Quantitative Approaches for Studying Cellular Structures and Organelle Morphology in Caenorhabditis elegans
Summary
This study outlines quantitative measurements of synaptic size and localization, muscle morphology, and mitochondrial shape in C. elegans using freely available image processing tools. This approach allows future studies in C. elegans to quantitatively compare the extent of tissue and organelle structural changes as a result of genetic mutations.
Abstract
Defining the cellular mechanisms underlying disease is essential for the development of novel therapeutics. A strategy frequently used to unravel these mechanisms is to introduce mutations in candidate genes and qualitatively describe changes in the morphology of tissues and cellular organelles. However, qualitative descriptions may not capture subtle phenotypic differences, might misrepresent phenotypic variations across individuals in a population, and are frequently assessed subjectively. Here, quantitative approaches are described to study the morphology of tissues and organelles in the nematode Caenorhabditis elegans using laser scanning confocal microscopy combined with commercially available bio-image processing software. A quantitative analysis of phenotypes affecting synapse integrity (size and integrated fluorescence levels), muscle development (muscle cell size and myosin filament length), and mitochondrial morphology (circularity and size) was performed to understand the effects of genetic mutations on these cellular structures. These quantitative approaches are not limited to the applications described here, as they could readily be used to quantitatively assess the morphology of other tissues and organelles in the nematode, as well as in other model organisms.
Introduction
The nematode Caenorhabditis elegans (C. elegans) is increasingly utilized as a model system to uncover the biological and molecular processes involved in human disease. An adult nematode has a body length of just over 1 mm, and can produce a large brood of up to 300 eggs1. After hatching, C. elegans only require 3-4 days to reach adulthood, and live for around 2 to 3 weeks2. Due to its ease of culturing, C. elegans is currently one of the most sought-after in vivo animal models for conducting cost-effective, rapid drug screening to identify therapeutics for human diseases. Additionally, its genetic conservation, well defined behavioral paradigms, transparent body for fluorescence or light microscopy, and ease of genetic manipulation make the study of cellular and molecular consequences of genetic mutations readily achieveable3. The C. elegans genome shares approximately 60-80% orthology with human genes, and about 40% of those genes are known to be disease-related. Some of human diseases that have been modelled and studied in C. elegans include neurodegenerative disorders (Alzheimer’s disease, Parkinson’s disease, amyotrophic lateral sclerosis, Charcot-Marie-Tooth disease), muscle-associated diseases (Duchenne muscular dystrophy), and metabolic diseases (hyperglycemia)2,4. In most human disorders, disease-induced cellular and organelle localization and morphological changes occur, which can readily be evaluated in the nematode model.
Fluorescent markers have been widely used to label tissues and organelles for dynamic visualization under the microscope. However, in C. elegans, conventional methods that assess morphological irregularities due to genetic mutations have largely relied on visual descriptions. While qualitative assessments can cover wider ranges of phenotypic descriptions (synaptic morphology, GFP clumping, specific axonal shape, muscle fiber thickness, etc.) and provide a bird’s eye view of the morphological changes, they are less well suited for comparing small variations across different groups. Furthermore, qualitative assessments are based on visual, subjective evaluation, which may lead to over- or under-estimations of morphological abnormalities. Finally, qualitative observations can also vary greatly between individuals, creating difficulties with data replication.
In recent years, a number of user-friendly, readily available computational algorithms that can quantitatively analyse images have been developed. However, the utilization of such image analysis software for some morphological studies, especially in relation to body wall muscles and mitochondria, in C. elegans research has lagged behind. To improve underlying structural analysis in C. elegans, some of the readily available, open-source image analysis software were trialed to quantitatively compare the effects of genetic mutations on muscle mitochondria, body wall muscle and synaptic morphology. These experimental procedures outline in detail how these programs (Fiji, ilastik, CellProfiler, SQUASSH) can be used to evaluate changes in synaptic size and synaptic protein localization, body wall muscle area and fiber length, and mitochondrial size and circularity as a result of genetic mutations in the nematode.
Protocol
1. Growth and maintenance of C. elegans strains
- Seed Nematode Growth Medium (NGM, see Table of Materials) agar plates with 300 µL of the slow-growing E. coli strain OP50 in a laminar flow cabinet.
- Leave the NGM agar plates in the laminar flow cabinet to dry.
NOTE: In the absence of laminar flow cabinet, plates can be left to dry on the bench but they are more prone to contamination. - Transfer at least 20 animals to each of two OP50-seeded NGM agar plates to have working stocks. There are two main ways to transfer animals.
- Picking: Gently lift the animals using a heat-sterilized pick. This method is effective for selecting individual or multiple animals from an agar plate. Having a small scrape of bacteria at the tip when picking helps the transfer.
NOTE: A pick is made of a small piece of platinum wire about 4 cm long that is embedded in a glass pipette, with the wire tip flattened to be ‘spear-like’. Heat-sterilize the pick between picking to prevent cross-contamination. - Chunking: Using a sterilized spatula, cut a small square of agar containing the animals from one plate and transferred upside-down to a new plate.
NOTES: This method is generally useful for transferring large numbers of worms, and to remove contamination from plates through multiple rounds of chunking non-contaminated agar. Avoid cross-contamination by flaming the spatula for a few seconds and immersing the spatula in 100% ethanol between transfers. Transferring from starved plates is not recommended as starvation and overcrowding can affect the health and morphology of cellular and subcellular structures.
- Picking: Gently lift the animals using a heat-sterilized pick. This method is effective for selecting individual or multiple animals from an agar plate. Having a small scrape of bacteria at the tip when picking helps the transfer.
- Maintain the worms at standard growth temperature (20 °C). To ensure continuous supply of well-fed worms, transfer the worms to new OP50-seeded plates every 2 to 3 days.
2. Age-synchronizing C. elegans
- Maintain the worms on 3-4 NGM plates until the population is crowded but not starved.
- Gently wash the worms off the NGM plate with M9 solution. The eggs will remain on the plate since they stick to the E. coli OP50. Repeat with four additional wash steps to remove all hatched animals.
- Allow the worms to hatch for 40 min.
- Add 1-2 mL of M9 solution to the plate and gently swirl to loosen recently hatched worms.
- Collect the M9 solution with recently born worms and transfer the solution with worms to a fresh NGM plate.
- Allow the M9 solution to evaporate by exposing the NGM plate to air. Do this adjacent to an open flame to avoid contamination of the NGM plate.
- Grow the worms for 27 h at 20 °C to allow development into the third larval (L3) stage. For synchronized 3-day old adult animals, the worms are picked at larval stage 4, and grown for a further 3 days to reach 3-day old adult stage.
3. Preparation of slides for imaging
- Prepare 3-4% w/v agarose stock in a microwave-safe bottle (3-4 g in 100 mL of ultrapure water).
NOTE: Agar can be used in place of agarose at the same concentration. - Melt the agarose carefully in a microwave oven, taking care not to over boil, until all agarose dissolves (clear solution). Keep the solution at 70 °C to prevent solidification.
NOTE: Agarose stock can be used several times. Reheat before use if the agarose has solidified. - Using a Pasteur pipette, place a drop of melted agarose onto a microscope slide.
NOTE: Avoid having air bubbles in the drop of agar as these disrupt the integrity of the pad. - Immediately press down the agarose droplet with another microscope slide, gently flattening the agarose into a pad between the two slides.
NOTE: Any leftover bubbles should be removed at this stage. If not, new slides should be made as animals can easily get stuck in the bubbles. Avoid overflow of agar to the side when pressing on the slide. - Leave the agarose to dry for 1 min.
- Carefully separate the two slides to avoid damage to the agarose, while leaving the solidified pad on one of the slides.
NOTE: Slides with agarose pads should always be prepared fresh just before the experiment as they can dry out. Make sure the agarose pad has a consistent thickness. There should be no air bubbles or cracks in the pad as this may interfere with imaging. - Add a small drop (5-10 µL) of 0.05% tetramisole hydrochloride (anesthetic) to the center of the agarose pad.
- Using heat-sterilized pick, quickly transfer about 10 – 15 animals from the stock plate to the anesthetic droplet before the solution dries out. Avoid transferring too much OP50 bacteria as this makes the solution cloudy, which can interfere with imaging.
NOTE: If the anesthetic solution dries out while picking, simply pipette a second drop of 0.05% tetramisole hydrochloride onto the animals. The animals tend to lay on their lateral side in the anesthetic solution. - Apply a coverslip by positioning it just above the agarose pad and gently dropping it down. This will prevent the formation of bubbles. Do not apply pressure to the coverslip as this may crush the animals.
- Label the slides with the strain name.
4. Assessing synaptic morphology
NOTE: The effects of MEC-17 overexpression on synapse integrity in the posterior lateral microtubule (PLM) touch receptor neurons were studied by quantifying the synaptic size and localization using line scanning confocal microscopy. The PLM neurons (including the synaptic regions) were visualized using the uIs115(Pmec-17::tagRFP) transgene (strain: TU40655) and the synaptic region was specifically labeled with jsIs37(Pmec-7::snb-1::GFP) (strain: NM6646). This study was performed in synchronized L3 non-transgenic and transgenic animals of the extrachromosomal MEC-17 overexpression strain BXN507 [cjnEx036(Pmec-4::mec-17, Pmyo-2::mCherry); jsIs37; uIs115]7. A complete list of strains used in this study is included in Table 1.
- Confocal imaging of touch receptor neuron synapses
- To image the synaptic regions, use a line scanning confocal microscope coupled to a 488 nm and 552 nm optically pumped semi-conductor diode laser and equipped with image capture software.
- Locate the worms using the lowest magnification.
- When the animals are successfully situated, switch to a 40X magnification and apply immersion medium (in this study: a 40X/1.10 HC PL APO CS2 multi-immersive objective with water immersion was used).
- Visualize the synaptic region by exciting the fluorophores with a 488 nm laser (2% power) for the GFP fluorophore and 552 nm (1% power) for the tagRFP fluorophore.
- Determine optimal x and y frame to capture the entire synaptic region. Make sure to consider the optimal pixel size when selecting the frame. In this study, a consistent pixel size of 56 nm was obtained.
- Capture images using a hybrid photo-detector and set the gains to prevent overexposure of the fluorophores (100% for 488 nm excitation and 15% for 552 nm excitation).
- Collect z-stack images to cover the entire synapse, using the optimal z-step size depending on the specific objective used. In this study, an optimal z-step size of 0.211 nm was used.
NOTE: Ensure all images are taken under stringent condition to allow integrated fluorescence quantification and comparison. This can be done by keeping microscopy parameters identical across all captured images (i.e., laser and detector settings, pixel size, scanning speed, and optimal z-step size). Optimal settings depend on the objective and can be calculated using accessible bioinformatics programs, such as the Nyquist calculator (https://svi.nl/NyquistCalculator). Label images systematically, this will be useful when using bioinformatics software that will organize and process files based on filename.
- Quantification of the synaptic region
- To analyze the synaptic region, export images as TIFF files. Java based imaging processing software such as ImageJ, can be used (in this study, the MRI image conversion macro in ImageJ was used).
- The size and the integrated fluorescence intensity levels of synapses were measured from sum intensity of the obtained z-stacks with CellProfiler-3.1.5. Load the images into the CellProfiler3.1.5 software. If required, images can be filtered based on image file names.
- Set up the Name and Type module. Optionally, extract the metadata from the image label to organize calculated data accordingly.
- Determine the synaptic region by hand definition of this region using the diffuse tagRFP expressed from the uIs115(Pmec-17::tagRFP) transgene. This can be done by adding the regarding module to the CellProfiler-3.1.5 pipeline.
- To measure the size and fluorescence of the hand defined synapse, add the Measure object size and shape and Measure object intensity module to the pipeline.
- Calculate the relative integrated fluorescence intensity by adding the Calculate Math module. Divide the integrated intensity units obtained from jsIs37(Pmec-7::snb-1::GFP) by the integrated units obtained for uIs115(Pmec-17::tagRFP).
- Export measurements and calculations.
5. Quantifying body wall muscle structure
- Fluorescence imaging of body wall muscle structure
- Obtain a strain (for example, RW1596) carrying the transgene stEx30(Pmyo-3::gfp::myo-3 + rol-6(su1006))8, which labels myosin fibers with GFP.
NOTE: Introduction of the transgene into animals can be achieved by either crossing a strain of interest with animals that already express the transgene, or microinjecting DNA into the distal arm of the gonad. The ‘rolling’ phenotype induced by rol-6 mutation in this transgene facilitates the visualization of the muscles. The body wall muscles run along the dorsal and ventral sides that are normally precluded from view in wild-type animals because they typically lie on one of their lateral sides. The rol-6(su1006) allele induces twisting of the animals, thereby exposing some muscles cells for imaging. - Image the animals using a fluorescence microscope coupled with a high resolution charged coupled device camera (CCD), 40X objective or higher, GFP filter and acquisition software.
- Adjust the exposure time and illumination intensity to prevent oversaturated images.
- Capture and save images of muscles (400X magnification) from a section of the animal containing at least one complete visible oblique muscle cell. Avoid regions with a single muscle cell that is incomplete or sections that are out of focus. Also exclude images from extreme anterior and posterior regions, and regions adjacent to the vulva.
NOTE: If the animal does not have a single visible complete muscle cell, gently slide the coverslip to turn the animal so as to expose a complete cell.
- Obtain a strain (for example, RW1596) carrying the transgene stEx30(Pmyo-3::gfp::myo-3 + rol-6(su1006))8, which labels myosin fibers with GFP.
- Measurement of muscle cell area
- Open the image in Fiji software9 (version 2.0.0 was used in this study).
- Use the polygon selection to carefully trace around a single oblique muscle cell. Adjust the line of the polygon at the end by dragging the anchor dot to improve tracing.
- Go to Analyze tab at the top of the software and click Measure to calculate the area of the selection. A separate results table containing the area and other measurements will be shown. If the area measurement is missing from the results table, go to Set Measurements under the Analyze tab and check that the area box is ticked. Likewise, untick other measurements that are not needed.
- For muscle cells with a degenerated/missing region, trace the missing region with the polygon selection tool and click Measure again. If there are multiple gaps within the cell, trace each gap separately.
- Calculate the ratio of gap area to the entire single muscle cell. A high ratio indicates a higher extent of muscle degeneration due to large gaps within the cell. If there are no missing regions, as commonly seen in wild-type animals, the ratio would be calculated as zero.
NOTE: As a control, score also for muscle structure defects visually and compare results with the quantitative measurement. This is especially useful if the strain is being studied for the first time in the lab. For phenotypic scoring, assess the animals as defective or not-defective based on the integrity of the filaments. For example, animals with loss of distinct filament striations, GFP clumping, or gaps within muscle cells due to structural breakdown, would be scored as defective.
- Segmentation and quantification of myosin fiber length
- If necessary, first convert files to .tif using Fiji or another software package. Save the TIFF files in desired location.
- Open ilastik10 (free software, version 1.3.2 can be downloaded from https://www.ilastik.org/).
- Once in ilastik, choose Pixel Classification under Create Nuovo Project. The software will prompt for the project to be saved. Rename the project and save to a desired location.
NOTE: A new project only needs to be created once. Subsequent segmentation can be done using the same project. To use the same project settings, go to Project tab at the top, and click on Open Project to access the project file. - There should be 5 tabs on the left panel (Input Data, Feature Selection, Training, Prediction Export and Batch Processing). In the Input Data tab, click on Add Nuovo and then Add separate image(s). Direct the pop-up window to the folder containing the TIFF files and select the image to open.
- In the next tab below (Feature Selection), click on Select Features to select all pixel features, indicated by green ticked boxes. The pixel selection in this step is used to distinguish the different classes of pixels in the next step.
NOTE: Pixel feature selection depends on the image resolution. Hence, the developers suggest to select all or as many features as possible, if the processing power and time permit. - The following Training panel allows the classification of different object classes based on pixels. In this case, the two classes are the individual GFP-expressing myosin filaments and unwanted background or blurred edges. Click on Add Label to have the first label. Scrawl over a number of filaments to begin classifier training.
NOTE: Double-clicking the label allows for name change (for example, renaming ‘label 1’ to ‘filament’). Double-clicking the colored box allows colors for drawing and probability displays to be altered. The default size of the paintbrush is 1, although the size can be increased to 61. If the wrong pixel has been drawn, simply use the eraser tool to remove the drawing. Keyboard controls (-) and (Shift +) can be used to zoom in and out of image, respectively. Arrow keys are used to navigate around the image. - Add a second label and rename it to Background. Scrawl over unwanted backgrounds and spaces between the filaments.
- Click on Live Update and make sure that classification has been done by the software correctly. Add more scrawls and fine tune the training if necessary.
NOTE: It is important to keep an eye out for merged fibers and blurry edges (such as body line) in this step. Improve the prediction here as much as possible to increase the accuracy of measurements. The prediction accuracy also depends on image resolution, as pixels in low quality images are more difficult to tease apart. It is also optional but recommended to run the Filter Method in the Suggest Features function next to Live Update control. - Once training classifier has been adequately performed, go to Prediction Export panel. Click on Choose Export Image Settings with Probabilities as the source.
- Use the following settings. Cut-out subregion x and y ticked, and c unticked. The start and stop values for c should be 0 and 1, respectively. Tick and convert data type in Transformations to unsigned 8-bit, tick renormalize [min, max] and use the default values. Change the format of the output file video to PNG, and rename the file and directory as desired.
- Close the export settings window and export the specific image that has just been segmented.
- Open the saved PNG image in Fiji software. The image should appear in black and white.
- Adjust the threshold of the image using the default settings.
- Apply the skeletonization plugin (Skeletonize 2D/3D, and then Analyze Skeleton). A separate table of results will be displayed, which includes branch length. The branch length represents the fiber length. Other data in the results could also be useful, such as Euclidean distance.
NOTE: Omit fibers with lengths of 0 µm or more than 250 µm, as these sizes are not physiologically possible.
6. Quantifying mitochondrial morphology
NOTE: For the quantification of mitochondrial morphology, this study used the BXN0387 strain containing the uIs115(Pmec-17::tagRFP)5 transgene to visualize the PLM neurons and jsIs609(Pmec-4::MLS::GFP)11 to visualize mitochondria specifically within the PLM neurons. This study was performed in synchronized 3 day old adult worms, but has successfully been performed at other ages, as well as in other tissues7.
- Fluorescence imaging of mitochondria within the PLM neurons
- To measure size and shape changes of mitochondria within the PLM neurons, visualize both the background neuron and mitochondria using fluorescent transgenes.
- To image the PLM neuron, use a confocal microscope coupled to a 488 nm and 552 nm optically pumped semi-conductor diode laser and equipped with image capture software.
- Image the worm as per steps 4.1.1-4.1.4 above, ensuring non-saturating exposure conditions.
- In order to visualize the entire neuron, a tiled image may be needed. To achieve this, use software with a tile scan option to locate and drop a marker at one corner of the neuron and then locate the opposite end and drop the other marker. The software will calculate the optimal number of tiles needed to capture the required area.
NOTE: To reduce scanning time, it is often easier to locate neurons that follow one single plane, resulting in fewer tiles. - To couple the tile scan with a Z-stack, set the lower and upper limits prior to starting the tile scan, and ensure that the optimal slice size is set.
- Ensure resolution is optimized, as segmentation will be more refined with higher resolutions (here, a resolution of 3224 x 3224 pixels was used).
NOTE: Whilst the uIs115(Pmec-17::tagRFP) transgene is used to mark the PLM and successfully position the camera, it does not necessarily need to be imaged due to slight background fluorescence observed from the jsIs609(Pmec-4::MLS::GFP) transgene within the PLM axon itself. Thus, scanning and imaging time can be reduced by removing the 552 nm filter from the experiment.
- Object segmentation using SQUASSH
- Convert image files into .tiff images and generate maximum intensity projections from Z-stacks using Fiji imaging software. Crop images to the minimal size while still containing the complete neuron to reduce computational load and reduce background noise.
NOTE: For the representative results, only mitochondria within the long axonal processes were considered, so the cell body and any mitochondria within were excluded from each image. - Using the Mosaic Suite SQUASSH macro for ImageJ, perform Split Bregman segmentation12 on the maximum projections. A detailed guide to the installation and application of this macro is available on the Mosaic website (http://mosaic.mpi-cbg.de/) and described by Rizk et al. 201413. The process of image segmentation is described in short below.
NOTE: This segmentation identifies objects (in this case individual mitochondrion) over a threshold of fluorescence intensity, allowing for accurate measurement of each individual objects size and shape. - Remove background noise from an image using the rolling ball method, in which a user defined window moves across the image, imputing the local background intensity estimate from the most frequently occurring intensity in each window. This corrects uneven background intensities.
- Use object detection to roughly find regions of the image that contain objects (mitochondria) of interest, based on the fluorescence intensity and number of pixels. The parameters that govern this step are regularization and minimum object intensity.
NOTE: Regularization is used to avoid segmenting small intensity, noise-induced peaks, and the minimum object intensity threshold helps to define the subcellular organelles from background tissue fluorescence. These are the main two parameters that are manipulated to find an optimal segmentation of mitochondria, and the optimization process tends to be trial and error of different parameters for a test image, followed by manual inspection. - Use the software to estimate the local background intensities around each object to aid computation of optimal segmentation.
- Measure individual objects for size, perimeter, length via pixel number, as well as signal intensity and x/y location on the image. To calculate measurements in size (nm), factor in the pixel size of the original image.
NOTE: For each experiment, optimal parameters must first be calculated. To ensure accuracy of segmentation, it is advisable to perform a SQUASSH analysis on a test image and manually inspect the accuracy of segmentation. The macro provides output files showing the individual objects that were identified on the original image, and inspecting these closely and adjusting parameters to suit will ensure an accurate segmentation. The defined parameters must then be used for the entire experiment for comparability. The parameters used for the representative results are included in Table 2. - Open Fiji or ImageJ analysis software, navigate to the Mosaic macro container and open the SQUASSH menu.
- Open the Background Subtraction submenu and ensure Remove background? is checked. The default for window size is 10.
- Set the desired Regularization and Minimum object intensity, channel 1 values to best identify and segment objects (see step 6.2.4). Channel 2 values are not necessary for objects marked with a single transgene.
- Click the Estimate PSF from objective properties to estimate the Point Spread Function based on the microscope’s characteristics. This helps the segmentation of closely located objects by accounting for the influence of each pixel on its neighboring pixel.
NOTE: Subpixel segmentation was not used in this experiment due to its increased load on computer processing power. Cell masks were also not used as the axon is easily defined. These settings are useful for more complex tissues and to define objects on a cell by cell basis. - For visualization options, ensure Object outlines and Save object and image characteristics are checked. For use later in graphics and statistical software, ensure the correct number of conditions are input.
- Locate the folder with .tiff images and run the macro. This process can be performed on an individual image or on a batch of images per genotype located within a folder. Output files will be located in the folder along with the original images.
- Convert image files into .tiff images and generate maximum intensity projections from Z-stacks using Fiji imaging software. Crop images to the minimal size while still containing the complete neuron to reduce computational load and reduce background noise.
- Analysis and measurements of shape
- Use the output from SQUASSH to provide an itemized object data .csv file, which provides each individual object per row including its measurements as described above.
NOTE: While this macro process automates object measurements, it is always important to manually check output images after SQUASSH segmentation to ensure the macro has properly identified mitochondria. - To analyse the shape of each mitochondria, use a two-dimensional measure for sphericity, Circularity, to estimate the deviation of the object from a perfect circle.
NOTE: Circularity is a function of area and perimeter (Circularity = (4 x π) x (Area/Perimeter2) where 1 = a perfect circle and 0 = a straight line. This can be achieved using a formula in graphics and statistical software. - To determine the variance in size and shape across the pool of mitochondria, calculate histograms and a fitted Gaussian distribution using graphics and statistical software, with bins of 0.2 μm for mitochondrial size and 0.05 for mitochondrial circularity.
- Use the output from SQUASSH to provide an itemized object data .csv file, which provides each individual object per row including its measurements as described above.
Representative Results
C. elegans is an ideal model organism for studying morphology of different tissues and organelles due to its simplicity, known cell lineage, transparency, and available tools. Here, we provide quantitative approaches for studying organelles (e.g., mitochondria) and tissues, including synapses and muscles using live fluorescence imaging and free bio-image processing software.
Strict regulation of MEC-17 expression is required for normal synapse development
To further understand the function of the alpha-tubulin acetyl-transferase MEC-1714,15,16 in the nervous system, we studied synaptic morphology in the touch receptor neurons in C. elegans overexpressing the protein17. These cellular structures are essential for neuron function as they allow signals to be passed on to neighboring neurons. Images of the PLM synapse were collected using a laser scanning confocal microscope and quantitative morphology analysis was performed using accessible bio-image processing software (Figure 1A,B). The overexpression of MEC-17 results in significant reductions in the area of pre-synaptic accumulations in PLM and reduced relative integrated fluorescence intensities of the SNB-1::GFP marker (Figure 1C,D). Together, these results suggest that overexpression of MEC-17 disrupts normal synapse development in the PLM touch receptor neurons.
The size of the presynaptic site and relative integrated fluorescence were studied using the free bio-image processing software CellProfiler3.1.5. Due to their complex structure, synapses were defined by hand using the diffuse tagRFP protein expressed from the uIs115(Pmec-17::tagRFP) transgene. The area of the defined synapse was calculated by the number of pixels within the defined region. Compared to the wild-type control, animals overexpressing MEC-17 have significantly reduced pre-synaptic regions (P = 0.0025; n ≥ 10) (Figure 1C). In addition, to study synaptic integrity, relative integrated fluorescence intensity levels were compared. This was calculated by measuring integrated fluorescence intensity levels within the defined synapse for both SNB-1::GFP and the diffusible tagRFP. Subsequently, the SNB-1::GFP fluorescence values relative to tagRFP values were compared for which a significant decrease (P = 0.0479; n ≥ 10) was observed with the overexpression of MEC-17 (Figure 1D). Together, these results suggest that the correct regulation of MEC-17 is important for synapse development.
Quantitative measurement of body wall muscle cell area and fiber length reveals significant defects as a result of mutations in CMT2-related genes
Mutation in the genes MFN2, DNM2 and KIF5A have been shown to cause Charcot-Marie-Tooth type 2 (CMT2), an axonal form of the most common inherited peripheral neuropathy18,19. CMT2 is commonly associated with slowly progressive distal muscle weakness and atrophy, distal sensory impairment, foot deformities and secondary steppage gait. As a result, patients often suffer debilitating lifelong mobility impairments. There is currently no cure for the disease, partly due to genetic and clinical heterogeneity associated with CMT219, as well as a lack of animal models to study disease pathophysiology. Hence, using animal models to understand the underlying cellular defects can provide answers to the unknown CMT2 disease mechanisms.
Thus far, published studies evaluating body wall muscle defects in C. elegans have relied on visual scoring rather than quantitative measurements20. In order to avoid subjective bias that can occur during qualitative assessment, quantitative measurements of body wall muscle area and myosin fiber length were trialed using available image processing programs. We used these methods to assess muscle morphology in animals carrying mutations in the fzo-1/MFN2, dyn-1/DNM2 and unc-116/KIF5A genes in 3-day old adult C. elegans. For both measurements, a number of conditions were applied, such that at least a single complete oblique muscle cell was within view, muscle cells that were fuzzy or out of focus were excluded, and extreme anterior and posterior regions, as well as regions adjacent to the vulva were omitted. In addition to the mutations in the CMT2-associated genes, animals also carried the stEx30(Pmyo-3::gfp::myo-3 + rol-6(su1006)) transgene that labels a C. elegans myosin heavy chain subunit with GFP (Figure 2A-E). The simple polygon tool in Fiji software9 (version 2.0.0) was used to measure the area of a single muscle cell. If an animal experienced body wall muscle structure breakdown, leaving behind empty spaces within muscle cells, the ratio of gap area to total single muscle cell area was calculated and compared to wild-type control (Figure 3A). To obtain measurements of myosin fiber length, each image was first segmented using ilastik software10 (version 1.3.0) (Figure 3B). Segmentation is a process that allows the individual muscle fibers to be classified and separated from unwanted sections such as the background. Following segmentation, the image was exported as an unassigned 8-bit binary image. Fiji (version 2.0.0) was next used to Skeletonize the binary image to filter out border pixels, leaving behind fragments representing the original fibers. The branch lengths of the Skeletons, or individual myosin filaments, were then analyzed in Fiji. Length measurements of 0 or more than 250 μm, which is the maximum reasonable length for a filament even with muscle stretching, were excluded. Altogether, between 2000 to 3000 myosin fiber lengths were measured. A summary of the segmentation workflow is shown in Figure 3B. Results from the quantitative analysis were further compared with visual scoring of body wall muscle defects, where 3-day old animals were categorized as defective or not-defective based on muscle structure integrity.
Defects in body wall muscle morphology were observed between 2.5 to 3.5 times higher in fzo-1(cjn020), dyn-1(ky51) and unc-116(e2310) animals than in wild-type animals during visual assessment (Figure 4A). Most animals with mutations in fzo-1 and unc-116 experienced conspicuous loss of muscle striations, large GFP clumping due to accumulation of cellular debris, and muscle fiber degeneration that left gaping space within muscle cells (Figure 2C,E). In contrast, while more than 60% of dyn-1 mutants were scored visually defective, the extent of defects was minimal compared to the other two genetic mutants. There was only slight loss of striations, no GFP aggregations and only minor muscle degeneration when compared to the other two mutants (Figure 2D and Figure 4A). As with other phenotypic scoring, the extent of the defects could not be considered, and a tendency for bias might cause a higher proportion of defects to be registered. Therefore, quantitative measurements of muscle cell area and fiber length would provide a more accurate representation of the extent of muscle defects than visual assessment alone. In line with the large gaps observed with visual scoring, the ratio of gap area to total single cell area was shown to be 5 to 6 times higher in fzo-1 and unc-116 mutant animals compared to the wild-type group, which only experienced negligible muscle structure breakdown (Figure 4B). The lack of structural collapse in dyn-1 mutants resulted in a non-significant, 3-fold increase in gap to muscle cell area ratio (Figure 4B). This also indicates that experimenter bias might be a factor contributing to the high proportion of defects assigned by qualitative assessment. Nevertheless, the small gaps decreased the average fiber length from 24 µm in wild-type to 22 µm in dyn-1 mutants (Figure 4C). The average fiber length was dramatically shorter in the fzo-1 and unc-116 mutants, correlating with the presence of larger gaps within degenerating muscle cells in these animals (Figure 2C,E and Figure 4C). These results are consistent with our recent findings studying the effects of mutating CMT2-asscoiated genes in C. elegans20. Moreover, they highlight the more precise assessment of muscle morphology using these quantitative methods compared to visual scoring alone, and provide evidence that fzo-1 and unc-116 are required for normal muscle morphology.
Quantitative measurements of mitochondrial morphology using SQUASSH object segmentation
To understand how the absence of mitochondrial fission affects mitochondrial morphology in C. elegans neurons, we performed quantitative analyses in the PLM mechanosensory neurons. DRP-1, the C. elegans orthologue of the mammalian Dynamin-related protein 1 (DRP1), controls mitochondrial fission, where upon activation is recruited from the cytosol to form a spiral around mitochondria, constricting and eventually severing both the inner and outer mitochondrial membranes21,22. We fluorescently labeled mitochondria with a tissue specific transgene in the PLM neurons, which have a long axonal process that can be readily visualized using epifluorescence microscopy. A loss of fission is hypothesized to disrupt overall mitochondrial dynamics, and therefore decrease the creation of newer, smaller mitochondria through a process known as budding23.
We performed SQUASSH segmentation to compare mitochondrial morphology in wild-type and drp-1 mutant animals. We analyzed the mitochondria from 6 PLM neurons per genotype, resulting in an n value of greater that 200 mitochondria for each. Schematic and representative images are shown in Figure 5A,B. We found that mutants lacking DRP-1 had larger and more elongated mitochondria (Figure 5B-F). Mitochondria in the drp-1 mutant background were 32% larger than wild-type (P = 0.0006, Figure 5C), and 14% more elongated (further from a perfect circle) (P < 0.0001, Figure 5D). For a more detailed assessment of the variance in size and shape, we plotted histograms for each measure, fitting the data to Gaussian distributions (Figure 5E,F). We found that for both size and circularity, mutants lacking the fission protein had a larger variance (p < 0.0001 for both), highlighted by the presence of larger and more elongated mitochondria/mitochondrial networks (see Figure 5Bii for examples). Overall, we found evidence to suggest that mutation of drp-1 causes a lack of fission within PLM neurons, resulting in larger and more elongated mitochondria. We also demonstrated that disruption of mitochondrial fission causes an increase in the variance of mitochondrial morphology.
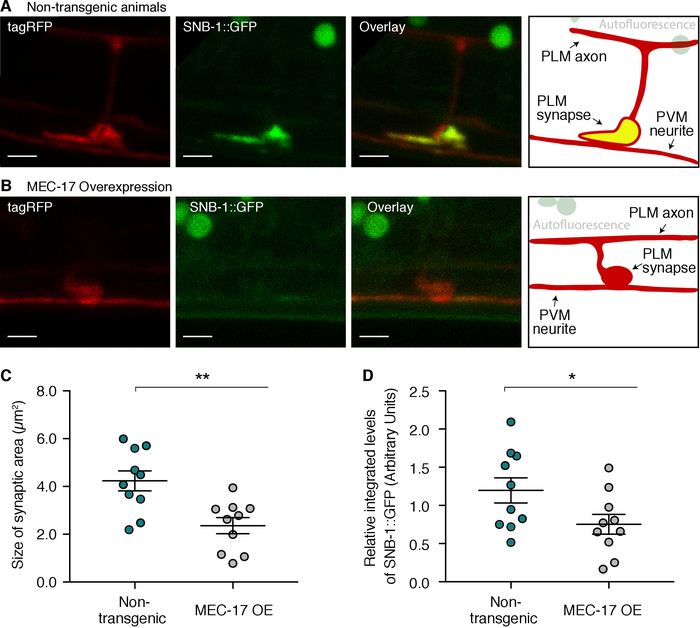
Figure 1: Quantification of synapse integrity in the PLM neurons. (A) Images are maximum z-projections of the PLM synaptic region in a non-transgenic animal. The left panel shows the PLM neuron labelled with tagRFP, the second panel displays the pre-synaptic region labelled with SNB-1::GFP, the third image is an overlay with co-localization represented in yellow, and the fourth panel is a schematic of the overlay image. (B) Maximum z-projection confocal images of the synaptic region in animals overexpressing MEC-17. Images are displayed as per panel A. (C) Size quantification of the pre-synaptic region in non-transgenic compared to transgenic animals overexpressing MEC-17. (D) Quantification of the relative integrated fluorescence levels of SNB-1::GFP in non-transgenics versus transgenic animals overexpressing MEC-17. For (C) and (D), individual synapses analyzed are represented by circles; n ≥ 10; mean ± S.E. shown in black. P values * < 0.05, ** < 0.01 from unpaired t-tests; scale bars represent 2 µm. Please click here to view a larger version of this figure.
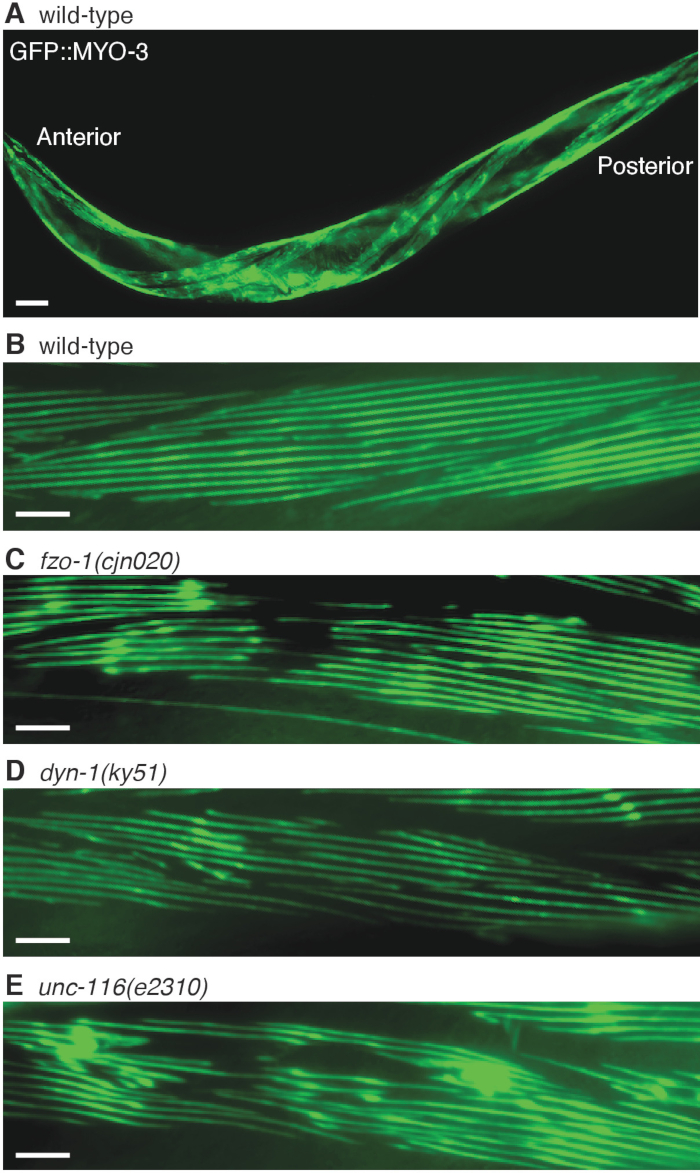
Figure 2: Visualization of the C. elegans body wall muscles. (A) An animal expressing the stEx30(Pmyo-3::gfp::myo-3 + rol-6(su1006)) transgene, which labels myosin heavy chain with GFP in the body wall muscles. The array also induces a rolling phenotype in the animal that facilitates visualization of the muscle cells arranged along the dorsal and ventral sides of the body. Representative magnified views of myosin fibers in (B) wild-type, (C) fzo-1(cjn020), (D) dyn-1(ky51) and (E) unc-116(e2310) animals. All animals were imaged at the 3-day old adult stage. Scale bar represents 60 µm in (A), and 30 µm in (B-E). Please click here to view a larger version of this figure.
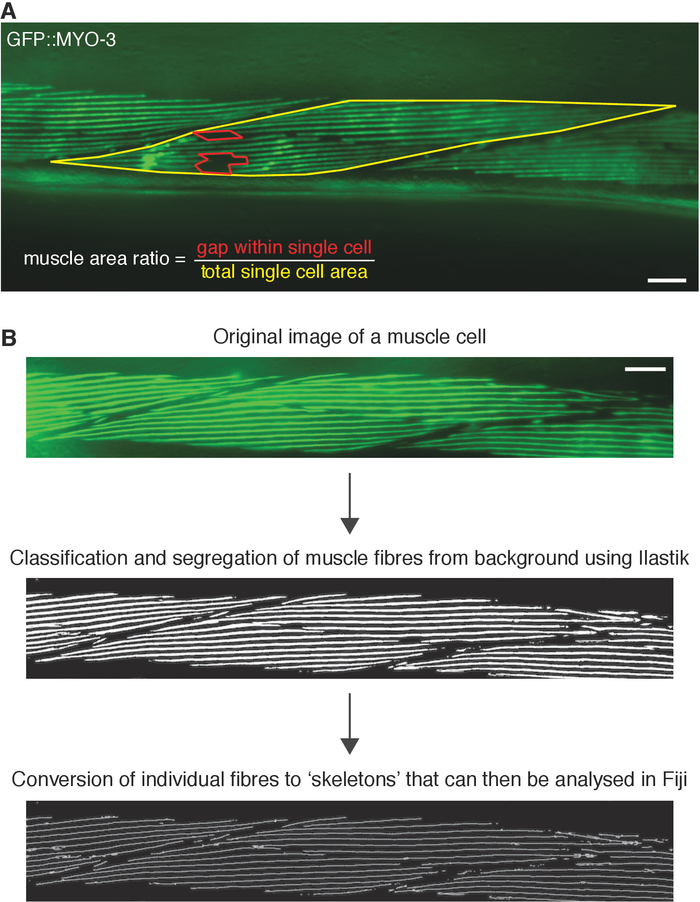
Figure 3: Measuring body wall muscle area and fiber length using available computational algorithms. (A) Example image demonstrating how the internal gap space within a single muscle cell (red-enclosed), and the area of the entire cell (yellow-enclosed) are drawn and calculated in Fiji to obtain the muscle area ratio. (B) Outline of segmentation protocol on a single example image using a combination of Fiji and ilastik. Scale bar represents 30 µm. Please click here to view a larger version of this figure.
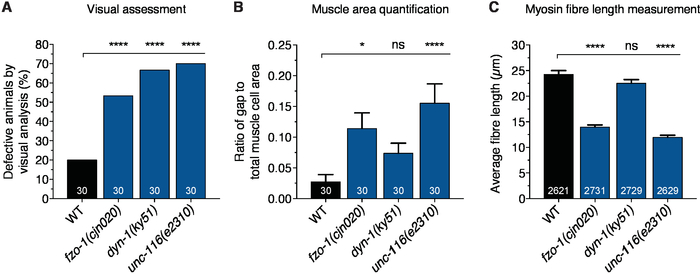
Figure 4: Qualification and quantification of body wall muscle defects. (A) Visual assessment of body wall muscle defects. (B) Comparison of blank space area to total muscle cell area ratio between WT and indicated mutants. (C) Measurement of myosin fiber length. For (B) and (C), only images with at least one complete visible oblique muscle cell were included for analysis. Fibers with length of 0 µm or longer than 250 µm were also excluded. Bars represent percentage defective animals in (A) and mean ± S.E.M. in (B) and (C); n values listed in each bar. Chi-square test with false discovery rate were performed to compare visual defects in (A), whereas one-way ANOVA with Dunnett’s post hoc tests for multiple comparison were used to analyze quantitative measurements in (B) and (C). *P < 0.05, ****P < 0.0001, ns = not significant. Please click here to view a larger version of this figure.
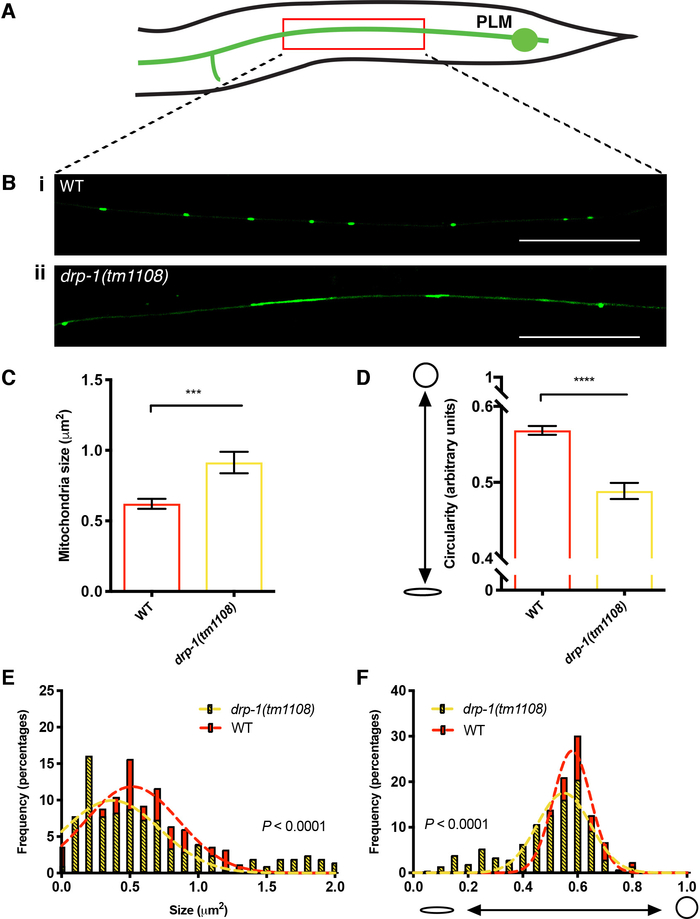
Figure 5: Mitochondrial morphology within the axons of the PLM neurons. (A) Schematic of a posterior lateral microtubule (PLM) mechanosensory neuron in the tail of C. elegans. The red box indicates the approximate location of the highlighted sections in (B), which show representative images of GFP labelled mitochondria (Pmec-4::MLS::GFP) within the PLM axon. Bright green puncta indicate mitochondria. Compared to the WT (i), drp-1(tm1108) mutants (ii) show larger, more elongated mitochondria. Images are representative of n = 6 PLM axons from 6 worms per genotype; scale bars = 15 μm. (C) Mean size (μm2) of individual mitochondrion as quantified by SQUASSH object segmentation in 3-day old adult (3DOA) worms. (D) Mean Circularity of individual mitochondrion within the PLM axon, as quantified from SQUASSH object measurements in 3DOAs. (E) Histogram and Gaussian distribution showing the variance of mitochondrial size. F test (P < 0.0001) shows that variances are significantly different between drp-1 and WT. (F) Histogram and Gaussian distribution showing the variance of mitochondrial circularity. Data is represented as mean +/- SEM. *** P < 0.001, **** P <0.0001 from Welch’s t-test; n ≥ 204 mitochondria for quantitative analysis from n = 6 worms. Please click here to view a larger version of this figure.
| Strain name | Genotype | Source | Reference # | ||
| BXN038 | uIs115(Pmec-17::tagRFP); jsIs609(Pmec-4::MLS::GFP) | Neumann lab | 7 | ||
| BXN366 | fzo-1(cjn020); myo-3(st386); zdIs5(Pmec-4::GFP); stEx30(Pmyo-3::gfp::myo-3 + rol-6(su1006)) | Neumann lab | 20 | ||
| BXN419 | drp-1(tm1108); jsIs609(Pmec-4::MLS::GFP); uIs115(Pmec-17::tagRFP) | Neumann lab | 7 | ||
| BXN507 | cjnEx036(Pmec-4::mec-17, Pmyo-2::mCherry); jsIs37(Pmec-7::snb-1::GFP); lin-15B&lin-15A(n765); uIs115(Pmec-17::tagRFP) | Neumann lab | 17 | ||
| BXN675 | unc-116(e2310); myo-3(st386); stEx30(Pmyo-3::gfp::myo-3 + rol-6(su1006)) | Neumann lab | 20 | ||
| BXN685 | dyn-1(ky51); myo-3(st386); stEx30(Pmyo-3::gfp::myo-3 + rol-6(su1006)) | Neumann lab | 20 | ||
| NM664 | jsIs37(Pmec-7::snb-1::GFP); lin-15B&lin-15A(n765) | Caenorhabditis Genetics Center | 6 | ||
| RW1596 | myo-3(st386); stEx30(Pmyo-3::gfp::myo-3 + rol-6(su1006)) | Caenorhabditis Genetics Center | 8 | ||
| TU4065 | uIs115(Pmec-17::tagRFP) | Martin Chalfie | 5 | ||
Table 1. List of C. elegans strains used in this study.
| PLM axon images | |
| Objective | 40x |
| Image resolution (per panel for tilescan) | 3224 x 3224 |
| Background removal window size | 20 |
| PSF xy | 0.78 |
| PSF z | 0.68 |
| Regularisation | 0.15 |
| Minimum fluorescence intensity of GFP | 0.4 |
Table 2. SQUASSH parameters for segmentation of mitochondria.
Discussion
Morphological variations have frequently been assessed via manual counting of noticeable differences or using arbitrary thresholds to determine defects in comparison to a wild-type phenotype. More recently, however, quantitative methods have been used for comparative studies of morphology to accurately measure and describe changes on a cellular and subcellular level in an unbiased fashion. The ability to identify subtle yet biologically relevant differences between phenotypes is a powerful means for understanding the underlying molecular mechanisms that control changes in morphology caused by environmental factors or genetic diseases. The continued increases in computation power and microscopy imaging resolution, coupled with the ease of growth and the versatility of C. elegans genetics has provided the perfect platform for the combination of sophisticated yet free imaging software with fine detailed confocal images. Here, we have demonstrated methods to evaluate and quantify synaptic size and protein localization, body wall muscle area and fiber length, and mitochondrial morphology in a range of genetic backgrounds.
While these analyses have demonstrated an ability to quantify subtle morphological differences, there are limitations. Although each of these analyses make use of open-source and readily available software, they rely on high quality, sharp images to accurately resolve cellular structures and subcellular organelles. This can often be a limiting step, as sensitive high-end equipment, such as confocal microscopes with advanced detectors, are required to capture images with satisfactory resolution. These are not always accessible to laboratories, and analysis with sub-par technology can lead to inaccurate and unrepeatable results. In addition, while each technique aims to reduce experimenter bias by using computer algorithms, there are still steps that can involve an element of human error. The manual definition of the synapse by hand, the outlining of the muscle cell area, the parameter optimizations of mitochondrial segmentation by eye are all areas that can be improved in this respect. To reduce the chance for human error, studies could be conducted in a blinded fashion or alternatively image analysis software could be used to define the region of interest itself. Using cell masks, in which the desired area for analysis is defined by the fluorescence at a different wavelength, may help this. This is used to limit the analysis to a section of an image displaying fluorescence above a set threshold, such as a cell’s nucleus or a transfected cell13,24. Furthermore, the use of single time point images limits the questions these techniques can address. The processes that govern organelle morphology, synapse formation and muscle structure are most likely dynamic, and differences can be seen within the same cell depending on the developmental or biological status. Analysis of these structures at a specific end-point can therefore be limiting and time-lapse imaging would be beneficial for monitoring structural changes over time. Finally, while morphological changes can be a good indicator of disruption in cellular processes, correlation is not always implicit. For example, recent findings in different species have shown that mitochondrial function and form can indeed be dissociated7,25. Therefore, additional assays must be taken into account when inferring functional changes as the result of morphological differences.
We have been able to demonstrate the use of these quantitative techniques in a particular set of tissues and genetic backgrounds; however, their usage can be applied more broadly. Fluorescence quantification via imaging software can be used to analyze a range of different subcellular organelles such as nuclei and endosomes, or different structures such as neurons and epithelial cells, all within the context of different mutant or disease backgrounds.
Divulgazioni
The authors have nothing to disclose.
Acknowledgements
We thank members of the Neumann lab for valuable discussions and input. Some strains were provided by the CGC, which is funded by NIH Office of Research Infrastructure Programs (P40 OD010440). The authors thank WormBase for its wealth of information on C. elegans, and acknowledge Monash Micro Imaging, Monash University, for the provision of instrumentation, training and technical support. This work was supported by CMTAA research grants (2015 and 2018), and NHMRC Project Grants 1101974 and 1099690 awarded to B.N.
Materials
| Agar-agar | Merck | 1.01614.1000 | |
| Agarose | Invitrogen | 16500-500 | |
| Confocal microscope | Leica | TCS SP8 | Inverted platform |
| Fluorescence microscope | Carl Zeiss AG | Zeiss Axio Imager M2 | |
| Glass coverslips #1 | Thermo scientifique | MENCS22221GP | |
| Glass coverslips #1.5 | Zeiss | 474030-9000-000 | Made by SCHOTT |
| Glass slides | Thermo scientifique | MENS41104A/40 | |
| Light LED | Schott | KL 300 LED | |
| Stereo Microscope | Olympus | SZ51 | |
| Tryptone (Peptone from casein) | Merck | 107213 | Ingredients for Lysogeny Broth (LB) medium |
| Yeast Extract | Merck | 103753 | Ingredients for Lysogeny Broth (LB) medium |
| Sodium chloride | Merck | 106404 | Ingredients for Lysogeny Broth (LB) medium |
| Peptone (Peptone from meat) | Merck | 107214 | Ingredients for Nematode Growth Media (NGM) agar |
| Agar | Sigma | A1296 | Ingredients for Nematode Growth Media (NGM) agar |
| Sodium chloride | Merck | 106404 | Ingredients for Nematode Growth Media (NGM) agar |
| Cholesterol | Sigma | C8667-25G | Ingredients for Nematode Growth Media (NGM) agar |
| Calcium chloride | Merck | 102382 | Ingredients for Nematode Growth Media (NGM) agar |
| Magnesium sulfate | Merck | 105886 | Ingredients for Nematode Growth Media (NGM) agar |
| Dipotassium phosphate | Merck | 105101 | Ingredients for Nematode Growth Media (NGM) agar |
| Potassium dihydrogen phosphate | Merck | 104873 | Ingredients for Nematode Growth Media (NGM) agar |
| Disodium phosphate | Merck | 106586 | Ingredients for M9 buffer |
| Sodium chloride | Merck | 106404 | Ingredients for M9 buffer |
| Potassium dihydrogen phosphate | Merck | 104873 | Ingredients for M9 buffer |
| Magnesium sulfate | Merck | 105886 | Ingredients for M9 buffer |
| Pasteur pipette | Corning | CLS7095D5X-200EA | |
| Petri dishes | Corning | CLS430589-500EA | |
| Platinum wire | Sigma | 267201-2G | |
| Spatula | Met-app | 2616 | |
| Tetramisole hydrochloride | Sigma | L9756-5G |
Riferimenti
- Brenner, S. The genetics of Caenorhabditis elegans. Genetica. , (1974).
- Markaki, M., Tavernarakis, N. Modeling human diseases in Caenorhabditis elegans. Biotechnology Journal. 5 (12), 1261-1276 (2010).
- Laranjeiro, R., Harinath, G., Burke, D., Braeckman, B. P., Driscoll, M. Single swim sessions in C. elegans induce key features of mammalian exercise. BMC Biology. , (2017).
- Alexander, A. G., Marfil, V., Li, C. Use of C. elegans as a model to study Alzheimer’s disease and other neurodegenerative diseases. Frontiers in Genetics. , (2014).
- Zheng, C., Jin, F. Q., Chalfie, M. Hox Proteins Act as Transcriptional Guarantors to Ensure Terminal Differentiation. Cell Reports. 13 (7), 1343-1352 (2015).
- Koushika, S. P., et al. Mutations in Caenorhabditis elegans cytoplasmic dynein components reveal specificity of neuronal retrograde cargo. Journal of Neuroscience. 24 (16), 3907-3916 (2004).
- Byrne, J. J., et al. Disruption of mitochondrial dynamics affects behaviour and lifespan in Caenorhabditis elegans. Cellular and Molecular Life Sciences. , (2019).
- Campagnola, P. J., et al. Three-Dimensional High-Resolution Second-Harmonic Generation Imaging of Endogenous Structural Proteins in Biological Tissues. Biophysical Journal. 82 (1), 493-508 (2002).
- Schindelin, J., et al. Fiji: An open-source platform for biological-image analysis. Nature Methods. 9 (7), 676-682 (2012).
- Sommer, C., Straehle, C., Kothe, U., Hamprecht, F. A. Ilastik: Interactive learning and segmentation toolkit. 2011 IEEE International Symposium on Biomedical Imaging: From Nano to Macro. , (2011).
- Mondal, S., Ahlawat, S., Koushika, S. P. Simple microfluidic devices for in vivo imaging of C. elegans, Drosophila and zebrafish. Journal of Visualized Experiments. , (2012).
- Paul, G., Cardinale, J., Sbalzarini, I. F. Coupling image restoration and segmentation: A generalized linear model/bregman perspective. International Journal of Computer Vision. , (2013).
- Rizk, A., et al. Segmentation and quantification of subcellular structures in fluorescence microscopy images using Squassh. Nature Protocols. 9, 586-586 (2014).
- Akella, J. S., et al. MEC-17 is an alpha-tubulin acetyltransferase. Nature. 467 (7312), 218-222 (2010).
- Neumann, B., Hilliard, M. A. Loss of MEC-17 leads to microtubule instability and axonal degeneration. Cell Reports. 6 (1), 93-103 (2014).
- Shida, T., Cueva, J. G., Xu, Z., Goodman, M. B., Nachury, M. V. The major alpha-tubulin K40 acetyltransferase alphaTAT1 promotes rapid ciliogenesis and efficient mechanosensation. Proceedings of the National Academy of Sciences of the United States of America. 107 (50), 21517-21522 (2010).
- Teoh, J. -. S., Wang, W., Chandhok, G., Pocock, R., Neumann, B. Development and maintenance of synaptic structure is mediated by the alpha-tubulin acetyltransferase MEC-17/αTAT1. bioRxiv. 84, 522904-522904 (2019).
- Bird, T. D. . Charcot-Marie-Tooth Neuropathy Type 2. , (1998).
- Chandhok, G., Lazarou, M., Structure Neumann, B. Structure, function, and regulation of mitofusin-2 in health and disease. Biological Reviews. 93 (2), 933-949 (2018).
- Soh, M. S., Cheng, X., Liu, J., Neumann, B. Disruption of genes associated with Charcot-Marie-Tooth type 2 lead to common behavioural, cellular and molecular defects in Caenorhabditis elegans. bioRxiv. , 605584 (2019).
- Kamerkar, S. C., Kraus, F., Sharpe, A. J., Pucadyil, T. J., Ryan, M. T. Dynamin-related protein 1 has membrane constricting and severing abilities sufficient for mitochondrial and peroxisomal fission. Nature Communications. , (2018).
- Zhu, P. -. P., et al. Intra- and Intermolecular Domain Interactions of the C-terminal GTPase Effector Domain of the Multimeric Dynamin-like GTPase Drp1. Journal of Biological Chemistry. 279 (34), 35967-35974 (2004).
- Osellame, L. D., et al. Cooperative and independent roles of the Drp1 adaptors Mff, MiD49 and MiD51 in mitochondrial fission. Journal of Cell Science. , (2016).
- Dima, A. A., et al. Comparison of segmentation algorithms for fluorescence microscopy images of cells. Cytometry Part A. 79 (7), 545-559 (2011).
- Trevisan, T., et al. Manipulation of Mitochondria Dynamics Reveals Separate Roles for Form and Function in Mitochondria Distribution. Cell Reports. , (2018).

