Remote Magnetic Actuation of Micrometric Probes for in situ 3D Mapping of Bacterial Biofilm Physical Properties
Summary
This paper shows an original methodology based on the remote actuation of magnetic particles seeded in a bacterial biofilm and the development of dedicated magnetic tweezers to measure in situ the local mechanical properties of the complex living material built by micro-organisms at interfaces.
Abstract
Bacterial adhesion and growth on interfaces lead to the formation of three-dimensional heterogeneous structures so-called biofilms. The cells dwelling in these structures are held together by physical interactions mediated by a network of extracellular polymeric substances. Bacterial biofilms impact many human activities and the understanding of their properties is crucial for a better control of their development — maintenance or eradication — depending on their adverse or beneficial outcome. This paper describes a novel methodology aiming to measure in situ the local physical properties of the biofilm that had been, until now, examined only from a macroscopic and homogeneous material perspective. The experiment described here involves introducing magnetic particles into a growing biofilm to seed local probes that can be remotely actuated without disturbing the structural properties of the biofilm. Dedicated magnetic tweezers were developed to exert a defined force on each particle embedded in the biofilm. The setup is mounted on the stage of a microscope to enable the recording of time-lapse images of the particle-pulling period. The particle trajectories are then extracted from the pulling sequence and the local viscoelastic parameters are derived from each particle displacement curve, thereby providing the 3D-spatial distribution of the parameters. Gaining insights into the biofilm mechanical profile is essential from an engineer's point of view for biofilm control purposes but also from a fundamental perspective to clarify the relationship between the architectural properties and the specific biology of these structures.
Introduction
Bacterial biofilms are communities of bacteria associated with biological or artificial surfaces1-3. They form by an adhesion-growth mechanism coupled with the production of polysaccharide-rich extracellular matrix that protects and stabilizes the edifice4,5. These biofilms are not simply passive assemblages of cells stuck to surfaces, but organized and dynamic complex biological systems. When bacteria switch from planktonic to biofilm lifestyle, changes in gene expression and cell physiology are observed as well as increased resistance to antimicrobials and host immune defenses being at the origin of many persistent and chronic infections6. However, the controlled development of these living structures also offer opportunities for industrial and environmental applications, such as bioremediation of hazardous waste sites, bio-filtration of industrial water or formation of bio-barriers to protect soil and groundwater from contamination.
While molecular features specific to biofilm way of life are increasingly described, the mechanisms driving the community development and persistence remain unclear. Using the recent advances on microscale measurements using scanning electrochemical or fluorescence microscopy, these living organizations have been shown to exhibit considerable structural, chemical and biological heterogeneity7. Yet, until now, biofilm mechanics have been mainly examined macroscopically. For instance, observation of biofilm streamers deformation due to variations in fluid flow rates8,9, uniaxial compression of biofilm pieces lift from agar medium or grown on cover slides10,11, shear of biofilm collected from the environment and then transferred to a parallel plate rheometer12,13, atomic force spectroscopy using a glass bead and coated with a bacterial biofilm attached to an AFM cantilever14 or a dedicated microcantilever method for measuring the tensile strength of detached biofilm fragments15,16 have been implemented during the ten last years, providing useful information on the viscoelastic nature of the material17. However, it seems likely that information on in situ biofilm mechanical properties is lost when the material is removed from its native environment, which was often the case in these approaches. Moreover, the treatment of the biofilm as a homogeneous material misses the information on the possible heterogeneity of the physical properties within the community. Therefore, the exact implications of the structure mechanics in the biofilm formation and biological traits such as gene expression patterning or chemical gradients can hardly be recognized. To progress towards a microscale description of the biofilm physical properties, new dedicated tools are required.
This paper details an original approach conceived to achieve measurement of local mechanical parameters in situ, without disturbing the biofilm and enabling drawing of the spatial distribution of the microscale material properties and then the mechanical heterogeneity. The principle of the experiment rests on the doping of a growing biofilm with magnetic microparticles followed by their remote loading using magnetic tweezers in the mature biofilm. Particle displacement under controlled magnetic force application imaged under the microscope enables local viscoelastic parameter derivation, each particle reporting its own local environment. From these data, the 3D mechanical profile of the biofilm can be drawn, revealing spatial and environmental condition dependences. The whole experiment will be shown here on an E. coli biofilm made by a genetically engineered strain carrying a derepressed F-like plasmid. The results detailed in a recent paper18 provide a unique vision of the interior of intact biofilm mechanics.
Protocol
1. Bacteria Culture and Suspension Preparation
- Pick a freshly grown colony from a Lysogeny Broth (LB) agar plate, inoculate it in 5 ml liquid LB medium containing 100 µg/ml ampicillin and 7.5 µg/ml tetracycline and incubate it for 5 to 6 hr at 37 °C on a shaking platform.
- Then, add 100 µl of the bacterial culture in 5 ml minimum medium (M63B1) supplemented with 0.4% glucose and the same antibiotic concentrations. Incubate this freshly diluted culture overnight at 37 °C on a shaking platform.
- After 16 hr of incubation, add 100 μl the overnight culture to 5 ml M63B1, 0.4% glucose. Keep the tube at 37 °C to the shaking platform until a 0.5 OD is reached. The suspension is then ready for injection into the experiment channel for biofilm formation.
2. Magnetic Particle Preparation
- Take 10 μl magnetic particles — 2.8 µm in diameter — from the stock and wash them 3x in 190 μl minimum medium with the aid of a magnetic sample rack.
- Adjust the particle concentration to 5 x 106 beads/ml. Typically 50 μl of the washed bead solution is mixed with a further 950 µl of M63B1 with 0.4% glucose.
3. Channel Preparation and Biofilm Growth
- Channel Mounting
- Cut two square (800 µm side length) borosilicate glass capillaries 10 cm long to obtain two 8 cm long pieces.
- Glue the two capillary pieces on two glass slides — cut in half first — 2 cm apart with 1 cm overhang on either end as in Figure 1 using a fast-acting cyanoacrylate glue (so called super-glue).
- Autoclave the entire setup and the necessary tubing required for further channel connection.
- Gather all the sterile materials under a laminar flow hood: i) the mounted channel and trap 1, ii) the tubing and connectors, iii) the two bubble traps — the bubble filter commonly used to secure children's drip-feeding (trap 1) and the home-made bubble trap as a 4 cm long tube of larger diameter (trap 2), iv) clamps, v) 30 ml syringes filled with M63B1, 0.4% glucose, and vi) the waste bottle.
- Connect the whole setup with Luer Lock connecters or junctions in the following order: 50 ml M63B1 syringe controlled by the syringe pump, pediatric bubble filter, homemade bubble trap, capillary (Figure 1, panel B), and the tube to the waste bottle. Then fill the setup with sterile M63B1, 0.4% glucose, turning on the syringe pump at a rate of about 10 ml/hr, higher than the experimental rates. Carefully track and eliminate all the bubbles in the circuit.
- Flow the medium through the system for 10-15 min; concurrently mix 1 ml of the bacteria suspension at OD 0.5 from Section 1.3 with 1 ml of the washed bead solution prepared in Section 2.2.
- Attach (but don't close) clamps to the tubing at two positions: before and after the capillaries. Switch the flow off.
- Introduce the bacterial-bead mixture into the capillary after the home made bubble trap using a 1 ml syringe, taking care to hold up the tube ends to prevent air entry. Re-attach the tubing and then close the clamps.
- Repeat the same procedure for the second capillary and check all the tubes for bubbles.
- Transfer the apparatus to the microscope and allow it to stand for 15-20 min to enable the bacteria to settle and attach to the surface of the capillary. Install the capillary onto the microscope stage with the waste container at a slightly higher level. Place the syringe pump on the counter top beside the microscope. Elevate the bubble trap before the capillary slightly higher than the capillary plane to capture bubbles.
- Biofilm Growth
- Adjust the flow rate on the syringe pump and start the flow, the biofilm will now develop on the capillary surface during the required period — usually 24 or 48 hr in these experiments.
- Focus on the capillary bottom plane and start the time-lapse recording of the sample images — usually an acquisition frequency of 2 images/min will adequately report the biofilm growth. This video monitors biofilm growth post-control (see extracts in Videos 1, 2 and 3).
4. Magnetic Tweezers Installation
- Screw the magnetic tweezers on manually controlled X-Y-Z micromanipulators and screw the micromanipulators on the microscope stage to adjust the position of the tweezers relatively to the capillary. Place the tweezers as in Figure 2 to ensure the appropriate magnetic field gradient is generated in the observation zone.
- Connect the tweezers to the function generator via the power amplifier to generate a 40 sec period of time made of 24 sec zero signal and 16 sec of 4 A direct current with a trigger sent to the bright field light shutter after 20 sec for signal synchronization providing a sequence of events as in Figure 3.
Note: These two operations can be achieved at any time between capillary installation and measurement beginning. See experiment overview sketch in Figure 4.
5. Creep Curve Acquisition
- Use the xy movement control of the microscope stage to bring the edge of the left-hand magnetic pole and the left-hand edge of the capillary in the same observation field. Take the origin of the analysis referential at the intersection of x- and y-axes defined by the edge of the capillary and the edge of the pole piece, respectively (see Figure 2).
- Adjust the vertical position of the capillary using fine focus control knob of the microscope position. Typically the first examined plane is located between 4 to 7 µm above the capillary bottom. Video 1 corresponds to the xy field that has its top left-hand corner at the origin of the spatial referential.
- Trigger the 40 sec sequence of events described in section 4.2 and Figure 3 by switching on the current generator and at the same time, manually trigger the image acquisition sequence of Video 1.
- Move the capillary to the neighbor right field by a 250 µm translation of the microscope stage to the left and operate as in section 5.3 to generate Video 2 of slice 2 and so on to obtain the required videos. Typically 3 to 4 fields of 250 x 250 µm2 are collected along the x-axis before changing the plane and repeating same operations for the new plane.
6. Force Calibration
- Prepare a glycerol solution by mixing 39.8 g of glycerol with 190 µl of bi-distilled water and 10 μl of magnetic particles at a 2 x 109 particles/ml and fill up an experimental channel with this mixture and place it on the microscope stage as described for the biofilm sample.
- After installing the magnetic tweezers as indicated in section 4, apply the magnetic force as indicated in section 5.4 and make time-lapse images to extract each single particle velocity (v) and its position in the capillary to derive the calibration file. This file should contain the applied force as a function of its position in the zone of analysis of the capillary according to stokes law, F = 6πRηv (R: radius of the particle).
7. Analysis
- Use a "particle tracker" software to obtain text files with the particle positions in each frame for all the stacks of images acquired as indicated in section 5. Using the image stack acquisition frequency, calculate the particle displacement as a function of time (e.g. Figure 5 and Video 4).
- Using the force calibration file, convert the displacement curves to compliance curves (total compliance of the material — J(t)— as a function of time) according to the compliance formula:

which gives the relation between material strain and applied stress for a probe particle of radius R embedded in an incompressible, homogeneous viscoelastic medium as previously established by Schnurr and co-workers19. - Adjust the creep compliance curves to the general Burgers model for viscoelastic materials and derive the viscoelastic parameters, J0, J1, η0, η1 for each particle (Figure 6) according to the Burgers' formula:

Note: This phenomenological analysis has been previously employed for a wide range of materials including biological materials such as biofilms to interpret macroscopic rheology data20-22.
Representative Results
A typical analysis will provide the spatial distribution of the viscoelastic parameters at the micron scale on a living biofilm without disturbing its original arrangement. Typical results are shown in Figure 7 where the values of J0 — the elastic compliance — are given as a function of the z-axis along the depth and of the y-axis along a lateral dimension of the biofilm. Each point corresponds to a bead which creep curve analysis has provided a J0 value. The data revealed that local compliance varied along the depth of the biofilm over almost three orders of magnitude but also that strong lateral heterogeneity took place at all biofilm heights.
The data also gave access to the distribution of the compliance values which exhibited here a widespread and asymmetric shape (Figure 8), providing strong indications that mechanical properties of the biofilm were supported by highly cross-linked polymer gels. Indeed, analogous behavior has been previously demonstrated on concentrated and highly cross-linked actin gels23.
In addition, these in situ measurements of biofilm local properties performed under different environmental conditions such as lower flow rate enable to demonstrate the effect of such a variation on the biofilm internal organization. In Figure 9, the compliance values of a biofilm grown at 0.1 ml/hr exhibited a still highly heterogeneous mechanical profile but no higher rigidity of the deeper biofilm layer as compared to the biofilm grown at 1 ml/hr. These results demonstrated the significant impact of the external conditions on the biofilm organization.
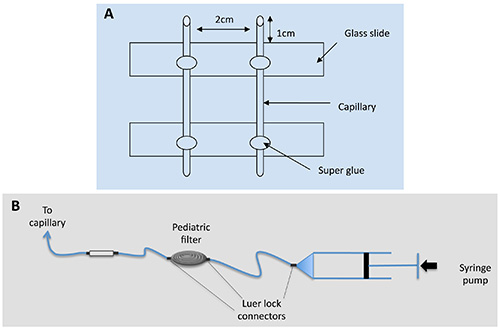
Figure 1. Capillary mounting sketch.
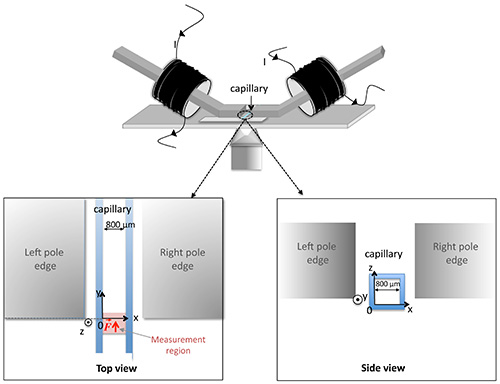
Figure 2. Positioning of the magnetic tweezers with respect to the capillary on the microscope stage and definition of the spatial referential. The top view indicates the position of the measurement zone in the capillary.
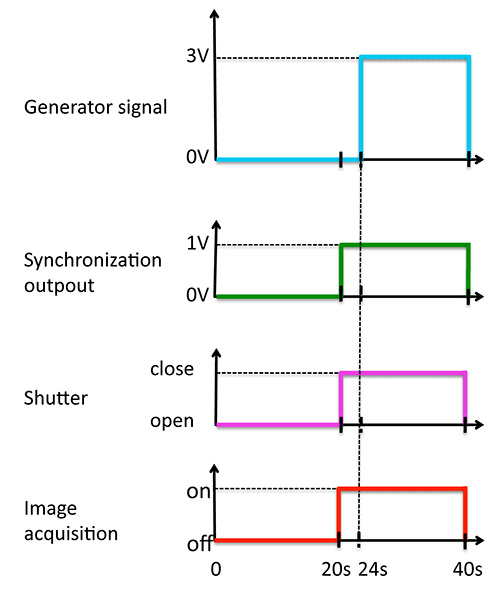
Figure 3. Synchronization of the signals over the 40 sec period of time.
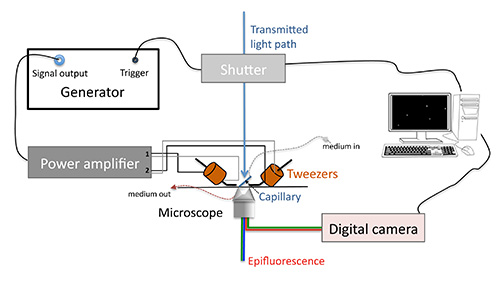
Figure 4. Experiment overview sketch.
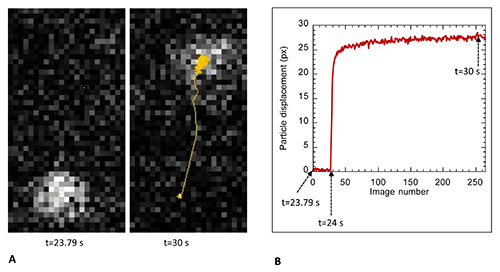
Figure 5. Extracts from particle trajectory analysis. Panel A shows on the left the image of a typical particle at its initial position just before magnetic force application (time 23.79 sec of the 40 sec period) and on the right the position of the particle at time 30 sec together with the display of the trajectory found using ImageJ particle tracker. Panel B shows the plot of the displacement curve derived from these data; here we show a trajectory extract starting at time 23.79 sec up to time t=31.29 sec of the 40 sec period.
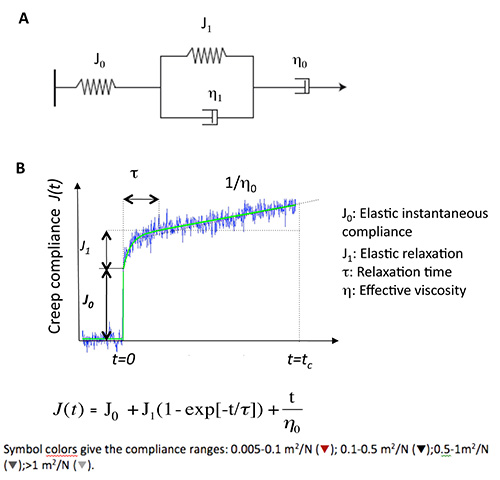
Figure 6. Viscoelastic parameter derivation on the basis of the phenomenological model of Burger. (A) Burgers mechanical model consisting in a combination of elastic Hookean spring (J0) and Newtonian dashpot (η0) in series with a Maxwell element (spring (J1) and dashpot (η1) in parallel). (B) Creep experimental trace in blue together with the curve (in green) adjusted to the Burgers equation.
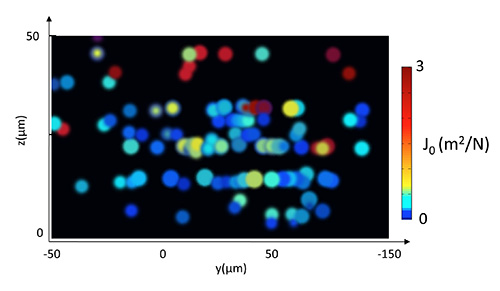
Figure 7. Spatial distribution of elastic compliance in a biofilm grown 24 hr at 37 °C under 1 ml/hr nutrients flow. Compliance values reported by each particle are represented here as a projection onto a (yz) plane using a cool to warm custom color map. Compliance values reported by particles located in the bottom layer of the biofilm are all lower than 0.2 m2/N while particles found in the upper 15 µm layer report values above 1 m2/N.
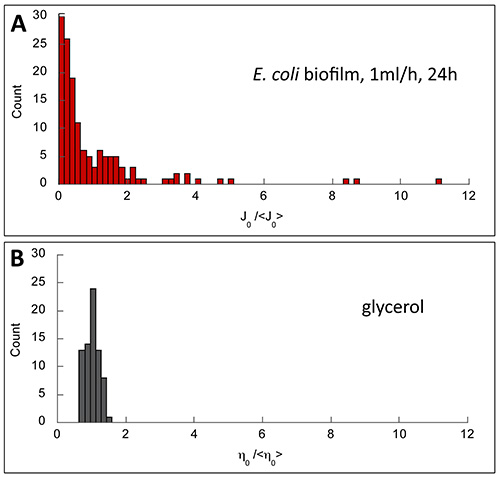
Figure 8. Normalized distributions of the viscoelastic parameters obtained in biofilm (A) and in glycerol (B), a homogeneous viscous liquid. Biofilm exhibited asymmetric and wide distribution (skewness of 3.6 and normalized variance 2.4) indicating material heterogeneity. By contrast, symmetric and narrow distribution characterizing a homogeneous medium was obtained in glycerol (skewness of 0.23 and normalized variance 0.03).
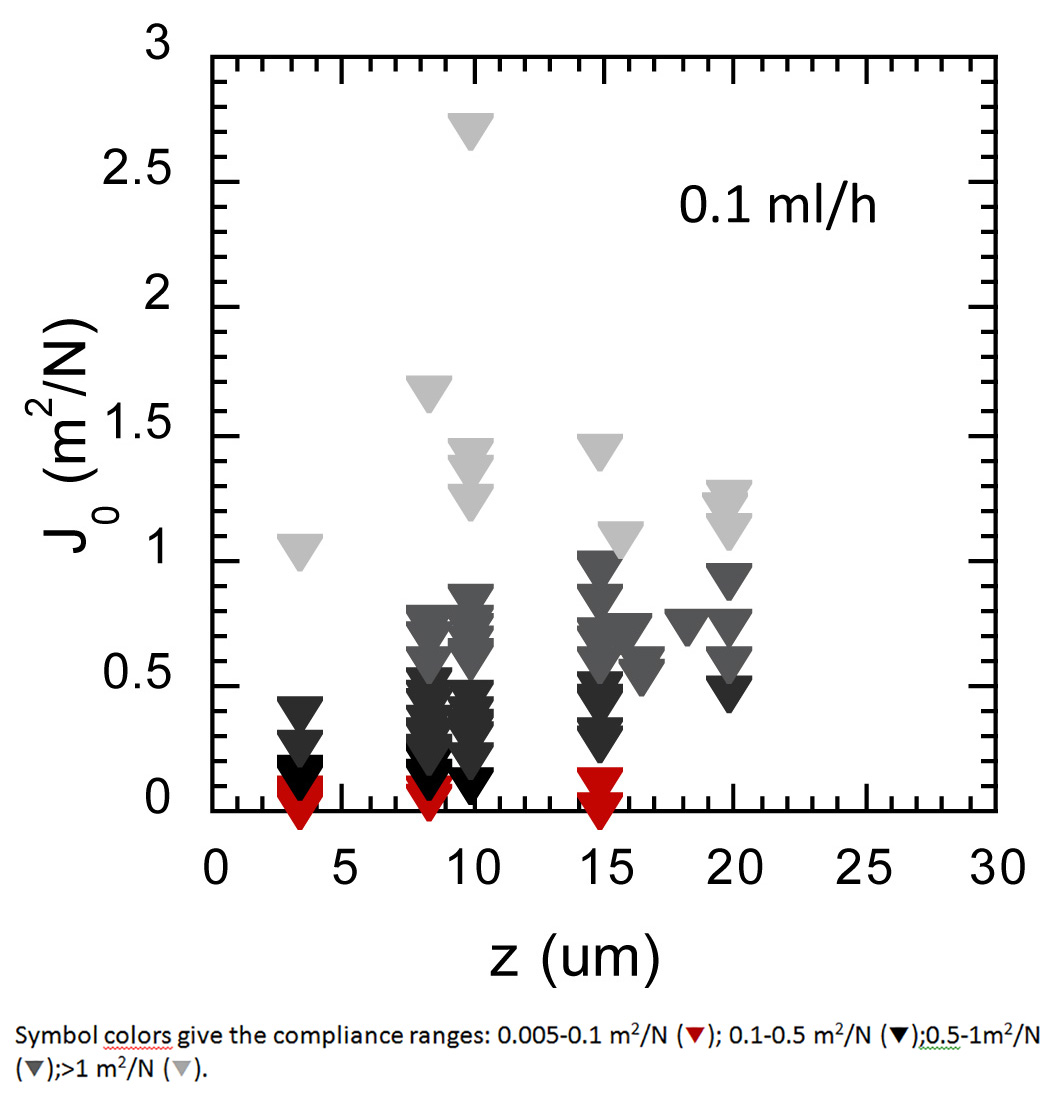
Figure 9. Spatial distribution of elastic compliance in a biofilm grown 24 hr at 37 °C under a lower flow of 0.1 ml/hr nutrients flow. Compliance values are represented according to the biofilm depth. Symbol colors give the compliance ranges.
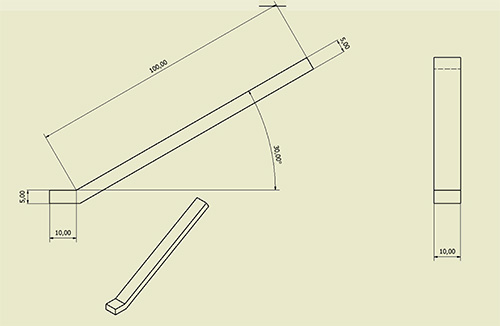
Figure 10. Magnetic pole blueprint. Dimensions are given in cm.
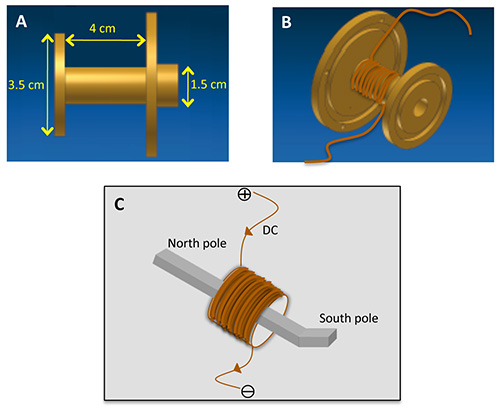
Figure 11. Magnetic tweezers wiring. A brass support — side view with dimensions in Panel A — is used to tightly wind the 2,120 turns of copper wire and build the magnetic coil (panel B) before inserting the soft magnetic alloy pole in its center. Direct current is supplied in the coil defining magnetic polarity as shown in panel C. See also experiment overview sketch in Figure 4.
Video 1: This movie shows the initial stage of the biofilm just after injection of the cell-particle mixture in the capillary. Acquisition frequency is 1 image/sec and the movie is played at 10 frames/sec. Please click here to view this video.
Video 2: This movie shows the biofilm after 4 hr growth. Particles are progressively detached from the surface to be distributed into the biofilm volume, blue arrows mark detachment events. Acquisition frequency is 2 image/min and the movie is played at 15 frames/sec. Please click here to view this video.
Video 3: Biofilm after 20 hr growth. The movie is taken in a middle plane located 20 µm over the capillary bottom. Images were acquired at 2 images/min frequency and the movie is played at 15 frames/sec. Please click here to view this video.
Video 4: Typical particle tracking using ImageJ. The movie shows an extract of the whole sequence starting at time 23 sec and ending at time 30 sec. Yellow trajectory shows particle displacement upon magnetic actuation. Acquisition frequency is 30 images/sec and the movie is played at 30 frames/sec. Please click here to view this video.
Discussion
This magnetic particle seeding and pulling experiment enabled in situ 3D mapping of the viscoelastic parameters of a growing biofilm in its original state. This approach revealed the mechanical heterogeneity of the E. coli biofilm grown here and gave clues to point out the biofilm components supporting the biofilm physical properties, strongly suggesting a fundamental implication of the extracellular matrix and more precisely its degree of cross-linking.
The recognition of mechanical properties patterns in bacterial biofilm is crucial for building a comprehensive representation of these complex materials. These findings open up the way to clarify the causal relationships linking mechanical patterns and biological heterogeneities such as gene expression microniches, which should help clarifying the driving forces underpinning biofilm development.
Up to now, the question of the biofilm mechanical properties had been addressed mainly from a macroscopic point of view and often by scraping biofilm from its initial site8,9,12,13, which represents an important risk of information loss. Our noninvasive method brings about the first in situ characterization of the biofilm 3D mechanical profile in its original environment.
The approach yet has a limitation residing in the characteristics of the magnetic remote actuation of colloidal probes. Indeed, it displays an intrinsic range of accessible forces given by the pole-particle distance, pole material properties and coil performance. The current configuration of the set-up enabled the probing of rigidity values up to 200 Pa, which was sufficient for the biofilms examined here. Nevertheless, further engineering of our set-up — involving, for instance, electromagnet cooling — should enable shifting of force and time limits by a factor of 2 to 3.
We are currently working on relating the physical and dynamical properties in a biofilm investigating the relation linking local viscoelastic parameters reported by the passive displacement of the probe on a larger time scale. In addition, we are exploring the impact of various chemicals targeting different components of the biofilm on its mechanical profile.
This new approach reveals the details of bacterial biofilm internal mechanics and contributes to a better understanding of these living structures, crucial from an engineer's point of view for biofilm control purposes but also from a fundamental perspective to clarify the relationship between the architectural properties and the specific biology of these structures.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
This work was in part supported by grants from the Agence Nationale pour la Recherche, PIRIbio program Dynabiofilm and from CNRS Interdisciplinary Risk program. We thank Philippe Thomen for his critical reading of the manuscript and Christophe Beloin for providing the E. coli strain used in this work.
Materials
| Table 1: Reagents and cells | ||||
| Magnetic particles | Life technologies | 14307D | Micrometric magnetic particle, 2.8 µm diameter | |
| Ampicillin (Antibiotic) | Sigma-Aldrich | A9518 | ||
| Tetracycline (Antibiotic) | Sigma-Aldrich | 87128 | ||
| Bacterial strain MG1655gfpF | UGB, Institut Pasteur, France | produces F pili at its surface, resistant to Ampicilllin and tetracycline | ||
| Table 2: Capillaries and tubing | ||||
| Filters for pediatric perfusion | Prodimed-Plastimed | 6932002 | ||
| Hollow Square Capillaries | Composite Metal Scientific | 8280-100 | Manufactured in Borosilicate glass. Square 0.8mm x0.8mm | |
| Tubing silicone peroxyde | VWR international | 228-0512 | Diameter 1mm | |
| Tubing silicone peroxyde | VWR international | 228-0700 | Diameter 3mm | |
| Table 3: Biofilm growth | ||||
| Lysogeny Broth (LB) solution | Amresco-VWR | J106-10PK | standard medium used to grow bacteria | |
| M63B1 solution | Home-made | Standard minimum medium used to grow bacteria | ||
| Glucose | Sigma-Aldrich | G8270 | Used to make M63B1 medium with 0.4% glucose | |
| Table 4: Electronics | ||||
| Camera EMCCD | Hamamatsu | C9100-02 | ||
| Heater controller | World precision instruments | 300354 | ||
| Function generator | Agilent technologies | 33210A | ||
| Power amplifier | Home-made | It gives a current signal with amplitudes up to 4 A. | ||
| Syringe pumps | Kd Scientific | KDS-220 | ||
| Shutter | Vincent Associates | Uniblitz T132 | ||
| Magnetic tweezers | Home-made | Two electromagnetic poles, each made of a copper coil with 2,120 turns of 0.56 mm in diameter copper wire and soft magnetic alloy cores (Supra50-Arcelor Mittal, France) square shaped according to the blueprint shown in Fig. 10. The two cores are mounted north pole facing south pole, in order to generate a magnetic force in one direction along the length of the capillary. See coil wiring details in Figure 11. | ||
| Table 5: Optics | ||||
| Inverted microscope | Nikon | TE-300 | ||
| S Fluor x40 Objective (NA 0.9, WD0.3) | Nikon | This a long working distance ojective enabling observation of the biofilm in the depth | ||
| Epifluorescence filters: 1) for green fluorescence: Exc 480/20 nm; DM 495; Em 510/20 2) for Red fluorescence: Exc 540/25 nm; DM 565; Em 605/55 | Chroma | 1)#49020 2)#31002 | Particle displacement upon force application is recorded using the red fluoresecnce filter block. | |
| Table 6: Image analysis | ||||
| ImageJ | NIH – particle tracker plugin | |||
Riferimenti
- Hall-Stoodley, L., Costerton, J. W., Stoodley, P. Bacterial biofilms: from the natural environment to infectious diseases. Nat Rev Microbiol. 2, 95-108 (2004).
- Donlan, R. M. Biofilms: microbial life on surfaces. Emerg Infect Dis. 8, 881-890 (2002).
- Costerton, J. W., Stewart, P. S. Battling biofilms. Scientific American. 285, 74-81 (2001).
- Branda, S. S., Vik, S., Friedman, L., Kolter, R. Biofilms: the matrix revisited. Trends Microbiol. 13, 20-26 (2005).
- Flemming, H. C., Wingender, J. The biofilm matrix. Nat Rev Microbiol. 8, 623-633 (2010).
- Costerton, J. W., Stewart, P. S., Greenberg, E. P. Bacterial biofilms: a common cause of persistent infections. Science. 284, 1318-1322 (1999).
- Stewart, P. S., Franklin, M. J. Physiological heterogeneity in biofilms. Nat Rev Microbiol. 6, 199-210 (2008).
- Stoodley, P., Lewandowski, Z., Boyle, J. D., Lappin-Scott, H. M. Structural deformation of bacterial biofilms caused by short-term fluctuations in fluid shear: an in situ investigation of biofilm rheology. Biotechnology and bioengineering. 65, 83-92 (1999).
- Klapper, I., Rupp, C. J., Cargo, R., Purvedorj, B., Stoodley, P. Viscoelastic fluid description of bacterial biofilm material properties. Biotechnol Bioeng. 80, 289-296 (2002).
- Korstgens, V., Flemming, H. C., Wingender, J., Borchard, W. Uniaxial compression measurement device for investigation of the mechanical stability of biofilms. Journal of microbiological. 46, 9-17 (2001).
- Cense, A. W., et al. Mechanical properties and failure of Streptococcus mutans biofilms, studied using a microindentation device. Journal of microbiological methods. 67, 463-472 (2006).
- Shaw, T., Winston, M., Rupp, C. J., Klapper, I., Stoodley, P. Commonality of elastic relaxation times in biofilms. Physical Review Letters. 93, (2004).
- Towler, B. W., Rupp, C. J., Cunningham, A. B., Stoodley, P. Viscoelastic properties of a mixed culture biofilm from rheometer creep analysis. Biofouling. 19, 279-285 (2003).
- Lau, P. C., Dutcher, J. R., Beveridge, T. J., Lam, J. S. Absolute quantitation of bacterial biofilm adhesion and viscoelasticity by microbead force spectroscopy. Biophysical journal. 96, 2935-2948 (2009).
- Poppele, E. H., Hozalski, R. M. Micro-cantilever method for measuring the tensile strength of biofilms and microbial flocs. Journal of microbiological methods. 55, 607-615 (2003).
- Aggarwal, S., Poppele, E. H., Hozalski, R. M. Development and testing of a novel microcantilever technique for measuring the cohesive strength of intact biofilms. Biotechnology and bioengineering. 105, 924-934 (2010).
- Guélon, T., Mathias, J. -. D., Stoodley, P. Biofilm Highlights. Series on Biofilms (eds Hans-Curt Flemming, Jost Wingender, & Ulrich Szewzyk). 5, (2011).
- Galy, O., et al. Mapping of Bacterial Biofilm Local Mechanics by Magnetic Microparticle Actuation. Biophysical journal. 103, 1-9 (2012).
- Schnurr, B., Gittes, F., MacKintosh, F. C., Schmidt, C. F. Determining Microscopic Viscoelasticity in Flexible and Semiflexible Polymer Networks from Thermal Fluctuations. Macromolecules. 30, 7781-7792 (1997).
- Aggarwal, S., Hozalski, R. M. Effect of Strain Rate on the Mechanical Properties of Staphylococcus epidermidis Biofilms. Langmuir. 28, 2812-2816 (2012).
- Towler, B. W., Cunningham, A., Stoodley, P., McKittrick, L. A model of fluid-biofilm interaction using a Burger material law. Biotechnol Bioeng. 96, 259-271 (2007).
- Jones, W. L., Sutton, M. P., McKittrick, L., Stewart, P. S. Chemical and antimicrobial treatments change the viscoelastic properties of bacterial biofilms. Biofouling. 27, 207-215 (2011).
- Apgar, J., et al. Multiple-particle tracking measurements of heterogeneities in solutions of actin filaments and actin bundles. Biophysical journal. 79, 1095-1106 (2000).

