Fast and Simplified Method for High Through-put Isolation of miRNA from Highly Purified High Density Lipoprotein
Summary
MicroRNAs play an important regulatory role and are emerging as novel therapeutic targets for various human diseases. It has been shown that miRNAs are carried in high density lipoproteins. We have developed a simplified method to rapidly isolate purified HDL suitable for miRNA analysis from human plasma.
Abstract
Small non-coding RNAs (miRNAs) have been implicated in a variety of human diseases including metabolic syndromes. They may be utilized as biomarkers for diagnosis and prognosis or may serve as targets for drug development, respectively. Recently it has been shown that miRNAs are carried in lipoproteins, particularly high density lipoproteins (HDL) and are delivered to recipient cells for uptake. This raises the possibility that miRNAs play a critical and pivotal role in cellular and organ function via regulation of gene expression as well as messenger for cell-cell communications and crosstalk between organs. Current methods for miRNA isolation from purified HDL are impractical when utilizing small samples on a large scale. This is largely due to the time consuming and laborious methods used for lipoprotein isolation. We have developed a simplified approach to rapidly isolate purified HDL suitable for miRNA analysis from plasma samples. This method should facilitate investigations into the role of miRNAs in health and disease and in particular provide new insights into the variety of biological functions, outside of the reverse cholesterol transport, that have been ascribed to HDL. Also, the miRNA species which are present in HDL can provide valuable information of clinical biomarkers for diagnosis of various diseases.
Introduction
MicroRNAs are endogenous non-coding tiny RNA species that are highly conserved and are considered key players in the regulation of various biological processes by degrading or repressing specific target messenger RNAs1. Because miRNAs act intracellularly they have been explored as tissue-derived biomarkers which led to the discovery of tissue-specific functions of these miRNA. However, miRNAs are also found extracellularly either associated with proteins or in exosomes/micro vesicles that effectively can shield them from degradation by extracellular RNases2. More recent studies have shown that the protective effect of HDL may not be closely linked to its capability to promote cholesterol efflux but rather to its non-cholesterol cargo, in particularly as a circulating miRNAs carrier 3, 4. These miRNAs may not only modulate lipid metabolism but are also associated with anti-inflammatory, antioxidant and antithrombotic effects of the HDL-miRNA complex 5, 6.
To further explore the role of miRNAs carried in HDL particles, a simple and easy protocol needs to be established for miRNA extraction from isolated highly purified HDL for use in clinical routine. Numerous methods have been described to isolate HDL. These methods are either very time consuming or require large volume of plasma that may require sample pooling, extensive dialysis for desalting isolated lipoproteins and they do not completely remove exosomes as a source of miRNAs3, respectively. Here we describe a simple and rapid method that can isolate miRNA from highly purified HDL utilizing small volume of blood samples on a larger scale. We believe that this method may serve as good reference to promote research into the role of circulating miRNAs and in particular the role of HDL in facilitating communication between various cells and organs.
Protocol
1. Collection of Blood Samples
- Collect fasting peripheral venous blood samples into 10 ml plastic tubes containing anticoagulant Ethylenediaminetetraacetic acid (EDTA) (which has several advantages over other anticoagulants) by standard venipuncture of a prominent vein in the antecubital fossa.
- Centrifuge the blood samples at 1,600 x g for 20 min at 4 °C in a tabletop centrifuge to obtain plasma free of red blood cells and small amounts of RNA.
- Sequentially centrifuge the supernatant at 3,000 g (4 °C) in a swinging bucket rotor for 10 min to remove WBC & Platelets and then additional 15 min to remove remaining cell debris respectively.
- Measure the density of the plasma using a densitometer at RT as per manufacture instructions.
NOTE: Adjustment of the density (d = 1.023 g/ml) with 0.9% saline solution may be required after removal of exosomes but prior to density gradient ultracentrifugation.
2. Exosome Removal from Plasma
- Remove the circulating exosomes that have a density similar to HDL and represent a quantitatively significant source of miRNA3.
- Do this by adding 252 µl exosome precipitation solution to 1 ml plasma followed by incubation for 30 min at 4 °C. To pellet out the exosomes, centrifuge the mixture for 30 min at 1,500 g at 4 °C.
- To isolate HDL, transfer 1 ml of the resulting supernatant to a polycarbonate thick-walled ultracentrifuge tube for further processing with density gradient ultracentrifugation (see below).
3. Density Gradient Ultracentrifugation (Figure 1).
- To seperate HDL use a 3-step process employing a floor ultracentrifuge with a fixed-angle rotor operating at 448,811 x G and 8 °C, respectively.
- Prepare three different density solutions sequentially and fresh for each isolation.
- Prepare Solution A (isolation of VLDL, d =1.006 g/ml) by dissolving 11.4 g NaCl (NaCl: 0.195 mol), 0.1 g EDTA2Na and 1 ml 1N NaOH in 1,000 ml of autoclaved-distilled water. Then add an additional 3 ml of autoclaved-distilled water.
- Prepare Solution B (isolation of LDL, d = 1.182 g/ml) by adding 25.2 g NaBr to 100 ml solution A (NaCl 0.195 mol, NaBr 2.44 mol).
- Prepare Solution C (isolation of HDL, d=1.470 g/ml) by mixing 78.8 g NaBr with 100 ml of solution A (NaCl 0.195 mol, NaBr 7.7 mol). Confirm the appropriate density at RT using a densitometer. Keep all solutions at 4 °C until further use.
4. Isolation of VLDL
- Mix 1 ml of plasma (average density = 1.023 g/ml) and nuclease free 200 µl of Fat Red 7B in a 6.5 ml polycarbonate thick-walled ultracentrifuge tube.
- Then carefully layer 5 ml of solution A on top of the mixture. If needed, add additional Fat Red 7B on top of solution A to balance the weight of each tube. Centrifuge for 2 hr (acceleration – 5), (deceleration – 7).
NOTE: During centrifugation, the lipoproteins are accumulated as a band at their equilibrium density regions. - At the end of the run observe 2 layers. Remove 1.5 ml of the VLDL fraction representing the top layer and store at 4 °C.
- Finally, using a pipette transfer 4 ml from the bottom of the tube containing the LDL, HDL, albumin and fatty acid fraction to a new polycarbonate tube for LDL isolation.
5. Isolation of LDL
- Mix 2 ml of solution B and 100 µl nuclease free Fat Red 7B into the tube containing the LDL and HDL fraction (section 4), respectively.
- Then centrifuge out for 3 hr (acceleration 9, deceleration 7). Thereafter, remove 1.5 ml of the LDL fraction representing the top layer and keep at 4 °C or store at -80 °C. Finally, transfer 4 ml from the bottom of the tube containing the HDL fraction to a new polycarbonate tube.
6. Isolation of HDL
- Mix 2 ml of solution C, 100 µl nuclease free Fat Red 7B and 15 µl of 98% β-mercaptoethanol into the tube containing the HDL fraction, respectively.
- Centrifuge for 3 hr (acceleration 9, deceleration 7). Thereafter remove 2 ml of the HDL fraction representing the top layer and either keep at 4 °C or store at -80 °C.
7. Desalting and Concentration of Lipoprotein Fractions
- To avoid interference with subsequent agarose gel electrophoresis and PCR, remove excessive salt added during density gradient ultracentrifugation using centrifugal filter devices with the appropriate molecular weight cutoff (3K tube for VLDL and 10K tube for LDL/HDL) as described by the manufacturer's instructions.
- Briefly, after adding 2.5 ml cold PBS (137 mM NaCl, 2.7 mM KCL, 8 mM Na2HPO4, 2 mM KH2PO4; pH 7.4) centrifuge the entire VLDL fraction collected-during density gradient ultracentrifugation at 4 °C for 60 min using a swinging bucket rotor.
- Desalt the LDL fraction twice with 10 ml ice cold PBS for 30 min each. Next, Use 13 ml ice cold PBS twice for desalting the HDL fraction. The higher PBS volume is necessary to improve mobility with agarose gel electrophoresis. After centrifugation, remove the lipoprotein containing solutes and keep at 4 °C or stored at -80 °C.
8. Agarose Gel Electrophoresis
- Perfrom lipoprotein agarose gel electrophoresis employing the kit with minor modifications of the manufacturer's instructions as follows.
NOTE: This step is just to assess the quality and purity of the concentrated lipoprotein samples.- Briefly, obtain 6 µl of the desalted lipoprotein fraction with density gradient ultracentrifugation and load onto a pre-cast lipoprotein gel. Use human lipoprotein standards for VLDL, LDL and HDL as size reference. Carry out electrophoresis at RT at 100 V for 60 min using Rep Prep buffer.
- Dry the gel for 10 min and then stain for 10 min at RT with Fat Red 7B. Destain the gel in a mixture of methanol-water 75:25 (v/v) and dry again for 5 min.
9. RNA Extraction and Purification
- Perfrom isolation of miRNA by purified human HDL using the serum/plasma miRNA isolation and purification kit.
- Briefly, add 1 ml of RNA lysis reagent to 200 µl of purified HDL, mix with a vortexer and then incubated for 5 min at RT to ensure complete dissociation of nucleoprotein complexes and inactivation of RNases.
- Then spike 3.5 µl of synthetic Caenorhabditis elegans microRNA (cel-miR-39; 1.6 x 108 copies/µl) into the mixture. Then carry out RNA extraction according to the manufacturer's instructions.
- Perform purification of extracted-miRNA with elute spin columns as per manufacturer's instructions. Measure the concentration of miRNA from purified HDL with a spectrophorometer.
NOTE: Elution of miRNA from the spin columns employed 16 µl of RNase-free water.
10. Reverse Transcription (RT-PCR)
- Isolate 100 ng of the miRNA from HDL spiked with synthetic miRNA (cel-miR-39) and reverse-transcribed in a 20 µl reaction volume employing the reverse transcription kit and according to the manufacturer's instructions.
- Perform appropriate controls without template miRNA (NTC) and without reverse transcriptase enzyme mix (NRT).
11. Real-time PCR (qRT-PCR)
- Perfrom Real-time PCR in a total volume of 20 µl with 2 µl of a 1:2 dilution of the cDNA, 10 µl PCR mix, 2 µl universal primer, 2 µl of miRNA primers and 4 µl RNase-free water.
- Run the reaction in 96-well plates at 95 °C for 15 min, followed by 45 cycles of 94 °C for 15 sec and 55 °C for 30 s and an extension phase at 70 °C for 30 s.ec Perfrom all reactions in triplicates.
- Next, Calcuate relative quantities of miRNA by using the 2-ΔΔCt method after normalization to the synthetic housekeeping gene as per manufacturer's instructions .
Representative Results
Isolation of High Density Lipoprotein After Removal of Exosomes
To obtain miRNA from highly purified HDL it is necessary to remove exosomes that represent a source of miRNA contamination7. This was done prior to density gradient ultracentrifugation with a commercially available kit. For practical purposes a three step standard density gradient ultracentrifugation protocol developed by commercial company was modified (Figure 1). This protocol requires centrifugation with a fixed-angle rotor at a speed of 140,000 rpm which is substantially faster than commonly used protocols with centrifugation forces up to 54,000 rpm3, 8, 9, and 10. Employing a widely available T-1270 rotor with a maximum force of 70,000 rpm, centrifugation was initially carried out with polyallomer tubes which have failed to resist the centrifugation forces. To avoid collapse of tubes, polycarbonate tubes were successfully used. Centrifugation time is critical for lipoprotein oxidation and potentially miRNA degradation11. Therefore several different centrifugation times, ranging from a total of 8 to 96 hr were tested. Furthermore temperature at which centrifugation was carried out was adjusted based on centrifugation time and force, respectively.
Purity of the High Density Lipoprotein Fractions
Purity of isolated high density lipoprotein fractions were checked with agarose gel electrophoresis. Figure 2. illustrates a typical electrophoresis result for the HDL subtraction isolated by gradient ultracentrifugation. It clearly showed that HDL was devoid of any contamination of VLDL and exhibits typical α-mobility. The absence of any α-migrating lipoproteins in the VLDL and LDL fractions demonstrates the complete recovery from HDL during the ultracentrifugation step. The HDL fraction, however, also showed some trace of β-mobility, a known electrophoretic banding pattern due to contamination with lipoproteins (Lp(a))12.
Elimination of Lp(a) from Isolated HDL
Studying miRNAs carried in HDL requires isolation of HDL of highest purity. Interference of Lp(a) with HDL potentially contributes to cross-contamination of HDL with miRNAs carried in LDL. To minimize this contamination of HDL, 15 μl of β-mercaptoethanol 10 was added to solution C during the last centrifugation step (Figure 1). Addition of β-mercaptoethanol has been shown not to change HDL density properties10. As displayed in Figure 3., addition of β-mercaptoethanol resulted in the absence of any b-migrating lipoproteins consistent with very effective removal of Lp (a).
Purification of miRNA from HDL
Isolation of miRNA was initially attempted employing lysis reagent which is commonly used for RNA extraction from blood. Although this method resulted in a very good RNA yield (91.45 ng/µl) but after spectrophotometric analysis showed unsatisfactory RNA purity. Additional purification of the isolated RNA by removing phenol improved the purity but the RNA yield was now significantly low. Next, another kit was tested which showed acceptable RNA purity (260/280 nm ratio of 1.7) but the RNA yield was 26-fold lower compared with the lysis reagent (3.45 vs. 91.45 ng/µl). The best results for acceptable RNA purity with a good yield were obtained with the miRNeasy serum/plasma kit (48.2 ng/µl; 260/280 nm ratio of 1.6) and therefore this extraction procedure was subsequently employed for the detection of miRNA from the isolated HDL lipoprotein fraction.
To demonstrate the feasibility to quantify miRNAs carried in human HDL purified with this method, we chose to amplify miR-223. This miRNA was chosen because miR-223 was identified in cargo of HDL3 and was shown to repress HDL cholesterol uptake5. The Figure 4A. shows a typical log plot of amplification curves comparing baseline threshold and threshold cycle values after optimization of the PCR reaction for the miR-223 and the spiked-in Cel-mir-39. This synthetic gene was used as internal control as there are no reports of known normalization control for miRNA in plasma. In addition, several potential established endogenous housekeeping genes like RNU6-2, RNU-48, HY3 could not be detected and SNORD95 could be detected in purified HDL, but, the difference in Ct values of miR-223 and SNORD95 is less than five (data not shown). Melting curve analysis illustrated in Figure 5. clearly shows a distinct single peak consistent with amplification of a selective miRNA in the preceding PCR. As illustrated in Figure 4B., both miR-223 and the reference Cel-mir-39 could consistently be detected in all six pro bands. Relatively small variations among all individuals were observed for Cel-mir-39 compared with miR-223. These findings support isolation of HDL at high purity to allow detection of its miRNA cargo.
Detection of miR-223 in Purified HDL
Real time quantitative PCR method was used to quantify double hairpin-structure miRNA precursors to single strand miRNA. This method needs a forward/reverse gene-specific primers and a thermostable reverse transcriptase to convert the hairpin structure of the miRNA to cDNA. The cDNA was subsequently amplified and quantified using real-time qPCR with the help of SYBR green detection. MicroRNA from serum, plasma and purified HDL plasma can be accurately profiled using the miRNA RT PCR system.
Our experimental product of mean miR-223 gene expression showed a Ct value of 30.9 in purified HDL and its corresponding negative controls were above cut-off 35 (NTC = 38.1 Ct value, NRT and NAC were not amplified). In this experiment, we saw in the purified HDL samples the formation of Primer-dimer with low melting temperature (73 °C in miR-223 and 75.5 °C in Cel-miR-39) and higher Ct value than the specific miRNA assays (Table 1). The Primer-dimers in Table 1. occur probably because of high concentration of primers in the solution. This occurs mostly when primer molecules attach to each other after 30 PCR cycles 13. This allows them to anneal to other primer molecules and in effect, become linear mini-templates and it will cause higher background and may lead to a generation of Ct value < 40 for NTC (No template control) samples.
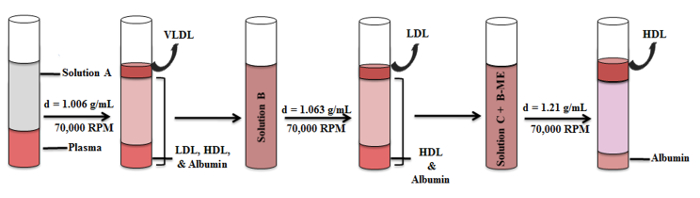
Figure 1. Schematic Representation of the HDL Isolation Procedure. HDL was prepared by density gradient ultracentrifugation in a series of three centrifugation steps. Distribution of lipoprotein bands and intermediate fractions in the density gradient are illustrated. Please click here to view a larger version of this figure.
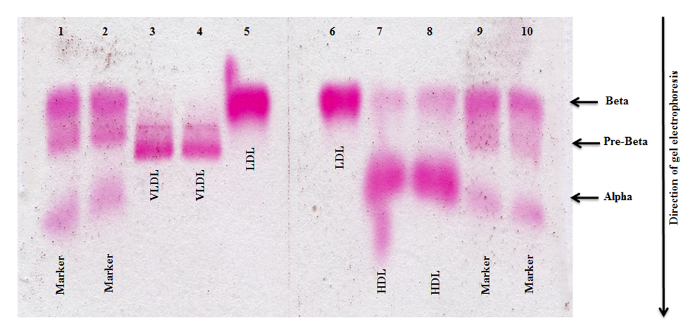
Figure 2. Agarose Gel Electrophoresis of Lipoproteins Isolated Without β-mercaptoethanol. VLDL, LDL and HDL fractions isolated by density gradient ultracentrifugation without adding β-mercaptoethanol to solution C demonstrate the presence of Lp (a) in the HDL fraction. Lanes 1, 2, 9, and 10 each represent the size marker; Lanes 3/4, 5/6 and 7/8 represent VLDL, LDL and HDL respectively. Please click here to view a larger version of this figure.
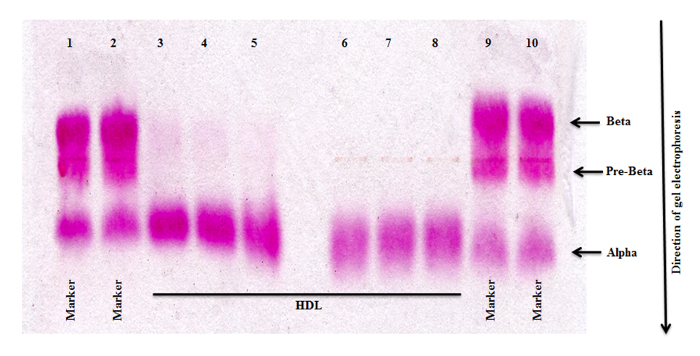
Figure 3. Agarose Gel Electrophoresis of HDL Isolated with β-mercaptoethanol. Adding β-mercaptoethanol to solution C before the last centrifugation step resulted in the removal of Lp(a) from the HDL fraction. Lanes 1, 2, 9 and 10 each represent the size marker; Lanes 3-8 represent HDL. Please click here to view a larger version of this figure.
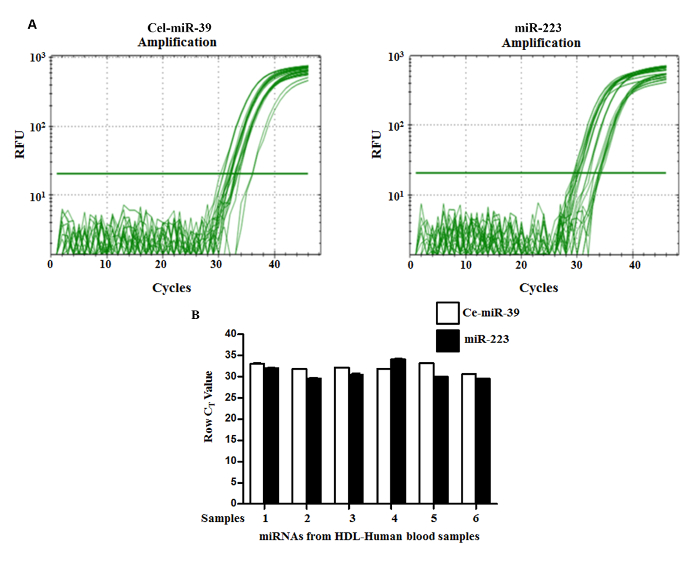
Figure 4. Amplification Plot of Two Different miRNAs and Expression of Cel-miR-39 and miR-223. (A) Real-time quantitative PCR employing isolated HDL was carried out as described. The PCR was run for 45 cycles and the point at which the curve intersects the threshold is the CT-Value. The data show the expression of Cel-miR-39 (left panel) and miR-223 (right panel), respectively. The horizontal line represents the detection threshold. A cycle number of 35 was set as cutoff for positive amplification. (B) Representative real-time PCR data from isolated HDL obtained from 6 samples are shown. Raw mean Ct values are shown for Cel-miR-39 and miR-223, whose expression levels vary less than 2 Ct value in purified HDL plasma fraction between 6 samples, each from a different donor. Expression profiling was performed using the PCR System and the Human Serum & Plasma miRNA qRT-PCR. Please click here to view a larger version of this figure.
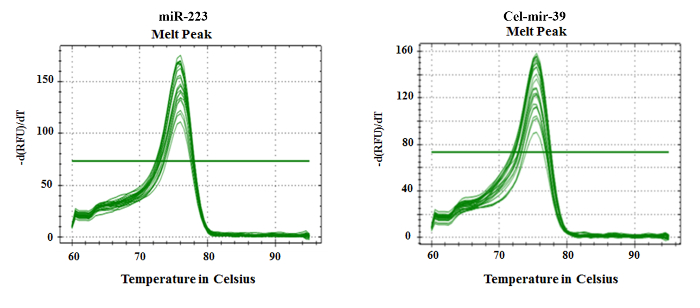
Figure 5. Melting Curves for Cel-miR-39 and miR-223. As illustrated, the presence of a single peak indicates specific amplification of Cel-miR-39 (right panel) and miR-223 (left panel), respectively. Please click here to view a larger version of this figure.
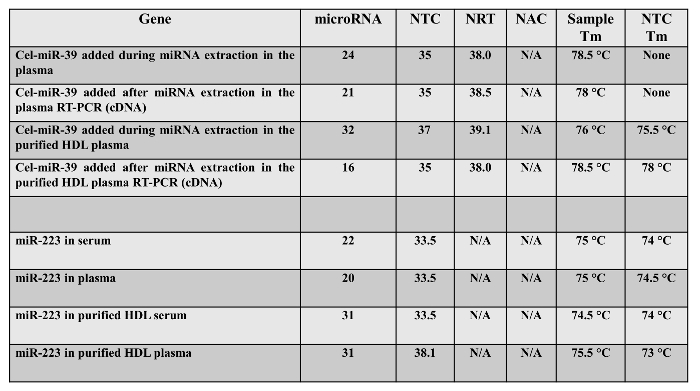
Table 1. Row Ct value of negative and positive control of synthetic Cel-miR-39 and miR-223 characterized in serum, plasma, cDNA, and purified HDL fraction. NTC (No template control), NRT (No reverse transcription enzyme), NAC (No amplification control, only water and reagents of qRT-PCR).
Discussion
Identification of novel biomarkers from blood will aid in the clinical diagnosis and prognosis of various diseases. MicroRNAs have known to possess all the qualities of biomarkers and have been shown in various studies 14-17. In this study we have demonstrated rapid and simple easy method to isolate miRNA from plasma HDL. Conventional density gradient ultra-centrifugation method of isolation of VLDL, LDL and HDL depends on accurate sampling of plasma, precise preparation of the buffer solution, measurement of density and quantitative transfer of bottom lipoprotein fractions8. There are numerous methods that have been described to isolate HDL8 -11. These methods are often either very laborious or require large volume of plasma and extensive dialysis for desalting isolated lipoproteins and do not completely remove exosomes and lipoproteins as a source of miRNAs 3. To address these shortcomings we have established this simple and fast method of isolation of miRNA from plasma HDL.
The key goal and primary purpose of this study was to separate and isolate HDL-miRNA complex by ultracentrifugation without RBCs, buffy coat, exosomes, secretory vesicles, VLDL, LDL and Lipoproteins (a) interference, then isolate miRNAs from purified HDL. Contamination and interference of any of these components with HDL will alter the expected experimental results and miRNA profiling data. Isolation of DNA, RNA and miRNAs from blood is a very difficult task due to its complex nature. It has always been posed with lot of technical challenges to nucleic acids (including miRNAs) extraction and purification compared to cells, tissues and organ samples18. Also there are relatively less miRNA genes present in circulating blood cells. Blood is a known carrier for exosomes, secretory vesicles and HDLs along with free circulating miRNAs, which are relatively less18. Therefore, larger amount of blood plasma sample volume is required for processing and isolation of HDL-miRNA complex. In addition, the short nature of the miRNAs and their target sequences, makes it even difficult to achieve sufficient specificity with standard PCR oligonucleotide technologies. The cellular components of blood are always know to secrete miRNA and mRNA in response to change in external environment including temperature, microorganisms and toxins or stress. Also presence of nucleases, RNases, circulating proteins and other enzymes in blood will affect these miRNA isolation.
The circulating miRNA amplification also lacks a well-established known housekeeping genes like β-actin or GAPDH for data normalization while using the ΔΔCt method of qRT-PCR. This again poses one of the major hurdle for miRNA profiling from circulating blood serum or plasma19. Commonly used short non coding snoRNAs and snRNAs are rarely expressed in blood and this makes it further difficult in utilizing them as internal control genes. To solve these demerits of snoRNAs and snRNAs and to avoid any technical difficulties we used Serum/Plasma synthetic Spike-In control in the assays as internal control gene as followed according to the manufacturer protocol. It helps in normalization for any nonspecific amplification and changes occurred during miRNA purification and PCR reaction18. Even in the case of degradation of miRNA or absence of any specific miRNA amplification signals, the spike-in control should amplify in qRT-PCR reactions and give an expected signal with a reasonably constant and uniform Ct value. Based on the spike in control amplification along with three negative controls and positive controls we got good optimal data validation for miR-223 gene. This confirmed that we have got good yield of miR-223 from circulating plasma HDL from this simple method.
In conclusion, we have established a method of density gradient ultracentrifugation (DGUC) with the addition of β-mercaptoethanol for purification of miRNA from plasma HDL. The method has several advantages: (a) VLDL, LDL and HDL are separated within a short period of time by floor ultracentrifuge compare to other methods that needs 24 – 96 hr 8-11; (b) LDL and HDL dialysis, desalting/concentrating, take place within 1 hr compare with other methods that took 24 or more hours8,9; (c) Contamination of Lipoproteins is removed by β-mercaptoethanol in HDL fraction, but is not intended to be used in the LDL fraction; (d) The sample volume required is only 1 ml for all stepwise VLDL, LDL and HDL isolations compare with the other methods that needs more specimen volume; (e) It minimized oxidative damage to HDL during isolation11; (f) A maximum of 12 samples can be analyzed in a single lipoprotein separation analytical run; (g) The 250 µl sample volume of extracted HDL is enough to analyse miRNA concentration/purity, agarose gel electrophoresis, protein concentration, microarray and RT-qPCR procedures. The limitations of our method is that small percentage of HDL-miRNA may lose during the exosome precipitation step and the required plasma sample volume should be more than 200 µl to isolate HDL-miRNA.
Finally, the plasma microRNAs are not only derived from damaged or renegade blood cells in the circulation but also from healthy normal blood cells and cells from other parts of the body which includes various healthy and diseased tissues and organs affected by ongoing health status of the body 20. This present study has systematically purified mature miR-223 from isolated HDL of human plasma. This study clearly demonstrates that levels of mature miR-223 in the highly purified HDL plasma are detectable, stable, reproducible and consistent among individuals sample, thereby greatly facilitating clinical use of futurity tests for lipoproteins, liver, cardiovascular and other metabolic syndromes. Surprisingly, mature miRNAs, particularly miR-223 from HDL plasma are probably resistant to nuclease digestion and other harsh conditions which potentially explain the stability of miR-223 in purified HDL. The mechanism of resistance of miR-223 to RNases requires further study. Obviously, studying miRNA expression profiles in these purified HDL plasma would shed light on future miRNA markers in studying various maladies. These HDL associated microRNAs in the plasma can be used as a potential clinical diagnostic biomarkers and personalized medicine in the future.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
This work was supported, in whole or in part, by NIH Grants R01 AA 020758-04, U01DK 061731-13 and T32 DK 007150-38 to AJS and T32 DK 007150-38 to AA. This is original work and is not under consideration elsewhere for publication.
Materials
| Plastic Vacutainer Lavender K2EDTA tubes | Becton, Dickinson and Company | 366643 | |
| Centrifuge | Thermo Scientific, Sorvall Legend X1R | 75004261 | |
| Densito 30PX densitometer | Mettler Toledo | MT51324450 | |
| ExoQuick solution | Invitrogen | 4484451 | |
| Polycarbonate thick-walled ultracentrifuge tube | Thermo Scientific | O3237 | |
| Sorvall WX100 ultracentrifuge | Thermo Scientific | 46902 | |
| Fat Red 7B | Sigma-Aldrich | 201618 | |
| β-mercaptoethanol | Sigma-Aldrich | ||
| Amicon Ultra-15 Centrifugal filter devices 10K | Millipore | UFC901008 | |
| Amicon Ultra-centrifugal filter devices 3K | Millipore | UFC800308 | |
| QuickGel Lipo kit | Helena Laboratories | 3344,3544T | |
| Human lipoprotein standards for VLDL, LDL and HDL | LipoTrol; Helena Laboratories | 5069 | |
| Rep Prep buffer | Helena Laboratories | 3100 | |
| RNeasy MinElute spin columns | Qiagen | ||
| NanoDrop 1000 analyzer | Thermo Scientific | ||
| miScript II RT Kit | Qiagen | 218161 | |
| CFX96 Touch real-time PCR detection system | BioRad | ||
| miRNeasy Serum/Plasma Kit | QIAGEN | 217184 | |
| miScript Primer Assays | QIAGEN | 141078139 | |
| miScript SYBR Green PCR Kit | QIAGEN | 218073 | |
| miRNeasy Serum/Plasma Spike-In Control | QIAGEN | 219610 |
Riferimenti
- Bartel, D. P. MicroRNAs: genomics, biogenesis, mechanism, and function. Cell. 116 (2), 281-297 (2004).
- Arroyo, J. D., et al. Argonaute2 complexes carry a population of circulating microRNAs independent of vesicles in human plasma. Proc Natl Acad Sci U S A. 108 (12), 5003-5008 (2011).
- Vickers, K. C., et al. MicroRNAs are transported in plasma and delivered to recipient cells by high-density lipoproteins. Nat Cell Biol. 13 (4), 423-433 (2011).
- Wagner, J., et al. Characterization of levels and cellular transfer of circulating lipoprotein-bound microRNAs. Arterioscler Thromb Vasc Biol. 33, 1392-1400 (2013).
- Wang, L., et al. MicroRNAs 185, 96, and 223 repress selective high-density lipoprotein cholesterol uptake through posttranscriptional inhibition. Mol Cell Biol. 33 (10), 1956-1964 (2013).
- Rayner, K. J., Moore, K. J. MicroRNA control of high-density lipoprotein metabolism and function. Circ Res. 114 (1), 183-192 (2014).
- Raposo, G. Exosomes: endosomal-derived vesicles shipping extracellular messages. Curr Opin Cell Biol. 16 (4), 415-421 (2004).
- Redgrave, T. G., Roberts, D. C., West, C. E. Separation of plasma lipoproteins by density-gradient ultracentrifugation. Anal Biochem. 65, 42-49 (1975).
- Foreman, J. R., et al. Fractionation of human serum lipoproteins by single-spin gradient ultracentrifugation: quantification of apolipoproteins B and A-1 and lipid components. J Lipid Res. 18, 759-767 (1977).
- Dong, J., et al. Serum LDL- and HDL-cholesterol determined by ultracentrifugation and HPLC. J Lipid Res. 52, 383-388 (2011).
- Tong, H., Knapp, H. R., VanRollings, A. low temperature flotation method to rapidly isolate lipoproteins from plasma. J Lipid Res. 39, 1696-1704 (1998).
- Fless, G. M., ZumMallen, M. E., Scanu, A. M. Physicochemical properties of apolipoprotein (a) and lipoprotein (a-) derived from the dissociation of human plasma lipoprotein (a). J Biol Chem. 261, 8712-8718 (1986).
- Brownie, J., et al. The elimination of primer-dimer accumulation in PCR. Nucleic Acids Res. 25, 3235-3241 (1997).
- Alton, E., Inyoul, L., Leroy, H., David, G., Kai, W. Extracellular microRNA: a new source of biomarkers. Mutat Res. 717 (1-2), 85-90 (2011).
- Stefanie, S. J. Cancer biomarker profiling with microRNAs. Nature Biotechnology. 26, 400-401 (2008).
- Prasun, J. M. MicroRNAs as promising biomarkers in cancer diagnostics. Biomarker Research. 2 (19), (2014).
- Creemers, E. E., Tijsen, A. J., Pinto, Y. M. Circulating microRNAs: novel biomarkers and extracellular communicators in cardiovascular disease?. Circ Res. 110, 483-495 (2012).
- Jonathan, S., Martin, S., Eric, L. . miRNA profiling from blood -challenges and recommendations. , (2015).
- Francesco, M., Paola, D. C., Anna, T., Jesper, T., Sergio, A., Riccardo, L. R. Normalization of circulating microRNA expression data obtained by quantitative real-time RT-PCR. Brief Bioinform. 3, 1-9 (2015).
- Chen, Y., et al. Circulating microRNAs, novel biomarkers of acute myocardial infarction: a systemic review. World J Emerg Med. 3, 257-260 (2012).

