Single-step Purification of Macromolecular Complexes Using RNA Attached to Biotin and a Photo-cleavable Linker
Summary
RNA/protein complexes purified using botin-streptavidin strategy are eluted to solution under denaturing conditions in a form unsuitable for further purification and functional analysis. Here, we describe a modification of this strategy that utilizes a photo-cleavable linker in RNA and a gentle UV-elution step, yielding native and fully functional RNA/protein complexes.
Abstract
For many years, the exceptionally strong and rapidly formed interaction between biotin and streptavidin has been successfully utilized for partial purification of biologically important RNA/protein complexes. However, this strategy suffers from one major disadvantage that limits its broader utilization: the biotin/streptavidin interaction can be broken only under denaturing conditions that also disrupt the integrity of the eluted complexes, hence precluding their subsequent functional analysis and/or further purification by other methods. In addition, the eluted samples are frequently contaminated with the background proteins that nonspecifically associate with streptavidin beads, complicating the analysis of the purified complexes by silver staining and mass spectrometry. To overcome these limitations, we developed a variant of the biotin/streptavidin strategy in which biotin is attached to an RNA substrate via a photo-cleavable linker and the complexes immobilized on streptavidin beads are selectively eluted to solution in a native form by long wave UV, leaving the background proteins on the beads. Shorter RNA binding substrates can be synthesized chemically with biotin and the photo-cleavable linker covalently attached to the 5' end of the RNA, whereas longer RNA substrates can be provided with the two groups by a complementary oligonucleotide. These two variants of the UV-elution method were tested for purification of the U7 snRNP-dependent processing complexes that cleave histone pre-mRNAs at the 3' end and they both proved to compare favorably to other previously developed purification methods. The UV-eluted samples contained readily detectable amounts of the U7 snRNP that was free of major protein contaminants and suitable for direct analysis by mass spectrometry and functional assays. The described method can be readily adapted for purification of other RNA binding complexes and used in conjunction with single- and double-stranded DNA binding sites to purify DNA-specific proteins and macromolecular complexes.
Introduction
In eukaryotes, RNA polymerase II-generated mRNA precursors (pre-mRNAs) undergo several maturation events in the nucleus before becoming fully functional mRNA templates for protein synthesis in the cytoplasm. One of these events is 3' end processing. For the vast majority of pre-mRNAs, 3' end processing involves cleavage coupled to polyadenylation. This two-step reaction is catalyzed by a relatively abundant complex consisting of more than 15 proteins1. Animal replication-dependent histone pre-mRNAs are processed at the 3' end by a different mechanism in which the key role is played by U7 snRNP, a low abundance complex consisting of U7 snRNA of ~60 nucleotides and multiple proteins2,3. The U7 snRNA base pairs with a specific sequence in histone pre-mRNA and one of the subunits of the U7 snRNP catalyzes the cleavage reaction, generating mature histone mRNA without a poly(A) tail. 3' end processing of histone pre-mRNA also requires Stem-Loop Binding Protein (SLBP), which binds a conserved stem-loop located upstream of the cleavage site and enhances the recruitment of the U7 snRNP to the substrate2,3. Studies aimed at identifying individual components of the U7 snRNP have been challenging due to the low concentration of the U7 snRNP in animal cells and the tendency of the complex to dissociate or undergo partial proteolysis during purification as a result of using mild detergents4,5,6, high salt washes and/or multiple chromatographic steps7,8,9.
Recently, to determine the composition of the U7-dependent processing machinery, a short fragment of histone pre-mRNA containing biotin at either 3' or 5' was incubated with a nuclear extract and the assembled complexes were captured on streptavidin-coated agarose beads5,6,10. Due to the exceptionally strong interaction between biotin and streptavidin, proteins immobilized on streptavidin beads were eluted under denaturing conditions by boiling in SDS and analyzed by silver staining and mass spectrometry. While this simple approach identified a number of components of the U7 snRNP, it yielded relatively crude samples, often contaminated with a large number of background proteins nonspecifically bound to streptavidin beads, potentially masking some components of the processing machinery and preventing their detection on silver stained gels5,6,10. Importantly, this approach also precluded any functional studies with the isolated material and its further purification to homogeneity by additional methods.
A number of modifications were proposed over time to address the virtually irreversible nature of the biotin/streptavidin interaction, with most of them being designed to either weaken the interaction or to provide a chemically cleavable spacer arm in the biotin-containing reagents11,12. The downside of all these modifications was that they significantly reduce the efficiency of the method and/or often required non-physiological conditions during the elution step, jeopardizing either the integrity or activity of the purified proteins.
Here, we describe a different approach to resolve the inherent problem of the biotin/streptavidin strategy by using RNA substrates in which biotin is covalently attached to the 5' end via a photo-cleavable 1-(2-nitrophenyl)ethyl moiety that is sensitive to long wave UV13,14. We tested this approach for the purification of the limiting U7-dependent processing machinery from Drosophila and mammalian nuclear extracts15. Following a short incubation of histone pre-mRNA containing biotin and the photo-cleavable linker with a nuclear extract, the assembled processing complexes are immobilized on streptavidin beads, thoroughly washed and gently released to solution in a native form by exposure to ~360 nm UV light. The UV-elution method is very efficient, fast and straightforward, yielding sufficient amounts of the U7 snRNP to visualize its components by sliver staining from as little as 100 µL of the extract15. The UV-eluted material is free of background proteins and suitable for direct mass spectrometry analysis, additional purification steps and enzymatic assays. The same method may be adopted for the purification of other RNA/protein complexes that require relatively short RNA binding sites. Biotin and the photo-cleavable linker can also be covalently attached to single- and double-stranded DNA, potentially extending the UV-elution method for the purification of various DNA/protein complexes.
Chemical synthesis of RNA substrates containing covalently attached biotin and the photo-cleavable linker is practical only with the sequences that do not exceed ~65 nucleotides, becoming expensive and inefficient for significantly longer sequences. To address this problem, we also developed an alternative approach that is suitable for much longer RNA binding targets. In this approach, RNA of any length and nucleotide sequence is generated in vitro by T7 or SP6 transcription and annealed to a short complementary oligonucleotide that contains biotin and the photo-cleavable linker at the 5' end (trans configuration). The resultant duplex is subsequently used to purify individual binding proteins or macromolecular complexes on streptavidin beads following the same protocol described for the RNA substrates containing photo-cleavable biotin attached covalently (cis configuration). With this modification, the photo-cleavable biotin can be used in conjunction with in vitro generated transcripts containing hundreds of nucleotides, extending the UV-elution method for the purification of a broad range of RNA/protein.
Protocol
1. Substrate Preparation
NOTE: RNA substrates shorter than ~65 nucleotides can be synthesized chemically with biotin (B) and the photo-cleavable (pc) linker (together referred to as photo-cleavable biotin or pcB) covalently attached to the RNA 5’ end (cis configuration). RNA substrates containing significantly longer binding sites need to be generated in vitro by T7 (or SP6) transcription and subsequently annealed to a short complementary adaptor oligonucleotide containing pcB moiety at the 5’ end (trans configuration) (Figure 1).
- For binding sites consisting of fewer than ~65 nucleotides, use a commercial manufacturer (Table of Materials) to chemically synthesize RNA of interest with pcB covalently attached to the RNA 5’ end (Figure 1A and Figure 2A). Dissolve in sterile water to achieve desired concentration and store in small aliquots at -80 °C.
- For binding sites significantly longer than ~65 nucleotides, form a partial duplex of an in vitro generated RNA of interest and a complementary adaptor oligonucleotide containing the pcB moiety at the 5’ end (Figure 3A).
- Perform T7 transcription on either a linearized plasmid DNA or an appropriate PCR template to generate RNA with the binding site of interest extended at the 3’ end by a ~20-nucleotide arbitrary sequence.
- Use a commercial manufacturer (Table of Materials) to chemically synthesize an adaptor oligonucleotide that is complementary to the 3’ extension in the T7-generated RNA and contains pcB at the 5’ end (Figure 3A).
NOTE: Two 18-atom spacers and 2-3 non-complementary nucleotides can be placed at the 5’ end of the oligonucleotide to reduce potential steric hindrance between the bound complex and streptavidin beads. It is also desirable that the oligonucleotide is uniformly modified with a 2’O-methyl group to enhance the strength of the duplex and to provide resistance against various nucleases. - Anneal the T7-generated RNA to the pcB adaptor oligonucleotide.
- Mix 20 pmol of the RNA and 100 pmol of the adaptor pcB oligonucleotide (1:5 molar ratio) in a 1.5 mL tube containing 100 µL of binding buffer with the following composition: 75 mM KCl, 15 mM HEPES pH 7.9, 15% glycerol, 10 mM ethylenediaminetetraacetic acid (EDTA).
NOTE: It is important to use a minimal amount of the adaptor oligonucleotide that is sufficient to form a duplex with most (if not all) pre-mRNA substrate used in the annealing reaction. This could be conveniently determined by labeling RNA substrate at the 5’ end with 32P, annealing the labeled substrate with increasing amounts of the adaptor oligonucleotide using various buffer conditions and monitoring the retention of the radioactive signal on streptavidin beads after several washes of the beads with the binding buffer. - Place the tube in boiling water for 5 min.
- Allow the water to cool down to room temperature.
- Mix 20 pmol of the RNA and 100 pmol of the adaptor pcB oligonucleotide (1:5 molar ratio) in a 1.5 mL tube containing 100 µL of binding buffer with the following composition: 75 mM KCl, 15 mM HEPES pH 7.9, 15% glycerol, 10 mM ethylenediaminetetraacetic acid (EDTA).
2. Complex Assembly
- Supplement 1 mL of a mouse nuclear extract (or another extract of choice) with 80 mM EDTA pH 8 to a final concentration of 10 mM to block nonspecific metal-dependent nucleases that are present in the extract.
NOTE: This will result in adjusting buffer in the extract to the same composition as that in the binding buffer (see Step 1.2.3.1). EDTA does not affect in vitro processing of histone pre-mRNAs but may be harmful in purifying proteins or protein complexes that assemble in a magnesium-dependent manner. In these cases, EDTA should be avoided. - Add 5-10 pmol of the RNA substrate tagged with the pcB moiety in cis (see Step 1.1) or trans (see Step 1.2.3.3) to 1 mL of the extract containing 10 mM EDTA.
NOTE: The amount of RNA needs to be carefully evaluated in a series of trial experiments. Using too much RNA substrate for the purification of a limiting complex maybe counterproductive, significantly increasing the background of nonspecific RNA binding proteins without having any effect on the yield of specific proteins. - Incubate the RNA substrate with the extract for 5 min on ice, occasionally mixing the sample.
NOTE: Both the time and temperature of the incubation need to be established empirically and may vary significantly, depending on the specific nature of the RNA/protein complex. - Spin the incubation mixture in a pre-cooled microcentrifuge for 10 min at 10,000 x g to remove any potential precipitates and carefully collect the supernatant avoiding transferring the pellet.
3. Immobilization of the RNA/protein Complexes on Streptavidin Beads.
- Transfer ~100 µL of streptavidin agarose bead suspension from a commercial supplier (Table of Materials) to a 1.5 mL tube and increase the volume with the binding buffer (75 mM KCl, 15 mM HEPES pH 7.9, 15% glycerol, 10 mM EDTA).
NOTE: This buffer should have the same composition as that in the annealing mixture and in the extract used for complex assembly (Step 2.1). - Collect the beads at the bottom of the tube by spinning for 2-3 min at 25 x g and aspirate the supernatant.
- Repeat the same washing/spinning procedure 2-3 times to equilibrate the beads with the binding buffer.
NOTE: At the end of this step, the pellet of the beads should have a volume of ~ 30 µL. - Load the supernatant containing the assembled complex (Step 2.4) over the equilibrated streptavidin agarose beads.
- Rotate 1 h at 4 °C to immobilize RNA and the bound complexes on the beads.
- Collect the beads at the bottom of the tube by spinning for 2-3 min at 25 x g.
NOTE: Use a swing out rotor to avoid substantial loss of the beads if they tend to adhere to the side of the tube in angular rotors. - Aspirate the supernatant and rinse the beads twice with 1 mL of the binding buffer, using the same centrifugation conditions.
- Add 1 mL of the binding buffer and rotate the sample 1 h at 4 °C.
NOTE: This step can be shortened if the complex formed on the substrate tends to dissociate. - Spin down the beads for 2-3 min at 25 x g, add 1 mL of the buffer and transfer the suspension to a new tube.
- Rotate for an additional 1 h or shorter, if the complex is not stable, spin down for 2-3 min at 25 x g and transfer the beads to a 500 µL tube in 200 µL of binding buffer.
4. UV-Elution.
- Turn on a high intensity UV lamp emitting 365 nm UV light for 5-10 min before reaching full brilliance.
NOTE: This is essential to achieve maximum energy of UV emission at the start of irradiation. - Fill up the bottom of a Petri dish (100 mm x 15 mm) with tightly packed ice and stack it over the dish’s lid.
- Briefly vortex the tube containing the immobilized complex, place it horizontally on ice and cover with the pre-warmed lamp, ensuring that the sample is located within the distance of 2-3 cm of the surface of the bulb.
NOTE: If the distance is too large, use additional Petri dishes or other suitable objects to bring the sample closer to the bulb. - Irradiate for a total of 30 min, frequently inverting and vortexing the tube to ensure uniform exposure of the suspension to UV and to prevent overheating.
NOTE: To provide additional cooling, the UV-elution step can be carried out in a cold room. Switch to a Petri dish with fresh ice in case of excessive ice melting. - Spin down the beads for 2-3 min at 25 x g and collect the supernatant.
- Re-spin the supernatant using the same conditions and collect the supernatant.
NOTE: Leave a small amount of supernatant at the bottom to avoid transferring residual beads.
5. Sample Analysis by Silver Staining Followed by Mass Spectrometry.
- Use SDS/polyacrylamide gel electrophoresis to separate a fraction of the UV-eluted supernatant and the same fraction of the material left on the beads following UV elution.
NOTE: The analysis may also include an aliquot of the beads withdrawn from the sample before UV-elution. - Stain the gel containing separated proteins using a commercially available silver staining kit.
- Evaluate the efficiency of UV-elution by comparing the intensities of proteins present in the UV-eluted supernatant and those left on the beads following UV-irradiation.
- Excise the protein bands of interest and determine their identities by mass spectrometry using standard protocols10,15.
6. Global Analysis of UV-eluted Sample by Mass Spectrometry.
- Directly analyze a fraction of the UV-eluted supernatant by mass spectrometry to determine the entire proteome of the purified material in an unbiased manner.
NOTE: This can be done by in-solution treatment of the purified samples with trypsin followed by standard mass spectrometry protocols to determine the identity of the generate peptides10,15.
Representative Results
The UV-elution method was tested with two chemically synthesized RNA substrates covalently attached at the 5' end to the pcB moiety (cis configuration): pcB-SL (Figure 1) and pcB-dH3/5m RNAs (Figure 2). The 31-nucleotide pcB-SL RNA contains a stem-loop structure followed by a 5-nucleotide single stranded tail and its sequence is identical to the 3' end of mature histone mRNA (i.e., after the cleavage of histone pre-mRNA by U7 snRNP). This unique sequence is a known binding site for two proteins present in the mammalian cytoplasmic fraction: SLBP and 3'hExo16,17,18. pcB-dH3/5m is a 63-nucleotide fragment of Drosophila H3 histone pre-mRNA and in addition to the stem-loop contains a sequence that binds Drosophila U7 snRNP (Figure 2).
dH3 Ext (125 nucleotides) is an example of longer RNA substrates that can be generated by T7 transcription and provided with biotin and a photo-cleavable spacer in trans by annealing to pcB/22mer, a chemically synthesized adaptor oligonucleotide (trans configuration). pcB/22mer contains the two groups at the 5' end and consists of 22 2'O-methyl-modifed nucleotides (Figure 3A). 19 nucleotides at the 3' end of the pcB/22mer(underlined in the sequence in Figure 3A) are complementary to the last 19 nucleotides of the dH3 Ext pre-mRNA. This pre-mRNA besides being longer does not significantly differ from the synthetic pcB-dH3/5m, containing the same two key processing signals: stem-loop and U7-binding site.
pcB-SL RNA was incubated with 1 mL of S100 extract prepared by ultracentrifugation (100,000 x g for 1 h) of a cytoplasmic fraction obtained from mouse myeloma cells19 and the effect and efficiency of UV-elution were first analyzed by silver staining. A number of proteins were detected on streptavidin beads before UV-elution (Figure 1B, lane 1). Irradiation with long wave UV released only some of these proteins to the supernatant (Figure 1B, lane 3), leaving a nonspecific background on the beads (Figure 1B, lane 2), hence emphasizing the importance of the UV-elution step. Major UV-eluted proteins were identified by mass spectrometry as 3'hExo and SLBP, with SLBP being represented by full length protein (FL) and a number of shorter degradation products (DP). Smaller amounts of other proteins, identified as various RNA binding proteins (RBPs), were also selectively released to solution by UV irradiation (Figure 1B, lane 3).
pcB-dH3/5m pre-mRNA (Figure 2) or dH3 Ext pre-mRNA annealed to pcB/22mer (Figure 3) were incubated with 1 mL of a nuclear extract from Drosophila Kc cells to form processing complexes20,21. To better evaluate the specificity of proteins that bind to each histone pre-mRNA, a negative control was prepared in parallel by adding two competitors to the nuclear extract: SL RNA that sequesters SLBP, and a short antisense oligonucleotide that base pairs with the U7 snRNA and prevents the interaction of the U7 snRNP with its site on the pre-mRNA.
The UV-elution step of the immobilized pcB-dH3/5m pre-mRNA resulted in a selective release of only a small number of proteins to the supernatant (Figure 2B, lane 1), with an intense background of non-specific proteins remaining on the beads (Figure 2B, lane 3). These proteins, labeled A-I, were identified by mass spectrometry as components of the U7 snRNP15. The sample also contained partially degraded SLBP that migrates at 10 kDa and can only be visible on higher concentration SDS/polyacrylamide gels. All these proteins were not detected in the presence of the two processing competitors that block binding of U7 snRNP to histone pre-mRNA (Figure 2B, lane 2).
The same components of Drosophila U7 snRNP were released to solution by UV irradiation of an immobilized duplex consisting of dH3 Ext pre-mRNA and pcB/22mer (Figure 3B, lane 1). This sample additionally contained multiple RNA binding proteins that interacted with the pcB/22mer oligonucleotide used in excess to form a duplex with dH3 Ext pre-mRNA. In contrast to the subunits of the U7 snRNP, these contaminating proteins, as well as all the background proteins that remained on the beads, persisted in the presence of the processing competitors (Figure 3B, compare lanes 1 and 2, and lanes 3 and 4).
A small fraction of the UV-supernatant containing processing complexes and the same fraction of the UV-supernatant from a negative control (prepared in the presence of the two competitor oligonucleotides and hence lacking processing complexes) can be directly analyzed by mass spectrometry without a prior separation of the eluted proteins by gel electrophoresis15. This method is very sensitive in detecting all eluted proteins regardless of their size and abundance and in conjunction with the negative sample provides a complete and unbiased list of proteins that specifically associate with histone pre-mRNA to conduct 3' end processing reaction.
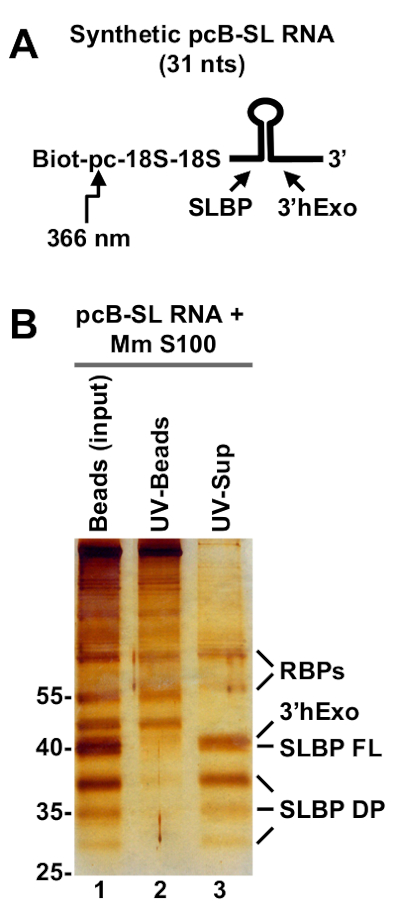
Figure 1: Purification of mouse cytoplasmic proteins bound to pcB-SL RNA. (A) A diagram of chemically synthesized pcB-SL RNA (31-nucleotides). Biotin (Biot) at the 5' end is followed by a photo-cleavable moiety (pc) sensitive to long wave UV (366 nm), two 18 atom spacers and 31 nucleotides that form the conserved stem-loop structure found at the 3' end of mature histone mRNAs. (B) pcB-SL RNA was incubated with S100 cytoplasmic extract from mouse myeloma cells. The RNA and bound proteins were purified on streptavidin beads, extensively washed, UV eluted and analyzed by silver staining (lane 3). Proteins immobilized on streptavidin beads before UV elution and left on the beads after UV elution are shown in lanes 1 and 2, respectively. Position of protein size markers (in kDa) is indicated to the left.
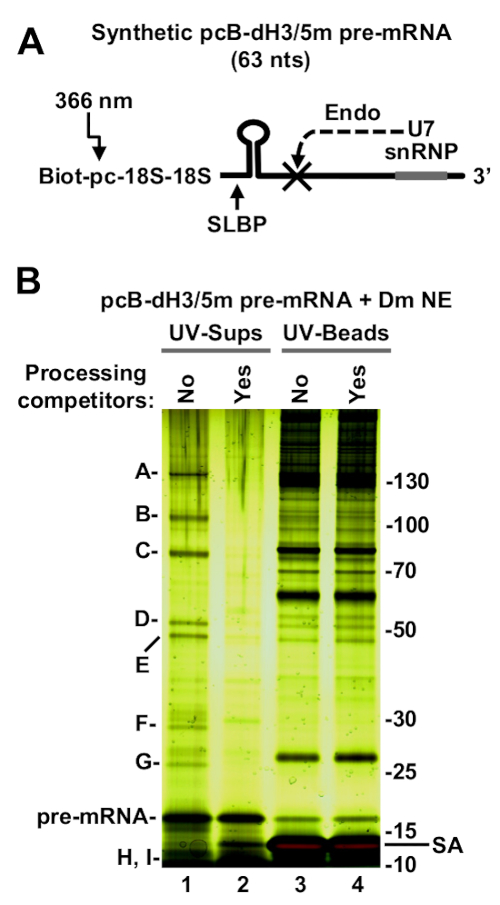
Figure 2: Purification of Drosophila processing complexes assembled on pcB-dH3/5m pre-mRNA. (A) A diagram of chemically synthesized Drosophila-specific pcB-dH3/5m pre-mRNA (63-nucleotides). Biotin (Biot) at the 5' end is followed by a photo-cleavable moiety (pc), two 18-atom spacers and 63 nucleotides that contain the two sequence elements essential processing: stem-loop structure and U7-binding site. Five nucleotides around the major cleavage site located between the two elements are modified with a 2'O-methyl group to block the cleavage by the U7 snRNP during complex assembly (crossed lines). (B) pcB-dH3/5m was incubated with a Drosophila nuclear extract to assemble processing complexes. In the negative control, the nuclear extract contains two processing competitors to block binding of SLBP and U7 snRNP to histone pre-mRNA. The assembled complexes were immobilized on streptavidin beads, extensively washed and released to solution by the exposure of the sample to long wave UV. The same fractions of the UV-eluted material (UV-sups) and the beads following UV-elution (UV-beads) were analyzed by silver staining. Position of protein size markers (in kDa) and streptavidin (SA) is indicated to the right.
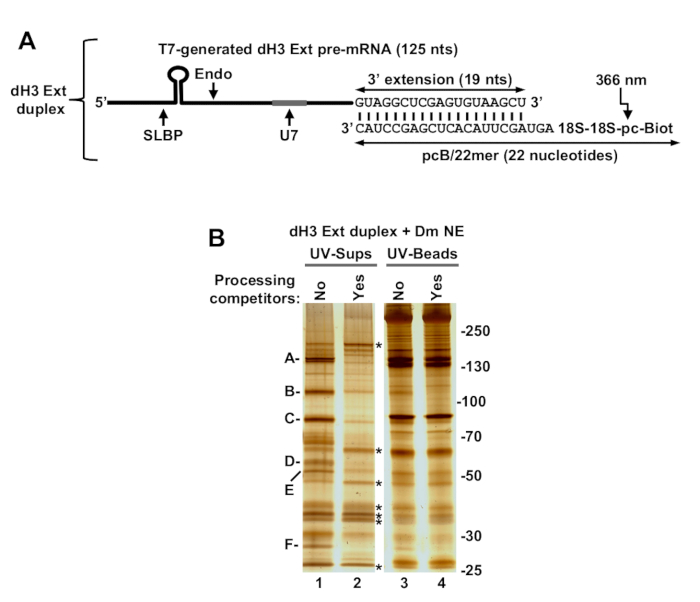
Figure 3: Purification of Drosophila processing complexes assembled on dH3 Ext pre-mRNA attached to the photo-cleavable group in trans. (A) A diagram of the dH3 Ext duplex generated by annealing T7-generated dH3 Ext pre-mRNA and chemically synthesized pcB/22mer oligonucleotide with the following sequence: 5'Biot/pc/18S/18S/mAmGmUmAmGmCmUmUmAmCmAmCmUmCmGmAmGmCmCmUmAmC. In the oligonucleotide, biotin (Biot) is placed at the 5' end and is followed by the photo-cleavable (pc) linker. The last 19 nucleotides (underlined in the sequence above) are complementary to the 3' extension added to the dH3 Ext pre-mRNA. The presence of 2'O-methyl modifications (not indicated in the figure) serves two purposes: it stabilizes the oligonucleotide against extract nucleases and increases the strength of the duplex formed with the dH3 Ext pre-mRNA. (B) dH3 Ext duplex was incubated with a Drosophila Kc nuclear extract either in the absence or in the presence of processing competitors, immobilized on streptavidin beads, extensively washed and UV-eluted along with the bound proteins. The same fractions of the UV-eluted material (UV-sups, lanes 1 and 2) and the beads following UV-elution (UV-beads, lanes 3 and 4) were analyzed by silver staining. Specific componenst of the processing complexes (those that are eliminated by processing competitors) are indicated with A to F letters. Major non-specific RNA binding proteins (those that are UV-eluted but persist in the presence of the two processing competitors) are indicated with asterisks. Position of protein size markers (in kDa) is indicated to the right. Please click here to view a larger version of this figure.
Discussion
The method described here is straightforward and besides incorporating a photo-cleavable linker and the UV-elution step does not differ from the commonly used methods that take advantage of the extremely strong interaction between biotin and streptavidin. The UV-elution step is very efficient, typically releasing more than 75% of the immobilized RNA and associated proteins from streptavidin beads, leaving behind a high background of proteins that non-specifically bind to the beads. By eliminating this background, the UV-elution step yields remarkably pure samples suitable for direct analysis by silver staining, mass spectrometry and functional assays.
As a result of involving a relatively few and simple steps, the UV-elution method is highly reproducible, providing sufficient amounts of the U7 snRNP for silver staining from as little as 100 µL of mouse and Drosophila nuclear extracts15. The method is an attractive alternative to other experimental strategies previously used to purify RNA/protein complexes, including tagging RNA substrates with an MS2 binding site and immobilizing associated complexes on amylose beads via an MS2-MBP fusion protein. The MS2 strategy, while successful in isolating relatively abundant spliceosomes and canonical 3' end processing complexes22,23, failed to yield sufficient amounts of processing complexes containing U7 snRNP (ZD, unpublished results), which is approximately 100 times less concentrated in animal cells than the major U1 spliceosomal snRNP.
Many aspects of the protocol need to be modified to meet various individual research goals and to produce optimal results with other RNA/protein complexes. First, it is important to match the amount of the RNA substrate used for the complex assembly with the amount of this complex that exists in the extract. For limiting complexes, including U7 snRNP, using too much substrate is counterproductive, only increasing the background of non-specific RNA-binding proteins in the UV-supernatant. Second, it is also important to optimize the length and the temperature of incubation to promote vigorous assembly of the complex, preventing at the same time its dissociation due to potential proteolysis or excessive RNA degradation.
A key difficultly in working with the U7-dependent processing complexes and similar RNA/protein complexes that carry enzymatic activities is that after being fully assembled they may dissociate as a result of catalysis. Cleavage of histone pre-mRNAs occurs rapidly even at low temperatures, significantly reducing the yield of the purified processing complexes. To prevent this undesirable effect, the major cleavage site and four nearby nucleotides in pcB-dH3/5m pre-mRNA were modified during chemical synthesis with 2'O-methyl groups (5m)10. This pre-mRNA is almost completely resistant to cleavage by U7 snRNP and can be incubated with a nuclear extract at room temperature for 60 min without causing detectable complex disruption. The T7-generated dH3 Ext annealed to pcB/22mer lacks these modifications and its incubation with a nuclear extract is carried out on ice for 5 min or shorter to limit catalysis. pcB-SL RNA contains mature 3' end of histone mRNA and in cytoplasmic extracts does not undergo any addition processing. 3'hExo, which is magnesium-dependent 3'-5' exonuclease is capable of removing 2-3 nucleotides of the single stranded tail in vivo24,25 but in vitro its activity is inhibited by the presence of EDTA16. As a result, pcB-SL RNA can be incubated in cytoplasmic extracts at room temperature for an extended time without any adverse effects, although typically a short incubation on ice is sufficient to form a ternary complex of the RNA with SLBP and 3'hExo.
Of the two alternative variants of the UV-elution method, using RNA substrates with biotin and the photo-cleavable linker in cis (covalently attached to the 5' end of RNA) is overall simpler and yields relatively pure samples. An important limitation of this approach is that the two groups can be covalently attached to RNA substrates not exceeding ~65 nucleotides. Longer RNA binding targets can be attached to the pcB moiety in trans via an adaptor oligonucleotide. However, this approach may produce higher background of non-specific proteins that tend to bind excess of the adaptor oligonucleotide. It is therefore important to use only a minimal molar excess of the oligonucleotide sufficient to bind all the pre-mRNA substrate used in the experiment.
The sequence, length and chemical nature of adaptor oligonucleotides can be varied to improve their ability to base pair with the complementary sequences in the RNA substrates, while reducing their ability to interact with non-specific RNA binding proteins. Finally, longer pre-mRNA substrates duplexed with adaptor oligonucleotides can be immobilized on streptavidin beads prior to being used for complex formation. One advantage of this pre-selection approach is that it eliminates from the sample RNA molecules that failed to anneal to the oligonucleotide. These molecules retain a normal ability to assemble into a complex in the extract but are unable to bind streptavidin beads, partially reducing the yield of purification.
In addition to purifying complexes assembled on single-stranded RNA substrates, the UV-elution method in a similar manner can be used to purify proteins or protein complexes that bind single- and double-stranded DNA sequences.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
We thank our colleagues and collaborators for their contribution to our work. This study was supported by the NIH grant GM 29832.
Materials
| Streptavidin-Agarose | Sigma | S1638-5ML | |
| High-intensity, long-wave UV lamp | Cole-Parmer | UX-97600-00 | |
| Replacement bulb | Cole-Parmer | UX-97600-19 | 100 Watts Mercury H44GS-100M bulb emitting 366 nm UV light (Sylvania) |
| RNAs and oligonucleotides containing biotin and photo-cleavable linker in cis | Dharmacon (Lafayette, CO) or Integrated DNA Technologies, Inc. (Coralville, IA) | Requst a quote in Dharmacon | |
| Beckman GH 3.8 swing bucket rotor | Beckman | ||
| RiboMAX Large Scale RNA Production Systems (T7 polymerase) | Promega | P1300 | |
| RiboMAX Large Scale RNA Production Systems (SP6 polymerase) | Promega | P1280 | |
| Pierce Silver Stain Kit | ThermoFisher Scientific | 24612 |
Riferimenti
- Mandel, C. R., Bai, Y., Tong, L. Protein factors in pre-mRNA 3′-end processing. Cellular and Molecular Life Sciences. 65 (7-8), 1099-1122 (2008).
- Dominski, Z., Carpousis, A. J., Clouet-d’Orval, B. Emergence of the beta-CASP ribonucleases: highly conserved and ubiquitous metallo-enzymes involved in messenger RNA maturation and degradation. Biochimica et Biophysica Acta. 1829 (6-7), 532-551 (2013).
- Dominski, Z., Marzluff, W. F. Formation of the 3′ end of histone mRNA: getting closer to the end. Gene. 396 (2), 373-390 (2007).
- Yang, X. C., Torres, M. P., Marzluff, W. F., Dominski, Z. Three Proteins of the U7-Specific Sm Ring Function as the Molecular Ruler To Determine the Site of 3 ‘-End Processing in Mammalian Histone Pre-mRNA. Molecular and Cellular Biology. 29 (15), 4045-4056 (2009).
- Sabath, I., et al. 3 ‘-End processing of histone pre-mRNAs in Drosophila: U7 snRNP is associated with FLASH and polyadenylation factors. RNA. 19 (12), 1726-1744 (2013).
- Skrajna, A., et al. U7 snRNP is recruited to histone pre-mRNA in a FLASH-dependent manner by two separate regions of the stem-loop binding protein. RNA. 23 (6), 938-951 (2017).
- Smith, H. O., et al. Two-step affinity purification of U7 small nuclear ribonucleoprotein particles using complementary biotinylated 2′- O-methyl oligoribonucleotides. Proceedings of the National Academy of Sciences, USA. 88, 9784-9788 (1991).
- Pillai, R. S., et al. Unique Sm core structure of U7 snRNPs: assembly by a specialized SMN complex and the role of a new component, Lsm11, in histone RNA processing. Genes and Development. 17 (18), 2321-2333 (2003).
- Pillai, R. S., Will, C. L., Luhrmann, R., Schumperli, D., Muller, B. Purified U7 snRNPs lack the Sm proteins D1 and D2 but contain Lsm10, a new 14 kDa Sm D1-like protein. EMBO Journal. 20 (19), 5470-5479 (2001).
- Yang, X. C., et al. A Complex Containing the CPSF73 Endonuclease and Other Polyadenylation Factors Associates with U7 snRNP and Is Recruited to Histone Pre-mRNA for 3 ‘-End Processing. Molecular and Cellular Biology. 33 (1), 28-37 (2013).
- Shimkus, M., Levy, J., Herman, T. A chemically cleavable biotinylated nucleotide: usefulness in the recovery of protein-DNA complexes from avidin affinity columns. Proceedings of the National Academy of Sciences of the United States of America. 82 (9), 2593-2597 (1985).
- Hirsch, J. D., et al. Easily reversible desthiobiotin binding to streptavidin, avidin, and other biotin-binding proteins: uses for protein labeling, detection, and isolation. Analytical Biochemistry. 308 (2), 343-357 (2002).
- Olejnik, J., Krzymanska-Olejnik, E., Rothschild, K. J. Photocleavable biotin phosphoramidite for 5′-end-labeling, affinity purification and phosphorylation of synthetic oligonucleotides. Nucleic Acids Research. 24 (2), 361-366 (1996).
- Olejnik, J., Krzymanska-Olejnik, E., Rothschild, K. J. Photocleavable affinity tags for isolation and detection of biomolecules. Methods in Enzymology. 291, 135-154 (1998).
- Skrajna, A., Yang, X. C., Dadlez, M., Marzluff, W. F., Dominski, Z. Protein composition of catalytically active U7-dependent processing complexes assembled on histone pre-mRNA containing biotin and a photo-cleavable linker. Nucleic Acids Research. 46 (9), 4752-4770 (2018).
- Dominski, Z., Yang, X. C., Kaygun, H., Dadlez, M., Marzluff, W. F. A 3 ‘ exonuclease that specifically interacts with the 3 ‘ end of histone mRNA. Molecular Cell. 12 (2), 295-305 (2003).
- Yang, X. C., Purdy, M., Marzluff, W. F., Dominski, Z. Characterization of 3’hExo, a 3′ exonuclease specifically interacting with the 3′ end of histone mRNA. Journal of Biological Chemistry. 281 (41), 30447-30454 (2006).
- Tan, D., Marzluff, W. F., Dominski, Z., Tong, L. Structure of histone mRNA stem-loop, human stem-loop binding protein, and 3’hExo ternary complex. Science. 339 (6117), 318-321 (2013).
- Mayeda, A., Krainer, A. R. Preparation of HeLa cell nuclear and cytosolic S100 extracts for in vitro splicing. Methods in Molecular Biology. 118, 309-314 (1999).
- Dominski, Z., et al. 3′ end processing of Drosophila histone pre-mRNAs: Requirement for phosphorylated dSLBP and co-evolution of the histone pre-mRNA processing system. Molecular and Cellular Biology. 22, 6648-6660 (2002).
- Dominski, Z., Yan, X. C., Purdy, M., Marzluff, W. F. Differences and similarities between Drosophila and mammalian 3 ‘ end processing of histone pre-mRNAs. Rna-a Publication of the Rna Society. 11 (12), 1835-1847 (2005).
- Jurica, M. S., Licklider, L. J., Gygi, S. R., Grigorieff, N., Moore, M. J. Purification and characterization of native spliceosomes suitable for three-dimensional structural analysis. RNA. 8 (4), 426-439 (2002).
- Shi, Y., et al. Molecular architecture of the human pre-mRNA 3′ processing complex. Molecular Cell. 33 (3), 365-376 (2009).
- Thomas, M. F., L’Etoile, N. D., Ansel, K. M. Eri1: a conserved enzyme at the crossroads of multiple RNA-processing pathways. Trends in Genetics. 30 (7), 298-307 (2014).
- Lackey, P. E., Welch, J. D., Marzluff, W. F. TUT7 catalyzes the uridylation of the 3′ end for rapid degradation of histone mRNA. RNA. 22 (11), 1673-1688 (2016).

