A Fluorescence-based Exonuclease Assay to Characterize DmWRNexo, Orthologue of Human Progeroid WRN Exonuclease, and Its Application to Other Nucleases
Summary
Exonucleases play critical roles in ensuring genome stability. Loss of WRN exonuclease function results in premature aging. Studying substrates and other requirements of the nuclease in vitro can help elucidate its role in vivo. Here we demonstrate a rapid and reproducible fluorescence-based assay to measure its nuclease activity.
Abstract
WRN exonuclease is involved in resolving DNA damage that occurs either during DNA replication or following exposure to endogenous or exogenous genotoxins. It is likely to play a role in preventing accumulation of recombinogenic intermediates that would otherwise accumulate at transiently stalled replication forks, consistent with a hyper-recombinant phenotype of cells lacking WRN. In humans, the exonuclease domain comprises an N-terminal portion of a much larger protein that also possesses helicase activity, together with additional sites important for DNA and protein interaction. By contrast, in Drosophila, the exonuclease activity of WRN (DmWRNexo) is encoded by a distinct genetic locus from the presumptive helicase, allowing biochemical (and genetic) dissection of the role of the exonuclease activity in genome stability mechanisms. Here, we demonstrate a fluorescent method to determine WRN exonuclease activity using purified recombinant DmWRNexo and end-labeled fluorescent oligonucleotides. This system allows greater reproducibility than radioactive assays as the substrate oligonucleotides remain stable for months, and provides a safer and relatively rapid method for detailed analysis of nuclease activity, permitting determination of nuclease polarity, processivity, and substrate preferences.
Introduction
Nucleases serve a vital role in cells in removing damaged DNA, resolving nonduplex structures such as Holliday junctions and providing proof-reading capacity during DNA replication, both intrinsic within DNA polymerases and extrinsic to them1. Nucleases can act either by sequentially degrading DNA from free ends (exonucleases) or by cleaving internal phosphodiester bonds within a longer DNA molecule (endonucleases). Loss of nuclease activity can result in highly specific genome instability phenotypes. While mutation of the RecQ helicase family member BLM result in excessively high rates of sister chromatid exchange and globally elevated cancer rates (reviewed byPayne and Hickson2), mutation of the highly related WRN protein leads to premature aging3; the major significant difference between these two family members is the presence of a 3'-5' exonuclease domain with in WRN4. Evidence of a critical role of the WRN exonuclease in maintaining genome stability has accumulated from analysis of genotypes in WS patients5, together with point mutation and deletion studies in human cells, backed by crystallographic studies of the isolated exonuclease domain6. However, cooperation and cross talk between WRN's exonuclease activity and its central helicase activity7 makes it difficult to tease apart the functionality of each and their relative contributions to genome stability. In plants and lower metazoan animals, WRN exonuclease activity is present on a single polypeptide lacking helicase activity8-10 (reviewed in Cox and Boubriak11); it has been demonstrated biochemically in Arabidopsis that this exonuclease acts coordinately with the cognate WRN helicase, effectively reconstituting the combined enzyme activities observed in vertebrate WRN9. We have studied WRN exonuclease in Drosophila since the excellent genetic tools allow analysis of the impact of exonuclease mutation (without impacting on the presumptive cognate helicase) at the whole organism level and through development10,12. Moreover, we have cloned, expressed, and purified recombinant Drosophila WRN exonuclease (DmWRNexo) allowing full biochemical analysis of its enzyme properties13,14.
Nuclease analysis in vitro has traditionally been conducted using radiolabeled oligonucleotides, assessing degradation by looking for laddering of products on acrylamide gels4,8,15. While sensitive, such assays are not quantitatively reproducible day-to-day because of radioactive decay of the labeled substrates. Additionally, handling and disposal of radioactive reagents pose significant environmental and health issues; sourcing of radiolabel is also becoming increasingly problematic. An alternative recent method assesses the amount of the final degradation product by mass spectrometry16. However, it is time consuming (taking several days), requires specialized equipment, and the readout is the amount of end product (single nucleotide) so is not suitable for sensitive measurement of aspects such as enzyme processivity or for determining whether some nucleotides, sequences, or modifications lead to nuclease pausing or halt. To overcome these problems, we have adapted the traditional gel-based assays for use with fluorescent oligonucleotide substrates, generating stably labeled substrates that can be used reproducibly over long time periods and thus allow direct comparison of nuclease activities under different conditions.
Protocol
1. Preparation of Substrate Oligonucleotides
- Synthesize custom oligonucleotides.
- Design one single-stranded oligonucleotide with a 5'-conjugated fluorescein molecule; this will be the backbone for every substrate. (Note that this is for a 3'-5' exonuclease; for a 5'-3' polarity exonuclease use a 3'-conjugated backbone strand). If polarity is not known, both 3' and 5' should be tested.
- Design complementary strands such that when annealed to the fluorescently labeled oligonucleotide, the duplex substrate(s) of choice are formed. Note that the complementary strand is not conjugated to a fluorophore. When ordering, specify PAGE-purification (desalting is not sufficient).
- Store oligonucleotide stocks buffered in 10 mM at 1 nmol/μl at −20 °C and in the dark.
- Mix the backbone labeled oligonucleotide with each complementary unlabeled oligonucleotide in a 1:1.2 molar ratio of labeled:unlabeled in buffer (10 mM Tris-HCl pH 7.6, 1 mM EDTA pH 8.0, 50 mM NaCl). (Note: a 1:1 ratio is most desirable but it is more important that ALL of the labeled strand is bound. Ratios of 1:1 – 1:1.5 can work.)
- Heat a waterbath or heatblock to 95 °C and place the annealing mixture into this for 3 min then switch off and allow the whole to cool slowly (1 °C/min) to below 30 °C.
- Check full annealing of the labeled oligonucleotide by running the resulting duplex substrate on a native 10% polyacrylamide gel (10% 19:1 acrylamide:bis-acrylamide in 1x Tris-Borate-EDTA (TBE, 89 mM Tris base, 89 mM boric acid, 2 mM EDTA) polymerized with 0.05% w/v ammonium persulfate (APS) and 0.005% v/v TEMED).Visualize via PhosphorImager (see section 5 below).
2. Preparation of Acrylamide Gels
- Set up the gel casting system e.g. Hoefer SE400 gel rig with 16 cm x 18 cm plates and 1.5 mm spacers.
- Clean glass plates and spacers thoroughly using detergent and water. Wear clean gloves to avoid resoiling the plates with oil from fingers, which would hamper consistent gel formation.
- Rinse the plates with plenty of distilled (deionized) water to remove detergent residue, then wipe with lint-free tissue (e.g. Kimwipes).
- Finally, spray with 70% ethanol and wipe – glass plates should squeak if they are clean. Set aside upright to allow the last of the ethanol to evaporate.
- Clean the rest of the gel forming equipment with distilled water (use detergent first if necessary) and dry.
- Assemble the glass plates and spacers into the gel-forming rig, making sure the system is leakproof.
- Prepare the gel solution to the desired polyacrylamide percentage
- For nuclease assay, use denaturing 14% polyacrylamide gels with 8 M urea in 1x TBE.
- To make 100 ml (enough for 2 gels with plenty of excess for leakage) take 48.05 g urea in a wide beaker and add 35 ml 40% 19:1 acrylamide solution and 10 ml 10x TBE (890 mM Tris, 890 mM boric acid, 20 mM EDTA, pH 8.3 with NaOH). CAUTION: Wear gloves – acrylamide is a potent neurotoxin. Dissolve the urea using a stir bar and magnetic stir plate (this will take some time).
- When dissolved, add milliQ-deionized water to 100 ml. Filter-sterilize using a 0.22 μm membrane. If not to be used immediately, degas briefly using suction equipment and store in the dark (will keep a few days). Do not refrigerate as the urea will precipitate.
- Polymerize and cast the gel.
- Take the urea-acrylamide and add 0.5 ml 10% (w/v) ammonium persulfate (APS, made up in distilled water) and 50 μl TEMED. Swirl the container slowly to mix without imparting air (which will inhibit polymerization) and pour into the gel former. Act quickly or the gel may set before pouring is finished.
- Add the appropriate comb to form wells for sample loading (chosen according to the gel rig and sample number to be analyzed e.g. 15- or 29-tooth combs are suitable for use with the Hoefer gel rig). Check that no air bubbles are introduced under the comb teeth.
- Leave to set. Monitor for leakage. Full polymerization is retarded at the combs owing to contact with the atmosphere. When the wells are firm, the rest of the gel will be set.
- Remove the combs and rinse out unpolymerized acrylamide using water and a pipette.
- Set up the gel for running.
- Set the cast gel into the gel running system and add the appropriate amount of 1x TBE running buffer into the lower chamber and into the gel wells. The gel is now ready to be loaded.
3. Preparation of His-tagged Recombinant DmWRNexo Protein
- Bacterial expression of DmWRNexo protein
- Using an inoculation loop, collect bacteria by scraping over the surface of glycerol stocks maintained at -80 °C (bacteria are E. coli T7 Iq LysY (NEB) transformed with pIVEX2.3d-DmWRNexo). Streak out onto LB-agar plates (10 g tryptone, 5 g yeast extract, 5 g NaCl, 15 g agar made up to 1 L with distilled water and pH adjusted to 7.4) with 50 μg/ml ampicillin. Incubate inverted overnight at 37 °C.
- Pick a single colony from the plate into 50 ml LB (10 g tryptone, 5 g yeast extract, 5 g NaCl made up to 1 L with distilled water) supplemented with 50 μg/ml ampicillin (made up in water and filter-sterilized to 0.22 μm). Grow with shaking at 220 rpm 37 °C overnight.
- Inoculate 1.8 L (aliquots of 600 ml each in three 2 L conical flasks to ensure adequate aeration) of LB-ampicillin with the overnight 50 ml culture and grow to OD600 ~ 0.8. Add IPTG to a final concentration of 0.7 mM and incubate at 26 °C (to prevent protein aggregation) with shaking for 4-8 hr.
- Harvest the cell pellet by centrifugation at 8,000 rpm 4 °C for 10 min – use 6 x 500 ml sterile plastic centrifuge bottles with 300 ml cell suspension in each. Discard supernatant and resuspend the pellet in a small volume (10 ml) of the original LB-amp , transfer to a 15 ml Falcon tube and then respin at 5,000 rpm to pellet the cells – remove the supernatant again and freeze at -80 °C (this helps with lysis).
- Bacterial cell lysis
- Make up 3.2 ml lysis buffer containing 1 ml 5 M NaCl, 1 ml 1 M DTT, 200 μl 1 M benzamidine and 1 ml 10 mg/ml lysozyme (resuspended in distilled water).
- To each pellet derived from 300 ml bacterial suspension, add 320 μl lysis buffer. Resuspend as much as possible – use a 'sawn-off' p1000 tip and stir as well as pipetting up and down. Incubate on ice 20 min. Add a further 100 μl lysis buffer and mix well then transfer as much as possible of the resuspended lysate into a microfuge tube (it will be very viscous and hard to pipette).
- Centrifuge the lysate at 14,000 rpm in a precooled microfuge rotor at 4 °C for 30 min, then take the supernatant to a clean microfuge tube: this contains the recombinant protein.
- His-trap purification
- Preparation of reagents: Label 5 microfuge tubes for each of the following concentrations of imidazole: 40 mM, 60 mM, 100 mM, 300 mM, and 500 mM (e.g. label 40-1, 40-2, 40-3, etc.), which will be used to collect the column fractions. Store the tubes on ice in a rack with lids open (but cover loosely with clingfilm or similar to keep clean whilst preparing everything else). Make up 25 ml of 20 mM imidazole and 10 ml stocks of each of the other concentrations of imidazole in 25 ml universal tubes, using 1x PBS (8 g NaCl, 0.2 g KCl, 1.44 g Na2HPO4, 0.24 g KH2PO4, pH adjusted to 7.4 and made up to 1 L with distilled water) as the diluent.
- Adjust the bacterial lysate to 20 mM imidazole using an appropriate volume of 2 M imidazole stock solution, and adjust volume to at least 1 ml (<5 ml) using 20 mM imidazole.
- Wash a 1 ml His-Trap column (Amersham) with 10 ml MilliQ water by using a 5 or 10 ml syringe with Luer lock and gentle manual pressure, then equilibrate the column by passing through 10 ml of 20 mM imidazole made up in PBS (see step 3.3.1 above).
- Load the lysate onto the column. As the lysate is loaded onto the column, significant back-pressure may be experienced; this is perfectly normal. Do not be tempted to apply too much force to the syringe, and maintain a steady even pressure. It is usual to see a mild color change as the lysate enters the column. Collect the flow through in a Sterilin bijou bottle (7 ml capacity) and store on ice.
- Wash the column with 10 ml of 20 mM imidazole and collect this in a 25 ml universal tube on ice, labeled as 'wash'.
- Pass 5 ml of 40 mM imidazole through the column, collecting 1 ml fractions into each of the microfuge tubes (labeled 40.1, 40.2, 40.3, etc.): watch the plunger of the syringe carefully as this allows measurement of 1 ml.
- Now sequentially pass through 5 ml of each of 60 mM, 100 mM, 300 mM, and 500 mM imidazole, again collecting 5 x 1 ml fractions at each salt concentration.
- Analysis of column fractions
- Dot blot: Dot 2 μl of each column fraction (and also load, flow through and wash) onto a sheet of nitrocellulose - it is easier if this is preruled with a grid using a soft pencil). Allow to air dry. Process the dot blot by blocking for 15 min to 1 hr room temperature in 5% nonfat dried milk made up in PBS with 0.2% v/v Tween 20, then wash in PBS-Tween several times and incubate with HRP-anti-his antibody diluted 1:500 in PBS-Tween for 30-60 min. Wash extensively with PBS-Tween then PBS, and detect bound antibody using enhanced chemiluminescence (ECL), visualized on Hyperfilm MP. This dot blot gives an excellent idea of peak fractions prior to SDS-PAGE analysis, and can inform subsequent purification steps if necessary (e.g. if protein stability is an issue, it is possible to pool peak fractions based on dot blot results and load straight onto gel filtration columns – see step 3.5 below).
- SDS-PAGE: Collect 50 μl of each column fraction into a labeled microfuge tube containing 50 μl 2x SDS-PAGE loading dyes (4% w/v SDS, 20% v/v glycerol, 120 mM Tris-HCl pH 6.8 with 200 mM DTT). Boil these samples for 5 min at 95-100 °C (note if using a hot block without a heated lid, make a small hole in the lid of each tube with a narrow gauge syringe needle to prevent the tubes 'exploding' as they heat up). Run 10% SDS-PAGE minigels loading 10 μl of each column fraction and stain with Coomassie brilliant blue to verify peak fractions and % purity of the protein. If required, it is also possible to western blot for his-tagged protein by transferring proteins separated by SDS-PAGE onto nitrocellulose; block the membranes with 5% nonfat milk in PBS-0.2% Tween 20, then wash and probe with HRP-conjugated anti-his antibody (1:500) as for the dot blot.
- To increase purity, the peak fractions from the his-trap column can be pooled and loaded onto a Superdex 75 gel filtration column with elution in increasing salt (as previously described Boubriak et al.13 and Mason et al.14).
4. Nuclease Assay
- Make working stocks of substrates.
- Use 20 pmol substrate per reaction (though this can be varied according to the question to be addressed) so a convenient working stock is 200 pmol/μl (i.e. 10x). Working stocks are made up in small volumes of annealing buffer (10 mM Tris-HCl pH7.6, 1 mM EDTA pH 8.0, 50 mM NaCl) or in 10 mM Tris-HCl pH 7.0 (up to pH 8.5, according to nuclease) without salt or EDTA, according to enzyme sensitivity or assay to be performed .
- Keep at 4 °C and in the dark (wrapped in foil). Substrates last for many weeks with no loss of duplex integrity or of fluorescent signal.
- Assay exonuclease protein concentration by colorimetric analysis, or UV spectrometry at Abs260.
- Store in small aliquots at -80 °C.
- If samples are frozen/thawed once, assume a protein misfold rate of 5% (or define empirically for each protein to be assayed) and adjust concentrations accordingly.
- Discard proteins for assay of exonuclease activity after two freeze/thaw cycles, and note status of protein for each experiment.
- Design experiment and work out master mix (MM) quantities for samples.
- Make a MM containing all the components common to the samples being tested (usually everything except the enzyme or substrate) to decrease inter-sample variation.
- Allow for pipetting error when making up the MM e.g. calculate volumes for N samples by making sufficient for (N + 0.1N) samples.
- Set up reactions in a total volume of 20 μl/sample, containing 1 pmol of enzyme and 20 pmol of substrate (as default before optimization – obviously amounts vary according to what is being tested e.g. alter amount of enzyme if testing effect of enzyme concentration). For DmWRNexo, the optimal buffer for nuclease activity was found empirically to contain 40 mM Tris-HCl pH 8.0, 4 mM MgCl2, 5 mM dithiothreitol, with 0.1 mg/ml molecular biology grade BSA.
- Set up reactions.
- Keep all reagents at 4 °C (ice bucket or cryorack).
- Make up master mix and aliquot into the appropriate number of sterile DNase-free 0.5 ml microfuge tubes. Include a no-enzyme control to visualize the initial substrate (nondegraded control).
- Set a heat block to the appropriate temperature for enzyme activity (37 °C, with a final time point of 30 min was optimal for DmWRNexo though this may need to be determined empirically by comparing activity of the chosen enzyme at a range of temperatures and times). Potential enzyme inhibitors commonly present in buffers should be carefully tested for (e.g. EDTA inhibits DmWRNexo, as does excess ATP14, presumably by chelation of the Mg2+ critical for nuclease activity). Note that even high concentrations of imidazole (>300 mM) have no impact on either DmWRNexo enzyme activity nor does it quench fluorescence.
- Make up STOP solution (80% Formamide, 1x TBE). This is at 3x. This may contain dyes such as bromophenol blue if required. Store at RT.
- Set up the following near the heat block: the final reagent(s) to be added to the reaction mix (usually enzyme); a 1-10 μl pipettor and tips, a timer, cryorack or ice bucket, and STOP solution.
- Start the reaction.
- Set the timer going on addition of enzyme to the first tube. Place the tube in the heat block.
- Continue adding enzyme to the rest of the tubes at appropriate intervals. (15 sec is ample for simple reactions that only require STOP solution to complete). Place in heat block as soon as enzyme has been added.
- Stop the reaction.
- As the timer reaches the correct time (for example 15 min) add 10 μl STOP solution to the first sample and mix by pipetting. Place immediately on ice/in the cryorack.
- Continue to stop the reactions at the appropriate time intervals and in the same order as they were set up so that each reaction has had exactly the same incubation time.
- The samples are now ready to load on the prepared denaturing gel.
- Load half (or all) of each sample using gel-loading tips for accurate pipetting. At the levels of oligonucleotide used in the reaction, half the reaction will give good output for visualization and allows for a repeat to be run (e.g. if the initial gel does not run correctly).
- It is convenient to load 1x STOP solution containing dye (e.g. bromophenol blue) in one or more lanes to ascertain how the gel is running.
- Fluorescent oligonucleotides of known sizes may be run alongside the samples as a size marker.
- Run the gel.
- Add the top running buffer chamber to the gel apparatus and fill with running buffer (1x TBE), making sure the samples are not disturbed. Check for buffer leakage.
- Connect the gel rig to a power supply and run at 200 V (constant voltage) for 150-200 min or until the bromophenol blue dye (if using) is around two-thirds of the way down the gel (it runs around the level of 10nt DNA in this percentage gel). Monitor for buffer leakage, which can skew or prevent separation.
- Stop the gel and remove from the running apparatus.
5. Gel Analysis and Imaging
- Determine the sensitivity of the Imager to be used, e.g. the Fuji FLA-3000 PhosphorImager/Fluorescence Imager can detect fluorescence of fluorescein using a blue 473 nm laser, using its greatest sensitivity setting of F1000 and the smallest pixel read (resolution) of 50 μm.
- Scan the gel.
- Remove the spacers and one gel plate carefully. Rinse the gel remaining on the other gel plate thoroughly (but carefully) to remove traces of precipitated urea that will otherwise interfere with scanning.
- Set the plate (with the gel still attached) gel-down onto the glass face of the gel-holder and place it into the Imager. Note the coordinates of the gel on the glass (if it does not fill the whole surface).
- Open the appropriate software on the connected computer and scan the noted region of the scanning area where the gel is. Use the best conditions determined as in step 5.1. The resulting file will be relatively large (several MB) but the image can be cropped and compressed if needed.
- Save, then export as a file that can be read by analysis software e.g. export as a .tiff for analysis using ImageJ.
- Analyze the data.
- A successful exonuclease assay will cause degradation of the substrate sequentially along the oligonucleotide towards the fluorophore, resulting in a characteristic ladder-like pattern of fluorescent DNA fragments on the gel. Species with higher mobility on the gel (which run further) are smaller than the nondegraded starting substrate and therefore represent products of exonuclease degradation.
- Use the 'Gels' tool in ImageJ to quantify the relative intensities of species in each lane dependent upon their mobility, in order to determine the levels of degradation hence exonuclease activity in each sample.
Representative Results
Performing in vitro analysis of exonuclease activity requires a number of preparatory steps in addition to the actual analysis. An overview of the procedures is shown in Figure 1.
Prior to conducting fluorescence-based exonuclease assays, it is critical to optimize detection of the fluorescently labeled oligonucleotide substrate following separation on urea-acrylamide gels using a suitable fluorescence imaging system. Filter choice is extremely important as this can have a marked impact on the sensitivity of the overall assay – our oligonucleotide substrate is fluorescein-labeled so requires a filter that permits excitation at 470 nm and emission at 520 nm. Using this filter, it is possible to increase the ability to detect low concentrations of substrate (or product) by adjusting both the sensitivity and resolution of the imager (Figures 2A and 2B). Note there is some tolerance in terms of filter bandwidth, as a suboptimal filter choice (ex 520/em 580 nm) still allows partial detection of the substrate, though with much lower sensitivity (Figure 2C). We suggest using a combination of labeled oligonucleotide concentration and imager settings that allow robust detection of substrate (Figure 2B and dark grey bars on right panel of Figure 2) so that degradation products, which are present at lower amounts in each band as the substrate is sequentially fragmented, can be reliably detected.
A successful exonuclease assay will cause degradation of the substrate sequentially towards the fluorophore (according to the polarity of the exonuclease being tested, this can be either a 5' or 3' end label on one strand of a duplex), resulting in a characteristic ladder-like pattern of DNA on the gel representative of exonuclease degradation (e.g. a time course of degradation is shown in Figure 3). Single stranded oligonucleotides can also be used but note that nucleases may show a length requirement for cleavage of ss DNA15,17 so false negatives may result from using too short an oligonucleotide. The kinetics of the enzyme activity can be approximated by performing densitometry of the gel images of a time course experiment and quantifying amount of nondegraded substrate remaining compared with amount present in smaller fragments (i.e. products of degradation – see Figure 2 of Mason et al.14 for an example). One simple method is to divide the gel equally into four vertical sections then obtain densitometry readings of each quarter. This avoids issues with minor variation in mobility of the substrate across the gel, though of course does not permit sensitive measurements such as cleavage of one or a few nucleotides from the substrate. It is important to normalize densitometry measurements against a region of the gel lacking any DNA (e.g. leave at least one lane empty), and to calculate degradation against an oligonucleotide-only control that is loaded for every experiment (see Mason et al.14).
Once optimized for detection, the assay can be used to detect differential activities both qualitatively and quantitatively. For example, the presence or absence of activity can be determined for nuclease mutants compared with wild type protein (Figure 4), while different substrates may also be tested for their ability to be degraded by the nuclease under investigation (Figure 4). If processivity measurements are required, it is possible to add excess unlabeled oligonucleotide at a defined time point and determine cleavage activity post-addition. A highly processive enzyme will continue to cleave the labeled substrate, while a poorly processive enzyme will show greatly diminished capacity to cleave the labeled substrate as it dissociates from substrate and reassociates with the unlabeled oligonucleotide that is in molar excess (e.g. see Mason et al.14 Figure 2).
Similarly, the intensity and migration of bands of the ladder will provide data on the activity and processivity of a particular exonuclease, either intrinsically or under differential conditions such as cation availability or temperature. As described previously14, DmWRNexo is partially processive but does not readily cleave the 5' fluorescein-labeled 50 nucleotide template to completion under the conditions used here. Single nucleotide bands are thus rarely seen with this enzyme and substrate. However, the assay can readily detect single nucleotide cleavage products, as observed when DmWRNexo (a 3'-5' exonuclease) acts on a 3' fluorescein-labeled substrate (see Figure 3 of Mason et al.14). Indeed, determination of directionality of exonuclease activity is achieved by analysis using substrates with backbones labeled either at the 5' or 3' end – a 3' exonuclease will immediately cleave the fluorophore from a 3'-labeled strand, rendering the subsequent laddering activity invisible to analysis, and resulting in only a single very-high mobility species apparent on the gel (see Figure 3 of Mason et al.14). Endonucleases that cut internally will show products at specific mobilities rather than a ladder. Similarly, altered bases that are not susceptible to nuclease cleavage will result in strong pause or stop sites that can be detected as more intense bands representing cessation of cleavage at those modified bases (e.g. See Figure 7 of Mason et al.14).
Nonoptimal results include smearing i.e. nonspecific degradation. Other issues to be aware of are gel problems that cause poor separation e.g. where the gel has not set evenly (Figure 5), bubbles are present in the gel, or the buffer leaks during the electrophoresis run. It is also important that substrates are checked before use as they might have separated or degraded, in which case they should be reannealed or discarded. Loss of protein activity through freeze/thaw cycles, or insufficient concentrations, can lead to negative results, though it is important to optimize for buffer requirements and other requirements e.g. ATP at high concentrations can inhibit DmWRNexo, presumably by titrating out Mg2+ ions15.
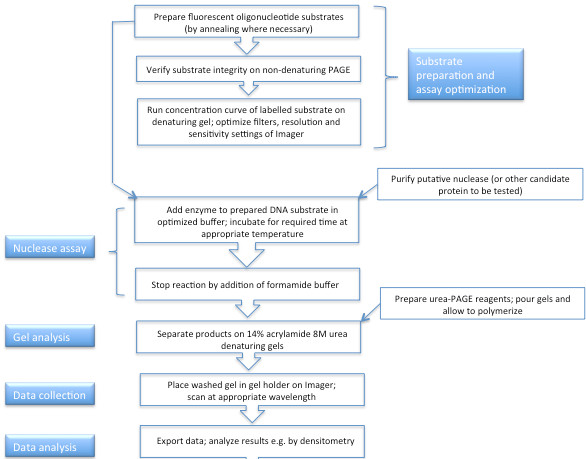
Figure 1. Flow chart of procedures undertaken when conducting a fluorescence-based exonuclease assay.
Click here to view larger image.
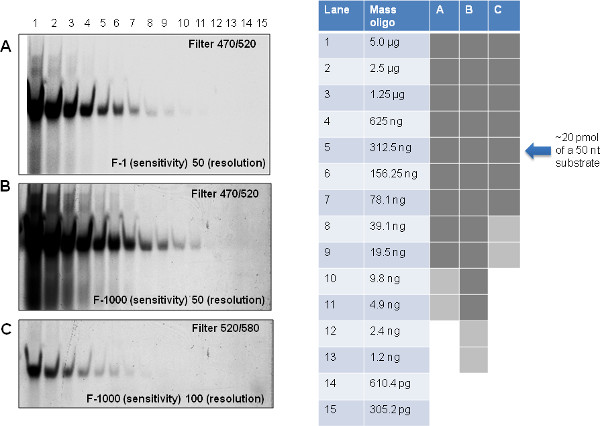
Figure 2. Optimizing detection of fluorescent substrate using the phosphorimager. A single gel of doubling dilutions of the 50-mer fluorescently labeled oligonucleotide substrate was exposed on the Phosphorimager using different filter conditions (A, B: 470 ex/520 em which is optimal for fluorescein, or C: 520 em/580 ex) and altering either sensitivity or resolution. Dark grey boxes indicate oligonucleotide concentrations that can be detected reliably with a high sensitivity and resolution; lighter grey boxes indicate those concentrations at which the signal can still be detected though the signal is much weaker. Unshaded areas of the table represent oligonucleotide concentrations that could not be detected using the filter, sensitivity and resolution settings indicated on the gel images A, B and C. Note that 20 pmol of a 50 nucleotide substrate is approximately equivalent to 312.5 ng. Click here to view larger figure.
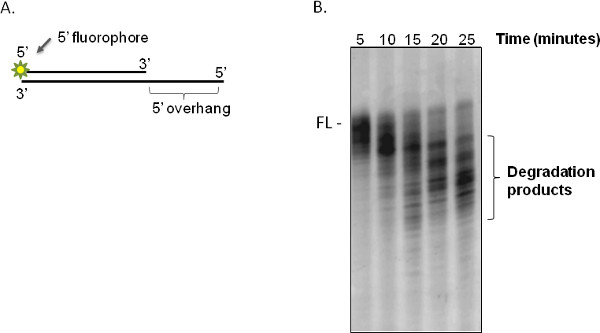
Figure 3. Typical fluorescently labeled substrate and exonuclease assay results. (A) Schematic representation of a duplex oligonucleotide substrate (5'OV) with 5' overhang on the unlabeled guide strand and 5' fluorescein label on the cleaved strand, as used in these assays. (B) Time course of degradation of the 5' overhang substrate by purified DmWRNexo at 37 °C. The enzyme binds to the 5' overhang of the guide strand and sequentially cleaves the labeled strand in a 3'-5' direction, resulting in a ladder of fragments of progressively smaller sizes that run more quickly on acrylamide gels. In this experiment, 10 pmol of DmWRNexo and 20 pmol oligonucleotide substrate were used in a 20 μl reaction. ('FL' represents the position of the nondegraded duplex 5'OV duplex fluorescent substrate).
Click here to view larger image.
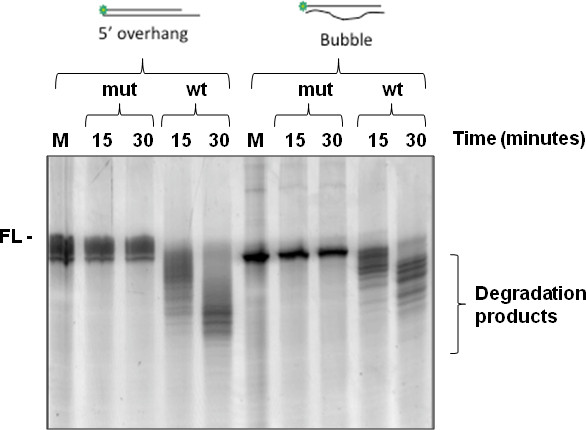
Figure 4. Testing the impact of point mutation of the exonuclease catalytic domain on ability of the enzyme DmWRNexo to cleave either duplex substrate with 5' overhang, or a bubble structure. In both cases, the substrate is labeled on one strand only at the 5' end with fluorescein. In a 20 μl reaction, 12.5 pmol purified DmWRNexo protein (wt or double point mutant D82A E84A) was incubated at 37 °C with 25 pmol oligonucleotide as shown in the schematic above the gels, and products analyzed on urea-acrylamide gels after 15 or 30 min. ('FL' represents the position of the nondegraded fluorescent substrate). 'wt' denotes wild type DmWRnexo protein, while 'mut' is a double point mutant version (D82A E84A). These results were useful in confirming the assignment of the enzyme's active site, which had been based on homology to human WRN exonuclease domain together with in silico structural modeling.
Click here to view larger image.
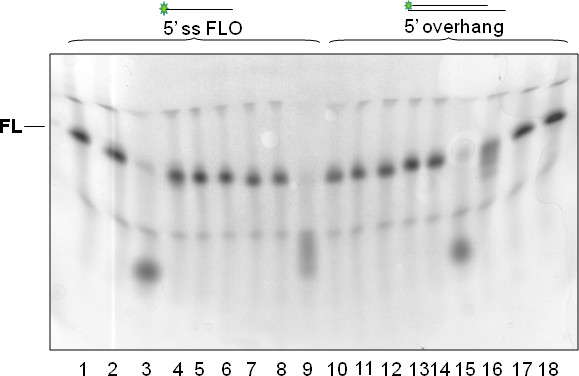
Figure 5. One example of a poor result. As a consequence of uneven polymerization of the gel, the fluorescently labeled substrate does not run at the same apparent mobility in all lanes. This can lead to incorrect interpretation of cleavage data, even though it is apparent that the substrates in lanes 3, 9 and 15 have been cleaved almost to completion, with partial cleavage in lane 16. Note that some lanes appear compressed in width in addition to the aberrant mobility leading to the 'smile' detected. Using identical conditions for polymerization (e.g. fresh APS, suitable concentrations of TEMED and a stable and consistent room temperature) can help to avoid such problems. 'FL' represents the position of the nondegraded fluorescent substrate.
Click here to view larger image.
Discussion
Determination of exonuclease activity of purified proteins requires the analysis of DNA cleavage products. Sequential cleavage of DNA by exonucleases can be visualized by separation of labeled cleavage products on acrylamide gels. Historically this involved end-labeling of the DNA substrate with a radiolabel (e.g. 32P or 35S), but with the disadvantages inherent in use of radiolabel (cost, safety issues, and instability over time). To overcome these problems, we have developed an exonuclease assay that employs fluorescently labeled DNA substrates, and show the utility of the assay system in determining nuclease activity of the WRN exonuclease orthologue from Drosophila melanogaster, DmWRNexo10. The key benefit of our assay is that the fluorescently labeled substrates are stable over time thus providing greater reproducibility and standardization e.g. when comparing different batches of purified recombinant exonuclease. We have used the assay to undertake detailed analysis of DmWRNexo nuclease activity, permitting determination of nuclease polarity, processivity, and substrate and cation preferences (see also13,14).
Whilst this assay was developed specifically to determine the activity of DmWRNexo and related orthologous 3'-5' DNA exonucleases, it is equally applicable to nucleases of opposite polarity (e.g. we have tested it on Lambda exonuclease14). Moreover, the substrates are suitable for analysis of cleavage sites by endonucleases. Excitingly, since the system employs a fluorescently labeled substrate, it should also be possible to conduct protein-DNA interaction studies +/- nuclease activity (e.g. in the presence of nuclease inhibitors such as EDTA, or comparing DNA binding between wild type and mutant proteins to explore the basis of lack of activity of nuclease-dead mutant forms), either in a band shift format or quantitatively using fluorescence polarization assays in multiwell plates – i.e. there is adaptability for high throughput screening, especially using plate readers that include injectors for addition of inhibitors/activators. Extension of such studies to helicases would be possible by including a quenching agent on the strand complementary to the fluorescently labeled strand: helicase-dependent unwinding of the substrate would result in release of the fluorescent signal (an excess of unlabeled complementary strand may be necessary to prevent reannealing of the original duplex substrate). While DmWRNexo is a relatively small protein (~50 kDa) and is readily expressed as a soluble protein in E. coli, larger proteins (including many helicases) may need to be expressed in eukaryotic systems such as from baculovirus vectors in insect Sf9 cells, or even in human cells (e.g. human embryonic kidney 293 cells). Such systems are more efficient than E. coli for expression of larger proteins; they also permit post-translational modification, which may be critical for activity of some enzymes.
As for any biochemical assay, there are limitations as to the sensitivity of this nuclease assay, both in terms of the fluorophore used and the detection system. To obtain a fluorescent signal the fluorophore must be excited, and while the sensitivity can be modulated to some extent by changing the setting on the phosphorimager (e.g. Figure 2), it is not possible to enhance a signal by increasing time of exposure, as is the case with X-ray film exposed to radiolabel. Since exonucleases may be inhibited by large moieties bound to the end of the DNA substrate18, care should be taken to choose a small fluorophore to minimize inhibition or modulation the enzyme being assayed. It is also important to test cleavage of substrates labeled at either end (5' or 3') and to quantify all data using ImageJ or similar to assess how much substrate is cleaved. The concentrations of both substrate and enzyme are higher in the fluorescence assay described here than in published radioactivity-based assays; however, the relative proportions of DNA to nuclease are the same, validating our assay. Given increasing global problems in sourcing radiolabel, together with safety issues in its use and disposal, we believe that the relative safety, ease of use and stability of the fluorescent substrate outweigh the minor point of using more protein and DNA: concentrations are still low at ≤20 pmol substrate and 1-20 pmol enzyme. There is also the added benefit that there is no need to regularly label and gel-purify substrate as labeling takes place at the time of synthesis and is stable over time.
Alternative methods to the fluorescence based assay described here include use of radiolabeled substrates (as discussed above e.g.4,8,15,18) and a relatively recent addition to the range of assays available, based on nonradioactive isotope dilution mass spectrometry (LC-MS/MS) analysis of the final single nucleotide product of cleavage16. However, this latter procedure requires access to highly specialized equipment, and measures only the end-point of the nuclease reaction. We believe that the use of fluorescently labeled DNA substrates permits reproducible analysis of, and comparison between, exonucleases from different sources and between different batches of the same nuclease, permitting robust biological replication of experiments. The stability of the labeled substrate together with increasingly sophisticated methods of detecting fluorescence (e.g. in fluorescence polarization-based interaction studies) should allow this assay to be developed in the future for moderate to high throughput screening, particularly for inhibitors of nucleases.
Divulgations
The authors have nothing to disclose.
Acknowledgements
We thank the cross-council New Dynamics of Ageing Programme for funding this work [ES/G037086/1] and Prof Dave Sherratt (Department of Biochemistry, University of Oxford) for access to the Fuji FLA-3000.
Materials
| Reagent/Material | |||
| Custom oligonucleotides | Eurogentec | It is necessary to obtain these at high purity e.g. with PAGE purification. | |
| 5'FLO | fluoresescein-5' GAACTATGGCTCTC GAGTGCTAGGACATGTCTGA CTACGTACAAGTCACC – 3' |
||
| bubble | 5'- GGTGACTTGTACGT AGTCAGACATGTCCTAGCAC TCGAGAGCCATAGTTC-3' |
||
| 40% 19:1 Acrylamide solution | Severn Biotech | 20-2400-05 | CAUTION: potent neurotoxin so gloves should be worn at all times |
| His-Trap columns (1 ml) | GE Healthcare | 17-5247-01 | |
| All other reagents | any reputable supplier | Molecular biology grade is necessary (DNase-free); microfuge tubes similarly should be DNase- and RNase-free | |
| Equipment | |||
| Hoefer SE400 gel apparatus | Hoefer | SE400-15-1.5 | |
| FLA-3000 (phosphor and fluorescence imager) | Fuji | ||
| Image Reader V2.02 | FujiFilm | ||
| Image Gauge V3.3 | FujiFilm |
References
- Mason, P. A., Cox, L. S. The role of DNA exonucleases in protecting genome stability and their impact on ageing. Age (Dordr. 34, 1317-1340 (2012).
- Payne, M., Hickson, I. D. Genomic instability and cancer: lessons from analysis of Bloom’s syndrome). Biochem. Soc. Trans. 37, 553-559 (2009).
- Yu, C. E., Oshima, J., Fu, Y. H., Wijsman, E. M., Hisama, F., Alisch, R., Matthews, S., Nakura, J., Miki, T., Ouais, S., et al. Positional cloning of the Werner’s syndrome gene. Science. 272, 258-262 (1996).
- Huang, S., Li, B., Gray, M. D., Oshima, J., Mian, I. S., Campisi, J. The premature ageing syndrome protein, WRN, is a 3′–>5′ exonuclease. Nat. Genet. 20, 114-116 (1998).
- Goto, M. Syndrome-causing mutations in Werner syndrome. Biosci. Trends. 2, 147-150 (2008).
- Perry, J. J., Yannone, S. M., Holden, L. G., Hitomi, C., Asaithamby, A., Han, S., Cooper, P. K., Chen, D. J., Tainer, J. A. WRN exonuclease structure and molecular mechanism imply an editing role in DNA end processing. Nat. Struct. Mol. Biol. 13, 414-422 (2006).
- Opresko, P. L., Laine, J. P., Brosh, R. M., Seidman, M. M., Bohr, V. A. Coordinate action of the helicase and 3′ to 5′ exonuclease of Werner syndrome protein. J. Biol. Chem. 276, 44677-44687 (2001).
- Plchova, H., Hartung, F., Puchta, H. Biochemical characterization of an exonuclease from Arabidopsis thaliana reveals similarities to the DNA exonuclease of the human Werner syndrome protein. J. Biol. Chem. 278, 44128-44138 (2003).
- Hartung, F., Plchova, H., Puchta, H. Molecular characterisation of RecQ homologues in Arabidopsis thaliana. Nucleic Acids Res. 28, 4275-4282 (2000).
- Saunders, R. D., Boubriak, I., Clancy, D. J., Cox, L. S. Identification and characterization of a Drosophila ortholog of WRN exonuclease that is required to maintain genome integrity. Aging Cell. 7, 418-425 (2008).
- Cox, L. S., Boubriak, I. DNA Instability in Premature Aging, in DNA Damage Repair, Repair Mechanisms and Aging. Thomas, A.E., Eds. Nova Science Publishers. , 1-34 (2010).
- Cox, L. S., Clancy, D. J., Boubriak, I., Saunders, R. D. Modeling Werner Syndrome in Drosophila melanogaster: hyper-recombination in flies lacking WRN-like exonuclease. Ann. N.Y. Acad. Sci. 1119, 274-288 (2007).
- Boubriak, I., Mason, P. A., Clancy, D. J., Dockray, J., Saunders, R. D., Cox, L. S. DmWRNexo is a 3′-5′ exonuclease: phenotypic and biochemical characterization of mutants of the Drosophila orthologue of human WRN exonuclease. Biogerontology. 10, 267-277 (2009).
- Mason, P. A., Boubriak, I., Robbins, T., Lasala, R., Saunders, R., Cox, L. S. The Drosophila orthologue of progeroid human WRN exonuclease, DmWRNexo, cleaves replication substrates but is inhibited by uracil or abasic sites : Analysis of DmWRNexo activity in vitro. Age (Dordr). , (2012).
- Machwe, A., Xiao, L., Orren, D. K. Length-dependent degradation of single-stranded 3′ ends by the Werner syndrome protein (WRN): implications for spatial orientation and coordinated 3′ to 5′ movement of its ATPase/helicase and exonuclease domains. BMC Mol. Biol. 7, 6 (2006).
- Mangerich, A., Veith, S., Popp, O., Fahrer, J., Martello, R., Bohr, V. A., Burkle, A. Quantitative analysis of WRN exonuclease activity by isotope dilution mass spectrometry. Mech. Ageing Dev. 133, 575-579 (2012).
- Xue, Y., Ratcliff, G. C., Wang, H., Davis-Searles, P. R., Gray, M. D., Erie, D. A., Redinbo, M. R. A minimal exonuclease domain of WRN forms a hexamer on DNA and possesses both 3′- 5′ exonuclease and 5′-protruding strand endonuclease activities. Biochimie. 41, 2901-2912 (2002).
- Machwe, A., Ganunis, R., Bohr, V. A., Orren, D. K. Selective blockage of the 3′–>5′ exonuclease activity of WRN protein by certain oxidative modifications and bulky lesions in DNA. Nucleic Acids Res. 28, 2762-2770 (2000).

