Vertical Immobilization Method for Time-Lapse Microscopy Analysis in Filamentous Cyanobacteria
Summary
We present a simple and accessible method for filamentous cyanobacterial visualization in the XY plane. A low-melting point agarose matrix was used, allowing the acquisition of images of proteins involved in the division, in a vertical orientation. Therefore, this methodology can be applied to any filamentous organism and different kinds of proteins.
Abstract
A main event in bacterial cell division is the septation process, where the protein FtsZ is the key element. FtsZ polymerizes forming a ring-like structure (Z-ring) in the middle of the cell that serves as a scaffold for other division proteins. Super-resolution microscopy in bacterial models Escherichia coli and Bacillus subtilis showed that the Z-ring is discontinuous, while live cell imaging studies demonstrated that FtsZ moves along the ring by a mechanism known as treadmilling. To study the dynamics of FtsZ in vivo, a special cell placement in a vertical position is necessary for imaging the complete structure of the ring in the XY plane. In the case of FtsZ imaging in multicellular cyanobacteria, such as Anabaena sp. PCC7120, maintaining the filaments in a vertical position is challenging because of the size of the cells and the filaments’ length. In this article, we describe a method that allows the vertical immobilization of Anabaena sp. PCC 7120 filaments using low melting point agarose and syringes, to record the Z-ring in a mutant that expresses a FtsZ-sfGFP fusion protein. This method is a rapid and inexpensive way to register protein dynamics at the division site using confocal microscopy.
Introduction
Bacterial cell division is the process in which a mother cell generates two daughter cells, in most cases by the mechanism known as binary fission. One of the earliest events in the septation process is the localization of FtsZ in the middle of the cell1. This protein, which is structurally homologous to tubulin2, is conserved and widely distributed in most bacteria, and its polymerization generates a contractile structure known as the Z-ring3. This ring serves as a scaffold for other division proteins and together they form a molecular machinery called a divisome. Several studies have shown that the Z-ring is highly dynamic and that FtsZ protofilaments move by treadmilling4,5,6. To study the Z-ring in time-lapse experiments, it is advisable to record the division site in the XY plane for better resolution and rapid sampling. In order to achieve this, it is necessary to develop vertical cell immobilization methods that commonly include the nanofabrication of microhole cell traps and complex microfluidic devices7.
Cyanobacteria are photosynthetic microorganisms that are classified as gram-negative by their cellular morphology. However, phylogenetically they are closer to gram-positive bacteria8. These organisms have cell division genes that are common within gram-positive and gram-negative bacteria, but their divisome also contains unique elements9. Anabaena sp. PCC 7120 (hereafter Anabaena sp.) is filamentous cyanobacteria with one division plane and is a model for the study of cell division in multicellular cyanobacteria. In this strain, it has been possible to determine the positioning of FtsZ in the middle of the cells10. Nevertheless, there are no studies showing the in vivo dynamics of FtsZ in this model. In our laboratory, through triparental mating and homologous recombination, we obtained a totally segregated mutant of Anabaena sp. that expresses the FtsZ protein fused to sfGFP, which replaced the complete endogenous ftsZ gene. We developed a rapid and simple cell immobilization method that favors the vertical orientation of the mutant strain filaments for time-lapse experiments to visualize the division proteins in filamentous cyanobacteria. This method does not need microfluidic devices that can be expensive and difficult to develop. As an example, we used this protocol to visualize the Z-ring in the FtsZ-sfGFP mutant by confocal microscopy.
Protocol
1. Considerations and selection of the cellular model
NOTE: Cyanobacteria have a strong autofluorescence due to the presence of photosynthetic pigments. This signal is in the red part of the spectrum, therefore, the fluorescent proteins suitable for imaging in cyanobacteria are the ones that far from the red emission. For example, GFP, YFP, Venus, Turquoise and BFP.
- Select a divisome component present in the target filamentous cyanobacterial strain. For the experiments, it is necessary to construct a filamentous cyanobacterial mutant that expresses the selected divisome component fused to a fluorescent protein. In this protocol, an Anabaena sp. mutant that expresses FstZ-sfGFP, as an example. This strain was obtained by triparental mating with E. coli11.
- Check the segregation level of the mutant that will be used. In the example, the segregation was determined by PCR amplification of the target gene. The FtsZ-sfGFP mutant used in this protocol is a fully segregated strain that lacks the wild-type ftsZ gene.
2. Growth conditions
- Make a fresh subculture of the mutant using 10 mL culture in stationary phase (grown for approximately 14 days in constant light, at 25°C , for Anabaena sp.) and adding to 90 mL of BG11 medium supplemented with antibiotics (Spectinomycin and Streptomycin 10 µl ml-1 for the FtsZ-sfGFP mutant).
- Grow the new cell culture in constant light and at the optimum temperature until reaching the exponential phase. The FtsZ-sfGFP mutant of Anabaena sp. is grown at 25 °C for 7 days before the sample preparation.
3. Sample preparation in an agarose matrix
- Take 2 mL of the homogenized growth culture.
- Centrifuge at 2,500 x g for 10 min at room temperature.
- In the meantime, in a 50 mL flask, heat 30 mL of 3% low melting point agarose in BG11 liquid medium. Use a microwave set to the medium power level and check the solution every 15 s until completely dissolved. Set aside to cool down to 37 °C.
NOTE: Autoclave the solution of agarose to avoid contamination. - Once centrifugation is done, discard 1.9 mL of the supernatant and resuspend the cells in the remaining volume by pipetting at least three times up and down.
- Mix the resuspended cells with 900 µL of the 3% agarose solution by pipetting up and down at least three times.
NOTE: The agarose solution must be liquid but not too hot to avoid cell damage. The next step must be performed fast and before the agarose solution forms a gel like consistency =. - Aspirate the agarose matrix in a 1 mL syringe carefully. It is important to avoid the generation of bubbles while the sample is being aspirated with the syringe.
NOTE: Before this step, make sure that the tip of the syringe is cut to eliminate the narrow entry hole. - Let the sample-agarose mixture solidify by placing the syringe in the horizontal position at room temperature for 2 h.
- Remove the agarose matrix carefully using the plunger and put it on a clean and flat surface.
- Cut the solidified sample into slices, in a thickness range of 0.5 – 1 mm, maintaining the integrity of the matrix.
NOTE: For better results, cut the agarose matrix using a scalpel or thin razor blades. - Place the slices side by side on a coverslip. The number of disks on a coverslip will depend on its dimensions.
NOTE: For time-lapse microscopy, it is recommended to use a cell chamber made for microscopy analysis (e.g., Attofluor or Chamlide Magnetic chambers). These chambers allow the acquisition of the sample avoiding dehydration. In this example the samples are prepared on rounded coverslips (25 mm) using the Attofluor chamber. - Add 2 mL of melted 3% agarose solution on the sample placed in the chamber to prevent dehydration.
- Wait until the agarose solution is completely solidified to form a gel.
4. Image acquisition
- Standardize the parameters to visualize the fluorescent protein of interest in the mutant strain in a confocal microscope. Make sure that the microscope can detect both the autofluorescence and the selected fluorescent marker.
- Visualize the sample in the bright field conformation with maximum magnification, and search a vertically oriented filament as shown in Figure 1.
- Find the division site of interest with an initial scanning using the autofluorescence signal along the Z plane.
- Use the parameters of the fluorescent protein marker to visualize the signal of the protein of interest at the division site.
- Perform time-lapse experiments according to the biological model and the protein of interest. For example, in this case time-lapse experiments of 10 s in 50 s were performed to record the dynamics of FtsZ along the Z-ring in Anabaena sp. (Figure 2).
Representative Results
Visualization of the Z-ring in Anabaena sp. using the vertical immobilization method
To study the dynamics of Z-ring components in bacteria, it is necessary to acquire images in vertically oriented cells. In this position, it is possible to visualize the Z-ring and the main proteins of the divisome to monitor protein dynamics by time-lapse microscopy. The classic sample preparation for bacteria does not work for filamentous cyanobacteria. In the case of Anabaena sp., the filaments are oriented in a position that does not allow to visualize the Z-ring. Additionally, it is not possible to keep the filaments immobilized for correct visualization of cell division proteins in a single cell over extended periods of time. Therefore, in order to visualize the Z-rings of the FtsZ-sfGFP mutant in Anabaena sp we developed and standardized a protocol using LMP agarose and horizontal incubation of the sample. This protocol allows obtaining a higher proportion of vertically oriented filaments, compared to previously used procedures (Figure 1). Therefore, it was possible to record the Z-ring in the XY plane in cells of the mutant strain on direct contact with the coverslip (Figure 2A).
Z-ring dynamics in the FtsZ-sfGFP Anabaena sp. mutant using the vertical immobilization method
After testing the vertical immobilization method in conventional confocal microscopy, samples of the FtsZ-sfGFP Anabaena sp mutant prepared using the same protocol were visualized by Airyscan microscopy. With this technique, we observed less photobleaching in the rings and, therefore, the experiments can be performed for longer periods without losing the fluorescence signal. In fact, we were able to obtain images for periods of approximately 16.5 min, with acquisition of images every 10 s with low photobleaching, which allowed us to detect variations in fluorescence intensity over time in different regions of the Z-ring (Figure 2B).
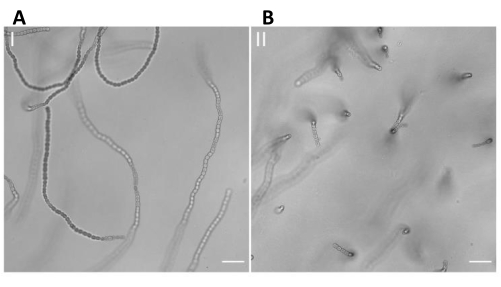
Figure 1. Visualization of Anabaena sp. filaments using the vertical immobilization method. A sample of Anabaena sp. was incubated horizontally in LMP agarose 3% in BG11 within a syringe. The sample was then cut into disks and finally placed in the cell chamber. The visualization of the cells was carried out using an inverted microscope. (A) Representative brightfield microscopy of a sample incubated in the agarose matrix without performing the vertical immobilization method. This image shows that most of the filaments are horizontally oriented. Scale bar = 20 µm. (B) Representative brightfield microscopy showing a high proportion of vertically oriented filaments of Anabaena sp. after using the vertical immobilization method. Scale bar = 20 µm. Please click here to view a larger version of this figure.
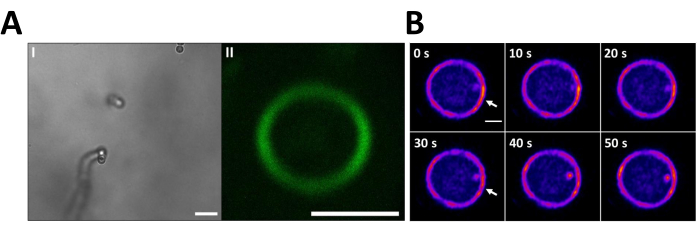
Figure 2. Visualization of the Z-ring in vertically oriented filaments of Anabaena sp. using the vertical immobilization method. To display the Z-rings in the XY plane, the cells of the sfGFP-FtsZ strain were incubated in LMP agarose 3% at 25 °C following the vertical immobilization method (A.I). Brightfield microscopy shows a filament of the mutant in a vertical position. Scale bar =10 µm. (A.II) The FtsZ-sfGFP signal in a Z-ring of the filament shown in (A.I). This image is the average intensity of 6 images taken every 10 s for 1 min. Scale bar = 2 µm (B) FtsZ-sfGFP signal of the Z-ring recorded every 10 s for 50 s in the cells of the FtsZ-sfGFP mutant oriented vertically in the BG11-agarose matrix. Time (seconds) is indicated on the upper left corner of each image. It is possible to observe the Z-ring and a cluster of FtsZ-sfGFP that changes its position over time (white arrows). Scale bar = 1 µm. Please click here to view a larger version of this figure.
Discussion
The study of the dynamics of divisome proteins is without doubt a challenge. Particularly, in filamentous cyanobacteria, one the challenges is the visualization the Z-ring in the horizontal plane, for which the cells must be vertically oriented. The method we described here allows the positioning of the Z-ring in the XY plane to perform the different analyzes. This is the first simple method for filamentous cyanobacteria that allows the complete visualization of the Z-ring.
Although this protocol is simple and fast, special care must be taken during the sample preparation step in the agarose matrix, since the low-melting point agarose must be at a suitable temperature to be fluid enough for mixing with the cell solution. At the same time, the agarose solution cannot be too warm since it could provoke cell damage. Another critical step to consider is the cutting of the discs because if it is not done carefully, the matrix can collapse and damage the sample.
Using this method, the proportion of filaments in the vertical position is higher than with the classical protocol. However, it is necessary to cut as many discs as possible to ensure the statistical analysis, i. e., several filaments in a vertical position.
Compared with other methods, where single cell studies in microfluidic systems can be carried out, our method is easier, faster, and cheaper, requiring only materials and reagents that are easily available at any laboratory. On the other hand, although microfluidic systems are being widely used in multicellular cyanobacteria studies, to our knowledge they have not solved the problem of cell positioning in a vertical position12.
A limitation of the technique is that the orientation is not always perfect. This is because we have seen tilted filaments in the agarose matrix, however, with the previous screening using autofluorescence signal it is possible to identify this artifact. Also, we could not visualize a full division event, because this protocol allows the visualization of the division site as much as 1 hour, while the division time of Anabaena sp. is around 12 hours. However, this protocol can be helpful to record a full division event in fast growing bacteria.
Although our method was standardized for Anabaena sp. it can be applied to all filamentous cyanobacteria or filamentous organisms, to study not only FtsZ but also other proteins associated with the divisome since they will be oriented in the same XY plane.
Divulgations
The authors have nothing to disclose.
Acknowledgements
We are grateful to National doctoral scholarships (ANID 21211333; 21191389) for the financing. Grant Fondecyt 1161232.
This work was supported by de Advanced Microscopy Facility UMA UC.
Materials
| 1 ml syringe | Qingdao Agna Medical Technology Co., Ltd. | – | The disposable medical plastic syringe with 1 ml needle generally used to pump liquid or injection liquid, in this experiment it is used to suck the sample and make it polymerize. |
| Attofluor Cell Chamber | ThermoFischer Scientific | A7816 | The Attofluor Cell Camera is a durable and practical coverslip holder designed for viewing live cell samples in upright or inverted microscopes. |
| Axygen MaxyGene II Thermal Cycler with 96 well block | CORNING | THERM-1001 | The new MaxyGene II Thermal Cycler increased speed and advanced features, providing the premium performance you have come to expect from Axygen brand products. Unique flexible programming. Rapid run times. Improved workflow over traditional gradient cyclers. Ramping rates up to 5°C/sec. Adjustable heated lid accommodates strips, tubes and microplates. |
| Fisherbrand Cover Glasses: Circles | ThermoFischer Scientific | 12-546-2P | Made of finest optical borosilicate glass, with uniform thickness and size. Circular shape. Corrosion-resistant. |
| Low Melting Point agarose | Promega | V3841 | Agarose, Low Melting Point, Analytical Grade, is ideal for applications that require recovery of intact DNA fragments after gel electrophoresis. |
| LSM 880 microscope from Zeiss Airyscan | Zeiss | – | The Zeiss Airyscan LSM 880 microscope is a laser scanning focal microscope that can acquire images under the resolution limit (lateral resolution ~ 120nm and axial resolution ~ 350nm). The detectors are highly sensitive making it possible to acquire super-resolution images at high speed. |
| Microcentrífuga Fresco 17 | ThermoFischer Scientific | 75002402 | Speed up routine sample preparation processes up to 17,000 × g with our standard microcentrifuge, available with refrigeration. These microcentrifuges offer productivity, versatility, safety and convenience in an easy-to-use, compact design laboratory instrument. |
| Nikon Timelapse Microscope | Nikon | – | The Nikon C2 laser scanning confocal microscope is an ideal microscope for long-term timelapse, because thanks to its incubation chamber it is possible to keep samples in optimal conditions for more than 8 hours. |
| Thin razor blades Schick Super Chromium | Farmazon | SKU: 401146 | The thin razor blades is used to cut the agarose matrix, allowing us to obtain the disks with the sample while maintaining the integrity of the matrix. |
References
- Löwe, J., Amos, L. A. Crystal structure of the bacterial cell-division protein FtsZ. Nature. 391 (6663), 203-206 (1998).
- Erickson, H. P. FtsZ, a prokaryotic homolog of tubulin. Cell. 80 (3), 367-370 (1995).
- Huang, K., Mychack, A., Tchorzewski, L. Characterization of the FtsZ C-terminal variable ( CTV ) region in Z-ring assembly and interaction with the Z-ring stabilizer ZapD in E. coli cytokinesis. , 1-24 (2016).
- Perez, A. J., et al. Movement dynamics of divisome proteins and PBP2x:FtsW in cells of Streptococcus pneumoniae. Proceedings of the National Academy of Sciences. 116 (8), 3211-3220 (2019).
- Bisson-Filho, A. W., et al. Treadmilling by FtsZ filaments drives peptidoglycan synthesis and bacterial cell division. Science. 355 (6326), 739-743 (2017).
- Yang, X., et al. GTPase activity-coupled treadmilling of the bacterial tubulin FtsZ organizes septal cell wall synthesis. Science. 355 (6326), 744-747 (2017).
- Whitley, K. D., et al. FtsZ treadmilling is essential for Z-ring condensation and septal constriction initiation in Bacillus subtilis cell division. Nature Communications. 12 (1), (2021).
- Mazón, G., et al. LexA-binding sequences in Gram-positive and cyanobacteria are closely related. Molecular Genetics and Genomics. 271 (1), 40-49 (2004).
- Mandakovic, D., et al. CyDiv, a conserved and novel filamentous cyanobacterial cell division protein involved in septum localization. Frontiers in Microbiology. 7 (FEB), 1-11 (2016).
- Camargo, S., et al. ZipN is an essential FtsZ membrane tether and contributes to the septal localization of SepJ in the filamentous cyanobacterium Anabaena. Scientific Reports. 9 (1), 1-15 (2019).
- Thiel, T., Peter Wolk, C. Conjugal Transfer of Plasmids to Cyanobacteria. Methods in Enzymology. 153 (C), 232-243 (1987).
- Moffitt, J. R., Lee, J. B., Cluzel, P. The single-cell chemostat: An agarose-based, microfluidic device for high-throughput, single-cell studies of bacteria and bacterial communities. Lab on a Chip. 12 (8), 1487-1494 (2012).

