VIGS-Mediated Forward Genetics Screening for Identification of Genes Involved in Nonhost Resistance
Summary
Virus-induced gene silencing is an useful tool for identifying genes involved in nonhost resistance of plants. We demonstrate the use of bacterial pathogens expressing GFPuv in identifying gene silenced plants susceptible to nonhost pathogens. This approach is easy, fast and facilitates large scale screening and similar protocol can be applied to studying various other plant-microbe interactions.
Abstract
Nonhost disease resistance of plants against bacterial pathogens is controlled by complex defense pathways. Understanding this mechanism is important for developing durable disease-resistant plants against wide range of pathogens. Virus-induced gene silencing (VIGS)-based forward genetics screening is a useful approach for identification of plant defense genes imparting nonhost resistance. Tobacco rattle virus (TRV)-based VIGS vector is the most efficient VIGS vector to date and has been efficiently used to silence endogenous target genes in Nicotiana benthamiana.
In this manuscript, we demonstrate a forward genetics screening approach for silencing of individual clones from a cDNA library in N. benthamiana and assessing the response of gene silenced plants for compromised nonhost resistance against nonhost pathogens, Pseudomonas syringae pv. tomato T1, P. syringae pv. glycinea, and X. campestris pv. vesicatoria. These bacterial pathogens are engineered to express GFPuv protein and their green fluorescing colonies can be seen by naked eye under UV light in the nonhost pathogen inoculated plants if the silenced target gene is involved in imparting nonhost resistance. This facilitates reliable and faster identification of gene silenced plants susceptible to nonhost pathogens. Further, promising candidate gene information can be known by sequencing the plant gene insert in TRV vector. Here we demonstrate the high throughput capability of VIGS-mediated forward genetics to identify genes involved in nonhost resistance. Approximately, 100 cDNAs can be individually silenced in about two to three weeks and their relevance in nonhost resistance against several nonhost bacterial pathogens can be studied in a week thereafter. In this manuscript, we enumerate the detailed steps involved in this screening. VIGS-mediated forward genetics screening approach can be extended not only to identifying genes involved in nonhost resistance but also to studying genes imparting several biotic and abiotic stress tolerances in various plant species.
Introduction
Nonhost resistance is the resistance of all plant species against races of a particular pathogen1,2. This imparts broad spectrum and durable disease resistance in plants2,3. However, its mechanism, particularly against bacterial pathogens, is not well understood4. Screening for mutants or silenced plants that compromise nonhost resistance, and high throughput transcript profiling for identification of differentially expressed genes during nonhost resistance5-9 are two major approaches previously used for dissecting bacterial nonhost resistance. Because nonhost resistance is controlled by a complex mechanism(s)4 with the involvement of many genes, a high throughput functional genomic approach for gene identification is critical for better understanding the nonhost resistance mechanism(s).
Virus-induced gene silencing (VIGS) has been successfully used to silence endogenous plant genes in many plant species10,11. Nicotiana benthamiana is one of the best suited plants for VIGS10,12 and its draft genome sequence is now available13. Tobacco rattle virus (TRV)-based VIGS has been widely used as reverse genetics tool to characterize genes involved in nonhost resistance2,4,14. This VIGS vectors and derivatives are now available through Arabidopsis Biological Resource Center (ABRC, http://www.arabidopsis.org/abrc/catalog/individ_cloned_gene_1.html). VIGS has also been used as a forward genetics tool for identifying genes involved in plant immunity15-17, especially nonhost resistance6,18. Assessing hypersensitive response (HR)-mediated cell death induced by plants against a specific nonhost pathogen and assessing the disease induced cell death are two major assays mainly used for identifying susceptible gene silenced plants. However, HR cell death is induced only against type-II nonhost pathogens and not against the type-I nonhost pathogens2. Hence, HR assays cannot be universally used to identify nonhost resistance strategies used by plants, especially against wide range of type-I nonhost pathogens. Also, partial loss of nonhost resistance in a gene silenced plant does not always lead to disease symptoms6 and hence disease scoring cannot be used for identifying plants compromising nonhost resistance. In contrary, assessing the growth of nonhost pathogens in the gene silenced plants is a better method for studying the loss of nonhost resistance in gene silenced plants.
Compared to conventional growth assay6,19, a faster method for assessing nonhost bacterial growth on the gene silenced plants can shorten the time required for forward genetics screening. We earlier reported a method for observing bacterial pathogen growth on leaves by naked eye under ultraviolet (UV) light using bacteria expressing green fluorescent protein (GFP)19. In this manuscript we demonstrate the usefulness of GFPuv expressing nonhost bacterial pathogens for easy identification of gene silenced plants that are compromised for nonhost resistance. This methodology is accurate for identification of susceptible plants and amenable for high throughput screening.
Protocol
1. Plant Growth and Target Gene Silencing
-
Plant growth conditions:
- Sow N. benthamiana seeds on soil-less potting mixture, Metro-Mix 350 and germinate the seeds in a growth chamber. Any other soil or soil-less medium can also be used instead of Metro-Mix.
- Transplant three-week old seedlings into individual pots and grow them in a greenhouse maintained at 21±2 °C along with other growth conditions as detailed in previous literature12. Two to three days after transplanting, plants can be used for TRV inoculation.
-
Growing TRV2 clones:
TRV is a bipartite virus and its genome consists of RNA1 and RNA2. RNA1 encodes an RNA-dependent RNA polymerase and a movement protein20,21. RNA2 encodes a coat protein (CP) and two nonstructural proteins from the subgenomic RNAs21. Both RNA1 and RNA2 are required for the formation of matured virus particles and their spread20,21. cDNA library construction in TRV2 vector is described in previous literature22,23. Briefly, VIGS library used in this study was constructed from the RNA extracted from leaf tissues exposed to various biotic and abiotic elicitors.- Take out Agrobacterium (GV2260 strain) containing the cDNA clones in TRV2 vector (a 96-well plate) from the freezer. Gradually thaw them and after the cultures reach room temperature, inoculate the individual Agrobacterium culture onto Luria-Bertani (LB) agar plate with rifampicin (10 μg/ml) and kanamycin (50 μg/ml) using a 96-pin replicator.
- Incubate the plates at 28 °C for two days. We usually grow four replicate colonies for each clone so that adequate Agrobacterium inoculum is available for prick inoculation of two plants. Perform all the steps under sterile conditions.
-
VIGS:
- Grow Agrobacterium (GV2260) carrying TRV1 at 28 °C in LB liquid medium with antibiotics mentioned above. Harvest cells by centrifugation from overnight grown cultures, re-suspend in the inoculation buffer (10 mM MES, pH 5.5; 200 μM acetosyringone), and incubate for 3 hr at room temperature on a shaker at 50 rpm.
- Harvest the cells by centrifugation and re-suspend in 5 mM MES buffer (pH 5.5) and inoculate (OD600 = 0.3) into abaxial side of 3-4 N. benthamiana leaves using a needleless syringe. Detailed inoculation procedure is demonstrated in previous literature24. Later, at the site of TRV1 inoculation, inoculate respective TRV2 colonies by pricking the leaves with a toothpick.
- Maintain the plants with adequate nutrition as vigorous growth is important for efficient VIGS12. About two weeks post inoculation the transcripts of targeted genes will begin to reduce and the plants are ready for pathogen inoculation at three weeks after TRV inoculation.
2. Preparation of Nonhost Pathogen Cultures and Plant Inoculation
-
Details of pathogens used in this study:
Pseudomonas syringae pv. tabaci, P. syringae pv. tomato T1, P. syringae pv. glycinea and Xanthomonas campestris pv. vesicatoria are used in this study. Grow P. syringae strains in King's B (KB) liquid medium supplemented with rifampicin (10 μg/ml), kanamycin (50 μg/ml) at 28 °C for 12 hr. Grow X. campestris pv. vesicatoria in LB liquid medium for 16 hr. All pathogens harbor a plasmid that can express GFPuv as described in previous literature19. -
Preparation of pathogen cultures for plant inoculation:
- Harvest bacterial cells by centrifugation and confirm the presence of green fluorescence using long wavelength UV lamp in the dark. Wash the cells twice with sterile water and re-suspend them to the desired concentration using sterile water.
- Inoculate the respective pathogen(s) on to abaxial side of target gene silenced leaves (5th to 8th) as spots (about 1.5 cm diameters). Several nonhost pathogens can be simultaneously tested for their growth in the target gene silenced plants. Simultaneously inoculate the host pathogen as this can be used as a positive control for viewing in planta bacterial growth. Also inoculate vector control (TRV::00) plants.
3. Observation of in planta Growth of Bacterial Pathogens
- Expose the inoculated leaves to a long wavelength UV light in the dark. Bacterial colonies can be viewed as green fluorescent dots in the abaxial side of the leaf in the background of red fluorescence emitted by leaf surface.
- Observe pathogen growth from 2 days post inoculation (dpi) to 5 dpi. Everyday observation during this time interval is necessary.
- After the first screen, short list the clones whose silencing results in growth of one or more nonhost pathogen(s).
- Repeat VIGS for the selected clones and again test the response of gene silenced plants to nonhost pathogen(s). This second level of screening is done to remove the false positives obtained during the first screen.
4. Confirmation of Shortlisted Plants for Compromised Nonhost Resistance
- Silence the cDNA clones selected from the screen as described above. Inoculate the whole plant with nonhost pathogen(s) by submerging the plants with respective pathogen cultures with 0.01% (v/v) Silwet L-77 and subjecting the submerged plants to vacuum for 3 – 5 min. Quantify the bacterial growth by conventional growth assay6,18,19 as described below.
- At 0 hr post inoculation (hpi), 3 dpi and 5 dpi, collect two leaf samples from five biological replicates for each nonhost pathogen(s) inoculated leaves using a leaf punch (0.5 cm2). Grind the leaf samples, serial dilute and plate the sap on KB agar medium supplemented with appropriate antibiotics and incubate at 28 °C for 2 days. Count the bacterial colonies using a colony counter and calculate the bacterial growth in the leaves as described in previous literature6.
- Plants susceptible to nonhost pathogen(s) should show higher number of pathogen growth compared to vector control.
5. Sequencing the Insert and Identification of Target Gene
- Perform colony PCR for the selected clone using attB1 and attB2 primers or using the primers flanking the cloning site in the TRV2 vector. Run the PCR product on the gel and check for amplification of the single band.
- Plant gene insert in the TRV2 vector can be sequenced using attB primers or primers flanking the cloning site.
- Perform BLAST using the sequence and identify the gene details.
- Obtain full length gene sequence along with the un-translated regions (UTRs).
- Select 300-400 bp fragments from three different regions of the sequence and clone them into TRV2 vector. Independently perform VIGS using all three VIGS constructs and confirm the compromise of nonhost resistance in all three gene silenced plants.
Representative Results
Major aim of this study is to demonstrate a method for easy and accurate identification of gene silenced N. benthamiana plants that are compromised for nonhost resistance. There are four major steps in this methodology. First step is to individually silence large number of genes using TRV-VIGS. We had silenced about 5,000 genes6,18 over a period of about 1.5 years using the protocol depicted in Figure 1. Some of the gene silenced plants showed various phenotypic alterations including stunted growth, yellowing and photo-bleaching phenotypes (Figure 2).
Second step is the identification of gene silenced plants showing susceptibility to nonhost pathogen(s). We used GFPuv expressing bacteria and tracked their growth in planta (Figures 3A-3C). In order to demonstrate the process of identification of the gene silenced plants susceptible to nonhost pathogen(s), we used three nonhost pathogens, P. syringae pv. tomato T1, P. syringae pv. glycinea and X. campestris pv. vesicatoria. As expected, non-silenced (vector control) N. benthamiana plants inoculated with these pathogens did not grow as they could not overcome the plant immunity. However, P. syringae pv. tomato T1 formed green fluorescent colonies on NbSGT1 gene (a gene implicated in nonhost resistance25) silenced plants (Figure 3D). These data demonstrate the process of identifying gene silenced plants compromised for nonhost resistance. This protocol can be directly applied to forward genetics screening. For example, the representative results taken from one of our previous forward genetics screen (Figure 4) shows two independent gene silenced plants with varying degree of P. syringae pv. tomato T1 growth (Figure 5).
Third step is the confirmation of bacterial growth in the gene silenced plants identified from the screen by quantifying the bacterial growth. An example for estimation of in planta bacterial growth in the NbSGT1 gene silenced plants is shown in Figure 3E. Results showed that in planta growth of P. syringae pv. tomato T1 was much higher in the NbSGT1 silenced plants compared to non-silenced (TRV::00) control plants (Figure 3E). Loss of nonhost resistance due to a particular gene silencing could be either partial or complete. Depending on this, the extent of nonhost pathogen growth on the gene silenced plants can vary.
Fourth step is sequencing the insert in TRV2 vector. BLAST is performed using this sequence to identify the gene and its details.
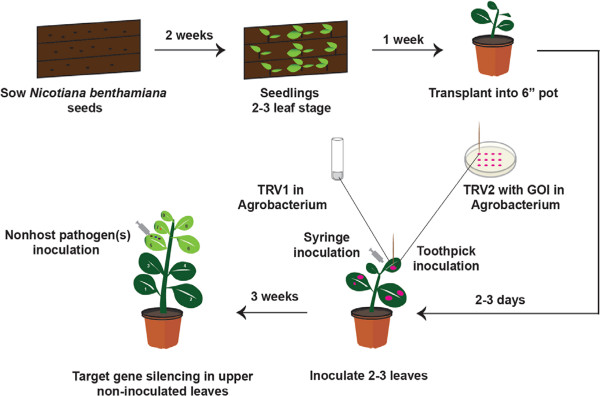
Figure 1. Cartoon showing the steps involved in silencing a large number of genes using TRV-based VIGS. Three week old N. benthamiana plants are transplanted into individual pots and allowed to grow for a few days. Agrobacterium carrying TRV1 construct is infiltrated into three to four leaves using needless syringe. Later, 96 cDNA clones grown on LB agar plate are individually taken using a toothpick and pricked on to the TRV1 inoculated spot of respective plants. About 3 weeks after inoculation, target gene silencing occurs in the top non-inoculated newly grown leaves. These leaves (5th to 8th) are used for testing the growth of nonhost pathogen(s).
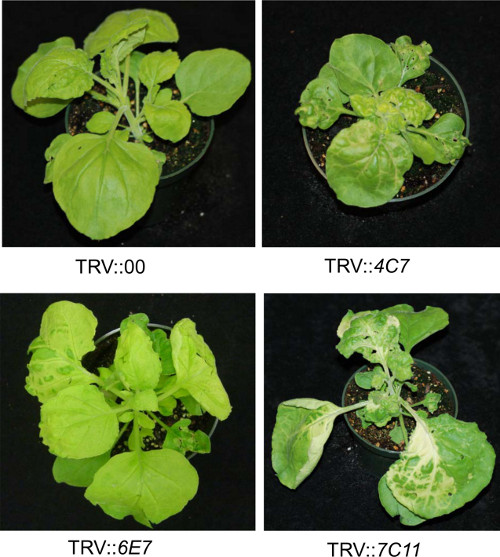

Figure 2. VIGS-mediated forward genetics screening in N. benthamiana shows various changes in plant phenotype. Three-week old N. benthamiana plants independently inoculated with respective TRV constructs as descried in Figure 1 as a part of forward genetics screen are shown. Photographs were taken at 21 days after TRV inoculation.
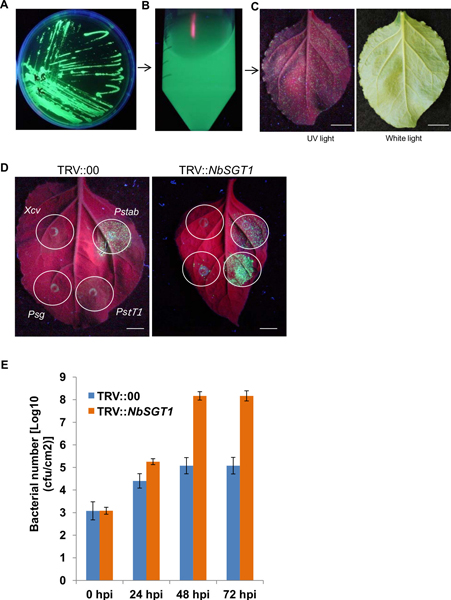
Figure 3. Green fluorescence from GFPuv expressing bacterial pathogens and its use for identification of genes involved in nonhost resistance. Pseudomonas syringae pv. tabaci, a host pathogen, carrying GFPuv construct streaked on KB-medium plate shows green fluorescence under long wave UV light in the dark (A). These bacteria are grown on KB medium and re-suspended in water (B) and inoculated on to N. benthamiana leaves (abaxial side, 1 x 104 cfu/ml). At 3 days post inoculation (dpi) bacterial colonies are viewed as green fluorescent spots by naked eye from abaxial side of leaf epidermal cells (C, left). The same leaf pictured under normal light (visible range) is also shown (C, right). GFPuv expressing P. syringae pv. tabaci (Pstab, 1 x 104 cfu/ml) and three nonhost pathogens, P. syringae pv. tomato T1 (PstT1, 1 x 104 cfu/ml), P. syringae pv. glycinea (Psg, 3 x 105 cfu/ml) and Xanthomonas campestris pv. vesicatoria (Xcv, 3 x 104 cfu/ml) are inoculated on to the abaxial side of leaves from vector control (TRV::00; D, left) and NbSGT1 gene silenced plants (D, right). Bacterial colonies of Pstab are seen on both leaves and those of PstT1 are seen only on the NbSGT1 gene silenced leaves. PstT1 growth in NbSGT1 gene silenced leaves quantified at 24, 24 and 72 hr post inoculation (hpi, E) is shown in the graph. Click here to view larger figure.
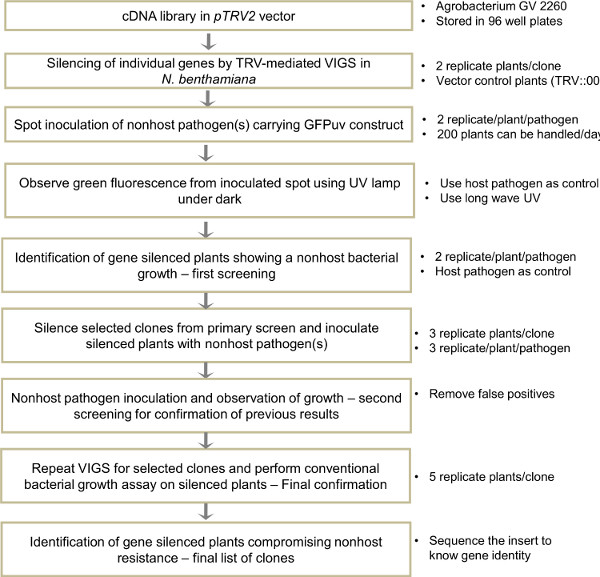
Figure 4. Overview of protocol followed for VIGS-mediated forward genetics screening for identification of genes involved in nonhost resistance. A cDNA library in TRV2 vector constructed using RNA from nonhost pathogen(s) inoculated N. benthamiana leaves can be used for screening. Individual clones from the cDNA library are silenced in N. benthamiana by TRV-VIGS. Respective GFPuv expressing nonhost bacterial pathogen(s) is inoculated on to the gene silenced plant leaves. Between 2 and 4 dpi the inoculated spots are observed using a UV lamp in the dark and the plants showing nonhost pathogen growth are identified. After this preliminary screening, the selected clones are silenced again and the nonhost pathogen growth is reconfirmed. This second level screen is done to remove the false positives. Later, confirmed clones are silenced for a third time and respective bacterial growth is estimated. Inserts of respective clones in the TRV2 vector are sequenced and gene information is identified. Simultaneously HR assay can also be performed on the gene silenced plants to identify genes contributing to HR induced against type-II nonhost pathogens.
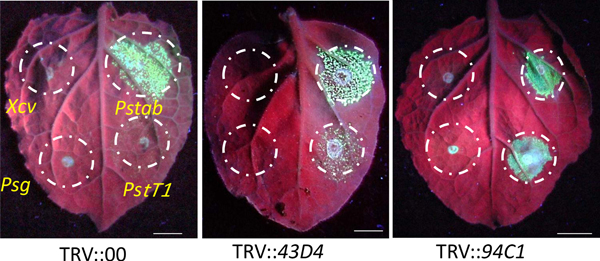
Figure 5. Nonhost bacterial pathogen growth in gene silenced plants that were identified as part of a forward genetics screen. Pictures from a forward genetics screening that identified genes involved in nonhost resistance of N. benthamiana plants against PstT1, Psg and Xcv. Photographs taken for two representative gene silenced plants at 3 dpi after inoculation are shown here. Bacterial concentrations used for inoculation are described in Figure 3. 43D4 or 94C1 indicates the 96-well plate number followed by the well number.
Discussion
Plant immunity limits the growth of nonhost pathogens and hence, little or no green fluorescence is emitted from vector control plant leaves inoculated with nonhost pathogen under long wavelength UV light (Figure 3D). However, when a gene involved in nonhost resistance is silenced, the gene silenced plants favor the growth of nonhost pathogen and green fluorescence is seen (Figure 3E). This is the basic principle involved in the method described in this manuscript. This methodology has two major advantages over the HR-based assay. One, identification of silenced plants susceptible to nonhost pathogen by GFPuv-based assay increases the accuracy as in planta growth can be viewed. Second, this method allows identification of genes involved in both type-I nonhost resistance2,26 (does not result in visible cell death) and type-II nonhost resistance2,26 (results in hypersensitive response cell death). For example, we previously identified a clone called 6C8 and the silenced plants were susceptible to both type-1 and type-2 pathogens18.
Also, it is possible that two or more independent clones used for VIGS resulted in gene silenced plants with similar phenotypes as they happen to silence a particular gene. This redundancy is useful as it confirms the silencing phenotype. Occurrence of such repeats depends on the library used for screening and type of gene (as some genes are highly expressed and hence occur repeatedly in a library). Further, cloning nearly all genes encoded by N. benthamiana genome into a cDNA VIGS library and silencing them independently is possible.
In order to effectively implement the forward genetics screening for identification of genes involved in nonhost resistance using VIGS followed by GFPuv fluorescence assay, the following tips are useful.
In order for successful VIGS, (1) use 3 – 4 leaf stage plants for TRV inoculation; (2) optimum TRV1 and TRV2 titer is important. Since TRV2 titer can be a limiting factor for initiation of VIGS, inoculate at least 4 fully grown bacterial colonies of Agrobacterium harboring TRV2; and (3) maintain optimum environmental conditions12. Environmental temperature plays a major role in inducing optimum VIGS.
Critical steps for efficient detection of green fluorescence from pathogens include: (1) always use fresh pathogen cultures harvested at exponential growth phase and do not store them before inoculation; and (2) HR cell death should be avoided as they can auto fluoresce under UV light and negatively influence the observations. This is especially important when type-II nonhost pathogen (for example, P. syringae pv. tomato T1) is used for the study. Use lower concentration of the nonhost pathogen if microscopic HR is suspected or evidenced.
Troubleshooting:
(1) Inconsistency in gene silencing and occurrence of variable silencing phenotype among replicates:
Change gloves for every clone during the TRV inoculation. Also, make sure that the TRV2 Agrobacterium colony does not contain multiple clones by colony PCR. If colony PCR reveals more than one band, it is possible that more than one clone is present in the colony. In order to separate the clones, streak them on the LB agar plate and screen the clones by colony PCR. After identification of individual clones, perform VIGS independently for each of them and identify the gene silenced plant compromising nonhost resistance.
(2) Severe growth defects in the silenced plants:
If the silenced target gene(s) is involved in basal metabolism of plants, growth defects are expected. However, some of the gene silenced plants may show higher multiplication of TRV compared to vector control plants and this may cause additive effect on the severity of the phenotype. Local lesion assay12 on Chenopodium amaranticolor plants may be performed to find out whether the silenced plants have higher TRV titer compared to vector control plants. Under these circumstances, plants about 4 weeks old can be used for VIGS to reduce the severity of the phenotype.
(2) Off-target gene silencing:
Non-specific gene silencing, often referred as off-target silencing can occur during VIGS. This occurs when a partial sequence homology allows siRNA generated for intended target gene to degrade mRNA for genes that are not the intended silencing targets27. Use UTR sequences as insert in the TRV2 vector. Study the endogenous gene downregulation of all expected off-target genes and make sure the construct silences a single gene of interest. To facilitate identification of gene fragments with minimum or no off-target, bioinformatics tools can be used (for example, http://plantgrn.noble.org/pssRNAit/).
(3) No green fluorescence from bacterial colonies in planta:
This occurs most likely by the use of over grown cultures. Another cause could be lesser bacterial inoculum. Although bacterial concentrations can be indirectly measured by measuring optical density OD600, dead bacterial cells can make up higher OD. It is advisable to calculate optimum bacterial concentration based on colony forming units (cfu) by counting live cells on the KB agar medium plate. OD values can be adjusted based on the cfu. Also, streak the pathogen as often as possible from the glycerol stock, grow it on the KB agar plate and inoculate single fresh colony for preparing cultures for inoculation.
(4) Issues with sequencing the insert:
One commonly occurring cause for difficulties with sequencing the insert in TRV2 vector (by colony PCR) is the presence of multiple colonies. In order to overcome this, clones harboring individual TRV2 construct should be isolated as mentioned above (#1).
Limitations of the GFPuv method and ways to overcome:
Pathogens may lose the GFPuv carrying plasmid during multiplication and hence, adversely influence the observations. Also, some of the gene silenced plants shows yellowing and photo-bleaching leaves and these leaves emit different fluorescence (than red). This makes the green fluorescence difficult to see. One of the ways to reduce the effect of auto fluorescence from the yellow or white leaves is to capture pictures of the green fluorescent bacteria using a camera fitted with filters that can remove or reduce the background effect. Pictures can be used to infer whether the bacteria have grown or not. Certain software (for example, Adobe Photoshop) can be used to reduce the background effect during analysis of such pictures.
Biosafety precautions like appropriate disposal of transgenic bacterial pathogens need to be followed during experimentation. Also, appropriate skin and eye protection shields should be used when handling the UV lamp.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
This project was funded by The Samuel Roberts Noble Foundation. Authors thank Mss. Janie Gallaway and Colleen Elles for excellent plant care; and Ms. Katie Brown for artwork. We also would like to thank Mss. Trina Cottrell, Pooja Uppalapati, Moumita Saha, Swetha Vinukonda and Mr. Isaac Greenhut for technical help during establishment of this protocol.
Materials
| Name of the reagent/equipment | Company | Catalogue number | Comments (optional) |
| 96-well U-bottom plates | Becton Dickinson Labware (Franklin Lakes, NJ, USA) | 35-3077 | |
| 96-pin replicator stainless steel | Nalge Nunc International (Naperville, IL, USA) | 250520 | |
| High Intensity UV Inspection Lamps, Model B-100ap | Thomas scientific (Swedesboro, NJ, USA) | 6283K50 | Manufacturer ID 95-0127-01 |
| Stuart SC6 colony counter | Bibby Scientific Limited, Staffordshire, UK | SC6PLUS | |
| Soil-less potting mixture, Metro-Mix 350 | SUNGRO Horticulture Distribution, Inc., (Bellevue, WA, USA) | ||
| Primers: attB1 (GGGGACAAGTTTGTACAAAAAAGCAGGCT) attB2 (GGGGACCACTTTGTACAAGAAAGCTGGGT) |
Integrated DNA Technologies, Inc (Coralville, IA, USA) | Custom synthesized | |
| MES, Monohydrate | VWR international (Radnor, PA, USA) | CAS No. 145224-94-8 | |
| Acetosyringone (Dimethoxy-4′-hydroxyacetophenone) | Sigma Aldrich (St. Louis, MO, USA) | D134406 | |
| Vac-In-Stuff (Silwet L-77) | Lehle Seeds (Round Rock, TX, USA) | VIS-30 |
Riferimenti
- Heath, M. C. Nonhost resistance and nonspecific plant defenses. Currrent Opinion in Plant Biology. 3, 315-319 (2000).
- Mysore, K. S., Ryu, C. -. M. Nonhost resistance: how much do we know. Trends in Plant Science. 9, 97-104 (2004).
- Ellis, J. Insights into nonhost disease resistance: can they assist disease control in agriculture. The Plant Cell. 18, 523-528 (2006).
- Senthil-Kumar, M., Mysore, K. S. Nonhost resistance against bacterial pathogens: retrospects and prospects. Annual Review of Phytopathology. 51, (2013).
- Lu, M., Tang, X., Zhou, J. -. M. Arabidopsis NHO1 is required for general resistance against Pseudomonas bacteria. The Plant Cell. 13, 437-447 (2001).
- Rojas, C. M., et al. Glycolate oxidase plays a major role during nonhost resistance responses by modulating reactive oxygen species mediated signal transduction pathways. The Plant Cell. 24, 336-352 (2012).
- Daurelio, L. D., et al. Transcriptome analysis reveals novel genes involved in nonhost response to bacterial infection in tobacco. Journal of Plant Physiology. 168, 382-391 (2011).
- Moreau, M., et al. EDS1 contributes to nonhost resistance of Arabidopsis thaliana against Erwinia amylovora. Molecular Plant-Microbe Interactions. 25, 421-430 (2012).
- Senthil-Kumar, M., Mysore, K. S. Ornithine-delta-aminotransferase and proline dehydrogenase genes play a role in non-host disease resistance by regulating pyrroline-5-carboxylate metabolism-induced hypersensitive response. Plant, Cell & Environment. 35, 1329-1343 (2012).
- Burch-Smith, T. M., Anderson, J. C., Martin, G. B., Dinesh-Kumar, S. P. Applications and advantages of virus-induced gene silencing for gene function studies in plants. The Plant Journal. 39, 734-746 (2004).
- Senthil-Kumar, M., Mysore, K. S. New dimensions for VIGS in plant functional genomics. Trends in Plant Science. 16, 656-665 (2011).
- Senthil-Kumar, M., Mysore, K. S. Virus-induced gene silencing can persist for more than 2 years and also be transmitted to progeny seedlings in Nicotiana benthamiana and tomato. Plant Biotechnology Journal. 9, 797-806 (2011).
- Bombarely, A., et al. A draft genome sequence of Nicotiana benthamiana to enhance molecular plant-microbe biology research. Molecular Plant-Microbe Interactions. 25, 1523-1530 (2012).
- Sharma, P. C., et al. Virus-induced silencing of WIPK and SIPK genes reduces resistance to a bacterial pathogen, but has no effect on the INF1-induced hypersensitive response (HR) in Nicotiana benthamiana. Mol Gen Genomics. 269, 583-591 (2003).
- Baulcombe, D. C. Fast forward genetics based on virus-induced gene silencing. Current Opinion in Plant Biology. 2, 109-113 (1999).
- Lu, R., et al. High throughput virus-induced gene silencing implicates heat shock protein 90 in plant disease resistance. EMBO Journal. 22, 5690-5699 (2003).
- Pozo, O., Pedley, K. F., Martin, G. B. MAPKKK[alpha] is a positive regulator of cell death associated with both plant immunity and disease. EMBO Journal. 23, 3072-3082 (2004).
- Wang, K., Senthil-Kumar, M., Ryu, C. -. M., Kang, L., Mysore, K. S. Phytosterols play a key role in plant innate immunity against bacterial pathogens by regulating nutrient efflux into the apoplast. Plant Physiology. 158, 1789-1802 (2012).
- Wang, K., Kang, L., Anand, A., Lazarovits, G., Mysore, K. S. Monitoring in planta bacterial infection at both cellular and whole-plant levels using the green fluorescent protein variant GFPuv. New Phytologist. 174, 212-223 (2007).
- Ratcliff, F., Martin-Hernandez, A. M., Baulcombe, D. C. Technical Advance: Tobacco rattle virus as a vector for analysis of gene function by silencing. The Plant Journal. 25, 237-245 (2001).
- MacFarlane, S. A. Molecular biology of the tobraviruses. Journal of General Virology. 80, 2799-2807 (1999).
- Anand, A., et al. Identification and characterization of plant genes involved in Agrobacterium-mediated plant transformation by virus-induced gene silencing. Molecular Plant-Microbe Interactions. 20, 41-52 (2007).
- Liu, E., Page, J. Optimized cDNA libraries for virus-induced gene silencing (VIGS) using tobacco rattle virus. Plant Methods. 4, 5 (2008).
- Velásquez, A. C., Chakravarthy, S., Martin, G. B. Virus-induced gene silencing (VIGS) in Nicotiana benthamiana and tomato. J. Vis. Exp. (28), e1292 (2009).
- Peart, J. R., et al. Ubiquitin ligase-associated protein SGT1 is required for host and nonhost disease resistance in plants. Proceedings of the National Academy of Sciences. 99. , 99-10869 (2002).
- Oh, S. -. K., et al. Insight into Types I and II nonhost resistance using expression patterns of defense-related genes in tobacco. Planta. 223, 1101-1107 (2006).
- Senthil-Kumar, M., Mysore, K., Kodama, H., Komamine, A. RNAi and Plant Gene Function Analysis. Methods in Molecular Biology. 744, 13-25 (2011).

