Glutamine Flux Imaging Using Genetically Encoded Sensors
Summary
This article will demonstrate how to monitor glutamine dynamics in live cells using FRET. Genetically encoded sensors allow real-time monitoring of biological molecules at a subcellular resolution. Experimental design, technical details of the experimental settings, and considerations for post-experimental analyses will be discussed for genetically encoded glutamine sensors.
Abstract
Genetically encoded sensors allow real-time monitoring of biological molecules at a subcellular resolution. A tremendous variety of such sensors for biological molecules became available in the past 15 years, some of which became indispensable tools that are used routinely in many laboratories.
One of the exciting applications of genetically encoded sensors is the use of these sensors in investigating cellular transport processes. Properties of transporters such as kinetics and substrate specificities can be investigated at a cellular level, providing possibilities for cell-type specific analyses of transport activities. In this article, we will demonstrate how transporter dynamics can be observed using genetically encoded glutamine sensor as an example. Experimental design, technical details of the experimental settings, and considerations for post-experimental analyses will be discussed.
Introduction
Due to remarkable progress in technologies that allows examination of the transcriptome and the proteome at a cellular level, it has now become clear that the biochemistry and the resulting flux of metabolites and ions are highly cell-type specific. For example, in the mammalian liver, sequential glutamine degradation and synthesis are carried out simultaneously by periportal cells and perivenous cells respectively, feeding ammonium to the urea cycle in the former cell type while consuming excess ammonia in the latter 1-3. In some cases, significant biochemical heterogeneity is detected even in a single “cell type” 4,5. In addition to such spatial specificity, the cellular levels of metabolites and ions are highly dynamic (e.g., signaling molecules such as Ca2+ and cyclic nucleotides). The spatiotemporal patterns of metabolites and ions often play critical roles in signal transduction. Monitoring cellular dynamics of metabolites and ions, however, pose unique challenge. In many cases the change in concentrations are rapid and transient, exemplified by the case of signaling molecules such as Ca2+, which decays within ~20 msec in dendritic spines 6. In addition, compartmentalization of biochemical pathways within and among the cells makes it difficult to quantify the dynamics of metabolites and ions using extraction and column chromatography/mass spectrometry techniques.
Genetically encoded sensors for biological molecules are now widely used due to the high spatiotemporal resolution that allows the experimenter to study short-lived and/or compartmentalized molecular dynamics (reviewed in 7,8). These genetically encoded sensors can roughly be divided into two categories; intensity-based sensors and ratiometic sensors. Intensity-based sensors typically consist of a binding domain and a fluorescent protein (FPs), and the solute binding to the binding domain changes the fluorescent intensity. Ratiometic sensors, on the other hand, often take advantage of Föster Resonance Energy Transfer (FRET) between two FPs that function as a FRET pair. These sensors consist of a binding domain and two FPs, and the solute binding induces the change in FRET efficiency between the two FPs. A large number of sensors for biologically important metabolites and ions have been developed in the past decade 8,9.
One of the exciting possibility offered by such genetically encoded sensors is their use in the high-resolution analysis of membrane transport processes, which previously was not easy to detect at the cellular level. Genetically encoded sensors facilitate the analysis of transport mechanisms such as substrate specificity and pH dependence 10,11. Moreover, in combination with the genetic resources such as the library of RNAi constructs for model organisms, it is now possible to conduct genome-wide searches for novel transport processes using genetically encoded sensors. Indeed, use of genetically encoded sensor lead to the discovery of previously uncharacterized transporters in multiple cases 12,13.
Recently, our laboratory has developed a series of FRET-based sensor for glutamine. We have demonstrated that cellular glutamine levels can be visualized using such FRET glutamine sensors 10. These sensors consist of a FRET donor (mTFP1) inserted into a bacterial glutamine binding protein glnH, and a FRET acceptor (venus) at the C-termini of glnH (Figure 1). FRET efficiency of these sensors decrease upon binding of glutamine, resulting in the decrease of acceptor/donor intensity ratio. Fine regulation of glutamine transport processes is important in biological processes such as neurotransmission 14,15 and the maintenance of urea cycle in the liver 1,16,17.
Here we show the methodology of analyzing transport activities with FRET sensors for glutamine, using a wide-field fluorescence microscope set-up. The goal of experiments shown here are to detect transporter activities in a single cell and to examine substrate specificity of a transiently expressed transporter.
Protocol
1. Sample Preparation
Note: In many cases perfusion experiments wash away a significant portion of cells, which can become a frustrating issue. Although not necessary for all cell lines, coating cover glass surfaces with poly-L-Lysine (add 1.0 ml/25 cm2 of 0.01% solution to the surface, incubate >5 min, wash twice with cell culture grade water, and dry in the biosafety cabinet) enhances cell adhesion. Also, be aware of the biosafety level (BSL) of the cell line used, and follow the standard operating procedure approved by the local environmental health and safety office. In this experiment, cos7 cells were used due to low endogenous glutamine transport activity (see Figure 4).
- Seed cos7 cells at ~70-80% confluency in Dulbecco’s Modified Eagle Medium (DMEM) + 10% cosmic calf serum and 100 U/ml penicillin/streptomycin, either on sterile 25 mm glass coverslips placed in the 6-well culture dish or 8-well glass-bottom slides. Incubate at 37 °C, 5% CO2, 100% humidity for 24 hr.
- Exchange the growth medium to DMEM +10% cosmic calf serum (no antibiotics), according to the manufacturer’s protocol. Grow O/N at 37 °C, 5% CO2 at 100% humidity.
- Add 0.4 mg each of plasmid DNA encoding the glutamine sensor (pcDNA3.1, carrying FLIPQTV3.0 sensor under CMV promoter 10) and the mCherry-tagged ASCT2 10 to 50 ml serum-free media (OPTI-MEM) and mix gently.
- Add 1 ml of transfection reagent to 50 ml serum-free media. Incubate at RT for 5 min.
- Mix the two solutions containing DNA (step 1.3) and transfection reagent (step 1.4) and incubate for 20 min.
- Gently add the transfection reagent on top of the cells and mix gently by rocking the chamber. Incubate for 24 hr at 37 °C, 5% CO2, 100% humidity.
- After 24 hr, exchange the medium to DMEM + 10% Cosmic Calf Serum (CCS) + 100 unit of penicillin/streptomycin.
- Cells are imaged 48-72 hr post-transfection.
2. Perfusion Experiment
Note: For cos7 cells used in this experiment, perfusion media and chamber were kept at RT and ambient CO2 concentration. However, if the cells being used require higher temperature and CO2 concentration control for survival, heated microscope stage and/or an environmental chamber should be used.
- Prepare perfusion buffer A-F (See List of Materials). Attach stopcocks to 50 ml syringes, close the stopcocks then fill them with the perfusion solutions. Open the stopcock for a brief period to fill the head space in the stopcock with the buffer.
- Switch on the 8-channel, gravity-feed perfusion system with a perfusion pencil, and then select “Manual Mode” under Select Function option. Place a waste container under the perfusion pencil.
- Manually activate the ports by pressing the corresponding numbers and fill the tubings using 10 ml syringe containing water, and then close the port. Once the ports are filled with water, set the syringes containing buffers A-F on the port 1-6 of perfusion system, and then open the stopcocks. Open the ports again to replace the water in the tubing with the buffers. The flow rate should be around 0.8 ml/min.
- Press cancel once on the perfusion controller to go back to the “Select Function” menu. Toggle to “Edit Program” option, then type in the perfusion protocol indicated in Table 1. Save the perfusion program.
- Take the cells out of the incubator. Wash twice with the perfusion buffer, and then set the cells on the stage. Connect the perfusion system to the chamber. If an open chamber is used, set up another pump that removes the perfusion solution from the chamber.
- Open Slidebook software. Start a new slide.
Note: The following procedure is specific to Slidebook software, but other software such as MetaFluor and Nis-Element can also be used for the type of imaging described here. Theoretically experiments can be performed using any software that allows time-course imaging, and the image sequences can be analyzed post-experimentally using software such as ImageJ (http://rsbweb.nih.gov/ij/) or Fiji (http://fiji.sc/Fiji). However, software that allows real-time monitoring of acceptor/donor ratio is highly useful. - Mount the slide onto the fluorescent microscope stage. Find the cells co-expressing the sensor and ASCT-mCherry using appropriate filter sets (see List of Materials) under “FOCUS” function. Adjust gain by clicking “Camera” tab and sliding the slide bar under “gain”. Also, adjust the neutral density filter setting from neutral density filter drop-down menu. Note: If the filter cube does not include an eye filter, do not use the eye pieces since short-wavelength excitation light is harmful to eyes. Use the computer screen to find the cells instead.
- Acquire images of cells with donor (mTFP1)exdonor(mTFP1)em , donor(mTFP1)exacceptor(venus)em and acceptor(mTFP1)exacceptor(venus)em m channels (filters are specified in the List of Materials). Click “Image Capture”, and then in the acquisition window, specify the filters that are going to be used. Click “TEST” button to adjust the exposure time by typing the desired exposure time in the box next to the filter settings. Note: All three channels need to be exposed for the same length. When adjusting the imaging parameters, take the expected FRET efficiency change and fold increase into account: for example, if the FRET efficiency is expected to go up 2-fold during the experiment, imaging parameters should be adjusted so that the intensity of DonorexAcceptorem at the resting state should be <50% of the maximal value that can be recorded by the instrument, so that channel does not become saturated during the experiment. Signal from the labeled cells should be at least three-times higher than that in the background region.
- Draw a region of interest using regions tool in the software. Alternatively, a software function to select pixels above certain intensity can be used, and then converted into regions. To do this, click Mask – Segment, then move the sliding bar to highlight the desired areas.
- Select “time lapse” in the capture type box, then specify the interval (10 sec in this experiment) and time points for the experiment.
- Draw a region in an area without any fluorescent cells. This area is used for background subtraction. Right click the region drawn, and then set it as a background. Note: Ideally, cells that are not expressing the sensors should be used as the background region to account for both the autofluorescence and the background florescence detected by the camera. If no such cells are available in the field of view, at least a region without the cells should be used to account for the background fluorescence.
- Select the desired program under “Select Function – Run programs” option. Toggle to the saved program (see step 2.3), and then hit ENTER on the perfusion controller. Now the program is loaded, and hitting any key will start the perfusion.
- Ensure perfusion and the imaging experiment start at the same time by clicking “Start” button in the capture window at the same time as pressing a key on the perfusion controller.
Note: It is recommended to record the timepoint where the solutions were exchanged (this can be done using “NOTES” function, found within the acquisition tab). - Perform the same perfusion protocol using cells expressing a sensor that is expected to be either saturated or unbound under the experimental condition.
Note: Here, FLIPQTV3.0_1.5μ, which is saturated under the experimental condition, is used as a control, Figure 6. Such control experiments are necessary to confirm that the FRET efficiency change is due to the actual change in the substrate and not due to the change in other parameters such as cellular pH.
3. Post-experiment Analysis
- Examine if sample drift has occurred during the experiment. Draw Regions Of Interest (ROI)s on the image taken at timepoint 0, then move the sliding bar at the top of the imaging window and check if the cells fall out of the ROIs in images taken later in the experiment.
- If yes, then use the tracking function to correct for the drift by first, creating masks around the cells by selecting Mask – Segment, then slide the line that specifies the cutoff to highlight the regions containing cells. Track the regions by clicking Mask – Particle tracking – Basic particle tracking.
- Re-draw the ROIs and background region when necessary.
- Export the mean intensity of the ROIs over the course of experiment. Click Statistics – Export – Ratio/timelapse data.
- When FRET efficiency and quantification of substrate concentration is not necessary, plot the time course of the intensity ratio (DonorexAcceptorem, / DonorexDonorem) as a proxy of the dynamics of the substrate. To determine the cytosolic concentration, calculate the maximal and minimal ratio change (i.e., at the saturating and absence of the substrate, respectively), Δrmax – Δrmin, by non-linear regression of Δratio (Δr) with the following equation:
Δr = [S]/(Kd+[S]) x (Δrmax – Δrmin) + Δrmin
where [S] is the substrate concentration in the cytosol. Using the Δrmax – Δrmin values obtained, saturation (S) of the sensor at a given Δr is calculated as
S = (Δr – Δrmin) / (Δrmax – Δrmin)
S can further be converted into the substrate concentration [S] using the following equation:
S = [S]/(Kd+[S])
Note: the intensity of AcceptorexAcceptorem channel should be also plotted to examine whether the experimental condition influenced the fluorescence of the acceptor directly. - When a baseline drift due to photo-bleaching is observed, perform a baseline correction. To do this, plot the intensities of donor and acceptor over time (Figure 5A). Then identify the timepoints used for the baseline, according to the delay in the perfusion system and the transport activity of the particular cell (Figure 5B).
- Calculate a polynominal fitting that best describes the baseline. Then, normalize the raw intensity from each timepoint to the baseline (Figure 5C). Calculate the intensity ratio (DonorexAcceptorem, / DonorexDonorem) from the normalized donor and acceptor intensities.
Representative Results
Typical time-course experiments are represented in Figure 2. In these experiments, FRET glutamine sensors with affinities of 8 mM (FLIPQTV3.0_8m, Figure 2A and 2B) and 100 μM (FlipQTV3.0_100 μ C and D) were co-expressed with an obligatory amino acid exchanger ASCT2 18 in cos7 cells 10. Influx of glutamine is detected as the change in fluorescence intensity ratios between the donor (mTFP1) and the acceptor (venus) (Figure 2A and 2C). Efflux of glutamine in the presence of another substrate (Ala) is also clearly demonstrated in these experiments. With both sensors, normalized fluorescent intensities change reciprocally (i.e., the donor intensity goes up as the acceptor intensity goes down, Figure 2B and 2D) in the presence of substrate, which suggest that the ratio change observed are indeed due to the change in FRET efficiency. These experiments show that the glutamine concentrations in these cells fluctuate very dynamically under the experimental condition; from the detection range of 100 μM sensor to the near-saturation concentration for 8 mM sensor.
Substrate specificities of transporters can also be examined using sensors, demonstrated in Figure 3. In these experiments the cells were pre-loaded with glutamine, and then various amino acids were added to the extracellular perfusate to examine whether those amino acids can induce glutamine efflux. As expected, ASCT2 substrates (Ala, Ser, Cys, Thr, D-serine) induced glutamine efflux (Fig. 3A), whereas non-substrate amino acids (Pro, His, Lys) did not (Fig. 3B), corroborating with previous studies 18. Usually, substrate specificity is measured by competition assays using a radio isotope-labeled substrate mixed with competing substrate, which is fairly laborious and requires a population of cells that are evenly expressing the transporter to be studied. Optical imaging exemplified here offers an alternative approach.
When no FRET efficiency changes are observed upon addition of the substrate, several reasons could be considered. One possible reason is the low uptake capacity for the substrate being tested and/or high activities of enzymes that maintain the concentration in the cells. For example, glutamine concentration change was minimal in cos7 cells that do not express ASCT2 transporter under the condition tested, even though this cell line can clearly grow in the media in which glutamine in the main nitrogen source (Figure 4). In addition, if the affinity of the sensor is either too high or too low compared to the intracellular concentration, most of the sensor proteins will remain constitutively bound or unbound respectively; hence no FRET efficiency change will be detected.
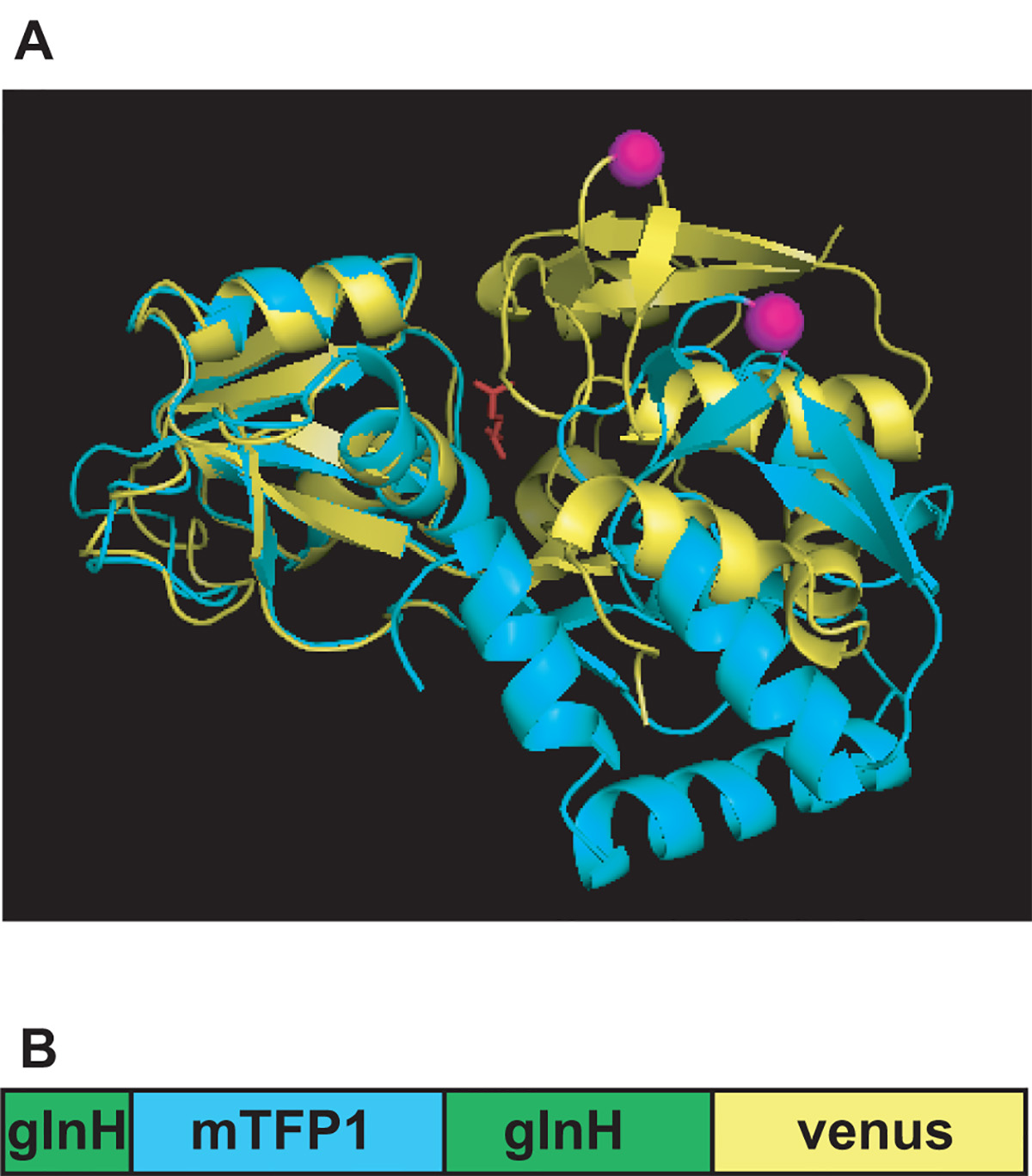
Figure 1. Configuration of a FRET glutamine sensor. (A) Open (cyan) and closed (yellow) conformation of glnH, glutamine binding protein from E.coli. The position where mTFP1 is inserted is marked in magenta. (B) Schematic representations of FLIPQTV_3.0 sensors.
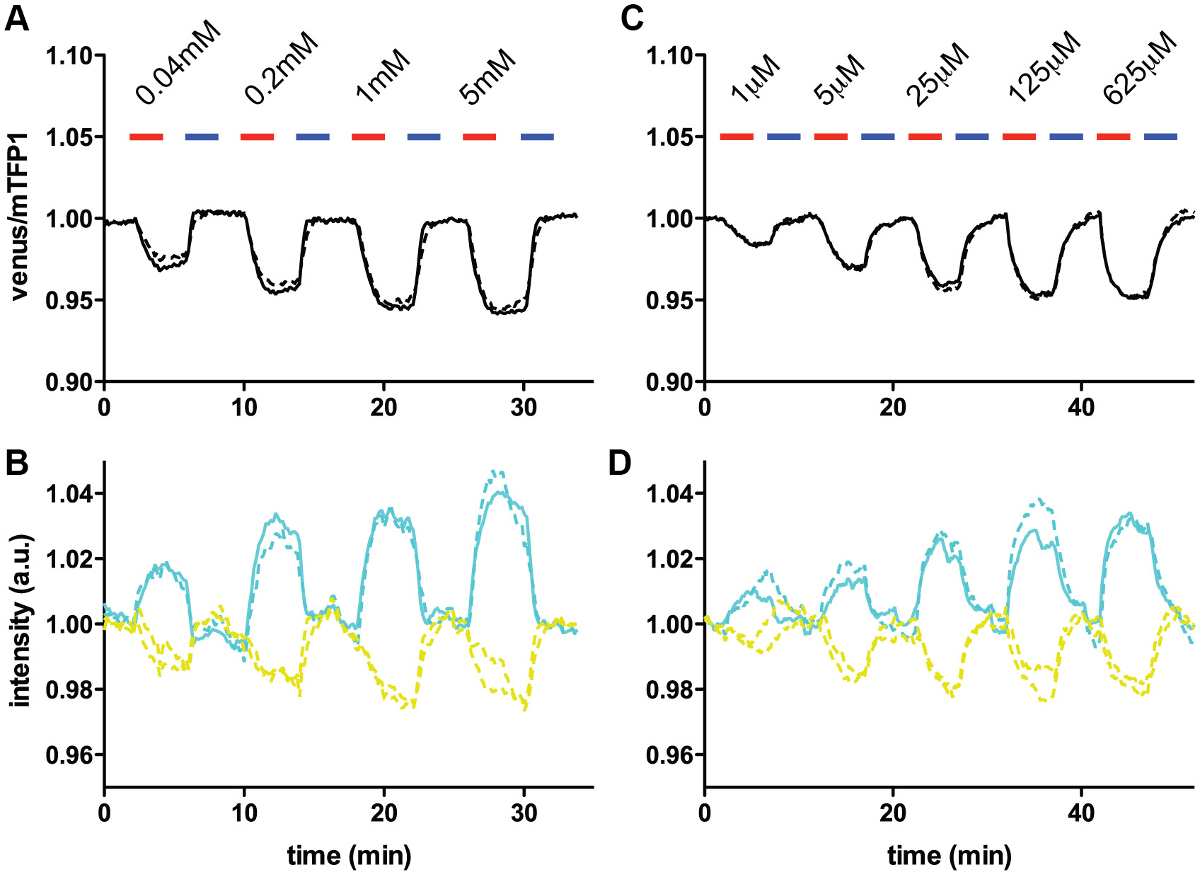
Figure 2. In vivo glutamine measurements using FLIPQ-TV3.0_8m and 100μ sensors. (A) The venus/mTFP1 ratio of cos7 cells co-expressing FLIPQ-TV3.0_8m sensor and ASCT2-mCherry. mCherry tag was used to identify the cells expressing the transporter without interfering the mTFP1 or venus emission channels. The cells were perfused with HEPES-buffered Hank’s buffer (pH 7.35). Timepoints when extracellular glutamine (red) and alanine (blue) was added to the perfusion media are indicated as boxes above the graph. Solid and dashed lines represent two individual cells measured in the same experiment. (B) The intensities of donor (mTFP1) and acceptor (venus) channels in the experiment shown in (A). The values were corrected for photobleaching and normalized to the baseline. (C) and (D) A similar experiment as in (A) and (B), performed with cos7 cells expressing the FLIPQ-TV3.0_100μ sensor. (Figure modified from 10).
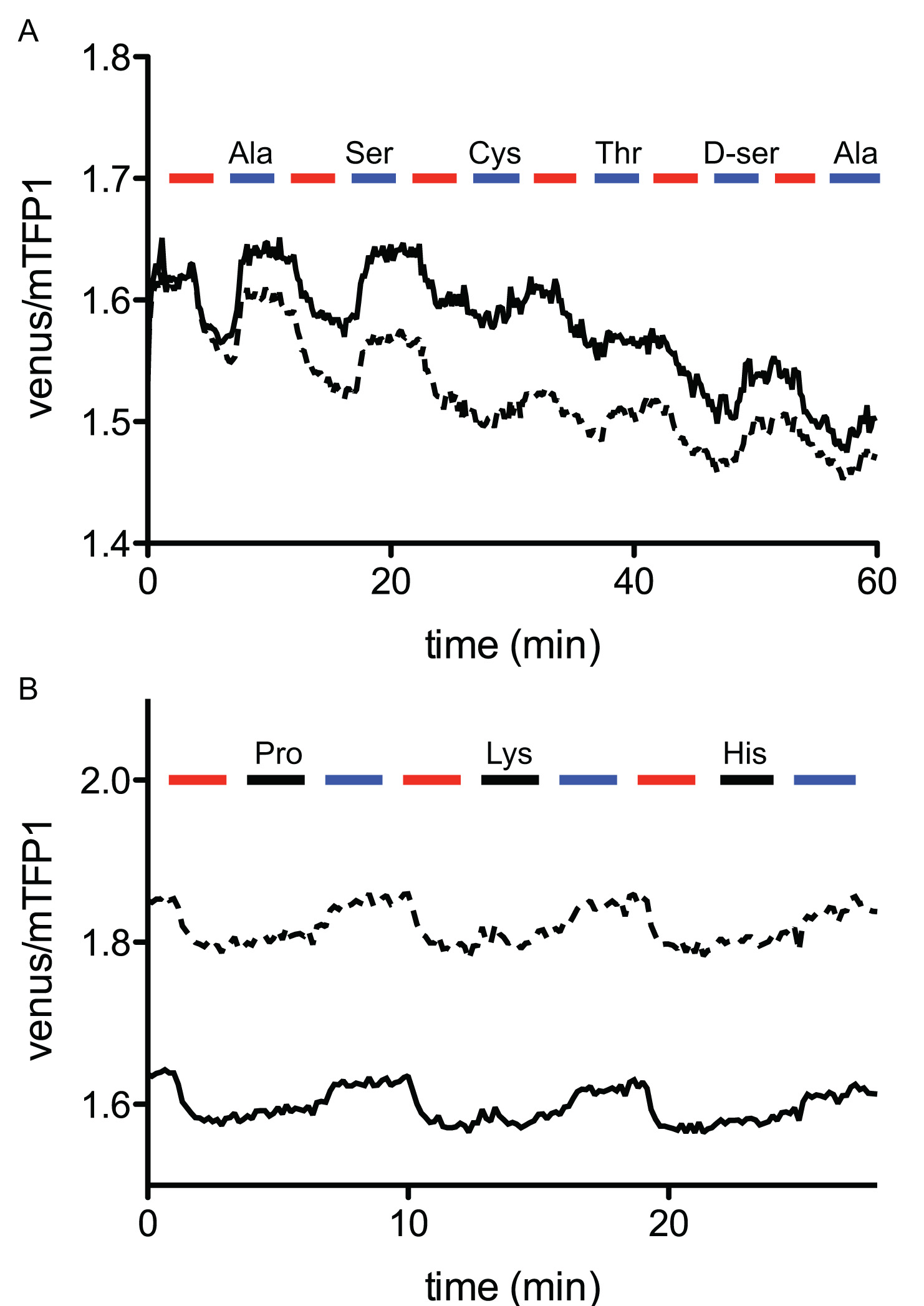
Figure 3. Elimination of cellular glutamine through the ASCT2 transporter in the presence of external amino acids, visualized using FLIPQ-TV3.0_8m sensor. (A) Cytosolic glutamine is exported by the addition of extracellular Ala, Ser, Cys, Thr, and D-ser. Timepoints when extracellular glutamine (red boxes) or other amino acids (blue boxes) were added to the perfusion media are indicated as boxes above the graph. (B) Addition of Pro, Lys, His (black boxes) does not alter cytosolic glutamine concentration, whereas the addition of Ala (blue boxes) promotes the export of glutamine. Solid and dashed lines represent two individual cells measured in the same experiment. All amino acids were added at 5 mM external concentrations. (Figure was originally published in 10).
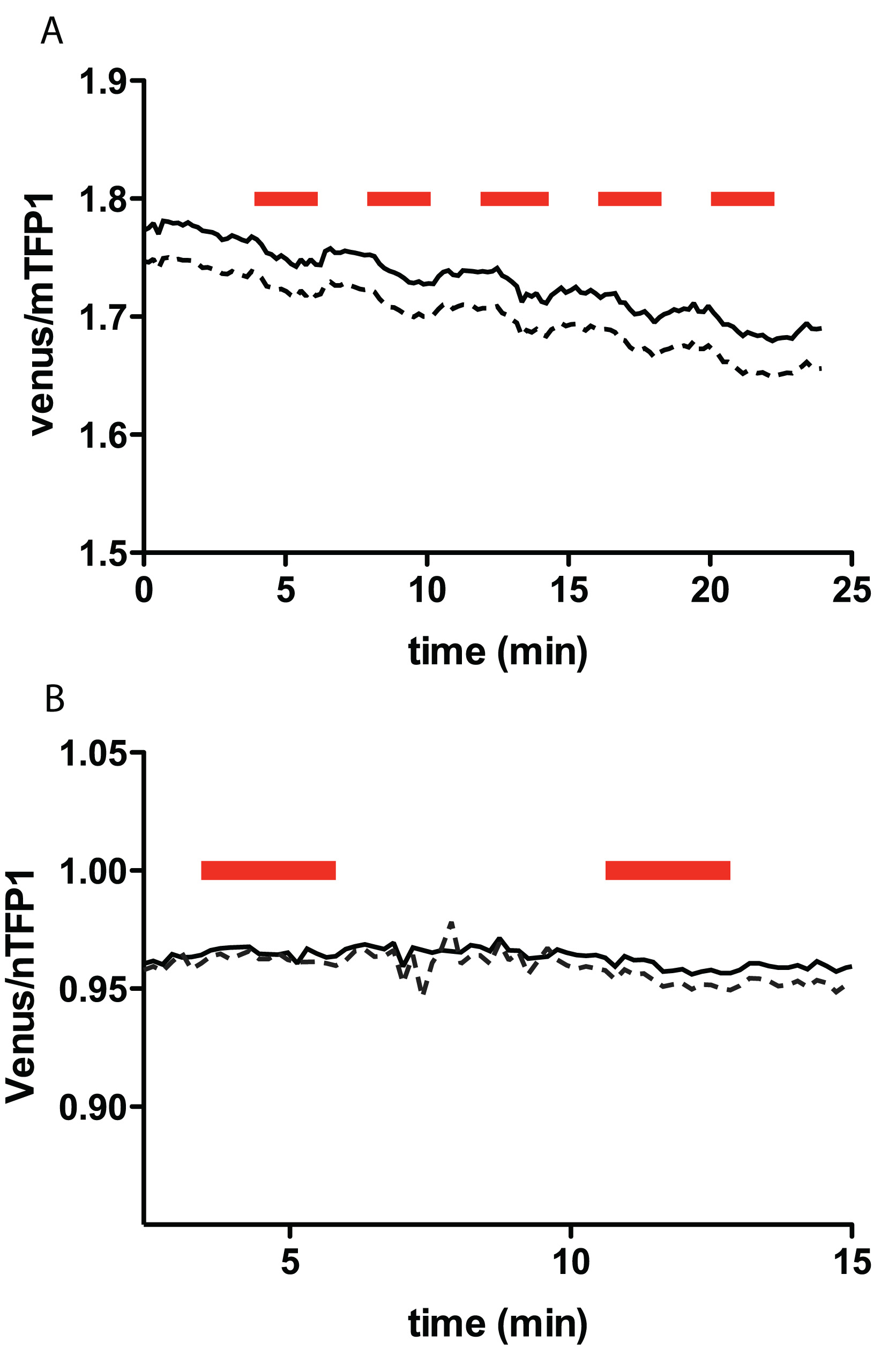
Figure 4. The venus/mTFP1 ratio of cos7 cells expressing FLIPQ-TV3.0_8m sensor (A) and 100μ sensor (B). The cells were perfused with HEPES-buffered Hank’s buffer. Timepoints when extracellular glutamine (5 mM) was added to the perfusion media are indicated as red boxes above the graph. Solid and dashed lines represent two individual cells measured in the same experiment.
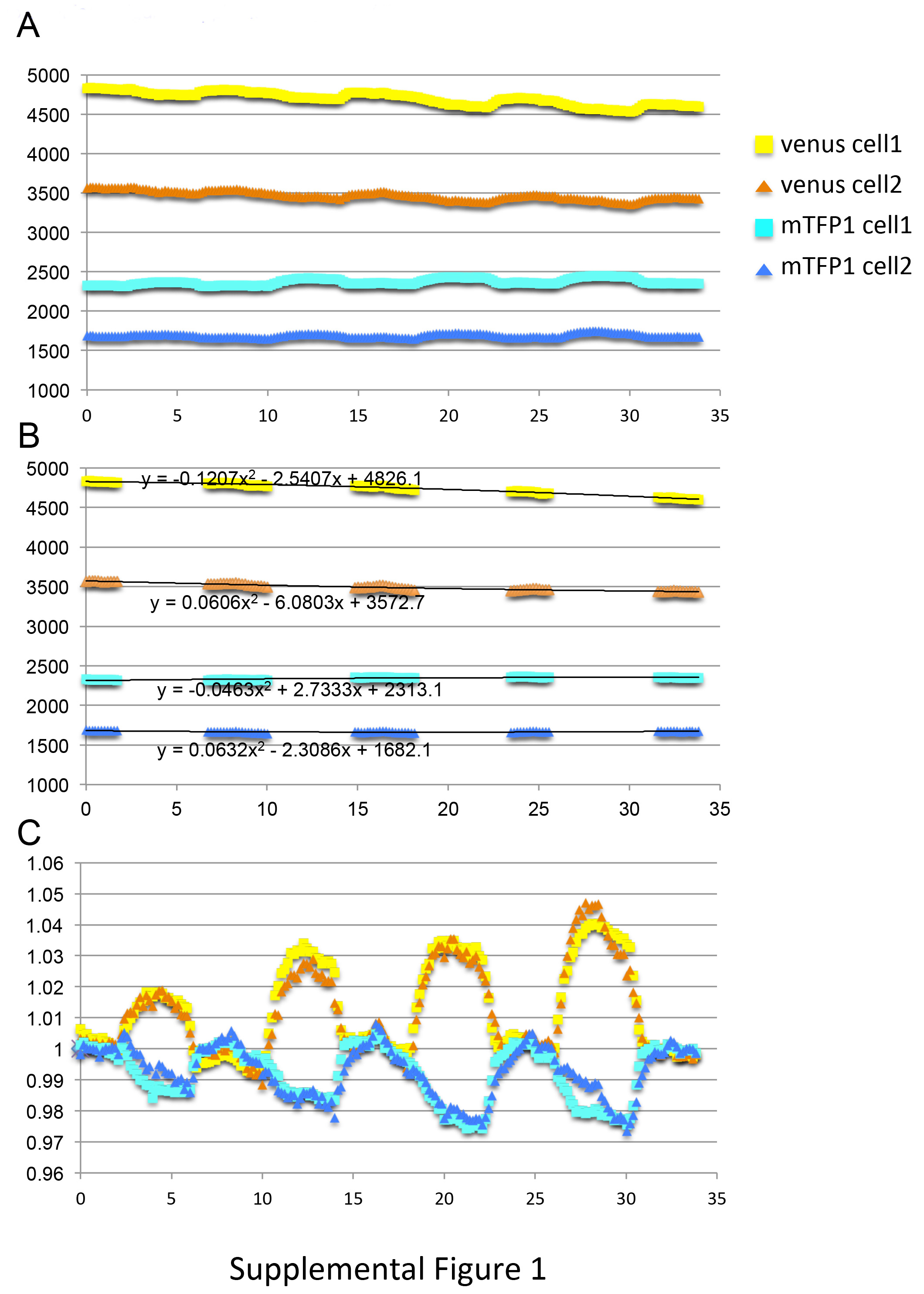
Figure 5. Correcting for photobleaching. (A) Raw intensities from two cells represented in Figure 2A and 2C. (B) Datapoints that were selected for calculating the baselines. Polynominal fitting curves are shown in the figure. (C) Intensities of channels shown in A, normalized against the baseline calculated in B. The dataset used in this figure is identical to the one shown in Figure 2A and 2C.
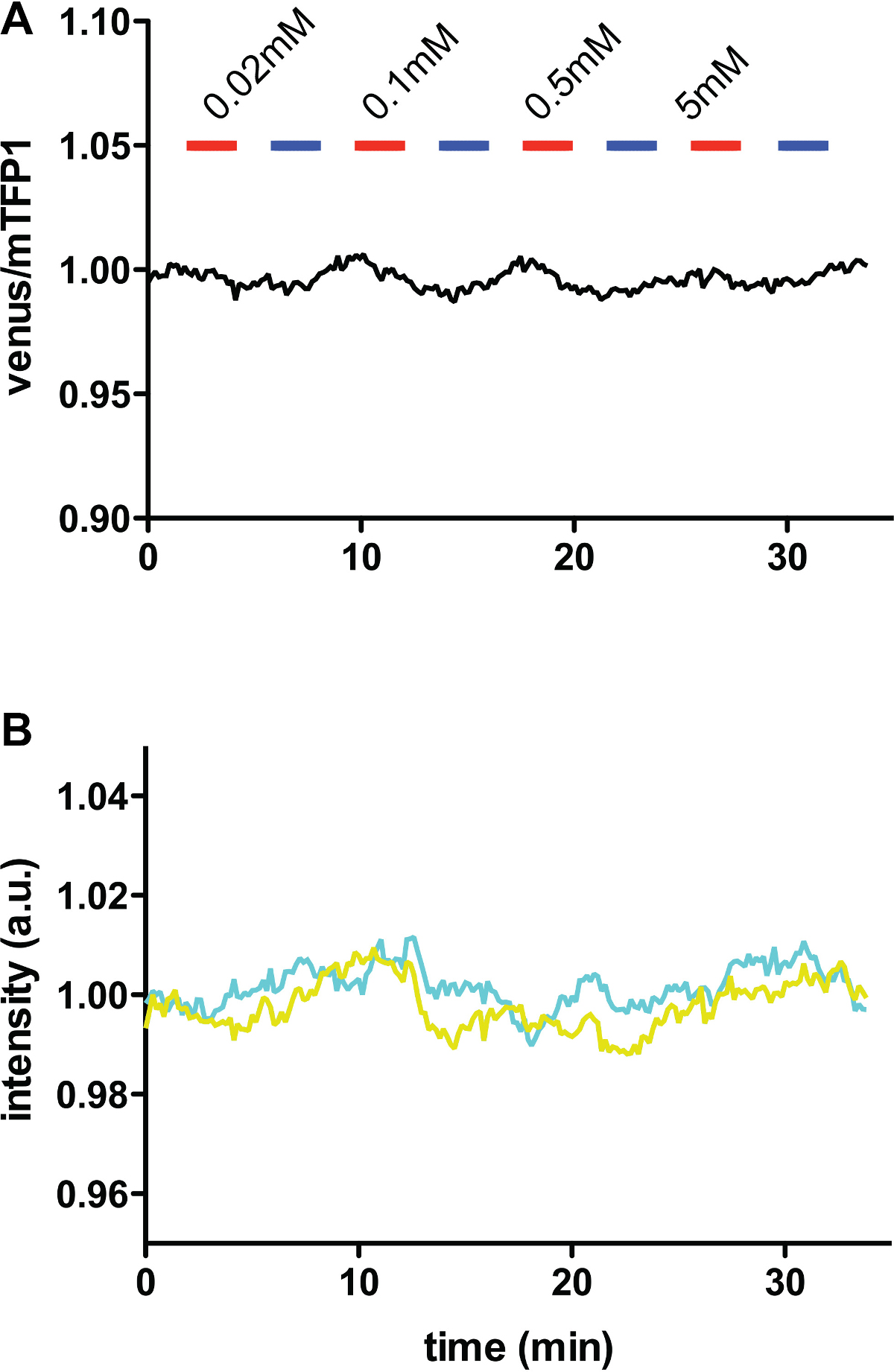
Figure 6. In vivo glutamine measurements using FLIPQ-TV3.0_1.5m sensor. (A) The venus/mTFP1 ratio of cos7 cells co-expressing FLIPQ-TV3.0_1.5m sensor and ASCT2-mCherry. mCherry tag was used to identify the cells expressing the transporter without interfering the mTFP1 or venus emission channels. The cells were perfused with HEPES-buffered Hank’s buffer (pH 7.35). Timepoints when extracellular glutamine (red) and alanine (blue) was added to the perfusion media are indicated as boxes above the graph. Solid and dashed lines represent two individual cells measured in the same experiment. (B) The intensities of donor (mTFP1) and acceptor (venus) channels in the experiment shown in (A). The values were corrected for photobleaching and normalized to the baseline.
| Time (min) | Solution A | Solution B | Solution C | Solution D | Solution E | Solution F |
| 0 | valve on | |||||
| 2 | valve off | valve on | ||||
| 4 | valve on | valve off | ||||
| 6 | valve off | valve on | ||||
| 8 | valve on | valve off | ||||
| 10 | valve off | valve on | ||||
| 12 | valve on | valve off | ||||
| 14 | valve off | valve on | ||||
| 16 | valve on | valve off | ||||
| 18 | valve off | valve on | ||||
| 20 | valve on | valve off | ||||
| 22 | valve off | valve on | ||||
| 24 | valve on | valve off | ||||
| 26 | valve off | valve on | ||||
| 28 | valve on | valve off | ||||
| 30 | valve off | valve on | ||||
| 32 | valve on | valve off | ||||
| 34 | valve off |
Table 1. Example of the perfusion protocol used in this experiment. Solution A: 9.7 g HANK salt (H1387), 0.35 g NaHCO3, 5.96 g HEPES to 1 L pH adjusted to 7.35 with NaOH. Solution B: Solution A + 0.04 mM Gln. Solution C: Solution A + 0.2 mM Gln. Solution D: Solution A + 1 mM Gln. Solution E: Solution A + 5 mM Gln. Solution F: Solution A + 5 mM Ala.
Discussion
The success of imaging experiments depends upon a few critical factors. One of these factors is the affinity of sensors used, as discussed above. Absolute concentration of the substrate in the subcellular compartment of interest, however, is often unknown. Therefore we recommend trying multiple sensors with staggered affinities to find the one that works best under the desired experimental condition. For example, in our case we transfected the cos7 cells with glutamine sensors with 1.5μ, 100μ, 2m, and 8m (Figure 3 and data not shown).
Another important factor is the expression level of sensor proteins. The level of expression required for the detection varies between, experiments, For example, when a short-lived event (e.g., single action potential) needs to be observed a higher expression level could become necessary 19. Therefore, in most cases, the level required needs to be empirically determined. If the target exists in very low concentration in the cell, perturbation of the endogenous processes due to target depletion can become an issue 20. An independent assay to assess such possibility, if available, is therefore desirable. In the experiments shown in this article, protein expression driven by CMV promoter was found to be sufficient. In situations where promoters with much weaker activities need to be used, adjustments to increase the signal intensities might become necessary. For example, compromise between the signal intensity and photo-bleaching during the exposure need to be reached. In addition to longer exposure time, factors as such as the brightness of the fluorophores, maximal intensity change of the sensor, binning, and the sensitivities of CCD camera can be changed to enhance the signal intensity.
When no ratio change is observed under the experimental condition even though the above two criteria are likely to be met, it is possible that the cellular homeostasis for the given metabolite or ion is strong enough to mask the transport activity (see the result section). In such cases, cell lines with different biochemical activities might be required as a background to detect the transporter activities 10,21.
While these sensors allow one to monitor concentration dynamics of the substrate at much higher spatiotemporal resolution than extraction-based methods, determination of absolute concentration requires further experimental steps and considerations. Usually, substrate concentration is calculated on the assumption that the sensor’s affinity, usually calculated by titrating purified sensor protein, is unchanged when expressed in the cell. The concentration is calculated using the following single binding isotherm,
[S] = Kd x (r – r min) / (r max – r)
[S] is the substrate concentration, Kd is the dissociation constant determined in vitro, r is the acceptor/donor ratio observed at a given timepoint, r min is the acceptor/donor ratio when sensors are in the apo form, and r max is the acceptor/donor ratio when all sensors are bound to the substrate. R min and r max are influenced by parameters such as the concentration of sensors, imaging settings used and the composition of the cellular milleu, and hence need to be measured in situ. To determine r min and r max values, experimental conditions that allows modification of intracellular substrate concentration becomes necessary. For example, selective ionophores (e.g., ionomycin for Ca2+) 22 or a perforating reagent such as digitonin 23 can be used to equilibrate intracellular substrate concentration with the external concentration.
One of the limitations in the type of experiments demonstrated in this protocol is the low throughput; the analysis is limited at analyzing one metabolite in a relatively small (<10) number of cells. Technological advances are being made, however, to overcome such limitations. For example, with the advance in automated, high-content screening, it is now possible to use genetically encoded sensors in combinations with small molecule or RNAi library; cells expressing a biosensor can be grown in a high-throughput format (e.g., 96- or 384-well), then individual wells can be treated with either siRNA or chemicals and imaged. Such experiments allow identification of siRNA constructs or chemicals that disrupt the biological process that can be observed by the biosensor 24 25. Another exciting recent advance includes the development of a time-resolved microfluidic flow cytometer, which allows high-throughput detection of FRET efficiency change at a cellular level 26. Such technical advances will aid discoveries of new drug candidates and new components in biological processes.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
This work was supported by NIH grant 1R21NS064412, NSF grant 1052048 and Jeffress Memorial Trust grant J-908.
Materials
| Inverted fluorescent microscope | Olympus | IX81F-3-5 | An equivalent inverted fluorescent microscope from other suppliers would also appropriate. |
| Exitation filter (mTFP1) | Omega Optical | 3RD450-460 | band width 450-460 nm |
| Exitation filter (Venus) | Chroma | ET500/20 | band width 490-510 nm |
| Emission filter (mTFP1) | Chroma | HQ495/30m | band width 475-505nm |
| Emission filter (Venus) | Chroma | ET535/30m | band width 520-550nm |
| Dichroic mirror (FRET channels) | Chroma | 470dcxr | long pass, 470nm cut off |
| Dichroic mirror (YFP channel) | Chroma | 89002bs | passes 445-490nm, 510-560nm, 590-680nm |
| mCherry filter set | Chroma | 49008 | excitation 540-580nm, long pass dichroic with 585nm cut off, emission filter 595-695nm |
| Light source | Olympus | U-LH100L-3-5 | LED- or halogen light source that produces stable light intensity, mercury lamps are not recommended |
| CCD camera | Qimaging | Rolera-MGi EMCCD | |
| apochromatic fluorescence objective | Olympus | ||
| perfusion system, ValveBank II | AutoMate Scientific | [01-08] | Other perfusion systems that allow fast solution exchange would also work |
| Laminar-flow chambers | C&L Instruments | VC-MPC-TW | Other larminar-flow chambers would also work |
| Chambered slide | Lab-Tek | 154534 | For open-chamber experiments |
| perfusion pump | Thermo Scientific | 74-046-12131 | For open-chamber experiments |
| Software supporting ratiometric measurements | Intellegenent Imaging Innovation | Slidebook 5.5 | |
| Laminar flow biosafety cabinet | ESCO | LA-3A2 | |
| Isotemp CO2 Incubator | Thermo Scientific | 13-255-25 | |
| Dulbecco's MEM (DMEM) | Hyclone | SH30243.01 | |
| Cosmic Calf Serum | Hyclone | SH3008703 | |
| penicillin/streptomycin | Hyclone | SV30010 | |
| Serum-free medium for transfection (OPTI-MEM I) | Invitrogen | 31985 | Used with Lipofectamine 2000 |
| Poly-L-Lysine solution | sigma | P4707 | |
| 25mm circular glass cover slips | Thermo Scientific | 12-545-102 | in case VC-MPC-TW is used |
| Lipofectamine 2000 | Invitrogen | 11668027 | Other transfection reagents can also be used |
| Solution A (Hank's buffer) | – | – | 9.7g HANK salt (Sigma H1387), 0.35g NaHCO3, 5.96g HEPES to 1L, pH adjusted to 7.35 with NaOH |
| Solution B | Solution A + 0.04mM Gln | ||
| Solution C | Solution A + 0.2mM Gln | ||
| Solution D | Solution A + 1mM Gln | ||
| Solution E | Solution A + 5mM Gln | ||
| Solution F | Solution A + 5mM Ala |
Riferimenti
- Haussinger, D. Regulation of hepatic ammonia metabolism: the intercellular glutamine cycle. Adv Enzyme Regul. 25, 159-180 (1986).
- Haussinger, D. Hepatic glutamine metabolism. Beitr Infusionther Klin Ernahr. 17, 144-157 (1987).
- Watford, M., Chellaraj, V., Ismat, A., Brown, P., Raman, P. Hepatic glutamine metabolism. Nutrition. 18, 301-303 (2002).
- Mustroph, A., et al. Profiling translatomes of discrete cell populations resolves altered cellular priorities during hypoxia in Arabidopsis. Proc Natl Acad Sci USA. 106, 18843-18848 (2009).
- Sasagawa, Y., et al. Quartz-Seq: a highly reproducible and sensitive single-cell RNA sequencing method, reveals non-genetic gene-expression heterogeneity. Genome Biol. 14, R31 (2013).
- Sabatini, B. L., Oertner, T. G., Svoboda, K. The life cycle of Ca(2+) ions in dendritic spines. Neuron. 33, 439-452 (2002).
- Okumoto, S., Jones, A., Frommer, W. B. Quantitative imaging with fluorescent biosensors. Annu Rev Plant Biol. 63, 663-706 (2012).
- Newman, R. H., Fosbrink, M. D., Zhang, J. Genetically encodable fluorescent biosensors for tracking signaling dynamics in living cells. Chemical reviews. , 111-3666 (2011).
- Okumoto, S. Imaging approach for monitoring cellular metabolites and ions using genetically encoded biosensors. Curr Opin Biotech. 21, 45-54 (2010).
- Gruenwald, K., et al. Visualization of glutamine transporter activities in living cells using genetically encoded glutamine sensors. PLoS One. 7, e38591 (2012).
- Chaudhuri, B., et al. Protonophore- and pH-insensitive glucose and sucrose accumulation detected by FRET nanosensors in Arabidopsis root tips. Plant Journal. 56, 948-962 (2008).
- Chen, T. W., et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature. 499, 295-300 (2013).
- Jiang, D. W., Zhao, L. L., Clapham, D. E. Genome-Wide RNAi Screen Identifies Letm1 as a Mitochondrial Ca2+/H+ Antiporter. Science. , 326-147 (2009).
- Barnett, N. L., Pow, D. V., Robinson, S. R. Inhibition of Muller cell glutamine synthetase rapidly impairs the retinal response to light. Glia. 30, 64-73 (2000).
- Rothstein, J. D., Tabakoff, B. Alteration of Striatal Glutamate Release after Glutamine-Synthetase Inhibition. J Neurochem. 43, 1438-1446 (1984).
- Gu, S., Villegas, C. J., Jiang, J. X. Differential regulation of amino acid transporter SNAT3 by insulin in hepatocytes. J Biol Chem. 280, 26055-26062 (2005).
- Palmada, M., Speil, A., Jeyaraj, S., Bohmer, C., Lang, F. The serine/threonine kinases SGK1, 3 and PKB stimulate the amino acid transporter ASCT2. Biochem Bioph Res Co. 331, 272-277 (2005).
- Bröer, A., et al. The astroglial ASCT2 amino acid transporter as a mediator of glutamine efflux. J Neurochem. 73, 2184-2194 (1999).
- Wallace, D. J., et al. Single-spike detection in vitro and in vivo with a genetic Ca2+ sensor. Nature. 5, 797-804 (2008).
- Haugh, J. M. Live-cell fluorescence microscopy with molecular biosensors: what are we really measuring. Biophys J. 102, 2003-2011 (2012).
- San Martin, A., et al. A genetically encoded FRET lactate sensor and its use to detect the Warburg effect in single cancer cells. PLoS One. 8, e57712 (2013).
- Palmer, A. E., Tsien, R. Y. Measuring calcium signaling using genetically targetable fluorescent indicators. Nat Protoc. 1, 1057-1065 (2006).
- Ponsioen, B., et al. Detecting cAMP-induced Epac activation by fluorescence resonance energy transfer: Epac as a novel cAMP indicator. Embo Rep. 5, 1176-1180 (2004).
- Joseph, J., Seervi, M., Sobhan, P. K., Retnabai, S. T. High throughput ratio imaging to profile caspase activity: potential application in multiparameter high content apoptosis analysis and drug screening. PLoS One. 6, e20114 (2011).
- Jiang, D., Zhao, L., Clapham, D. E. Genome-wide RNAi screen identifies Letm1 as a mitochondrial Ca2+/H+ antiporter. Science. 326, 144-147 (2009).
- Ma, H., Gibson, E. A., Dittmer, P. J., Jimenez, R., Palmer, A. E. High-throughput examination of fluorescence resonance energy transfer-detected metal-ion response in mammalian cells. J Am Chem Soc. 134, 2488-2491 (2012).

