Synthesis of Infectious Bacteriophages in an E. coli-based Cell-free Expression System
Summary
A new generation of cell-free transcription-translation platforms has been engineered to construct biochemical systems in vitro through the execution of gene circuits. In this article, we describe how bacteriophages, such as MS2, ΦΧ174, and T7, are synthesized from their genome using an all E. coli cell-free TXTL system.
Abstract
A new generation of cell-free transcription-translation (TXTL) systems, engineered to have a greater versatility and modularity, provide novel capabilities to perform basic and applied sciences in test tube reactions. Over the past decade, cell-free TXTL has become a powerful technique for a broad range of novel multidisciplinary research areas related to quantitative and synthetic biology. The new TXTL platforms are particularly useful to construct and interrogate biochemical systems through the execution of synthetic or natural gene circuits. In vitro TXTL has proven convenient to rapidly prototype regulatory elements and biological networks as well as to recapitulate molecular self-assembly mechanisms found in living systems. In this article, we describe how infectious bacteriophages, such as MS2 (RNA), ΦΧ174 (ssDNA), and T7 (dsDNA), are entirely synthesized from their genome in one-pot reactions using an all Escherichia coli, cell-free TXTL system. Synthesis of the three coliphages is quantified using the plaque assay. We show how the yield of synthesized phage depends on the biochemical settings of the reactions. Molecular crowding, emulated through a controlled concentration of PEG 8000, affects the amount of synthesized phages by orders of magnitudes. We also describe how to amplify the phages and how to purify their genomes. The set of protocols and results presented in this work should be of interest to multidisciplinary researchers involved in cell-free synthetic biology and bioengineering.
Introduction
Over the past decade, the cell-free expression technology has been engineered to address novel applications in emergent multidisciplinary research areas related to synthetic and quantitative biology. Originally used to express proteins independent of a living organism, new cell-free TXTL systems have been developed for both basic and applied sciences1,2, broadening considerably the scope of this technology. The new generation of TXTL platforms has been designed to be user friendly, more efficient (reaching 2 mg/mL of protein synthesis in batch mode3), more versatile at the level of transcription4, and modular so as to easily integrate novel natural or synthetic functions that expand the capabilities of existing biological systems5,6. In particular, cell-free TXTL systems have become handy for the rapid prototyping of genetic programs, such as regulatory elements or small genetic circuits7,8,9, by reducing the design-build-test cycle to a few days. Remarkably, the new TXTL systems are also capable of processing large DNA programs such as the complete synthesis of coliphages10,11, demonstrating strong enough performances to support the reconstitution of active genomic DNA encoded living entities.
TXTL systems present many technical advantages compared to traditional in vitro constructive biochemical assays. Cell-free TXTL links the process of gene expression to the final product in a reduced and open environment, as opposed to the complex cytoplasm of a living cell. TXTL uses DNA to reconstruct biochemical systems in vitro, which, with modern DNA assembly techniques, is affordable and fast in addition to not requiring fastidious protein purification steps. Cell-free expression provides direct access to most of the components in the biochemical reactions, thus allowing a deeper dissection of the molecular interactions12. In a TXTL reaction, one can change the biochemical and biophysical settings at will, which is almost impossible in a living cell. Given these advantages and recent improvements, the TXTL technology is becoming more and more popular as an alternative platform for synthetic and quantitative biology. While the research community using TXTL is rapidly growing and TXTL is becoming a standard technology in bioengineering, it is essential to understand how to use such platforms so as to develop adequate practices related to the execution of TXTL reactions and to the interpretation of the results.
In this article, we describe how to use an all E. coli TXTL system to synthesize, in one-pot reactions, bacteriophages from their genome11, such as MS2 (RNA, 3.4 kb), ΦΧ174 (ssDNA, 5.4 kb), and T7 (dsDNA, 40 kb). We show how the amount of phages synthesized changes with respect to some of the biochemical settings of the reactions (magnesium and potassium concentrations). Molecular crowding, emulated through a range of PEG 8000 concentrations, has a dramatic effect on phage synthesis over several orders of magnitude. The realization of such large biochemical systems in single test tube reactions that recapitulate concurrently the processes of transcription, translation, and self-assembly, is interesting for addressing basic questions related to biology and biophysics10 (gene regulation, self-assembly), as well as for developing applications, such as repurposing phage functions to build new nanostructures13. In addition to a practical on TXTL, we provide methods for phage amplification, genome extraction and purification, and phage quantification by the plaque assay. The methods presented in this manuscript are appropriate for researchers who use E. coli extract based cell-free systems and are interested in bacteriophages.
The protocols presented in this work can be summarized as follow: 1) Phage amplification (Day 1: prepare inoculation cells, day 2: single plaque, multiple phage growth, and concentration, and day 3: purification of phage), 2) Double-stranded genome DNA extraction (phenol/chloroform extraction), and 3) Cell-free phage reaction and titer experiment (Day 1: plate host cells and make agar plates, day 2: cell-free reaction and host cell pre-culture, and day 3: host cell culture and phage titer).
Protocol
NOTE: The following amplification steps and extraction method are largely generalizable for many double-stranded DNA phage, e.g., bacteriophage T7, enterobacteria phage T4, or enterobacteria phage λ (L). They are predominantly used for a phage whose genome is not readily available for purchase from commercial sources.
1. Phage Amplification
NOTE: The single-plaque, multi-cycle (SPMC) phage production technique is well described for the T4 phage in Chen, et al.14 The following phage amplification and DNA extraction method is a generalization for use with E. coli double-stranded DNA phages, e.g., T7, T4, or L. The ultimate success of the protocol heavily depends upon the selected host cell strain's ability to withstand superinfection conditions. If the host cells are unstable under superinfection, or superinfection is never reached, and lysis is never reached during this critical phase, it is best to proceed with a confluent lysis protocol. This includes separating the cellular debris through high-speed centrifugation, and phage pelleting at ultracentrifugation speeds. All following conditions and parameters are intended as a general starting point. The optimum conditions for a local host cell line may be different; determine and adhere to the appropriate conditions.
- Prepare Inoculation Cells
- Inoculate 10 mL Luria-Bertani (LB) media in a 15 mL culture tube with E. coli host cells, e.g., B or K12.
- Place in a shaker set to 250 rpm and 37 °C. Leave overnight to grow to saturation.
- Single-plaque, multi-cycle phage growth and concentration
- To prepare a plate of competent phage plaques, warm several LB-agar plates at 37 °C for 1 h, or overnight.
- Prepare several dilutions of phage stock (e.g., T7, T4, or L) of known concentration in LB media, aiming for ~102-103 phage/mL.
- Add 100 µL of overnight growth of concentrated host cells to each plate.
- Choose volumes of the prepared phage dilutions corresponding to a range from 10-100 phage/plate. Add the required volume of phage dilutions to each plate (e.g., 100 µL of 103 phage/mL; this should produce ~100 plaques). Incubate plates at 37 °C and begin a count-up timer (+ 0 h). Incubate for 4-5 h.
- Prepare 49 mL LB media in a 250 mL flask and place in rotary shaker set to 250 rpm and 37 °C.
- At + 2.5 h, dilute cells 50x by adding 1 mL of saturated cell culture from overnight growth into a pre-warmed flask containing 49 mL LB medium. Once the cells reach log growth (+ 4-5 h), measure absorbance at 600 nm (OD600). Determine the concentration of cells using the conversion OD600 of 1.0 = 8 x 108 cell/mL.
- Dilute to 2 x 107 cell/mL using additional LB growth media. Remove the agar plate containing the phage plaques from the incubator. Immediately use the end of a sterile Pasteur pipette to core and remove a single plaque. Blow the plaque into the diluted cells and incubate at 37 °C for an additional 2 h (+ 6-7 h).
- Test for full infection by taking a 500 µL sample into a 1.5 mL tube, adding 10 µL chloroform (CHCL3), and immediately vortexing at high speed. Observe if the solution has clarified. Once cells are fully infected, incubate for another 2 h (+ 8-9 h).
NOTE: If the cells are fully infected, they will rapidly lyse (<2 min) and clarify the solution. Other outcomes are considered in the Discussion Section below. The cells will be superinfected but will not lyse up to this point. If lysis begins, the addition of DNase and harvesting by centrifugation must begin immediately to prevent phage degradation. - Add DNase to 5 µg/mL and centrifuge cells at 8,000 x g at 4 °C to pellet. Discard the supernatant. Resuspend the pellet in 10 mL of 1x TRIS magnesium chloride (TM) (50 mM Tris at pH 7.8, 10 mM MgCl2) buffer with 5 µg/mL DNase. Add 500 µL of CHCL3 and vortex at high speed to lyse the cells. Clarify by centrifugation at 12,000 x g for 10 min at 4 °C. Decant the supernatant into a 15 mL conical tube and store at 4 °C.
2. Purification of Phage
NOTE: Sucrose purification is largely dependent on the size of the phage. Considerations for the mass of the phage to be isolated will have to be made and adjustments to gradient conditions performed. Final viral titers of over 1013 phage/mL are easily achievable with this method.
- Prepare 12 mL of 5% and 45% w/v sucrose in 1x TM buffer.
- Prepare 4x 5-45% sucrose gradients by pipetting 2.5 mL of 45% sucrose into 4 ultracentrifuge tubes. Top with ~ 2.6 mL of 5% sucrose, stopping when liquid reaches 2 mm from the rim of the tube.
NOTE: Place the pipette tip into contact with surface of the sucrose solution and very slowly dispense liquid. The lighter sucrose solution will float on top of heavier, forming a distinct boundary. A properly layered solution will have ~1 mm interface if held against background light. - Mix the gradients by tilt-tube rotation using a gradient forming instrument for 43 s at 86° at 23 rpm. If not immediately needed, cover with paraffin film and store at 4 °C.
- Remove 500 µL from the top of each prepared sucrose gradient with a 1 mL pipettor and balance to within ± 0.002 g.
- Pool phage suspensions from all tubes (step 1.2.9). Using a 1 mL pipette, add 500 µL of the suspension by bringing the tip into contact with liquid surface and very slowly dispensing the volume onto the top of the sucrose gradient, being careful to not mix through rapid pipetting. Centrifuge at 70,000 x g for 20 min at 4 °C.
NOTE: Balance the prepared gradients to within ± 0.002 g. Smaller phages may take up to 1 h. - Remove the phage bands using a sterile syringe and blunt cannula.
NOTE: When viewed from the side, the phage band will be thick and milky compared to the surroundings, approximately 5 mm in height, and situated half way down the centrifuge tube.- Submerge a blunt cannula into the solution until the tip is centered on the very upper edge of the phage band. Remove the band by drawing on the syringe plunger until the majority of phage band has been removed.
- Add the sucrose volume with the suspended phage to 3-4 ultracentrifuge tubes. Add no more than 1.5 mL/tube. Fill to ~3-4 mm from the top of the tube with cold 1x TM. Cover with paraffin film and invert to mix. Place back in an ultracentrifuge and spin at 145,000 x g for 1 h at 4 °C to pellet. Smaller phages may take up to 2 h.
- Quickly pour off the supernatant and drain upside down on disposable wipes. Wipe the inside of tube with sterile cotton swab to remove excess supernatant.
- Divide 200-400 µL cold 1x TM across all pellets and resuspend overnight at 4 °C; no shaking is required.
NOTE: If the pellet is not clean, e.g., stringy bits of membrane, DNA, etc., pool and perform additional centrifugation in a microcentrifuge at 17,000 x g. - The day before making titers, prepare a culture of E. coli host cells in 10 mL LB growth media and leave in a shaking incubator set to 250 rpm and 37 °C overnight. Incubate the appropriate amount of culture plates at 37 °C for at least 1 h before making titers of the phage stock.
NOTE: The number of plates to incubate depends on the number of dilutions of phage stock to be plated: each unique dilution of phage stock requires two culture plates (e.g., four different dilutions of phage stock require eight culture plates). - Prepare 100 mL of top agar by adding 2.5 g LB and 0.6 g bacto-agar to a 100 mL bottle. Dissolve the solids in deionized water in a volume of 100 mL and autoclave. After autoclave, slowly invert the bottle 6-8 times to homogenize the agar throughout the solution. Place the bottle in 45 °C water bath for 15-20 min to equilibrate the temperature.
- Prepare serial dilution of phage stocks with the LB solution.
- Add 990 µL LB to 10 µL phage stock for a 100-fold dilution. Vortex to homogenize. For a dilution factor of 10, add 100 µL of phage stock to 900 µL LB. Vortex to homogenize.
- Repeat the above dilution series as many times as needed to obtain a countable number of plaques (10-100 plaques).
NOTE: To achieve this, dilute the phage stock 2-3 orders of magnitude less than the expected phage yield in terms of phage/mL reaction. For example, if a particular phage stock contains 1011 infectious phage/mL, plate the phage stock at a 108-fold or 109-fold dilution.
- Prepare an area for phage titers by assembling 14 mL culture tubes (the number of culture tubes equals the number of phage stock dilutions to be plated). Place the diluted phage stock samples from step 2.12 and the host bacterial cells from step 2.10 on ice.
- Prepare a master mix by pipetting 5.25 mL top agar (step 2.11), 220 µL diluted phage samples (step 2.12), and 50 µL host bacterial cells (step 2.10) to a 14 mL culture tube. Cap the culture tube and vortex at high speed.
- Store the remaining phage solution at 4 °C.
- Retrieve two culture plates (step 2.10) from the 37 °C incubator. Add 2.5 mL of master mix slowly to the center of the first plate (without bubbles). Gently rotate the plate by hand to distribute the master mix evenly so that it spans the entire culture plate. Repeat with the second plate. Wait 20 min to ensure solidification of the top agar. Incubate plates at 37 °C for 4-7 h.
- Count the plaques and determine the phage concentration for each reaction sample.
NOTE: Plaques will appear as 1-2 mm clear circles in the opaque carpet of host cells. See Figure 1 for representative results. A successful phage production will result in final concentrations of 1012-1013 phage/mL.
3. Double-stranded Genome DNA Extraction
NOTE: Dilution, shaking, and centrifugation steps must be optimized to develop a thick, solid protein boundary layer between the phenol and aqueous phases. This generates the highest purity genomes that are free of protein contaminants. The initial dilution is dependent on the final phage titer. An extremely high titer phage stock (≥1013 phage/mL) may need a 10-20-fold dilution before extraction, while low titer stocks (~1010-1011 phage/mL) may only need a 2-fold or none. If it is difficult to form a solid protein layer in subsequent steps or the aqueous suspension is too sticky to be workable due to the high DNA concentration, consider a higher dilution of phage stocks before continuing extraction. During any genome-handling step, it is important to use wide-bore pipette tips when pipetting any aqueous phases. Many phage genomes are quite large and easily fragmented through pipette shearing. Additionally, any vortexing should be expressly avoided as this will severely shear the genomes.
- Dilute a portion of phage stock from step 2.9 to 400 µL with 1x TM buffer in a fresh 1.5 mL tube.
- Add an equal volume of Tris:Phenol:Chloroform to the dilution. Shake the mixture gently on a laboratory rocker, or by hand, for 5 min. Centrifuge the resultant emulsion at 17,000 x g in a benchtop centrifuge for 5-10 min.
NOTE: A distinct, white protein layer on the surface of the underlying phenol phase is present. - Remove the upper aqueous phase to a new tube taking care not to disturb the interphase boundary.
- Repeat steps 3.2-3.3 an additional 2 times for a total of 3 phenol extractions. Transfer the aqueous phase to a fresh tube and add an equal volume of CHCl3. Shake and centrifuge (step 3.2). Transfer the aqueous DNA sample to a clean tube.
- Estimate the volume of purified DNA. Add 0.4 volumes of 3 M sodium acetate (CH3COONa) and 3 volumes of 95% ethanol (EtOH). Place in -20 °C freezer for overnight precipitation. Centrifuge at 17,000 x g in a benchtop centrifuge for 10-15 min.
NOTE: The DNA will immediately dehydrate and become visible as a mass in suspension in the tube. The proper resulting pellet is white and well-packed against the bottom of the tube. - Remove and discard the supernatant either by decanting or pipetting. Add 500 µL 70% EtOH and gently shake the tube until the pellet floats free from the bottom of tube. Centrifuge at 17,000 x g for 5 min. Remove and discard EtOH, taking care to not disturb the pellet.
- Repeat step 3.6. Air dry the pellet on benchtop for 30-60 min. Add 50 µL ddH2O to the pellet and incubate for 1 h at RT or overnight at 4 °C to resuspend. Determine the DNA concentration using absorption measurements at 280 nm.
NOTE: The expected concentrations are 0.5-5 µg/µL using this technique. Divide by the total genomic molecular weight to determine the final genome concentration.
4. Cell-free Phage Reaction and Phage Titer Experiment
- Prepare 25 g LB medium and 15 g bacto-agar solid, pour into a 1 L bottle, and add deionized water to 1 L. Autoclave.
- After sterilization, invert the bottle slowly 6-8 times taking care to avoid formation of bubbles. The bottle inversion will homogenize the agar solid throughout the 1 L solution. Place the bottle in a 58 °C water bath for 20 min before dispensing onto culture plates.
- Once the temperature of the LB-agar solution has equilibrated, prepare a flame next to the culture plates to sterilize the environment near the open culture plates. Keep the 1 L bottle in the water bath to avoid solidification of agar while making the aliquots. Add 25 mL into a 100 mm x 15 mm culture plate. 1 L will produce 40 culture plates.
- Set one culture plate aside and let solidify for at least 1 h at RT; this plate will be used for plating the host cells. Let the other 39 culture plates solidify for 2 days at RT. Store at 4 °C or use immediately for making titers.
- Plate the host cells by streaking the appropriate bacterial strain (e.g., host B for T7) that was stored at -80 °C, onto LB-agar culture plate. Streak next to an open flame to avoid environmental contamination. Incubate overnight at 37 °C. The host cells have no antibiotic resistance so working in a sterile environment is essential.
- Cell-free reaction
NOTE: The cell-free TXTL reactions are composed of 33% crude E. coli extract (8.9-9.9 mg/mL protein) with the other 67% comprised of reaction buffer and phage genome. The crude extract is prepared as previously described15,16. Final reaction conditions are: 8.9-9.9 mg/mL protein (from crude extract), 3-6 mM Mg-glutamate, 40-100 mM K-glutamate, 2-4% PEG 8000, 3-4 mM of each amino acid, prepared as previously described17, and an energy mix solution, described previously15, composed of 0.33-3.33 mM DTT, 50 mM HEPES, 1.5 mM ATP and GTP, 0.9 mM CTP and UTP, 0.2 mg/mL tRNA, 0.26 mM CoA, 0.33 mM NAD, 0.75 mM cAMP, 0.068 mM folinic acid, 1 mM spermidine, and 30 mM 3-PGA. DNA-type phages require genome concentrations of 0.5-10 nM and RNA-type phages require a range of 50-150 nM. Final reaction concentrations are unique to the particular phage synthesized and will fall within the ranges described above. For optimal oxygenation of the reactions, final reaction volumes should be between 10-20 µL. There are small variations in the reaction protocol, which depend on the form of the genomic molecule. For example, linear genome molecules require an additional component in the reaction to inhibit the digestion of linear DNA pieces by the recBCD enzyme present in the crude extract.- Complete the "reaction details" section in Table 1 (PhageTXTL_JOVE) by entering the total number of reactions and final volume of reaction.
- Design the experiment by determining the constant components and variable components of the reaction. Enter the fractional volume percentage of the master mix, which is the ratio of the total volume of the constant reaction components to the product of the final reaction volume and number of samples. Enter the stock and final concentrations of the reaction reagents in the "Master Mix Reaction Recipe" section in PhageTXTL_JOVE. Enter the stock and final concentrations of the variable reagent(s) to be tested.
NOTE: The volumes of reaction components will automatically be calculated based on the master mix volume and the final volume of each individual reaction. - Remove the necessary amount of tubes (indicated in the "Tubes to thaw" section of PhageTXTL_JOVE.xlsx) of crude cell extract, energy buffer mix, and amino acid mix from -20 °C or -80 °C and thaw on ice.
NOTE: Once thawed, combine multiple aliquots of like components (if necessary). - Aliquot the indicated volume of crude extract, 33% of the final reaction volume, into a microcentrifuge tube.
- Prepare the master mix as per Table 1. Homogenize all components under "Master Mix Reaction Recipe" by vortexing. Add the appropriate volume of each component to the crude extract.
- If working with a phage with linear DNA genome (e.g., T7), add 1 µM of the gam protein of bacteriophage lambda to the reaction11. Vortex to homogenize the solution, and place the reaction on ice for 5 min. This inhibits digestion of the linear DNA pieces by the recBCD complex, which is endogenous in the crude extract.
- After adding the last component as per Table 1, homogenize the reaction by vortexing. Split the master mix into n microcentrifuge tubes.
NOTE: The volume of each split is the product of the fractional master mix volume percentage and the final volume of reaction. For example, for a fractional master mix volume percentage of 90% and the final reaction volume of 12 µL, the volume of each split is 10.8 µL. - Add the indicated volumes of the variable components to the master mix array (see Table 1). Add water to each reaction to reach the desired final reaction volume. Homogenize each reaction by vortexing. Incubate the microcentrifuge tubes at 29 °C for at least 8 h or overnight.
- Host cell pre-culture
NOTE: The overnight pre-culture is diluted 50:1 to ensure that the host cells used for the titer experiment are in the mid-log phase, which increases infection efficiency. The mid-log cells are then re-concentrated by 10-fold and stored on ice.- Using a sterile pipette tip, inoculate a culture tube of 5 mL LB with 3-5 healthy host cell colonies. Work next to an open flame to avoid contamination.
- Incubate the culture tube in a shaking incubator at 37 °C and 250 rpm for 16 h or overnight. Cells can reach saturation phase by the next day.
- Host cell culture and sample preparation for phage titer
- Dispense 50 mL LB into a 500 mL Erlenmeyer flask and cover with aluminum foil. Warm the flask at 37 °C for 20 min.
- Aliquot 1 mL of the host cell overnight pre-culture into the pre-warmed LB. Incubate for 3-4 h at 37 °C and shaking at 250 rpm.
- Centrifuge the 50 mL host cell culture for 10 min at 5,000 x g. Discard the LB.
- Resuspend the pellet with cold (4 °C) 5 mL LB and keep on ice.
- One hour before starting the titer experiment, incubate the culture plates at 37 °C.
NOTE: Each unique phage reaction (and corresponding dilution factor) will utilize two plates. Additionally, incubate 4 plates for experimental controls (see Figure 1 for representative positive and negative controls). - Prepare 100 mL of top agar (step 2.11).
- Prepare a serial dilution of the cell-free reaction with LB solution as described in section 2.12.
- Phage titer
NOTE: Begin to plate after culture dishes have incubated for at least 1 h and the top agar was in the 45 °C water bath for 15-20 min after removal from the autoclave.- Titer the final cell-free reaction-LB solution as described in section 2, steps 2.14-2.16.
Representative Results
We show four representative results. In Figure 1, we present a set of negative controls to ensure that the cell-free TXTL system and the phage DNA stocks are not contaminated with living E. coli cells. We verify that the cell-free TXTL system is free of intact E. coli cell contamination by plating both a non-incubated and incubated reaction solution void of genomic DNA (Figure 1A and Figure 1B). If the cell-free TXTL system was contaminated with E. coli cells, growth would be observed on the plate. Furthermore, we verify that the cell-free reaction, and therefore not any possible contaminant cells, are responsible for the synthesis of phage by plating non-incubated and incubated reactions with the phage genome (Figure 1C and Figure 1D). The plate with the non-incubated reaction (Figure 1C) shows zero plaque while the plate with the incubated reaction (Figure 1D) shows plaque, indicating the cell-free TXTL system was responsible for phage synthesis.
In Figure 2, we compare a homogeneous versus inhomogeneous plaque assay. Figure 2B illustrates a gradient of plaque density across the culture plate, which is sub-optimal in comparison to a homogeneous plaque density of the plate in Figure 2A. The source of this observed result is from the rapid cooling of master mix of top agar, sample and host bacterial cells when dispensed onto a culture plate during the phage titer portion of the experiment. To avoid an inhomogeneous spread of phage plaques, ensure that the culture plates are equilibrated to 37 °C before starting the phage titer experiment.
A critical step to properly count phages is to dilute the phage solution sufficiently to obtain between 50 and 200 plaques per plate (Figure 3). Below this range, it is difficult to determine true statistics, and above this range, the plaques overlap, preventing accurate counts. The number of phages synthesized in TXTL depends on the biochemical settings, such as the concentration of salts (potassium and magnesium), of molecular crowders, and of genomes. In Figure 4, we show a few examples of tests for the phages T7 and ΦΧ174. For a phage that replicates its DNA, such as T7, we observe a high efficiency of phage synthesis in the cell-free reaction: three infectious phages are synthesized per genome molecule in the reaction. One can expect a lower efficiency of reaction for a phage that does not replicate its DNA, such as ΦΧ174: the ratio of synthesized infectious phage to genome molecule is 1:5.
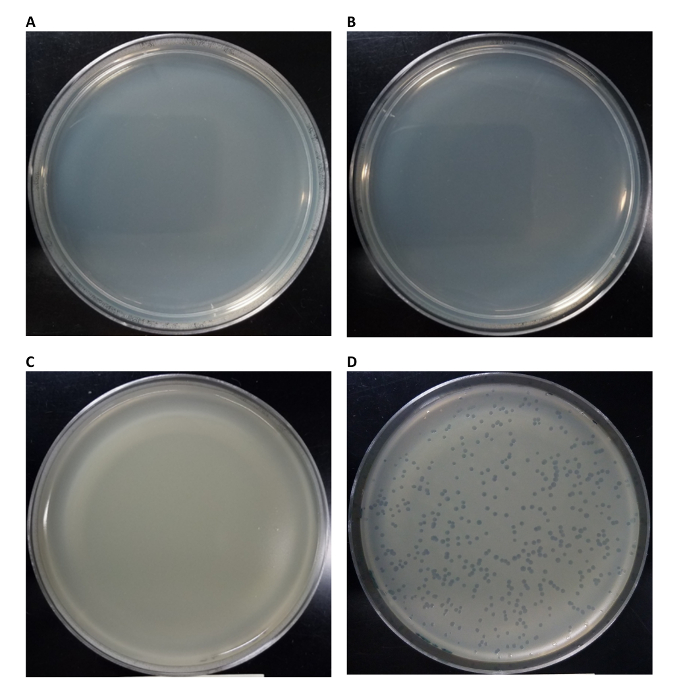
Figure 1: Plaque assay controls and successful infectious phage production using a cell-free reaction (CFR) system. (A) No genome was added and no incubation (0 min as incubation time) of CFR was performed. The reaction was then plated without host E. coli. Result shows no viable host cell contamination of CFR. (B) No genome was added and CFR was incubated 12 h at 29 °C and plated without host E. coli. Results show no viable host cell contamination of CFR, even after incubation. (C) 1 nM genome included with no incubation of CFR (0 min as incubation time) and plated with host E. coli. Result shows no phage contamination of the genome solution. (D) 1 nM genome included and CFR incubated 12 h at 29 °C. and plated with host E. coli. Results show successful infectious phage replication in CFR. Please click here to view a larger version of this figure.
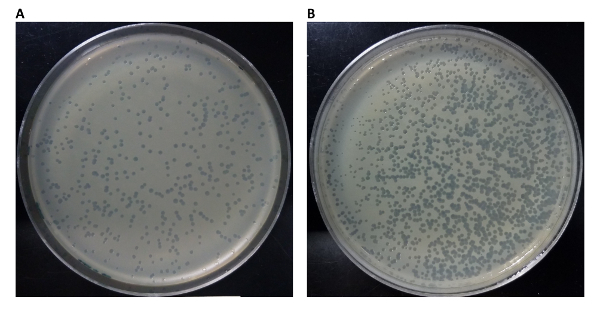
Figure 2: Homogeneous versus inhomogeneous spread of phage reaction during titer experiment. A possible effect of performing a phage titer on culture plates not equilibrated to a temperature of 37 °C is the inhomogeneous spread of phage reaction across the culture plate. A good result displaying a homogeneous spread of phage reaction is shown in (A), where the plaques are evenly distributed across the surface of the plate. A sub-optimal result is shown in (B), where the plaques are concentrated in the lower-left portion of the culture plate. This can occur with the top-agar, phage reaction, and host cell master mix are dispensed onto a culture plate that is not at a temperature of 37 °C. The majority of the master mix solidifies in the lower-left region, and a homogeneous spread is not possible. Please click here to view a larger version of this figure.
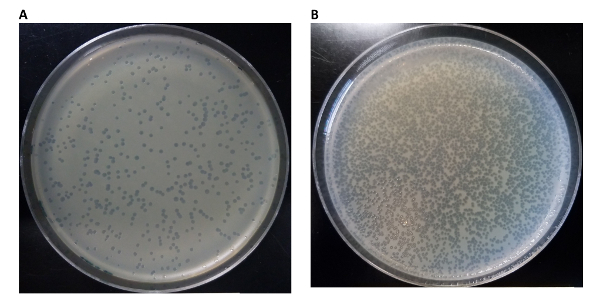
Figure 3: Setting an appropriate dilution factor for titers of a cell-free phage reaction. (A) This plate displays a reasonably countable number of plaques. The dilution factor used for the titer portion of the experiment diluted the yield of the phage reaction enough to leave a countable number of plaques on the plate. Conversely in (B) the dilution factor was too low with respect to the yield of synthesized phage, and needs to be adjusted. Please click here to view a larger version of this figure.
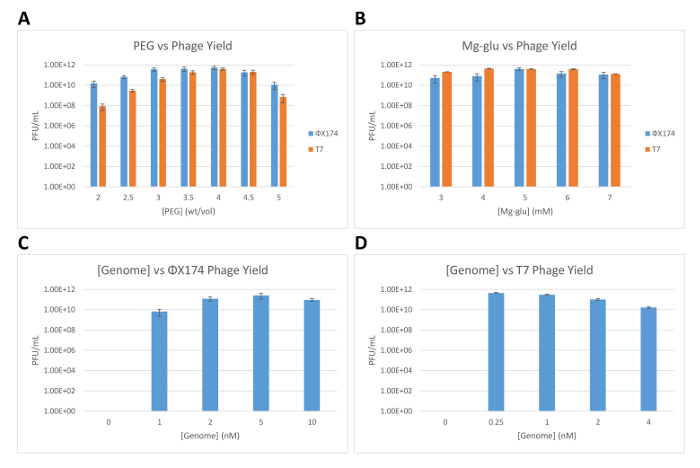
Figure 4: Characterization of the cell-free synthesis of phages ΦΧ174 and T7. (A) Number of synthesized ΦΧ174 and T7 phages (PFU/mL: plaque forming units per milliliter) as a function of PEG 8000 concentration in the cell-free reaction, measured after 16 h of incubation at 29 °C. (B) Number of synthesized ΦΧ174 and T7 phages as a function of magnesium-glutamate concentration in the cell-free reaction, measured after 16 h of incubation at 29 °C. (C) Number of synthesized ΦΧ174 phage as a function of ΦΧ174 DNA genome concentration in the cell-free reaction, measured after 16 h of incubation at 29 °C. (D) Number of synthesized T7 phage as a function of T7 DNA genome concentration in the cell-free reaction, measured after 16 h of incubation at 29 °C. All error bars displayed are standard deviations of three repeated trials. Please click here to view a larger version of this figure.
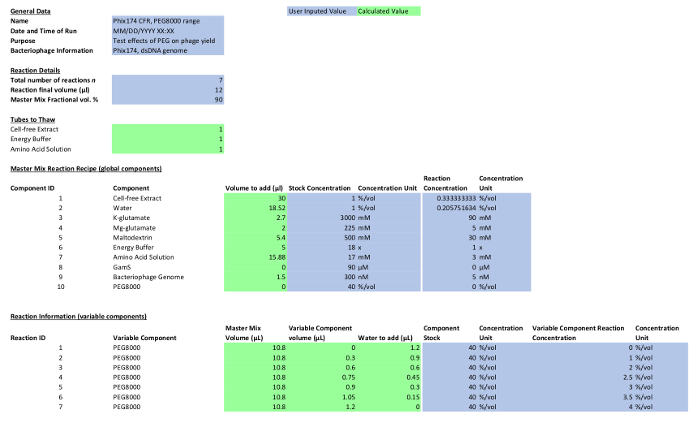
Table 1: Reaction composition. Please click here to view a larger version of this figure.
Discussion
Following the technique in Chen, et al.14 for SPMC, a critical step is reached when determining the proper conditions for superinfection. The parameter that most closely controls a host strain's ability to withstand superinfection is frequently the initial concentration of the infecting phage. The host cells must be in logarithmic growth phase before initial infection with a very small amount of phage. Eventually, the phage will also reach logarithmic growth and the goal is to allow the phage numbers to rapidly outstrip the cell count, allowing for multiple copies of phage in each host cell. This forces the cell into a quasi-stable state where it will not undergo lysis. If this state is not reached, the cells will undergo cascading, confluent lysis, and release their viral load. If superinfection cannot be reached using the above initial viral concentrations, lower the initial multiplicity of infection by a factor of ten before making a subsequent attempt. If the host cells do lyse before harvest and superinfection is not reached, proceed immediately with a separate purification procedure. Add DNase I to 5 µg/mL and 500 µL CHCl3 to finalize all cellular lysis. Continue to swirl for an additional 5 min and then centrifuge at 10,000 x g to remove cellular debris. Decant and centrifuge the supernatant at 75,000 x g for 1-2 h to pellet the phage and resuspend overnight in 1 mL total 1x TM buffer at 4 °C without shaking. When testing for superinfection using a small volume of cells, it is possible to gauge how close cells are to lysis by how quickly they lyse when exposed to the added chloroform. If the cells lyse and the solution clarifies almost instantly, the cell walls are very unstable and cells should be harvested immediately to prevent large-scale cascade lysis. If the test lyses within 1-3 min, the larger volumes may continue to superinfect for up to 2 h, depending on the local strain of E. coli in use. If the cells do not lyse within 5 min, phage growth has not caught up to the E. coli growth and primary incubation should continue. Perform the test again in 30-60 min.
When purifying phage on a prepared sucrose gradient, the phage will appear as a very thick, pearlescent band between one third and two thirds of the way down the tube. Assuming a relatively successful lysis and cellular remnant purification step, there may be other, light bands but none of similar density or size. The large, opaque band contains the phage and is to be removed from the solution with a sterile syringe and blunt cannula.
When starting the third section of the protocol, the cell-free reaction and phage titer, it is best to use fresh culture plates (less than one week old) for the host bacteria plate (from step 3.5) and also the culture plates used for the titer analysis (from step 3.21). This will promote stronger growth of host bacteria cells and higher infection efficiency for the phage titer. Phage culture plates must be pre-incubated at 37 °C for a minimum of 1 h before starting the phage titer portion of the protocol. Without pre-incubation of the phage culture plates, the top-agar solution (containing the synthesized phage and host cell culture) will not homogeneously distribute across the surface of the culture plate. Instead, the top-agar solution will homogeneously solidify on the surface (Figure 2b). This increases the probability of multiple phages localizing in the same spot, which can cause an underestimate of phage yield in the cell-free reaction.
Each plaque represents the event of one synthesized phage infecting a host cell, replicating within the host cell, and lysing the host cell. Plaques grow in size during the incubation period from the progeny phage infecting neighbor host cells from the initial infection event. If no plaques are observed, it is possible that the dilution factor made in Protocol section 4.6 is too high resulting in a low probability that any synthesized phage infects a host cell. To remedy this, decrease the dilution factor by 3-4 orders of magnitude (or as necessary) and repeat the titer experiment. Conversely, the dilution factor could be too high resulting in plaques spanning the entire surface of the culture plate, making it impossible to count the phage yield. In this case, increase the dilution factor by 3-4 orders of magnitude (or as necessary). For a well-characterized phage, the yield of the cell-free reaction will be known and only one dilution factor per reaction point will be necessary for the titer experiment. When characterizing a new phage, it is best to include 2-3 different dilution factors per reaction point, ensuring that one of the dilutions will yield a countable plaque (see Figure 3 for differences between a countable and uncountable number of plaques on a plate).
If no plaque is observed after modifying the dilution factor of the phage titer, this could indicate that the cell-free reaction was unsuccessful in that gene expression did not occur. A possible explanation for this could be structural damage of reagents/phage DNA due to the homogenization of the cell-free reaction by vortexing. To remedy this issue, repeat the reaction and homogenize by gently tapping the reaction tube five to seven times with a finger, or by slowly mixing the reaction with a pipette five to seven times.
The cell-free reaction system originates from E. coli and endogenously contains both E. coli RNA polymerase and the primary sigma factor, sigma 70, which combine to form the holoenzyme necessary to recognize the main promoter consensus sequences of E. coli genes. Additionally, the cell-free reaction system can recapitulate the entire sigma factor transcription scheme by exogenously supplying six other E. coli sigma factor genes under the promoter consensus sequence of sigma 70. This allows the capability to synthesize all bacteriophages whose genomes are compatible with E. coli's transcription/translation machinery. This leaves out a subset of bacteriophage that cannot be studied or synthesized with the cell-free reaction system utilized here.
The cell-free transcription/translation platform allows for an impressive level of control and flexibility of reaction conditions compared to in vivo methods of study. We can finely tune many biochemical and genetic components of a phage reaction and directly observe the effects in less than 24 h, as shown in Figure 4 for molecular crowding. The effects of molecular crowders, such as PEG, on the self-assembly of macromolecular complexes have been theoretically described and demonstrated in vitro18,19,20. Molecular crowding is an entropy driven mechanism that increases the association constant of large biomolecular structures. The cell-free approach allows the researcher to probe complicated biological processes like the biophysics of self-assembly with model systems, like bacteriophages, without the intrinsic limitations of in vivo work.
This platform also allows for the investigation of the utility of phages in applied fields. Components of phage genomes can be repurposed and quickly tested in a cell-free reaction system with the goal of incorporating results to novel nanostructures. Bacteriophages can be modified in order to increase the selectivity of host cell infection, which would promote the utility of phage therapy as an alternative for antibiotic-resistant bacteria.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
This material is based on work supported by the Office of Naval Research award number N00014-13-1-0074 (to V.N.), the Human Frontier Science Program grant number RGP0037/2015 (to V.N.), and the Binational Science Foundation grant 2014400 (to V.N.).
Materials
| Ultracentrifugation tubes | Beckman Coulter | 344057 | |
| Conical tubes | Falcon | 352070 | |
| Gradient maker | BioComp Gradient Master | see Anal Biochem. 1985 Jul;148(1):254-9. | |
| Syringe and Blunt Cannula | Monoject | 8881513918 and 888202017 | |
| Wide-bore pipette tips | Fischerbrand | 02-707-134 | |
| Plaque counter | New Brunswik Scientific | Colony Counter Model C-110 | |
| Culture tubes | Fischerbrand | 14-961-33 | |
| Cell-free system | Mycroarray Inc | Mytxtl | |
| BioComp Gradient Master | BioComp Instruments | Model 105ME | |
| LB agar plate recipe | 25 g/L Luria-Bertani medium (LB Broth, Miller – Fisher BioReagents product number BP1426) and 15 g/L Bacto-Agar solid (Brenton, Dickenson and Company – product number 214010). |
Riferimenti
- Carlson, E. D., Gan, R., Hodgman, C. E., Jewett, M. C. Cell-free protein synthesis: Applications come of age. Biotechnol Adv. , (2011).
- Hodgman, C. E., Jewett, M. C. Cell-free synthetic biology: Thinking outside the cell. Metab Eng. , (2011).
- Caschera, F., Noireaux, V. Synthesis of 2.3 mg/mL of protein with an all Escherichia coli cell-free transcription-translation system. Biochimie. 99, 162-168 (2014).
- Shin, J., Noireaux, V. An E. coli cell-free expression toolbox: application to synthetic gene circuits and artificial cells. ACS Synth Biol. 1 (1), 29-41 (2012).
- Chemla, Y., Ozer, E., Schlesinger, O., Noireaux, V., Alfonta, L. Genetically expanded cell-free protein synthesis using endogenous pyrrolysyl orthogonal translation system. Biotechnol Bioeng. , (2015).
- Hong, S. H., Kwon, Y. C., Jewett, M. C. Non-standard amino acid incorporation into proteins using Escherichia coli cell-free protein synthesis. Frontiers in Chemistry. 2, (2014).
- Noireaux, V., Bar-Ziv, R., Libchaber, A. Principles of cell-free genetic circuit assembly. Proc Natl Acad Sci U S A. 100 (22), 12672-12677 (2003).
- Sun, Z. Z., Yeung, E., Hayes, C. A., Noireaux, V., Murray, R. M. Linear DNA for Rapid Prototyping of Synthetic Biological Circuits in an Escherichia coli Based TX-TL Cell-Free System. Acs Synthetic Biology. 3 (6), 387-397 (2014).
- Takahashi, M. K., et al. Characterizing and prototyping genetic networks with cell-free transcription-translation reactions. Methods. , (2015).
- Shin, J., Jardine, P., Noireaux, V. Genome Replication, Synthesis, and Assembly of the Bacteriophage T7 in a Single Cell-Free Reaction. ACS Synthetic Biology. 1 (9), 408-413 (2012).
- Garamella, J., Marshall, R., Rustad, M., Noireaux, V. The All E. coli TX-TL Toolbox 2.0: A Platform for Cell-Free Synthetic Biology. ACS Synth Biol. , (2016).
- Karzbrun, E., Shin, J., Bar-Ziv, R. H., Noireaux, V. Coarse-grained dynamics of protein synthesis in a cell-free system. Phys Rev Lett. 106 (4), 048104 (2011).
- Daube, S. S., Arad, T., Bar-Ziv, R. Cell-free co-synthesis of protein nanoassemblies: tubes, rings, and doughnuts. Nano Lett. 7 (3), 638-641 (2007).
- Chen, X., et al. An immunoblot assay reveals that bacteriophage T4 thymidylate synthase and dihydrofolate reductase are not virion proteins. J Virol. 69 (4), 2119-2125 (1995).
- Sun, Z. Z., et al. Protocols for implementing an Escherichia coli based TX-TL cell-free expression system for synthetic biology. J Vis Exp. (79), e50762 (2013).
- Shin, J. N. V. Efficient cell-free expression with the endogenous E. coli RNA polymerase and sigma factor 70. J Biol Eng. , (2010).
- Caschera, F., Noireaux, V. Preparation of amino acid mixtures for cell-free expression systems. Biotechniques. 58 (1), 40-43 (2015).
- Minton, A. P. How can biochemical reactions within cells differ from those in test tubes. J Cell Sci. 119, 2863-2869 (2006).
- Minton, A. P. Implications of macromolecular crowding for protein assembly. Curr Opin Struct Biol. 10 (1), 34-39 (2000).
- Zimmerman, S. B., Minton, A. P. Macromolecular crowding: biochemical, biophysical, and physiological consequences. Annu Rev Biophys Biomol Struct. 22, 27-65 (1993).

