A Flow Cytometry Assay to Identify Immune Cell Subsets in Peripheral Blood Mononuclear Cells
Abstract
Source: Torchia, M. L. G., et al. Discrimination of Seven Immune Cell Subsets by Two-fluorochrome Flow Cytometry. J. Vis. Exp. (2019)
This video demonstrates a flow cytometry-based technique using two fluorochromes to identify CD4+ and CD8+ T cells, γδ T cells, B cells, natural killer (NK) cells, and monocytes in human peripheral blood mononuclear cells (PBMCs). Upon incubating the cells with fluorochrome-conjugated antibodies against the surface markers, the cell subsets are identified using an optimal flow cytometric gating strategy.
Protocol
All procedures involving sample collection have been performed in accordance with the institute's IRB guidelines.
NOTE: This protocol has been tested on freshly or frozen isolated peripheral blood cells and whole blood.
1. Cell Staining
NOTE: Choosing pairs of fluorochromes with virtually no spectral overlap is important to reduce the spread of data due to the high spillover of a fluorochrome in the other fluorochrome detector. To achieve an optimal identification of all the cell subsets, fluorochromes with a high quantum yield should be used such as antibody pairs PE-BV421 and PE-APC.
- Staining of fresh and frozen PBMC
- Transfer 100 µL of PBMC (1 x 106 cells) to a 96-well V-bottom plate.
NOTE: Any number of cells lower than 1 x 106 can be used with similar results. - Centrifuge at 350 x g for 3 min at RT and carefully aspirate the supernatant without disturbing the cell pellet. Add to each well 100 µL of phosphate-buffered saline (PBS) containing a live/dead fixable dye that reacts with free amine on proteins for 10 min to label dead cells.
- Prepare for each sample 30 µL of a mix containing all the antibodies (anti-CD3. -CD56, TCRγδ in fluorochrome A, and anti-CD4, CD8, CD19, CD14 in fluorochrome B). The concentrations of antibodies are indicated in Table 1 and Table 2. At this stage, titrated antibodies against different target molecules and in different fluorochromes can be added as well (e.g., Table 3).
NOTE: Antibodies concentration can vary depending on the manufacturer and lot number. Therefore, preliminary tests should be done to achieve the optimal signal. PBS, PBS/0.5% bovine serum albumin (BSA), PBS/0.2% sodium azide or PBS/0.5% BSA/0.1% sodium azide were used with similar results to dilute antibodies. Cells can also be stained in volumes different from 30 µL to easily integrate this methodology into already existing staining protocols. - Centrifuge at 350 x g for 3 min at RT and carefully aspirate the supernatant without disturbing the cell pellet. Add the antibody cocktail to each well and resuspend carefully without generating bubbles. Incubate for 30 min at RT in the dark.
NOTE: It is possible to stain the samples at 4 °C with similar results. - Add 150 µL of staining buffer and centrifuge at 350 x g for 3 min at RT and carefully aspirate the supernatant without disturbing the cell pellet. Resuspend the cells in 200 µL of PBS and acquire data on a flow cytometer. If staining volumes are changed, please ensure at least a 20-fold dilution of the original antibody mix used to wash the excess antibodies.
NOTE: Stained cells can be fixed within PBS/2% paraformaldehyde, kept in a refrigerator at 4 °C overnight, and then acquired on a flow cytometer the following day.
CAUTION: Paraformaldehyde is harmful if swallowed and can cause skin irritation
- Transfer 100 µL of PBMC (1 x 106 cells) to a 96-well V-bottom plate.
2. Antibody Titration
NOTE: Antibody titration is the most critical step for obtaining high-quality, reproducible data. Titration of anti-CD3, -CD8, -CD14, -CD19, and -TCR γδ follows the standard procedure by which the concentration of antibody to optimally separate positive and negative peaks is derived by maximum staining index. Dilutions at the peak or closer to the peak on the rising side of the stain index curve should be selected (Figure 1A-C). The anti-CD4 antibody is titrated to place the peak of the CD4 positive population between CD3 single positive populations and CD3+/CD8+ T cells, closer to the CD3 single-positive signal to better discriminate the CD8dim populations (CD8+ γδ T cells and NK T cells). Along the same line, CD56 titration aims to position NK CD56+ cells between the CD3+ and the CD3– population.
- Maximum stain index curve
NOTE: Titration of anti-CD3, -CD8, -CD14, -CD19, and -TCR γδ follows the standard procedure by which the concentration of antibody to optimally separate positive and negative peaks is derived by a maximum staining index curve. If antibodies against other markers are added to the panel, they also need to be titrated with a maximum staining index curve.- Prepare a 2-fold antibody dilution by filling 10 wells of a 96-well plate with 40 µL of staining buffer. In the first well, increase the final volume to 80 µL of staining buffer and add the antibody of interest at a concentration 4 times the concentration suggested by the manufacturer.
- Mix well and transfer 40 µL to the second well. Mix well and repeat this step for all the other wells.
- Stain 10 samples of PBMC or whole blood with 30 µL of the 10 different 2-fold dilutions of antibodies following the protocol described before.
- Acquire data with a flow cytometer and plot the signal from each dilution (Figure 1A).
- Gate on the negative and positive populations for each antibody concentration. Increasing the concentration of antibodies can lead to a higher background. Therefore, resize the negative gate accordingly.
- For each antibody concentration, extract information about the median and standard deviation for the fluorescent intensity of the negative population, and the median for the fluorescent intensity of the positive population. Calculate for each antibody concentration the stain index with this formula: (median fluorescent intensity of the positive population — median fluorescent intensity of the negative population) ÷ (2 x standard deviation of the fluorescent intensity of the negative population) (Figure 1B).
- Plot the stain index vs. the antibody concentration expressed as a fraction of the antibody dilution (e.g., 1:10 dilution = 0.1), and identify the concentration of the antibody with the maximum stain index value (Figure 1C).
- Anti-CD4 and -CD56 antibody titration
NOTE: Anti-CD4 and -CD56 antibody titration relies on the previous titration of the other markers in the two-fluorochrome panel. For the anti-CD4 antibody, the titration aims at placing the anti-CD4 signal between the double CD8+/CD3+ signal and the CD3 single positive population (Figure 1D).- Titrate the anti-CD4 and CD56 antibodies with a 2-fold dilution strategy as described before, adding additional concentrations in between to finely identify the range of concentration that allows to separate CD4+ T cells and NK cells from the other cell populations.
- Titrate the anti-CD4 antibody by placing the anti-CD4 signal between the double CD8+/CD3+ signal and the CD3 single positive population (Figure 1D).
NOTE: Special care should be done to clearly separate CD4+ T cells from CD8+ dim populations. - Titrate the anti-CD56 antibody following a strategy similar to the anti-CD4 antibody titration, by placing NK cells between the CD3-negative and the CD3-positive populations.
3. Gating Strategy
- Identify lymphocytic and monocytic cell populations and remove dead cells and most of the residual red blood cells from the analysis.
- Select the entire population containing lymphocytes and monocytes based on forward vs side scatter area (FSC-A vs SSC-A). Remove cell aggregates from the analysis via forward scatter height vs forward scatter width (FSC-H vs FSC-W) and side scatter height vs side scatter width (SSC-H vs SSC-W).
- Use a live/dead discrimination marker to exclude bright positive dead cells and residual red blood cells from the analysis. Gate on lymphocytes and monocytes based on the different FSC-A and SSC-A profiles.
- Two-fluorochrome seven-marker gating strategy of the lymphocytic populations.
NOTE: Within the CD3 positive subgroup, CD4+, CD8+, and γδ T cells can be separated using antibodies that solely target CD4, CD8, and the γδ receptor. In a comparable way, within the CD3 negative subgroup, B cells, NK cells, and monocytes can be uniquely identified using antibodies against CD19, CD56, and CD14, respectively.- Select the lymphocyte gate and create a dot plot with on each axis one of two fluorochromes used in this protocol (Figure 2B).
- Gate on CD8+ T cells identified as CD3+/CD8+ double positive cells at the top right corner of the dot-plot (Figure 2B). Exclude the dim CD8 population which might contain NKT cells. Gate on CD4+ T cells identified as population in between CD8+ T cells and the CD3 single positive populations. Gate on γδ T cells identified as high CD3 cells. Subdivide γδ T cells in CD8 positive and CD8 negative.
- Gate on NK cells identified as the population in between CD3-positive and CD3-negative cells. Subdivide NK cells in CD8 positive and CD8 negative. Gate on B cells identified as CD3-negative CD19+ population on the right lower corner of the dot plot.
- Select the monocyte gates and create a dot plot with on each axis one of two fluorochromes used in this protocol (Figure 2C). Gate on the CD3–/CD14+ population.
Table 1: Antibody panel used for the two-fluorochrome immune-cell staining of PBMC (BV421-PE combination).
| Target | Clone | Fluorochrome | Vendor | Concentration | Purpose |
| CD3 | UCHT1 | BV421 | BD | 1/20 | Lineage |
| CD56 | NCAM16.2 | BV421 | BD | 1/900 | |
| TCRγδ | B1 | BV421 | Bio | 1/30 | |
| CD4 | RPA-T4 | PE | BD | 1/450 | |
| CD8 | RPA-T8 | PE | BD | 1/20 | |
| CD14 | M5E2 | PE | BD | 1/15 | |
| CD19 | HIB19 | PE | BD | 1/300 | |
| Dead cells | L/D Blue | LT | 1/300 | Live/Dead discrimination | |
| BD = BD Biosciences, Bio = BioLegend, LT = Life Technologies | |||||
Table 2: Antibody panel used for the two-fluorochrome immune-cell staining of PBMC (APC-PE combination).
| Target | Clone | Fluorochrome | Vendor | Concentration | Purpose |
| CD3 | UCHT1 | APC | BD | 1/20 | Lineage |
| CD56 | NCAM16.2 | APC | BD | 1/60 | |
| TCRγδ | B1 | APC | Bio | 1/30 | |
| CD4 | RPA-T4 | PE | BD | 1/450 | |
| CD8 | RPA-T8 | PE | BD | 1/20 | |
| CD14 | M5E2 | PE | BD | 1/15 | |
| CD19 | HIB19 | PE | BD | 1/300 | |
| Dead cells | L/D Blue | LT | 1/300 | Live/Dead discrimination | |
| BD = BD Biosciences, Bio = BioLegend, LT = Life Technologies | |||||
Table 3: Antibody panel used to stain frozen PBMC from a patient with multiple myeloma.
| Target | Clone | Fluorochrome | Catalog | Vendor | Concentration | Purpose |
| CD3 | UCHT1 | BV421 | 562426 | BD | 1/20 | Lineage |
| CD56 | NCAM16.2 | BV421 | 562751 | BD | 1/900 | |
| TCRγδ | B1 | BV421 | 331217 | Bio | 1/30 | |
| CD4 | RPA-T4 | PE | 555347 | BD | 1/450 | |
| CD8 | RPA-T8 | PE | 555367 | BD | 1/20 | |
| CD14 | M5E2 | PE | 555398 | BD | 1/15 | |
| CD19 | HIB19 | PE | 555413 | BD | 1/300 | |
| CCR7 | G043H7 | AF647 | 353217 | Bio | 1/30 | Differentiation |
| CD45RA | HI100 | APC-H7 | 560674 | BD | 1/60 | |
| CCR4 | 1G1 | PE-Cy7 | 561034 | BD | 1/60 | Th subsets |
| CCR6 | G034-E3 | BV605 | 353419 | Bio | 1/30 | |
| CXCR3 | 1C6/CXCR3 | AF488 | 561730 | BD | 1/30 | |
| CD57 | NK-1 | PE-CF594 | 562488 | BD | 1/900 | Activation/Exhaustion |
| HLA-DR | G46-6 | BV510 | 563083 | BD | 1/30 | |
| CD16 | 3G8 | BUV395 | 563784 | BD | 1/30 | NK, Monocyte activation |
| Dead cells | L/D Blue | L-23105 | LT | 1/300 | Live/Dead discrimination | |
Representative Results
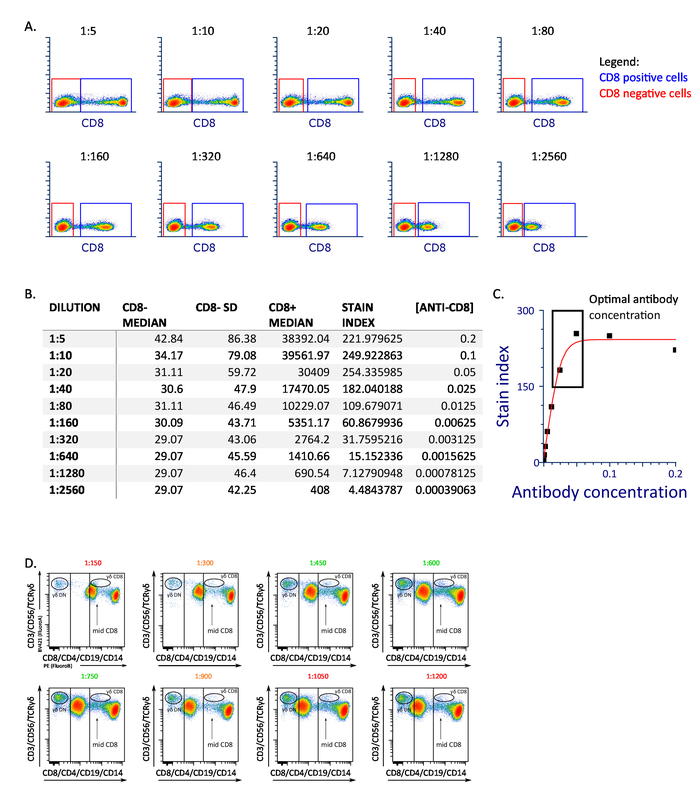
Figure 1: Representative antibody titration. (A) The dot plot shows CD8 expression on fresh PBMC stained with the indicated concentration of the antibody. (B) The table represents the median and standard deviation of fluorescent intensity of the CD8+, median fluorescent intensity CD8- population, and the derived stain index for each concentration tested. (C) The graph shows how to derivate the optimal concentration of the antibody as a function of stain index. (D) Representative titration of CD4 antibody. Panel D has been modified from Boin et al. 2017
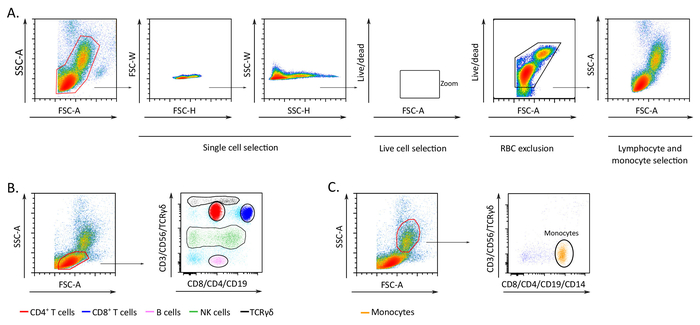
Figure 2: Representative gating strategy and results of subpopulation discrimination. (A) Schematic representation of doublet exclusion, live cell discrimination, and size-based gating of lymphocytes and monocytes. (B) Lymphocyte subpopulations identified with the two fluorochrome approach. (C) Monocytes identified with the two fluorochrome approach. The figure has been modified from Boin et al. 2017
Divulgazioni
The authors have nothing to disclose.
Materials
| CD3 | BD Biosciences | 562426 | Antibody for staining RRID: AB_11152082 |
| CD56 | BD Biosciences | 562751 | Antibody for staining RRID: AB_2732054 |
| TCRgd | BD Biosciences | 331217 | Antibody for staining RRID: AB_2562316 |
| CD4 | BD Biosciences | 555347 | Antibody for staining RRID: AB_395752 |
| CD8 | BD Biosciences | 555367 | Antibody for staining RRID: AB_395770 |
| CD14 | BD Biosciences | 555398 | Antibody for staining RRID: AB_395799 |
| CD19 | BD Biosciences | 555413 | Antibody for staining RRID: AB_395813 |
| CD3 | BD Biosciences | 555335 | Antibody for staining RRID: AB_398591 |
| CD56 | BD Biosciences | 555518 | Antibody for staining RRID: AB_398601 |
| TCRgd | BD Biosciences | 331211 | Antibody for staining RRID: AB_1089215 |
| CCR7 | Biolegend | 353217 | Antibody for staining RRID: AB_10913812 |
| CD45RA | BD Biosciences | 560674 | Antibody for staining RRID: AB_1727497 |
| CCR4 | BD Biosciences | 561034 | Antibody for staining RRID: AB_10563066 |
| CCR6 | BD Biosciences | 353419 | Antibody for staining RRID: AB_11124539 |
| CXCR3 | BD Biosciences | 561730 | Antibody for staining RRID: AB_10894207 |
| CD57 | BD Biosciences | 562488 | Antibody for staining RRID: AB_2737625 |
| HLA-DR | BD Biosciences | 563083 | Antibody for staining RRID: AB_2737994 |
| CD16 | BD Biosciences | 563784 | Antibody for staining RRID: AB_2744293 |
| Dead cells | Life technologies | L-23105 | Live/dead discrimination |
| 96-well V-bottom plate | Thermo fisher | 249570 | plate for staining |
| FACSAria IIu Cell Sorter | BD Biosciences | Flow cytometer | |
| FCS Express 6 | De Novo Software | FACS analysis | |
| Graphpad Prism | GraphPad software | Data analysis |

