Amplification of Escherichia coli in a Continuous-Flow-PCR Microfluidic Chip and Its Detection with a Capillary Electrophoresis System
Summary
This protocol describes how to build a continuous-flow-polymerase chain system based on a microfluidic chip and how to build a capillary electrophoresis system in the lab. It presents a simple method for the analysis of nucleic acids in the lab.
Abstract
Polymerase chain reaction (PCR) is a traditional method employed for the amplification of a target gene that has played an important role in biomolecular diagnostics. However, traditional PCR is very time-consuming because of the low-temperature variation efficiency. This work proposes a continuous-flow-PCR (CF-PCR) system based on a microfluidic chip. The amplification time can be greatly reduced by running the PCR solution into a microchannel placed on heaters set at different temperatures. Moreover, as capillary electrophoresis (CE) is an ideal way to differentiate positive and false-positive PCR products, a CE system was built to achieve efficient separation of the DNA fragments. This paper describes the process of amplification of Escherichia coli (E. coli) by the CF-PCR system built in-house and the detection of the PCR products by CE. The results demonstrate that the target gene of E. coli was successfully amplified within 10 min, indicating that these two systems can be used for the rapid amplification and detection of nucleic acids.
Introduction
Polymerase chain reaction (PCR) is a molecular biology technique used to amplify specific DNA fragments, thereby amplifying trace amounts of DNA hundreds of millions of times. It has been widely used in clinical diagnosis, medical research, food safety, forensic identification, and other fields. The PCR process mainly consists of three steps: denaturation at 90-95 °C, annealing at 50-60 °C, and extension at 72-77 °C. Thermal cycling is an important part of the PCR process; however, the traditional PCR thermal cycler is not only bulky but also inefficient, requiring approximately 40 min to complete 25 cycles. To overcome these limitations, a continuous-flow PCR (CF-PCR) system was built in-house, based on a microfluidic chip. CF-PCR can greatly save time by driving the PCR solution into microchannels placed on heaters at different temperatures1,2,3,4,5.
As capillary electrophoresis (CE) has many advantages, such as high resolution, high speed, and excellent reproducibility6,7,8,9,10,11, it has become a popular tool in the lab for the analysis of nucleic acids and proteins. However, most labs, especially labs in the developing world, cannot afford this technology because of the high price of the CE instrument. Herein, we have outlined protocols for how to fabricate the CF-PCR microfluidic chip and how to build a versatile CE system in the lab. We also demonstrate the process of amplification of E. coli by this CF-PCR system and the detection of the PCR products by the CE system. By following the procedures described in this protocol, users should be able to fabricate microfluidic chips, prepare PCR solutions, build a CF-PCR system for nucleic acid amplification, and set up a simple CE system, even with limited resources, to separate DNA fragments.
Protocol
NOTE: See the Table of Materials for details related to all materials, reagents, and equipment used in this protocol.
1. Fabrication of CF-PCR microfluidic chip
- Heat the silicon wafer at 200 °C for 25 min to remove the moisture.
- Dispense 1 mL of SU-8-2075 photoresist per inch of the wafer. Spin it on the silicon wafer using a spin coater at 500 rpm for 5-10 s with an acceleration of 100 rpm/s, and then at 2,000 rpm for 30 s with an acceleration of 500 rpm/s.
- Soft bake the silicon wafer at 65 °C for 3 min, then at 95 °C for 15 min.
- Set 150 – 215 mJ/cm² as the exposure energy for the photolithography machine, and engrave the designed pattern onto the photoresist with a photolithography mask. Place the silicon wafer and the mask ready for exposure.
- Post exposure, bake the silicon wafer at 65 °C for 2 min, and then at 95 °C for 7 min.
- Immerse the silicon wafer in developer solution to remove excess photoresist and take it out when the microchannels can be seen (Figure 1). Use isopropanol to rinse off the residual developer solution.
- Mix the polydimethylsiloxane (PDMS) prepolymer and curing agent at a ratio of 10:1. Pour the mixed PDMS solution into the replica mold and solidify it at 80 °C for 60 min.
- Bond the PDMS microfluidic chip on a slide after activation using a plasma cleaner, and solidify them at 80 °C for 30 min as soon as possible (Figure 2).
NOTE: The outer dimensions of the chip are 80.0 mm × 30.0 mm × 0.4 mm. There are 40 serpentine channels (100 µm × 1.46 mm × 100 µm, width × length × depth) in the microfluidic chip.
2. Preparation of the PCR solution
- Thaw the reagents completely before configuration. Use a vortex mixer and centrifuge to ensure the reagents are well mixed.
- Prepare a centrifuge tube, adding the following components in this order: 33.75 µL of water, 1.0 µL of DNA template, 0.5 µL of primer, 5.0 µL of buffer, 4.0 µL of dNTP mixture, 1.5 µL of Tween 20, 3.5 µL of PVP, and finally 0.25 µL of DNA polymerase.
NOTE: Here, the DNA template used is E. coli. The primers for E. coli and the components of the PCR solution are listed in Table 1 and Table 2, respectively. - Mix the solution by vortexing.
3. Construction of the CF-PCR system
- Prepare two positive temperature coefficient (PTC) ceramic heaters, two solid-state relays, two PID temperature controllers, two temperature sensors, and a power cord.
NOTE: The size of the PTC ceramic heaters depends on the size of the microfluidic chip; the length should be greater than the length of the chip-the length-width-height of the PTC ceramic heaters used here is 10 cm x 2 cm x 0.4 cm. - Connect the heater to the solid-state relay.
- Connect the solid-state relay to the PID temperature controller.
- Attach the temperature sensor probe to the bottom of the two heaters and connect the terminal to the PID temperature controller.
- Connect two solid-state relays in series and connect the power cord.
- 3D print a slot for the two heaters and keep the heaters on the same plane (see Supplemental File 1 for STL files required for 3D printing).
NOTE: Leave a 12 mm air gap between the two heaters. - Place the microfluidic chip on the two heaters.
- Prepare a syringe pump and a syringe, fix the syringe on the syringe pump, connect a silicone tubing (0.8 mm inner diameter [ID]) with a syringe, and connect a steel needle (0.7 mm ID) to the top of the silicone tubing (Figure 3).
- Insert the steel needle into the inlet of the microfluidic chip.
- Place a pipette tip at the outlet of the chip to collect the PCR products.
4. Construction of the CE system
- Use a high-voltage power supply to generate a pulsed-field electric field; note the positive and negative electrodes.
- Use a mercury lamp as the light source and filter the excitation wavelength from the mercury lamp through a filter.
- Place the capillary on the microscope stage.
- Collect the fluorescence emission with the objective, and then detect it using an R928 photomultiplier tube (PMT). Observe the microscope and the capillary. Turn on the light under dark room conditions; excitation light is collected by the objective.
- Use the self-developed LABVIEW software to control the power supply and complete the data acquisition (described in steps 6.3-6.6). See https://github.com/starliyan/labview/blob/main/CZE20170723.vi.
NOTE: All experiments were performed in a dark room, and the schematic of the CE system is shown in Figure 4.
5. Run the PCR solution
- Preset the temperature of the heaters of the CF-PCR system at 65 °C and 95 °C.
- Place the microfluidic chip on the two heating blocks.
- Insert the tip of the silicone tube into a centrifuge tube containing 50 µL of the PCR solution (from step 2.3). Pull the syringe plunger to slowly withdraw the solution. Fix the syringe on a syringe pump. Insert the steel needle into the inlet of the microfluidic chip.
- Set the flow rate of the pump to 10 µL/min and press the start button to push the solution in the microchannel at the inlet of the microchip.
- Collect the PCR products at the outlet of the microfluidic chip.
NOTE: Rinse the channel with ultrapure water before and after the PCR to remove impurities.
6. Detection of the PCR products by this CE system built in-house
- Prepare a capillary with an 8 cm total length and a 6 cm effective length.
- Prepare the separation buffer by mixing 100 µL of 1% hydroxyethylcellulose (HEC, w/v), 2 µL of 100x SYBR Green I, and 98 µL of ultrapure water to obtain 0.5% HEC (w/v) containing 1x SYBR Green I.
- Fill the capillary with the prepared separation buffer using a vacuum pump.
- Input the injection voltage (800 V) and the injection time (2.0 s) on the software interface, click the start button, and wait for the PCR products to be electrodynamically introduced into the capillary at 100 V/cm (2.0 s).
NOTE: In this step, the PCR products are placed on the negative electrode and the buffer is placed on the positive electrode. - Input the DC voltage (800 V) on the software interface, click the start button, and run the electrophoresis at 100 V/cm of electric field strength.
NOTE: In this step, both the positive and negative electrodes are placed with buffer. - Click the stop button after all DNA fragments are separated in the capillary.
- Flush the capillary with sterilized water for 1 min after each run.
Representative Results
Figure 5 represents the electropherogram of the PCR products and the DNA markers. Trace (Figure 5A) is the CE result of the CF-PCR amplified product, trace (Figure 5B) is the CE result of the product amplified by thermal cycling, and trace (Figure 5C) is the CE result of the 100 bp DNA ladder. We first amplified the target gene of E. coli in the CF-PCR system; the PCR solution took ~10 min, 30 s from the inlet to the outlet of the chip. The size of the target amplicon of E. coli was 544 bp. The amplified PCR products were analyzed in the CE system, and CE of a 100 bp DNA ladder was also performed under the same experimental conditions. The size of the PCR product can be evaluated according to the electropherogram of the DNA ladder. Each experiment was performed three times for reproducibility. The data in Figure 5 show that the peaks corresponding to the PCR products of E. coli were observed after separation, and the migration times of the PCR products in the microfluidic chip were consistent with the one in a thermal cycler.
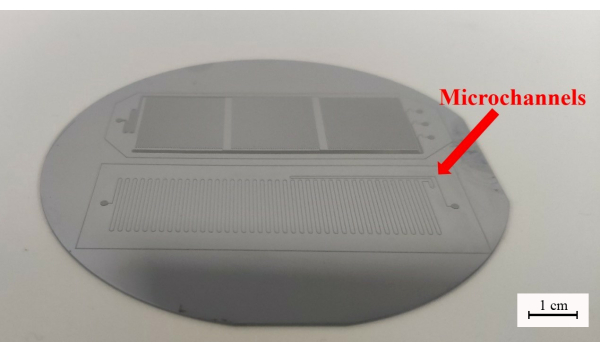

Figure 1: The cleaned silicon wafer. Scale bar = 1 cm. Please click here to view a larger version of this figure.
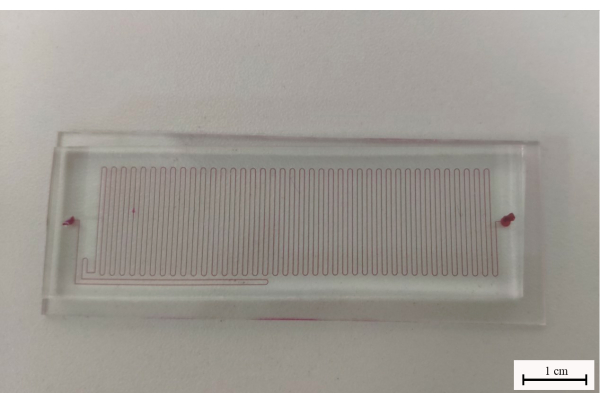

Figure 2: The CF-PCR microfluidic chip made using PDMS. The microchannels were filled with red ink for visualization. Scale bar = 1 cm. Abbreviations: CF-PCR = continuous-flow-PCR; PDMS = polydimethysiloxane. Please click here to view a larger version of this figure.
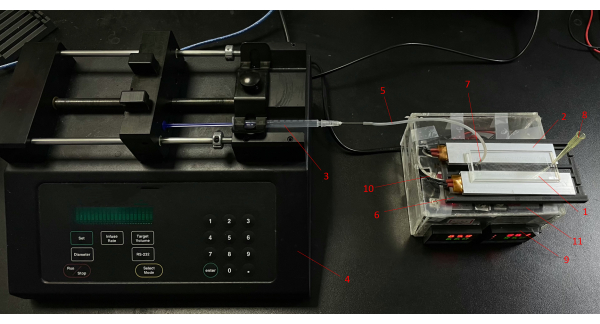

Figure 3: The CF-PCR system. (1) Microfluidic chip, (2) PTC ceramic heater, (3) syringe, (4) pump, (5) silicone tubing, (6) slot, (7) steel needle, (8) pipette tip, (9) PID temperature controller, (10) temperature sensor, (11) solid-state relay. Abbreviation: CF-PCR = continuous-flow-PCR. Please click here to view a larger version of this figure.
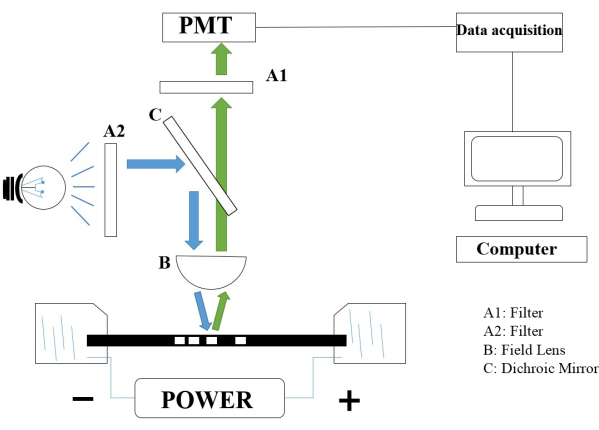
Figure 4: The construction of the CE system. Abbreviations: CE = capillary electrophoresis; PMT = photomultiplier tube; A1, A2 = filters; B = field lens; C = dichroic mirror. Please click here to view a larger version of this figure.
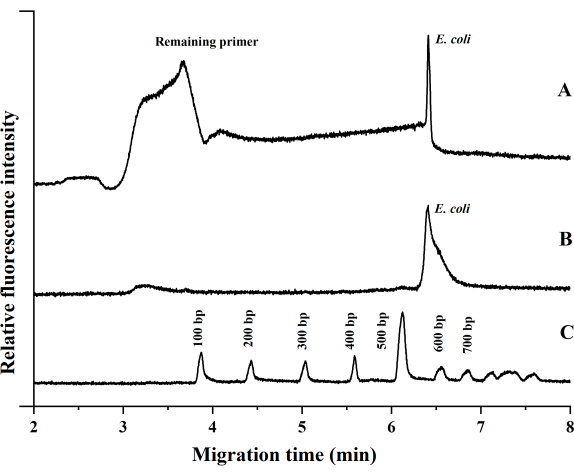
Figure 5: The electropherogram of Escherichia coli. Amplification by (A) CF-PCR microfluidic chip, (B) traditional PCR thermal cycler, and (C) DNA markers. Abbreviation: CF-PCR = continuous-flow-PCR. Please click here to view a larger version of this figure.
| Target | Sequence 5′……3′ | Amplicon (bp) | ||
| EC-F | GGAAGAAGCTTGCTTCTTTGCTGAC | 544 | ||
| EC-R | AGCCCGGGGATTTCACATCTGACTTA | |||
Table 1: Primers employed for Escherichia coli. Abbreviations: EC = Escherichia coli; F = forward; R = reverse.
| 10x Fast Buffer I | 5.0 μL |
| dNTP mixture (2.5 μM) | 4.0 μL |
| SpeedSTAR HS DNA Polymerase | 0.25 μL |
| polyvinyl pyrrolidone (PVP) | 3.5 μL |
| Tween 20 | 1.5 μL |
| template | 1.0 μL |
| EC-F | 0.5 μL |
| EC-R | 0.5 μL |
| ultrapure water | 33.75 μL |
Table 2: The components of the PCR solution. Abbreviations: EC = Escherichia coli; F = forward; R = reverse.
Supplemental File 1: STL file for 3D printing. Please click here to download this File.
Discussion
Both PCR and CE are two popular biotechnologies in the analysis of nucleic acids. This paper describes the amplification of E. coli and the detection of the PCR products using the CF-PCR and CE systems, both built in-house. The target gene of E. coli was successfully amplified within 10 min because of the high heat transfer rates. The DNA fragments smaller than 1,500 bp were separated within 8 min (Figure 5). The great advantage of these two techniques is that it can greatly save time compared to the traditional PCR and slab gel electrophoresis methods. Researchers should bear in mind that the CF-PCR microfluidic chip needs to be cleaned after use, and the CE system should be built in a clean and black house to avoid the contamination of samples. The CF-PCR system based on the microfluidic chip and the CE system introduced in this work are easy to fabricate and may offer a simple method for the analysis of nucleic acids in the lab. However, a limitation of CF-PCR is that it can only amplify and detect only one sample at a time, which may limit its wide application; to counter this, users can develop the integrated CF-PCR and CE array microfluidic chip to solve this problem. The platform reported in this work has great potential application in the field of clinical diagnosis of infectious diseases and nucleic acid research.
Divulgations
The authors have nothing to disclose.
Acknowledgements
This work was supported by the Science and Technology Commission of Shanghai Municipality, China (No. 19ZR1477500 and No.18441900400). We gratefully acknowledge financial support from the University of Shanghai for Science and Technology (No.2017KJFZ049).
Materials
| 100 bp DNA ladder | Takara Bio Inc. | 3422A | |
| 10x Fast Buffer I | Takara Bio Inc. | RR070A | |
| 10x TBE | Beijing Solarbio Science & Technology Co., Ltd. | T1051 | |
| developer solution | Alfa Aesar, USA | L15459 | |
| dNTP mixture (2.5 μM) | Takara Bio Inc. | RR070A | |
| EC-F | Sangon Biotech, Shanghai, China | ||
| EC-R | Sangon Biotech, Shanghai, China | ||
| HEC,1300K | Sigma-Aldrich, USA | 9004-62-0 | |
| isopropanol | Aladdin, Shanghai, China | 67-63-0 | |
| microscope | Olympus, Japan | BX51 | |
| photolithography | SUSS MicroTec, Germany | MJB4 | |
| photomultiplier tube | Hamamatsu Photonics, Japan | R928 | |
| photoresist | MicroChem, USA | SU-8 2075 | |
| PID temperature controllers | Shanghai, China | XH-W2023 | |
| plasma cleaner | Harrick Plasma | PDC-32G-2 | |
| polyvinyl pyrrolidone (PVP) | Aladdin, Shanghai, China | P110608 | |
| pump | Harvard Apparatus | PHD2000 | |
| silicone tubing | BIO-RAD,USA | 7318210 | |
| solid-state relays | KZLTD, China | KS1-25LA | |
| SpeedSTAR HS DNA Polymerase | Takara Bio Inc. | RR070A | |
| steel needle | zhongxinqiheng,Suzhou,China | ||
SYBR GREEN  |
Solarbio, Beijing, China | SY1020 | |
| temperature sensors | EasyShining Technology, Chengdu, China | TCM-M207 | |
| Template (E. coli) | Takara Bio Inc. | AK601 | |
| Tween 20 | Aladdin, Shanghai, China | T104863 | |
| voltage power supply | Medina, NY, USA | TREK MODEL 610E |
References
- Li, Z., et al. All-in-one microfluidic device for on-site diagnosis of pathogens based on an integrated continuous flow PCR and electrophoresis biochip. Lab on a Chip. 19 (16), 2663-2668 (2019).
- Crews, N., Wittwer, C., Gale, B. Continuous-flow thermal gradient PCR. Biomedical Microdevices. 10 (2), 187-195 (2008).
- Li, Z., et al. Design and fabrication of portable continuous flow PCR microfluidic chip for DNA replication. Biomedical Microdevices. 22 (1), 5 (2019).
- Kim, J. A., et al. Fabrication and characterization of a PDMS-glass hybrid continuous-flow PCR chip. Biochemical Engineering Journal. 29 (1-2), 91-97 (2006).
- Shen, K., Chen, X., Guo, M., Cheng, J. A microchip-based PCR device using flexible printed circuit technology. Sensors and Actuators B: Chemical. 105 (2), 251-258 (2005).
- Harstad, R. K., Johnson, A. C., Weisenberger, M. M., Bowser, M. T. Capillary Electrophoresis. Analytical Chemistry. 88 (1), 299-319 (2016).
- Redman, E. A., Mellors, J. S., Starkey, J. A., Ramsey, J. M. Characterization of intact antibody drug conjugate variants using microfluidic capillary electrophoresis-mass spectrometry. Analytical Chemistry. 88 (4), 2220-2226 (2016).
- Britz-Mckibbin, P., Kranack, A. R., Paprica, A., Chen, D. D. Quantitative assay for epinephrine in dental anesthetic solutions by capillary electrophoresis. Analyst. 123 (7), 1461-1463 (1998).
- Maeda, H., et al. Quantitative real-time PCR using TaqMan and SYBR Green for Actinobacillus actinomycetemcomitans, Porphyromonas gingivalis, Prevotella intermedia, tetQgene and total bacteria. FEMS Immunology and Medical Microbiology. 39 (1), 81-86 (2003).
- Hajba, L., Guttman, A. Recent advances in column coatings for capillary electrophoresis of proteins. TrAC Trends in Analytical Chemistry. 90, 38-44 (2017).
- Kleparnik, K. Recent advances in combination of capillary electrophoresis with mass spectrometry: methodology and theory. Electrophoresis. 36 (1), 159-178 (2015).

