanobacterial genome3,15,40. Therefore, the present method could stimulate new trials with other species. Because type IV pili are also relevant for motility, it is important to check for conditions under which cyanobacteria are motile.
Gene insertion is based on homologous recombination and results in a disruption of the homologous sites. Therefore, transformation is often used for gene knockout. The expression of the inserted gene will be induced if an active promoter and a coding sequence are integrated into the homologous site. In P. lacuna, promoter activity was dependent on the species. The cpc560, A2813, and psbA2 promoters of Synechocystis PCC sp 6803 or Synechococcus sp. PCC 7002 31 and the cpcB promoter of P. lacuna could drive sfGFP expression. Of these constructs, the endogenous cpcB promoter induced the strongest expression, although the sfGFP gene is located 3' of the phycocyanin ß gene. This indicates a more general use of endogenous promoters in cyanobacterial expression.
The combination of gene knockout and motion studies will shed light on molecular mechanisms of motility and phototaxis. LED light sources can provide light for phototaxis experiments. Almost any wavelength is available, and light intensity can be modulated by an adjustable power supply and potentiometers. LED holders can be built by 3D printers to easily realize combinations of different LEDs.
Divulgations
The authors have nothing to disclose.
Acknowledgements
The work was supported by the Karlsruher Institute of Technology.
Materials
| Autoclave 3870 ELV | Tuttnauer | 3870 ELV | |
| Bacto Agar | OttoNorwald | 214010 | |
| BG-11 Freshwater Solution | Sigma Aldrich | C3061 | |
| BG-11 medium | Merck | 73816-250ML | |
| Boric acid | Merck | 10043-35-3 | H3BO3 |
| Calcium chloride dihydrate | Carl Roth | 10035-04-8 | CaCl2 · 2 H2O |
| Cell culture flasks Cellstar with filter screw cap, sterile, 250 mL | Greiner | 658190 | |
| Cell culture flasks Cellstar with filter screw cap, sterile, 50 mL | Greiner | 601975 | |
| Centrifuge LYNX 4000 | Thermo Scientific | 75006580 | and rotor |
| Centrifuge microstar 17 | VWR International | N/A | for up to 13,000 rpm |
| Cetyltrimethylammonium Bromide (CTAB) | PanReac AppliChem | 57-09-0 | C19H42BrN |
| Chloroform : Isoamyl Alcohol 24 : 1 | PanReac AppliChem | A1935 |
|
| Cobalt(II) chloride hexahydrate | Merck | 7791-13-1 | CoCl2 · 6 H2O |
| Copper(II) sulphate pentahydrate | Merck | 7758-99-8 | CuSO4 · 5 H2O |
| D(+)-Biotin | Carl Roth | 58-85-5 | C10H16N2O3S |
| DNA ladder 1 kb | New England Biolabs | N3232 | |
| DNA ladder 100 bp | New England Biolabs | N3231 | |
| Electrical pipetting help accujet-pro S | Brand GmbH | 26360 | for pipetting 1-25 mL |
| Ethanol | VWR | 64-17-5 | C2H6O |
| Ethylenediamine tetraacetic acid disodium salt dihydrate | Carl Roth | 6381-92-6 | EDTA-Na2 · 2 H2O |
| Fluorescence microscope ApoTome | Zeiss | ||
| Fluorescence microscope Axio Imager 2 | Zeiss | ||
| French Pressure Cell Press | American Instrument Company | N/A | |
| Gel documation System Saffe Image | Invitrogen | ||
| Gelelctrophoresis system Mupid-One/-exu | ADVANCED | ||
| Glassware, different | |||
| Glycerol | Carl Roth | 56-81-5 | C3H8O3 |
| Iron(III) chloride hexahydrate | Merck | 10025-77-1 | FeCl3 · 6 H2O |
| Kanamycin | Sigma-Aldrich | 25389-94-0 | |
| Kanamycin sulphate | Carl Roth | 25389-94-0 | C18H36N4O11 · H2SO4 |
| Lauroylsarcosine, Sodium Salt (Sarcosyl) | Sigma Aldrich | 137-16-6 | C15H28NO3 · Na |
| LB Broth (Lennox) | Carl Roth | X964.4 | |
| Light source, fluorescent tube L18W/954 daylight | OSRAM | cultivation of cyanobacteria | |
| Light source, LED panel XL 6500K 140 W | Bloom Star | N/A | cultivation of cyanobacteria, up to 1,000 µmol m-2 s-1 |
| Magnesium chloride hexahydrate | Carl Roth | 7791-18-6 | MgCl2 · 6 H2O |
| Manganese(II) chloride tetrahydrate | Serva | 13446-34-9 | MnCl2 · 4 H2O |
| Microscope DM750 | Zeiss | ||
| Midi prep plasmid extraction kit NucleoBond Xtra Midi kit | Macherey-NAGEL GmbH & Co. KG | REF740410.50 | |
| Minicomputer Raspberry Pi 4 + | Conrad Electronics | 2138863-YD | for time-lapse recording |
| Ocular camera EC3 | Leica | for continuous recording up to 30 s | |
| Ocular camera MikrOkular Full HD | Bresser | for time-lapse recordings, coupled to Raspberry Pi minicomputer | |
| Petri dishes polystyrole, 100 mm x 20 mm | Merck | P5606-400EA | |
| Petri dishes polystyrole, 60 mm x 15 mm | Merck | P5481-500EA | |
| Photometer Nanodrop ND-1000 | Peqlab Biotechnologie | ||
| Photometer Uvikon XS | Goebel Instrumentelle Analytik GmbH | ||
| Pipetman 100-1,000 µL | Gilson | SKU: FA10006M | |
| Pipetman 10-100 µL | Gilson | SKU: FA10004M | |
| Plastic pipettes 10 mL, sterile | Greiner | 607107 | |
| Plastic tube, sterile, 15 mL | Greiner | 188271 | |
| Plastic tube, sterile, 50 mL | Greiner | 227261 | |
| Potassium bromide | Carl Roth | 7758-02-3 | KBr |
| Potassium chloride | Carl Roth | 7447-40-7 | KCl |
| Power supply Statron 3252-1 | Statron Gerätetechnik GmbH | ||
| Power supply Voltcraft PPS 16005 | Conrad Electronics | for LED | |
| Proteinase K | Promega | MC500C | from Maxwell 16 miRNA Tissue Kit AS1470 |
| Q5 polymerase | New England Biolabs | M0491S | |
| Sequencing kit NextSeq 500/550 v2.5 | Illumina | ||
| Sequencing system NextSeq 550 SY-415-1002 | Illumina | ||
| Shaker Unimax 2010 | Heidolph Instruments | for cultivation | |
| Sodium acetate | Carl Roth | 127-09-3 | NaCH3COO |
| Sodium chloride | Carl Roth | 7647-14-5 | NaCl |
| Sodium dihydrogen phosphate monohydrate | Carl Roth | 10049-21-5 | NaH2PO4 · H2O |
| Sodium fluoride | Carl Roth | 7681-49-4 | NaF |
| Sodium hydrogen carbonate | Carl Roth | 144-55-8 | NaHCO3 |
| Sodium molybdate dihydrate | Serva | 10102-40-6 | Na2MoO4 · 2 H2O |
| Sodium nitrate | Merck | 7631-99-4 | NaNO3 |
| Sodium sulphate | Carl Roth | 7757-82-6 | Na2SO4 |
| Strontium chloride hexahydrate | Carl Roth | 10025-70-4 | SrCl2 · 6 H2O |
| Thiamine hydrochloride | Merck | 67-03-8 | C12H17ClN4OS · HCl |
| TRIS | Carl Roth | 77-86-1 | C4H11NO3 |
| Ultrasonic device UP100H with sonotrode MS3 | Hielscher Ultrasound Technology | UP100H | |
| Ultraturrax Silent Crusher M | Heidolph Instruments | homogenizer | |
| Urea | Carl Roth | 57-13-6 | CH4N2O |
| Vitamin B12 | Sigma | 68-19-9 | C63H88CoN14O14P |
| Vitamin solution | 0.3 µM thiamin-HCl, 2.1 nM biotin, 0.37 nM cyanocobalamin | ||
| Water Stills, Water treatment | VEOLIA water technologies | ELGA_21001 | |
| Zinc sulphate heptahydrate | Sigma | 7446-20-0 | ZnSO4 · 7 H2O |
| software, URL | |||
| gatb-minia program for DNA assembly | https://github.com/GATB/gatb-minia-pipeline | makes large scaffolds from short DNA reads, Linux based | |
| ImageJ | software for immage processing (pixel intensities, circle diameter) | ||
| RAST annotation server | https://rast.nmpdr.org | input: genome DNA sequence, detects open reading frames, lists protein sequences and their functions | |
| Culture media | |||
| Artificial seawater | 0.41 M NaCl , 53 mM MgCl2,28 mM Na2SO4, 10 mM CaCl2 , 9 mM KCl , 2.4 mM NaHCO3 ,0.84 mM KBr, 0.49 mM H3BO3, 90 µM SrCl2, 72 µM NaF | ||
| f/2 -liquid medium | artificial seawater, 0.1 % (v/v) trace element solution, 0.05 % (v/v) vitamin solution, 0.88 mM NaNO3, 36 µM NaH2PO4 | ||
| f/2+ liquid medium | f/2-medium, with 10 times increased NaNO3 and NaH2PO4 (0.88 mM NaNO3, 36 µM NaH2PO4 | ||
| f/2+-agar | 3 % (w/v) bacto agar, artificial seawater, 0.1 % (v/v) trace element solution, 0.05 % (v/v) vitamin solution ,8.8 mM NaNO3, 0.36 mM NaH2PO4 | ||
| f/2-agar | 3 % (w/v) bacto agar, artificial seawater, 0.1 % (v/v) trace element solution, 0.05 % (v/v) vitamin solution ,0.88 mM NaNO3, 36 µM NaH2PO4 | ||
| Trace element solution | 0.36 mM NaH2PO4, 12 µM Na2EDTA, 39 nM CuSO4, 26 nM Na2MoO4 , 77 nM ZnSO4, 42 nM CoCl2, 0.91 µM MnCl2 | ||
| Vitamin solution | 0.3 µM thiamin-HCl, 2.1 nM biotin, 0.37 nM cyanocobalamin | ||
References
- Brasil, B., de Siqueira, F. G., Salum, T. F. C., Zanette, C. M., Spier, M. R. Microalgae and cyanobacteria as enzyme biofactories. Algal Research-Biomass Biofuels and Bioproducts. 25, 76-89 (2017).
- Gundolf, R., Oberleitner, S., Richter, J. Evaluation of New Genetic Toolkits and Their Role for Ethanol Production in Cyanobacteria. Energies. 12 (18), 3515 (2019).
- Wendt, K. E., Pakrasi, H. B. Genomics approaches to deciphering natural transformation in cyanobacteria. Frontiers in Microbiology. 10, 1259 (2019).
- Tandeau de Marsac, N., et al. A new approach for molecular cloning in cyanobacteria: cloning of an Anacystis nidulans met gene using a Tn901-induced mutant. Gene. 20 (1), 111-119 (1982).
- Porter, R. D. Transformation in cyanobacteria. Critical Reviews in Microbiology. 13 (2), 111-132 (1986).
- Joset, F. Transformation in Synechocystis PCC 6714 and 6803: preparation of chromosomal DNA. Methods in Enzymology. 167, 712-714 (1988).
- Tsinoremas, N. F., Kutach, A. K., Strayer, C. A., Golden, S. S. Efficient gene transfer in Synechococcus sp strains PCC-7942 and PCC-6301 by interspecies conjugation and chromosomal recombination. Journal of Bacteriology. 176 (21), 6764-6768 (1994).
- Matsunaga, T., Takeyama, H. Genetic engineering in marine cyanobacteria. Journal of Applied Phycology. 7 (1), 77-84 (1995).
- Frigaard, N. U., Sakuragi, Y., Bryant, D. A. Gene inactivation in the cyanobacterium Synechococcus sp. PCC 7002 and the green sulfur bacterium Chlorobium tepidum using in vitro-made DNA constructs and natural transformation. Methods in Molecular Biology. 274, 325-340 (2004).
- Iwai, M., Katoh, H., Katayama, M., Ikeuchi, M. Improved genetic transformation of the thermophilic cyanobacterium, Thermosynechococcus elongatus BP-1. Plant and Cell Physiology. 45 (2), 171-175 (2004).
- Vioque, A., Leon, R., Galvan, A., Fernandez, E. Transformation of cyanobacteria. Transgenic Microalgae as Green Cell. , 12-22 (2007).
- Nies, F., Mielke, M., Pochert, J., Lamparter, T. Natural transformation of the filamentous cyanobacterium Phormidium lacuna. Plos One. 15 (6), 0234440 (2020).
- Ravindran, C. R. M., Suguna, S., Shanmugasundaram, S. Electroporation as a tool to transfer the plasmid pRL489 in Oscillatoria MKU 277. Journal of Microbiological Methods. 66 (1), 174-176 (2006).
- El Semary, N. A. Optimized electroporation-induced transformation in Microcystis aeruginosa PCC7806. Biotechnologie Agronomie Societe et Environnement. 14 (1), 149-152 (2010).
- Nies, F., Mielke, M., Pochert, J., Lamparter, T. Natural transformation of the filamentous cyanobacterium Phormidium lacuna. PLoS One. 15 (6), 0234440 (2020).
- Sode, K., Tatara, M., Takeyama, H., Burgess, J. G., Matsunaga, T. Conjugative gene-transfer in marine cyanobacteria – Synechococcus Sp, Synechocystis Sp and Pseudanabaena Sp. Applied Microbiology and Biotechnology. 37 (3), 369-373 (1992).
- Stucken, K., Ilhan, J., Roettger, M., Dagan, T., Martin, W. F. Transformation and conjugal transfer of foreign gGenes into the filamentous multicellular cyanobacteria (Subsection V) Fischerella and Chlorogloeopsis. Current Microbiology. 65 (5), 552-560 (2012).
- Bellali, S., Khalil, J. B., Fontanini, A., Raoult, D., Lagier, J. C. A new protectant medium preserving bacterial viability after freeze drying. Microbiological Research. 236, 126454 (2020).
- Wallner, T., Pedroza, L., Voigt, K., Kaever, V., Wilde, A. The cyanobacterial phytochrome 2 regulates the expression of motility-related genes through the second messenger cyclic di-GMP. Photochemical & Photobiological Sciences. 19 (5), 631-643 (2020).
- Wilde, A., Fiedler, B., Borner, T. The cyanobacterial phytochrome Cph2 inhibits phototaxis towards blue light. Molecular Microbiology. 44 (4), 981-988 (2002).
- Bhaya, D., Takahashi, A., Grossman, A. R. Light regulation of type IV pilus-dependent motility by chemosensor-like elements in Synechocystis PCC6803. Proceedings of the National Academy of Sciences of the United States of America. 98 (13), 7540-7545 (2001).
- Guillard, R. R., Ryther, J. H. Studies of marine planktonic diatoms. I. Cyclotella nana Hustedt, and Detonula confervacea (cleve) Gran. Canadian Journal of Microbiology. 8, 229-239 (1962).
- Kester, D. R., Duedall, I. W., Connors, D. N., Pytkowicz, R. M. Preparation of artificial seawater. Limnology and Oceanography. 12 (1), 176-179 (1967).
- Sambrook, J., Russell, D. W. . Molecular Cloning. A Laboratory Manual. 3rd edition. , (2001).
- Cold Spring Harbor Protocols. CTAB DNA extraction buffer. Cold Spring Harbor Protocols. 2009 (3), (2009).
- Singh, S. P., Rastogi, R. P., Häder, D. -. P., Sinha, R. P. An improved method for genomic DNA extraction from cyanobacteria. World Journal of Microbiology & Biotechnology. 27, 1225-1230 (2011).
- French, C. S., Milner, H. W. Disintegration of bacteria and small particles by high-pressure extrusion. Methods in Enzymology. 1, 64-67 (1955).
- Sambrook, J., Fritsch, E. F., Maniatis, T. . Molecular cloning, a laboratory manual. 2nd ed. , (1989).
- Lamparter, T., et al. The involvement of type IV pili and phytochrome in gliding motility, lateral motility and phototaxis of the cyanobacterium Phormidium lacuna. bioRxiv. , (2021).
- Nies, F., et al. Characterization of Phormidium lacuna strains from the North Sea and the Mediterranean Sea for biotechnological applications. Process Biochemistry. 59, 194-206 (2017).
- Kachel, B., Mack, M. Engineering of Synechococcus sp. strain PCC 7002 for the photoautotrophic production of light-sensitive riboflavin (vitamin B2). Metabolic Engineering. 62, 275-286 (2020).
- Lukavsky, J., Cepak, V., Komarek, J., Kasparkova, M., Takacova, M. Catalogue of algal and cyanobacterial strains of culture collection of autotrophic organisms at Trebon. Algological Studies. 63, 59-112 (1992).
- Philbrick, J. B., Diner, B. A., Zilinskas, B. A. Construction and characterization of cyanobacterial mutants lacking the manganese-stabilizing polypeptide of photosystem-ii. Journal of Biological Chemistry. 266 (20), 13370-13376 (1991).
- Pinevich, A. V., Veprritskii, A. A., Gromov, B. V., Krautvald, K., Titova, N. N. Cellular and cultural properties and characterization of the pigment in Nostoc sp, a cyanobacterium unusually rich in c-phycoerythrin. Microbiology. 63 (5), 481-485 (1994).
- Schlosser, U. G., Friedl, T. Additions to the culture collection of algae at Gottingen since 1997. Nova Hedwigia. 71 (1-2), 243-262 (2000).
- Takano, H., Arai, T., Hirano, M., Matsunaga, T. Effects of intensity and quality of light on phycocyanin production by a marine cyanobacterium Synechococcus sp. 042902. Applied Microbiology and Biotechnology. 43 (6), 1014-1018 (1995).
- Carrer, H., Hockenberry, T. N., Svab, Z., Maliga, P. Kanamycin resistance as a selectable marker for plastid transformation in tobacco. Molecular and General Genetics: MGG. 241 (1-2), 49-56 (1993).
- Kaulen, H., Schell, J., Kreuzaler, F. Light-induced expression of the chimeric chalcone synthase-NptII gene in tobacco cells. EMBO Journal. 5 (1), 1-8 (1986).
- Cotlet, M., Goodwin, P. M., Waldo, G. S., Werner, J. H. A comparison of the fluorescence dynamics of single molecules of a green fluorescent protein: One- versus two-photon excitation. Chemphyschem. 7 (1), 250-260 (2006).
- Taton, A., et al. The circadian clock and darkness control natural competence in cyanobacteria. Nature Communications. 11 (1), 1688 (2020).

