Engineering Three-dimensional Epithelial Tissues Embedded within Extracellular Matrix
Summary
This manuscript describes a soft lithography-based technique to engineer uniform arrays of three-dimensional (3D) epithelial tissues of defined geometry surrounded by extracellular matrix. This method is amenable to a wide variety of cell types and experimental contexts and allows for high-throughput screening of identical replicates.
Abstract
The architecture of branched organs such as the lungs, kidneys, and mammary glands arises through the developmental process of branching morphogenesis, which is regulated by a variety of soluble and physical signals in the microenvironment. Described here is a method created to study the process of branching morphogenesis by forming engineered three-dimensional (3D) epithelial tissues of defined shape and size that are completely embedded within an extracellular matrix (ECM). This method enables the formation of arrays of identical tissues and enables the control of a variety of environmental factors, including tissue geometry, spacing, and ECM composition. This method can also be combined with widely used techniques such as traction force microscopy (TFM) to gain more information about the interactions between cells and their surrounding ECM. The protocol can be used to investigate a variety of cell and tissue processes beyond branching morphogenesis, including cancer invasion.
Introduction
The development of branched epithelial tissues, known as branching morphogenesis, is regulated by cell-derived, physical, and environmental factors. In the mammary gland, branching morphogenesis is an iterative process through which guided collective cell migration creates a tree-like architecture. The first step is primary bud formation from the milk ducts, followed by branch initiation and elongation1,2. Invasion of branches into the surrounding stroma is induced by the systemic release of steroid hormones at puberty. New primary buds then initiate from the ends of existing branches, and this process continues to create an epithelial tree3. Although many important biochemical signals have been identified, a comprehensive understanding of the cell biological mechanisms that guide this complex developmental process is currently lacking. Moreover, mechanistic studies on the influences of specific cues are difficult to deconstruct from experiments in vivo, as precise spatiotemporal perturbations and measurements are often not possible.
Three-dimensional (3D) culture techniques, such as whole organ culture, primary organoids, and cell culture models, are useful tools for systematically investigating the mechanisms underlying tissue morphogenesis4-6. These can be particularly useful for determining the influences of specific factors individually, such as mechanical forces and biochemical signals, on a variety of cell behaviors, including migration, proliferation, and differentiation.6 Engineered cell culture models, in particular, readily enable the perturbation of individual cells and their microenvironment.
One such culture model uses a microfabrication-based approach to engineer model mammary epithelial tissues with controlled 3D structure that consistently and reproducibly form branches that migrate collectively when induced with the appropriate growth factors. The major advantage of the model is the ability to precisely manipulate and measure the effects of physical and biochemical factors, such as patterns of mechanical stress, with high statistical confidence. This technique, together with computational modeling, has already been used to determine the relative contributions of physical and biochemical signals in the guidance of the normal development of mammary epithelial tissues and other branched epithelia7-11. Presented here is a detailed protocol for building these model tissues, which can be readily extended to other types of cells and extracellular matrix (ECM) gels, and which serves as a potential tool for the testing of therapeutics.
Protocol
1. Preparation of Solutions
- To prepare a 5 mg/ml solution of insulin, dilute the powdered insulin stock with 5 mM hydrochloric acid (HCl) in dH2O (500 mg insulin in 100 ml solvent). Prepare 100 ml solvent by adding 50 μl of concentrated HCl to 100 ml distilled water (dH2O).
- To make a 1x solution of PBS, dilute the 10x phosphate-buffered saline (PBS) stock solution to 1x with dH2O under sterile conditions.
- Prepare the polydimethylsiloxane (PDMS) elastomer solution as follows: mix the PDMS prepolymer together with the curing agent in a 10:1 (w:w) ratio.
- Prepare the cell culture medium as follows: to 500 ml of stock Dulbecco's Modified Eagle Medium: Nutrient Mixture F-12 (DMEM/F-12), add 10 ml of sterile fetal bovine serum (FBS), 500 μl of gentamicin reagent, and 500 μl of 5 mg/ml insulin.
- To prepare the 1% bovine serum albumin (BSA) in 1x PBS solution, add 1% (w:v) solution of BSA powder to 1x PBS and mix thoroughly.
- To prepare a 4 mg/ml solution of neutralized bovine type I collagen, add 50 µl 10x Hank's balanced salt solution (HBSS) buffer, 30 µl 0.1 N sodium hydroxide (NaOH), 30 µl cell culture medium, and 400 µl of stock collagen to a chilled 1.5 ml microcentrifuge tube. Mix slowly by pipetting up and down; avoid introducing bubbles. To remove any bubbles, briefly centrifuge the mixture at 4 °C.
- Prepare a fixative solution by diluting 16% paraformaldehyde 1:4 (v:v) in 1x PBS.
- Prepare a nuclei labeling solution by diluting the stock nuclei labeling solution 1:1,000 (v:v) in 1x PBS.
- Prepare a 0.3% phosphate-buffered saline and detergent (PBST) solution by mixing 1x PBS with 0.3% (v:v) detergent.
- Prepare blocking buffer by diluting goat serum 1:10 (v:v) in 0.3% PBST.
- Prepare a primary antibody solution for a particular protein of interest by diluting primary antibody (for example, rabbit anti-focal adhesion kinase (FAK)) 1:200 (v:v) in blocking buffer.
- Prepare a secondary antibody solution by diluting a fluorescent probe-conjugated antibody (for example, goat anti-rabbit) 1:1,000 (v:v) in blocking buffer.
2. Preparation of Elastomeric Stamps for 3D Micropatterning
Note: Elastomeric stamps are made with PDMS.
- Make a PDMS solution at the same ratio as described in step 1.3 and mix thoroughly. Place the mixture in a vacuum chamber for 15-30 min to remove any air bubbles that were introduced during the mixing process.
- Pour the degassed solution into a plastic weigh boat or Petri dish containing a silicon master that is lithographically patterned with features of desired geometry. For best results, cure the PDMS solution at 60 °C for ~12 hr.
Note: The silicon masters are highly customizable; this protocol uses rectangular structures with dimensions of 50 μm x 200 μm x 50 μm spaced 200 μm apart. The silicon masters can be made using standard photolithography techniques. Briefly, an epoxy-based photoresist is spin-coated on top of a silicon wafer. A mask detailing the desired stamp features is placed on top of the photoresist-coated wafer, which is then exposed to UV light. The desired features not blocked by the mask are exposed to the light and the photoresist at these locations becomes crosslinked, while the remainder of the photoresist coating is soluble and can be washed away. - After the PDMS has cured around the desired features on the silicon master, remove the PDMS and master from the plastic container. Carefully separate the PDMS from the silicon wafer and remove excess PDMS from around the imprinted features using a clean razor blade.
- Now cut the PDMS patterned with imprinted features into individual rectangular stamps (~8-mm x 5-mm) with a razor blade. Store these feature-side-up in a clean 100-mm-diameter Petri dish.
- Additionally, use the PDMS solution described in step 1.3 to make supports for the rectangular stamps.
- Using a spin coater, spread ~2-3 g of the PDMS solution evenly on a 100-mm-diameter Petri dish and cure the PDMS for ~12 hr at 60 °C as in step 2.1. Then, using a razor blade, cut the thin layer of PDMS into rectangles (~5 mm x 2 mm).
Note: Two supports are needed for each PDMS stamp made in step 2.3. The supports can be stored in the 100-mm-diameter Petri dish that contains the PDMS stamps. - Before the PDMS stamps and supports can be used for cell culture, sterilize by immersion in 70% ethanol and dry using an aspirator in a biosafety cabinet (cell culture hood).
3. Preparation of 3D Epithelial Tissues
- In a biosafety cabinet, add a drop of approximately 50 µl of 1% BSA in PBS to the top of each stamp. Place the droplet covered stamps at 4 °C for a minimum of 4 hr to ensure that the BSA adsorbs to the surface of the stamp.
- Aspirate BSA solution from the PDMS stamps.
- Wash the surfaces of the stamps twice with cell culture medium (50 µl per wash per stamp should be sufficient), aspirating after each wash.
- Handle the sterilized PDMS stamps and supports using curved stainless steel tweezers. In a biosafety cabinet, lay out one 35-mm-diameter tissue culture dish for each PDMS stamp. In each dish, lay down two PDMS supports separated by a distance slightly less than the length of the PDMS stamps.
- Using the tweezers, sterilize as many circular glass coverslips (15 mm in diameter, #1) as there are PDMS stamps using 70% ethanol. Aspirate the excess liquid from the coverslips while holding them with the tweezers. Store the washed coverslips in a separate 100-mm-diameter Petri dish.
- Dispense ~50 µl of the collagen mixture to evenly coat the surface of each PDMS stamp.
- Pick up each collagen-coated PDMS stamp with the tweezers and gently invert them. Lower the inverted stamps on top of the PDMS supports laid out in 35-mm-diameter tissue culture dished such that the collagen is between the stamp and the bottom of the tissue culture dish. Incubate the dishes at 37 °C for 30 min.
- Dispense ~50 µl of the remaining collagen mixture onto each of the circular coverslips (later, these will be placed on top of the engineered tissues to completely encapsulate them in collagen) and incubate them at 37 °C for 30 min.
- Obtain a 100-mm-diameter tissue culture dish containing epithelial cells (ex. EpH4 mouse mammary epithelial cells) at ~40% confluency. Aspirate the cell culture medium and wash once with 10 ml 1x PBS. Add 2 ml of trypsin to the cells and incubate at 37 °C for 5-10 min.
- Add 8 ml of fresh culture medium to the tissue culture dish containing trypsinized cells, detaching any remaining adherent cells from the dish. Mix gently by pipetting and move the cell mixture to a 15 ml conical tube and centrifuge at 100 × g for 5 min.
- Aspirate the supernatant from the conical tube and resuspend the cell pellet in cell culture medium. Using a hemocytometer, count the cells to determine the concentration of the suspension. Adjust the suspension volume to obtain a final concentration of ~106-107cells/ml.
- Remove the gelled collagen samples from the incubator. Using the tweezers, gently lift the PDMS stamps straight upwards to detach them from the molded collagen and discard them.
- Dispense ~30 µl of the concentrated cell suspension onto the surface of each collagen gel containing molded cavities of desired geometry. Observe the cells under a brightfield microscope with a 10X/0.25 NA objective while gently shaking the dishes side to side to promote cell settling within the cavities. The cavities should be filled within ~5 min.
- To remove excess cells from around the cavities, tilt each tissue culture dish on its side and gently dispense ~400 μl cell culture medium over the surface of the collagen gel. Aspirate the liquid and repeat the wash 1-2 more times, checking the collagen gels under the microscope in between each wash.
- After the excess cells have been cleared from around the cell-filled cavities in the collagen mold, place the tissue culture dishes into a cell culture incubator at 37 °C for 15 min. Then, using the tweezers, gently invert the collagen-coated class coverslips and place them on top of the cell-filled collagen molds such that the collagen from the coverslips forms a cap over the cell-filled cavities. Incubate the samples at 37 °C for 15 min.
- Once the collagen caps have adhered to the cell-filled collagen mold, dispense ~2-2.5 ml cell culture medium slowly over the glass coverslip on top of the gels. Culture the samples at 37 °C for 1-3 days.
Note: 24 hr after initial seeding, the epithelial tissues can be treated with growth factors such as epidermal growth factor (EGF) or hepatocyte growth factor (HGF) to induce branching.
4. Immunofluorescence and Image Analysis
- Aspirate the cell culture medium from the tissue culture dishes and add enough fixative solution to cover the cell-containing gels. Incubate the dishes at room temperature for 15 min on a shaker at 200 rpm.
- Aspirate the fixative, fill the tissue culture dishes with 1x PBS, and incubate at room temperature for 15 min on a shaker at 200 rpm. Repeat twice for three total washes.
- To label nuclei: Aspirate the PBS from the tissue culture dish and replace with Hoechst solution. Incubate at room temperature for 15-20 min. To stain for FAK or another marker that is detectable with antibodies, skip to Step 4.5.
- Aspirate the nuclear labeling solution and wash the cell-containing gels three times with 1x PBS as in Step 4.2. Stained samples can be stored in 1x PBS at 4 °C until further use.
- To stain for a protein of interest: Aspirate the PBS from the tissue culture dishes, add 300 μl of 0.3% PBST, and incubate the sample at room temperature for 15 min.
- Aspirate the PBS, cover the gels with blocking buffer, and incubate on a shaker (200 rpm) at room temperature for ~4 hr.
- Aspirate the blocking buffer, cover the gels with primary antibody solution, and incubate on a shaker at 200 rpm overnight at 4 °C.
- Aspirate the primary antibody solution, add 0.3% PBST solution, and incubate on a shaker at 200 rpm at room temperature for 30 min. Aspirate the PBST and repeat every 30 min for 3-4 hr.
- Repeat steps 4.7 and 4.8, this time incubating with the secondary antibody solution. Wrap the tissue culture dishes with aluminum foil to prevent photobleaching of the secondary antibody. After the final wash, stained samples can be stored in 1x PBS at 4 °C until further use.
- To visualize samples, use a 10X/0.30 NA objective focused on the midplane of the epithelial tissues.
- To visualize cell nuclei, image fixed samples labeled with a nuclear marker using a 10X/0.30 NA objective under UV illumination.
- To visualize samples stained for a protein of interest, image using a 10X/0.30 NA objective on an inverted fluorescence microscope.
- To create frequency maps of protein staining of multiple tissues (typically 50 or more) of identical initial geometry, first import each of the images using standard image analysis software (see Materials) and convert them to 8-bit grayscale.
- To threshold these images, convert them to binary (i.e., halftone/black and white) by defining a grayscale cutoff point; grayscale values below the threshold become black, and those above the threshold become white. Then, combine each of the individual binary images into a single image stack.
- Next, register the stacked images (i.e., align or match them) in the analysis software. Many image analysis software packages come with a freely available registration plugin.
- Use each image in a stack as a template for the alignment of the next image, such that the images in the entire stack are aligned by propagation. Finally, overlay (project) the images in the aligned image stack based on average intensity to form a single image with a pixel frequency map. This image can then be color-coded using image editing software of choice10,11.
Representative Results
General schematic of mammary epithelial tissue microfabrication
A general schematic of the microfabrication procedure outlining the experimental work flow is shown in Figure 1. The end result is an array of epithelial tissues of identical geometry and spacing that are completely embedded within an ECM gel. A representative experiment uses EpH4 mouse mammary epithelial cells cultured in a gel of bovine type I collagen at a concentration of 4 mg/ml. To ensure the highest quality of engineered tissues, the techniques outlined in the protocol should be followed closely. Figures 2A and 2B show low and high magnification views of arrays of rectangular wells that have been molded into a type I collagen gel prior to cell seeding. The shape of the wells is determined by the shape of the features on the silicon master. It is important to lift the PDMS mold straight up from the collagen so as not to distort the cavity geometry. Figure 2C shows rectangular wells in a type I collagen gel that have been filled with mammary epithelial cells (excess cells have been washed off the surface of the collagen). In this example, each 200 μm x 50 μm rectangular well contains approximately 80-100 cells.
Addition of growth factors induces morphogenesis
24 hours after seeding, the tissues may be treated with growth factors such as HGF or EGF and cultured over several days to model branching morphogenesis. Typically, branches begin to form as early as 4 hours after growth factor stimulation. Figure 3A shows representative results from rectangular mammary epithelial tissues within a type I collagen gel 24 hours after the microfabrication procedure, after which cells have adhered to the collagen and to each other. No branches are observed prior to growth factor addition. Figure 3B shows a representative rectangular tissue that has undergone branching 24 hours after the addition of HGF at 10 ng/ml. In this case, branches occur at the ends of the tissues (as opposed to the middle), where the cells experience the highest mechanical stress9. Multiple tissues of identical initial geometry in the same gel can then be imaged to determine population averages of branch location and branch length, enabling high-throughput analysis.
Immunofluorescence staining to visualize protein localization
Immunofluorescence staining of the tissue arrays in the culture model allows us to determine protein localization within a tissue with high statistical confidence. Figure 4A shows representative results from a branching rectangular mammary epithelial tissue stained for focal adhesion kinase (FAK). Creating frequency maps of tissues of identical geometry can be used to visualize the average spatial localization of proteins of interest within the tissues, which can be compared to the localization of other proteins as well as branching activity. Figure 4B shows a frequency map of average FAK staining for 50 tissues showing FAK enrichment at the short ends of rectangular tissues, where branching typically occurs.
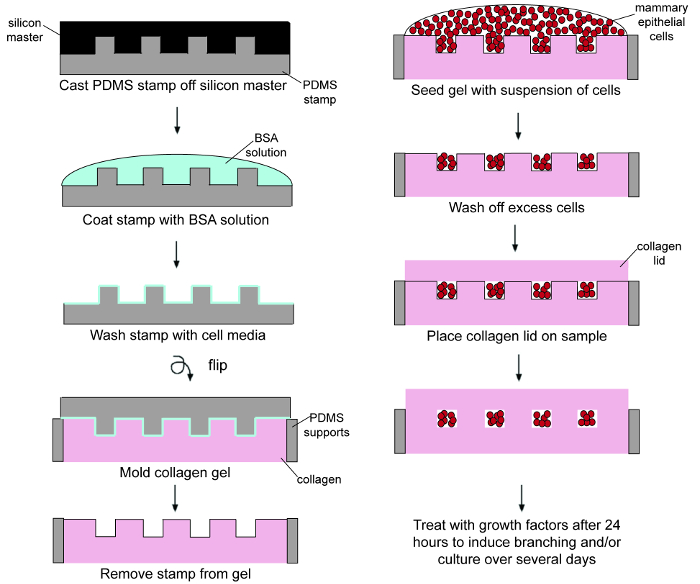
Figure 1. Schematic outlining the microfabrication procedure. Please click here to view a larger version of this figure.
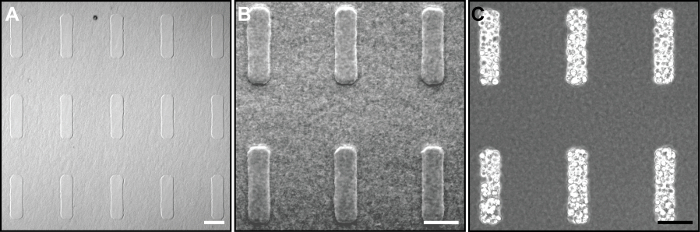

Figure 2. Images taken during the microfabrication process. (A) Low and (B) and high magnification phase-contrast images of rectangular cavities in type I collagen created using an elastomeric PDMS mold. (C) Cavities from (A) and (B) are filled with mammary epithelial cells. Scale bars, 100 μm. Please click here to view a larger version of this figure.
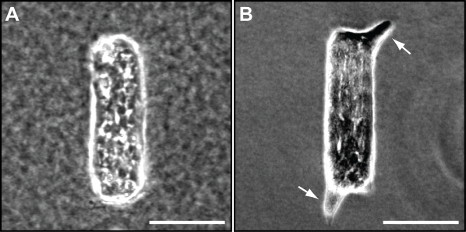
Figure 3. Microfabricated tissues undergo branching morphogenesis. (A) Phase-contrast image of a representative rectangular tissue 24 hr after microfabrication. (B) Phase-contrast image of a representative rectangular tissue that has started to undergo branching 24 hr after the addition of HGF at 10 ng/ml. White arrows indicate newly formed branches. Scale bars, 100 μm. Please click here to view a larger version of this figure.
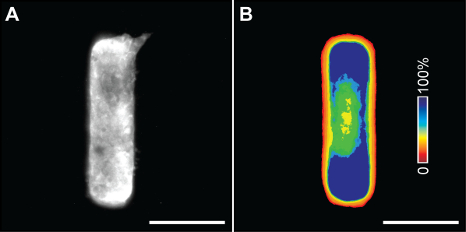
Figure 4. Immunofluorescence staining of microfabricated tissues. (A) Immunofluorescence staining for FAK in a mammary epithelial tissue after branch initiation. (B) A frequency map of average FAK staining in 50 tissues. Scale bars, 100 μm. Please click here to view a larger version of this figure.
Discussion
The protocol described above outlines a method to produce identical epithelial tissues of pre-defined shape, enabling spatial control of the mechanical stress experienced by cells in the tissue. An elastomeric mold is used to create cavities in type I collagen that are then filled with epithelial cells and covered with an additional collagen layer such that cells are completely encapsulated in a 3D collagen matrix environment. Further culture of these tissues and treatment with growth factors to induce branching from the initial architecture make this system amenable for the study of branching morphogenesis of epithelial cells. There are several critical steps in the protocol. The first is lifting the PDMS molds straight upwards from the gelled collagen prior to cell seeding. Any horizontal movement during this step will distort the cavities, and they will no longer retain the desired geometry. Then, after the cavities have been seeded with cells, it is important to carefully wash off any excess cells that remain on the surface, as these may interfere with the behavior of the patterned tissues, particularly if they are very close to the tissues. Washing too rigorously, however, may result in cells being washed out of the cavities. Lastly, in the final micropatterning step when the cell-containing collagen gels are being immersed in culture medium, it is important to add the medium slowly and directly on top of the coverslip. Adding medium to the side of the dish away from the coverslip may result in detachment of the coverslip and the collagen lid. Each of these steps needs to be performed with care to achieve the best results.
Over the past few decades, several culture models have emerged for the study of epithelial morphogenesis, many of which involve the ex vivo culture of intact organs or explants of organs. Major advantages of culturing tissue taken directly from an animal are that tissue architecture remains intact and that cell-cell interactions are maintained. However, these models are not readily amenable to studies investigating the specific effects of a particular cell type or physical microenvironment. Additionally, organs and organ explants can be fragile and difficult to culture ex vivo. Others used primary mammary epithelial cells12, in which gene expression can be more readily manipulated, though this method does not conserve in vivo tissue architecture. Moreover, like intact organs, these cells can also be difficult to culture for long periods of time5. There is a need, therefore, for more robust culture models of epithelial morphogenesis that can be more precisely controlled and readily perturbed to gather reproducible data.
A widely used 3D cell culture technique is the formation of epithelial cell clusters embedded in collagen or other ECM proteins11,13. Disadvantages of this technique, however, include the inability to precisely control or predict the location of clusters within the gel, cluster size and shape, and branching sites. In addition, it is difficult to manipulate or measure the mechanical cues experienced by cells in different regions of the tissue clusters. Engineered tissues enable better control of these parameters.
The engineered tissue protocol described here is an easy to use, versatile, and reproducible 3D cell culture method that eliminates the heterogeneity commonly found in ex vivo culture systems and 3D clusters. Such engineered cell culture models provide a platform for more precise control of physical and biochemical signals, and results can be easily combined with computational models to predict the effects of perturbations or measure cellular forces using techniques such as traction force microscopy (TFM)9,10. A caveat of the model system, however, is that it does not completely replicate the cellular, chemical, and physical microenvironment within the mammary gland in vivo. Nonetheless, essentially all parts of the system are modifiable: including the cells used (in which gene expression can also be altered by transfection), tissue geometry, ECM composition, and other biochemical factors (ex. growth factors). Primary mammary epithelial cells can also be cultured in this system; they form a lumen and undergo branching morphogenesis from the same locations as immortalized cell lines7. Moreover, the model can be modified to incorporate other cell types to more appropriately represent the native mammary gland microenvironment14, or to investigate the underlying mechanisms that give rise to any branched organ.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
This work was supported in part by grants from the NIH (HL118532, HL120142, CA187692), the David & Lucile Packard Foundation, the Camille & Henry Dreyfus Foundation, and the Burroughs Welcome Fund. A.S.P. was supported in part by a Charlotte Elizabeth Procter Honorific Fellowship.
Materials
| Polydimethylsiloxane (PDMS) | Ellsworth Adhesives | Sylgard 184 | |
| PDMS curing agent | Ellsworth Adhesives | Sylgard 184 | |
| Lithographically patterned silicon master | self-made | N/A | |
| Plastic weigh boat | Fisher Scientific | 08-732-115 | |
| 100-mm-diameter Petri dishes | BioExpress | D-2550-2 | |
| Ethyl Alcohol 200 Proof | Pharmco-Aaper | 111000200 | Make a 70% EtOH (v:v) solution by mixing with dH2O |
| Razor blade | American Safety Razor | 620179 | |
| 1:1 Dulbecco’s Modified Eagle’s Medium : Ham’s F12 Nutrient Mixture (DMEM/F12) (1:1) | Hyclone | SH30023FS | |
| Fetal Bovine Serum (FBS) | Atlanta Biologicals | S11150H | |
| 10x Hank’s balanced salt solution (HBSS) | Life Technologies | 14185-052 | |
| Insulin | Sigma Aldrich | I6634-500MG | |
| Gentamicin | Life Technologies | 15750-060 | |
| 10X Phosphate-buffered saline (PBS) | Fisher Scientific | BP399-500 | |
| Sodium hydroxide (NaOH) | Sigma Aldrich | 221465-500G | |
| Bovine type I collagen (non-pepsinized) | Koken | IAC-50 | |
| Albumin from bovine serum (BSA) | Sigma Aldrich | A-7906 | |
| Curved stainless steel tweezers | Dumont | 7 | |
| 35-mm-diameter tissue culture dishes | BioExpress | T-2881-6 | |
| 15 mL conical tubes | BioExpress | C-3394-2 | |
| 1.5 mL Eppendorf Safe-Lock Tube | USA Scientific | 1615-5500 | |
| Circular #1 glass coverslips, 15-mm in diameter | Bellco Glass Inc. | Special order | |
| 0.05% 1X Trypsin-EDTA | Life Technologies | 25300-054 | |
| Paraformaldehyde | VWR | 100503-916 | |
| Triton X-100 | Perkin Elmer | N9300260 | Detergent |
| HGF | Sigma Aldrich | H 9661 | Resuspended in dH2O at 50 mg/mL |
| Rabbit anti-mouse FAK antibody | Life Technologies | AMO0672 | |
| Goat anti-rabbit Alexa 488 antibody | Life Technologies | A-11034 | |
| Adobe Photoshop | Adobe | N/A | Used for color-coding pixel frequency maps. |
| FIJI (ImageJ) | NIH | N/A | Free image analysis software used for thresholding, registering, and overlaying images to create a pixel frequency map. The StackReg plugin was used for registering binary images. |
Riferimenti
- Affolter, M., et al. Tube or not tube: remodeling epithelial tissues by branching morphogenesis. Dev Cell. 4 (1), 11-18 (2003).
- Zhu, W., Nelson, C. M. PI3K signaling in the regulation of branching morphogenesis. Biosystems. 109 (3), 403-411 (2012).
- Sternlicht, M. D. Key stages in mammary gland development: the cues that regulate ductal branching morphogenesis. Breast Cancer Res. 8 (1), 201 (2006).
- Fata, J. E., et al. The MAPK(ERK-1,2) pathway integrates distinct and antagonistic signals from TGFalpha and FGF7 in morphogenesis of mouse mammary epithelium. Dev Biol. 306 (1), 193-207 (2007).
- Ip, M. M., Darcy, K. M. Three-dimensional mammary primary culture model systems. J Mammary Gland Biol Neoplasia. 1 (1), 91-110 (1996).
- Lo, A. T., Mori, H., Mott, J., Bissell, M. J. Constructing three-dimensional models to study mammary gland branching morphogenesis and functional differentiation. J Mammary Gland Biol Neoplasia. 17 (2), 103-110 (2012).
- Nelson, C. M., Vanduijn, M. M., Inman, J. L., Fletcher, D. A., Bissell, M. J. Tissue geometry determines sites of mammary branching morphogenesis in organotypic cultures. Science. 314 (5797), 298-300 (2006).
- Gjorevski, N., Nelson, C. M. Endogenous patterns of mechanical stress are required for branching morphogenesis. Integr Biol (Camb). 2 (9), 424-434 (2010).
- Gjorevski, N., Nelson, C. M. Mapping of mechanical strains and stresses around quiescent engineered three-dimensional epithelial tissues. Biophys J. 103 (1), 152-162 (2012).
- Gjorevski, N., Piotrowski, A. S., Varner, V. D., Nelson, C. M. Dynamic tensile forces drive collective cell migration through three-dimensional extracellular matrices. Sci Rep. 5, 11458 (2015).
- Zhu, W., Nelson, C. M. PI3K regulates branch initiation and extension of cultured mammary epithelia via Akt and Rac1 respectively. Dev Biol. 379 (2), 235-245 (2013).
- Barcellos-Hoff, M. H., Aggeler, J., Ram, T. G., Bissell, M. J. Functional differentiation and alveolar morphogenesis of primary mammary cultures on reconstituted basement membrane. Development. 105 (2), 223-235 (1989).
- Hirai, Y., et al. Epimorphin functions as a key morphoregulator for mammary epithelial cells. J Cell Biol. 140 (1), 159-169 (1998).
- Pavlovich, A. L., Manivannan, S., Nelson, C. M. Adipose stroma induces branching morphogenesis of engineered epithelial tubules. Tissue Eng Part A. 16 (12), 3719-3726 (2010).

